CLASSIFICATION OF AMINO ACIDS R group at neutral











































- Slides: 92




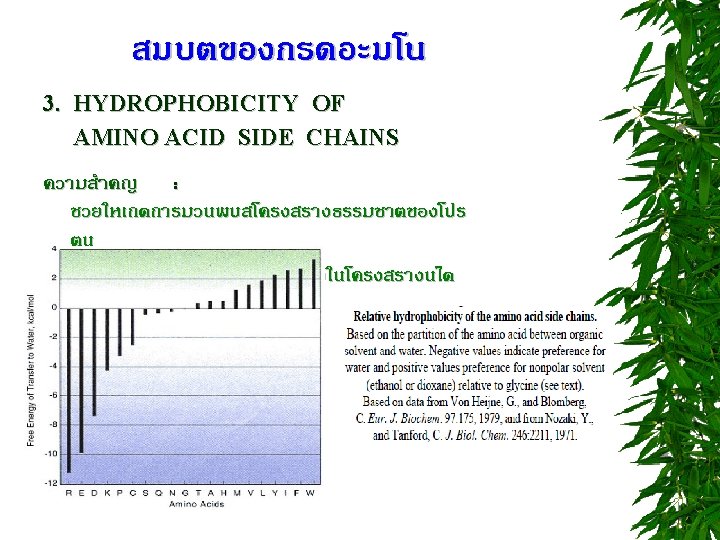
CLASSIFICATION OF AMINO ACIDS * R group at neutral p. H * Based on nutritional/physiological roles

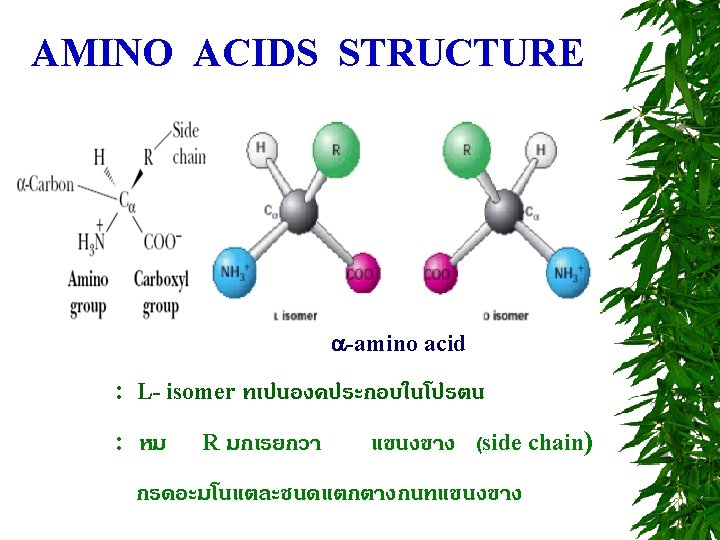
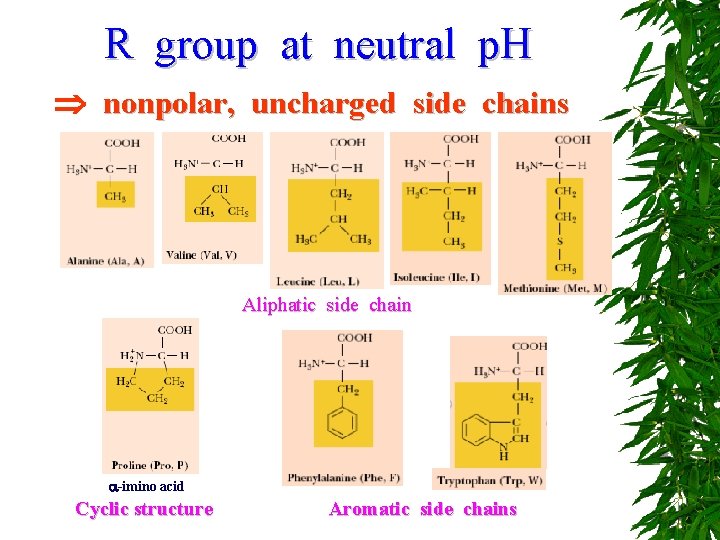
R group at neutral p. H nonpolar, uncharged side chains Aliphatic side chain -imino acid Cyclic structure Aromatic side chains

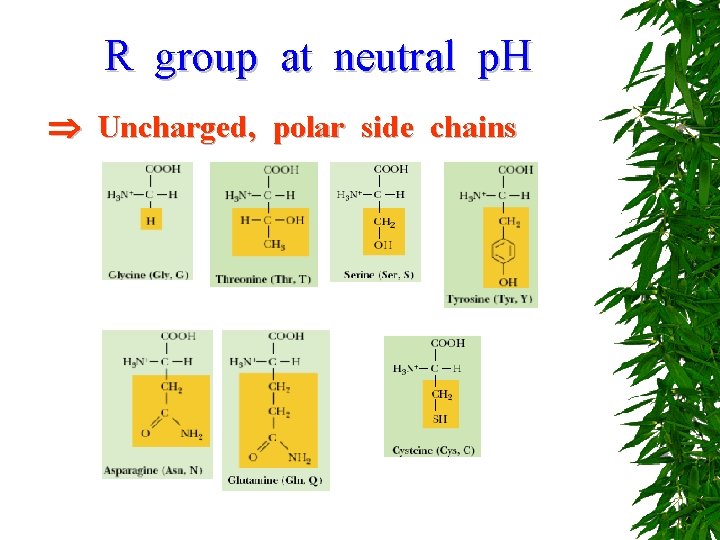
R group at neutral p. H Uncharged, polar side chains

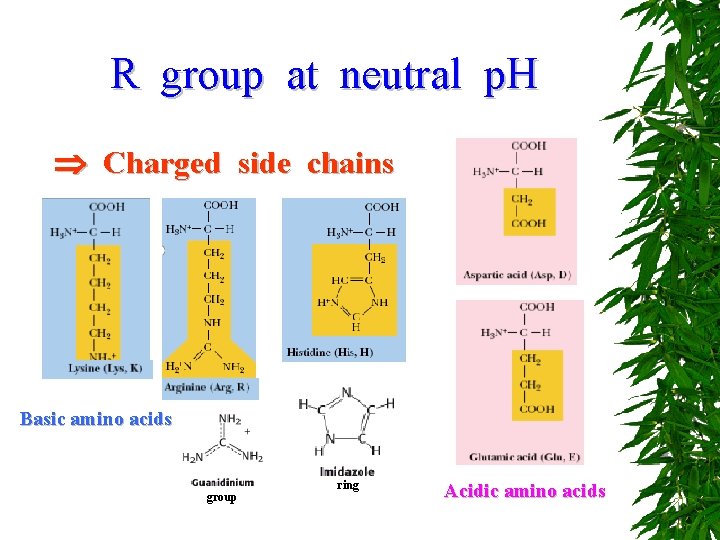
R group at neutral p. H Charged side chains Basic amino acids group ring Acidic amino acids




Based on nutritional/physiological roles Essential amino acids valine, leucine, isoleucine, phenylalanine, tryptophan, methionine, threonine, histidine (infant), lysine, arginine (semi-essential) Non-essential amino acids glycine, alanine, proline, serine, cysteine, tyrosine, asparagine, glutamine, aspartic acid, glutamic acid


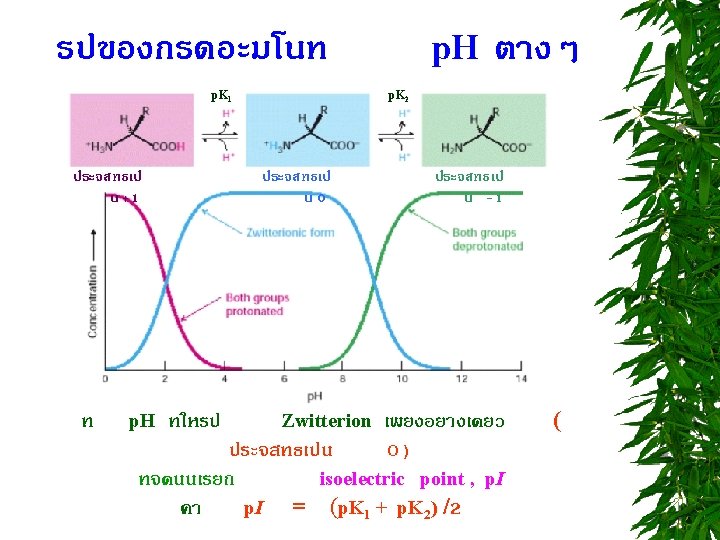
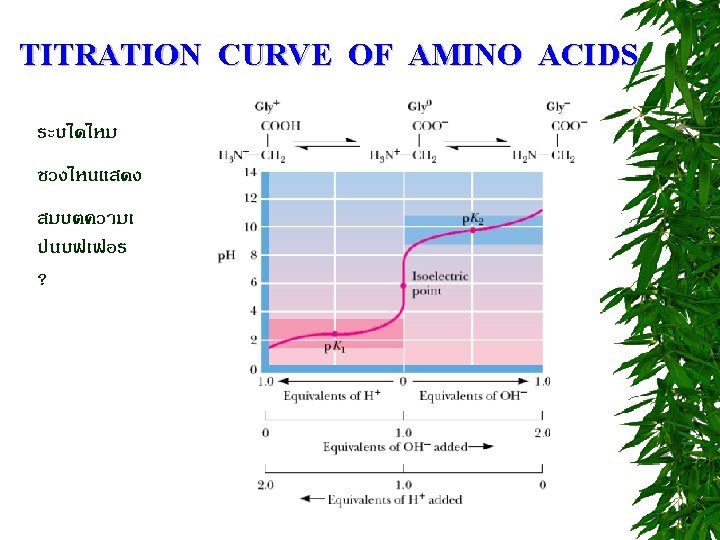
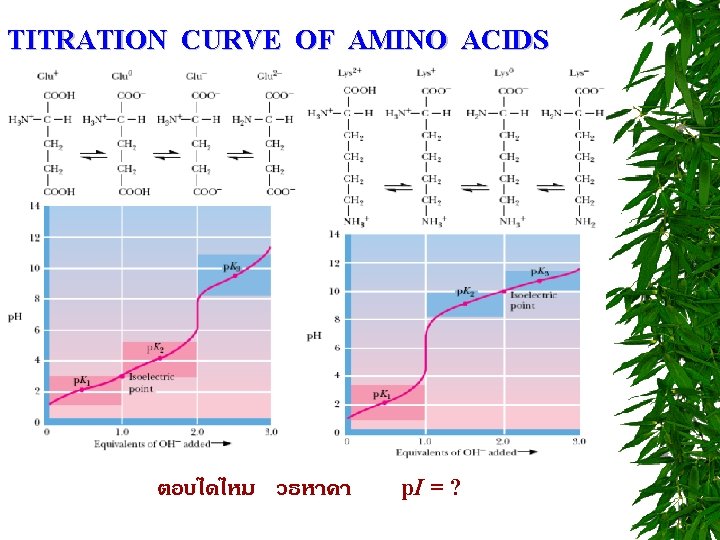
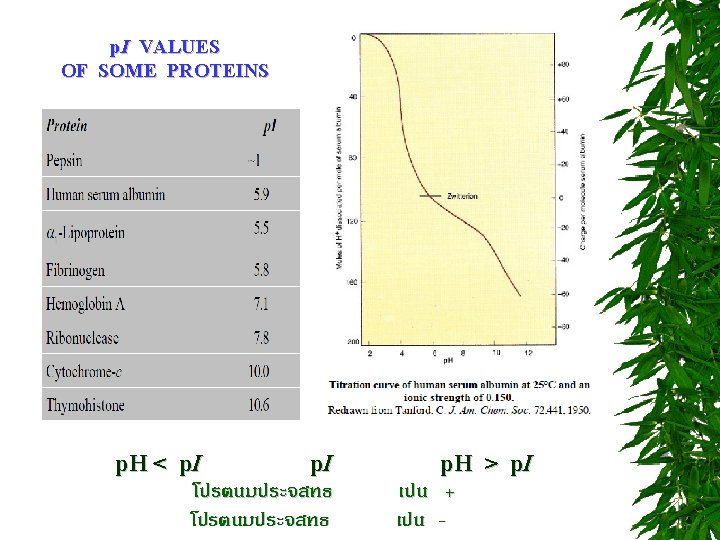
TITRATION CURVE OF AMINO ACIDS ตอบไดไหม วธหาคา p. I = ?


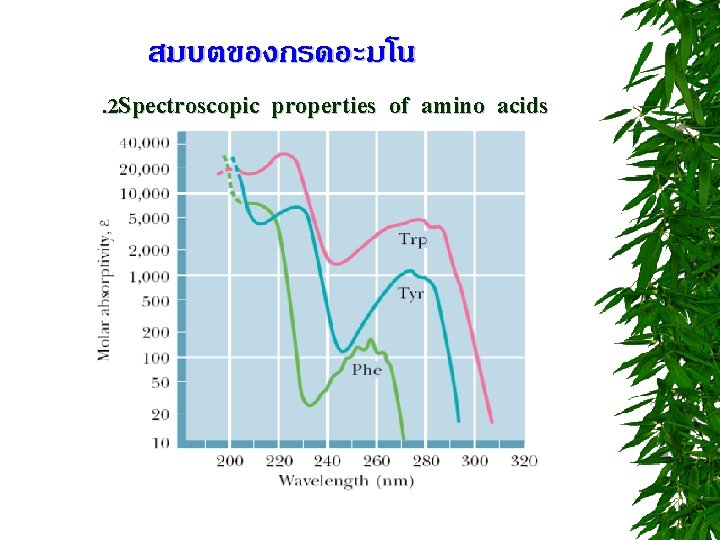
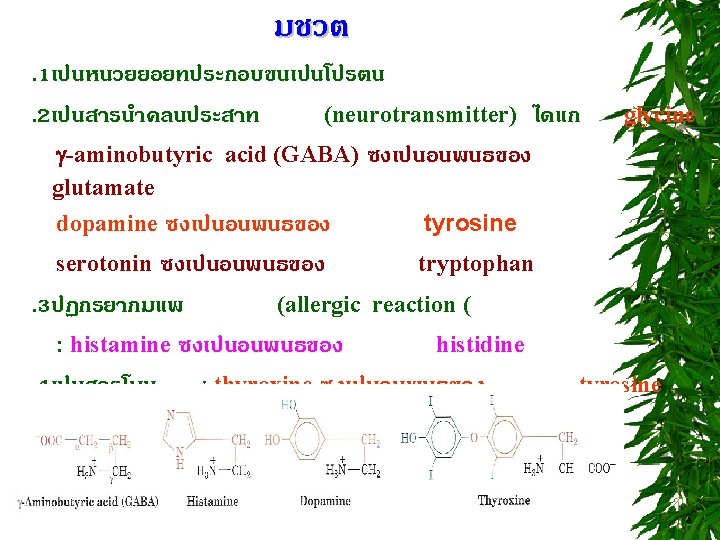
สมบตของกรดอะมโน. 2 Spectroscopic properties of amino acids





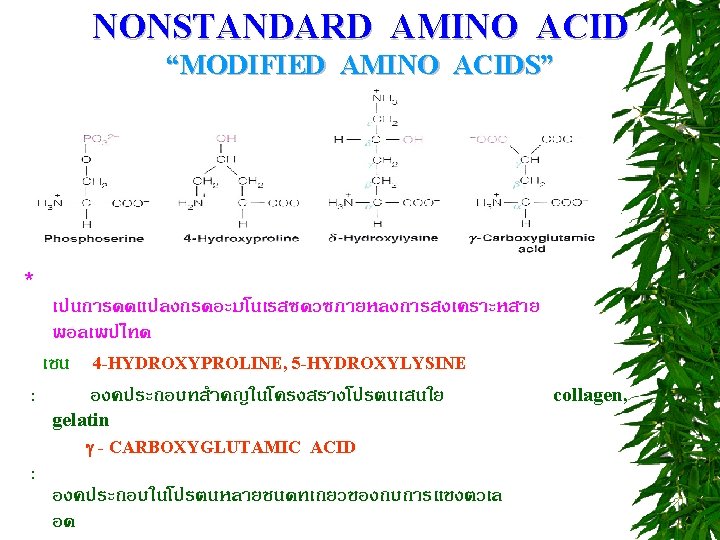
NONSTANDARD AMINO ACID “MODIFIED AMINO ACIDS” กรดอะมโนเรสซดวซบางชนดในโปรตน เชน histone อาจถกเตมหม methyl, หม acetyl, หม phosphate ตวอยาง o–phoserine, N–acetylserine, 3 -methylhistidine







PROTEINS






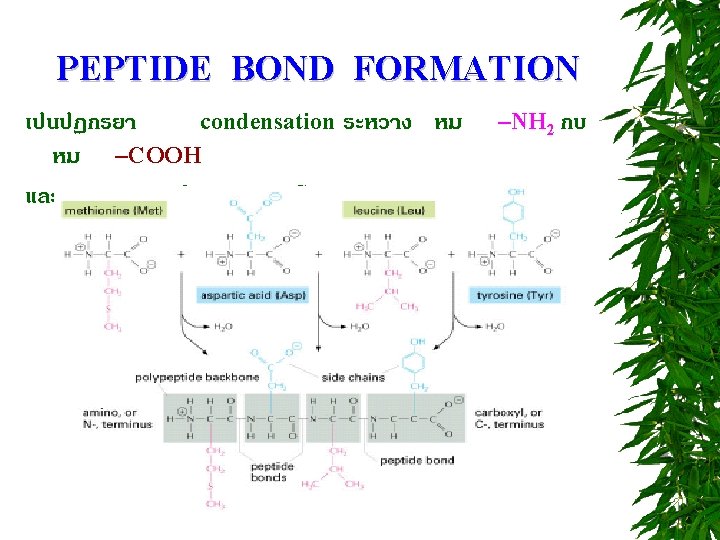
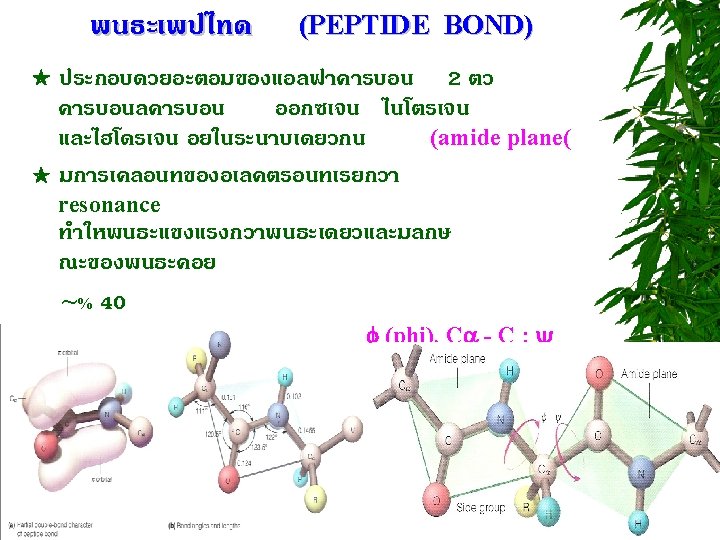
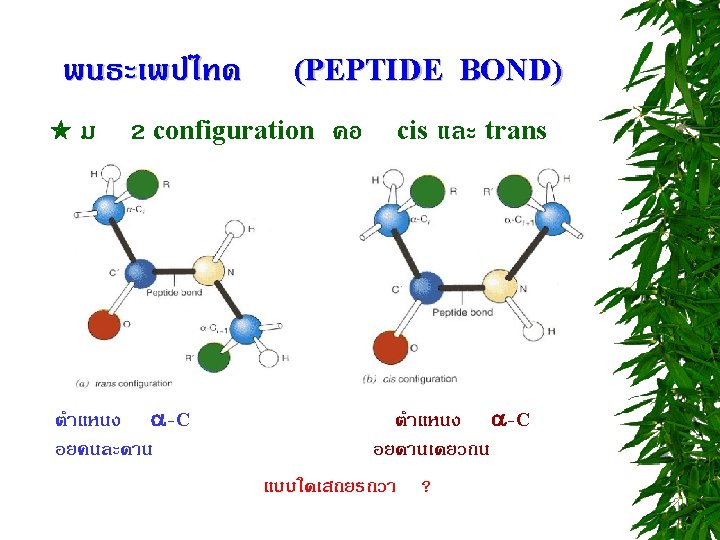
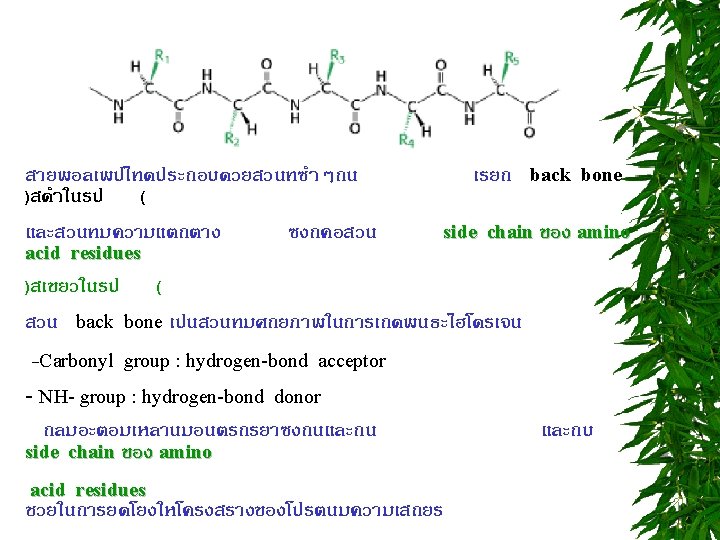
PEPTIDE BOND FORMATION N-terminus C-terminus









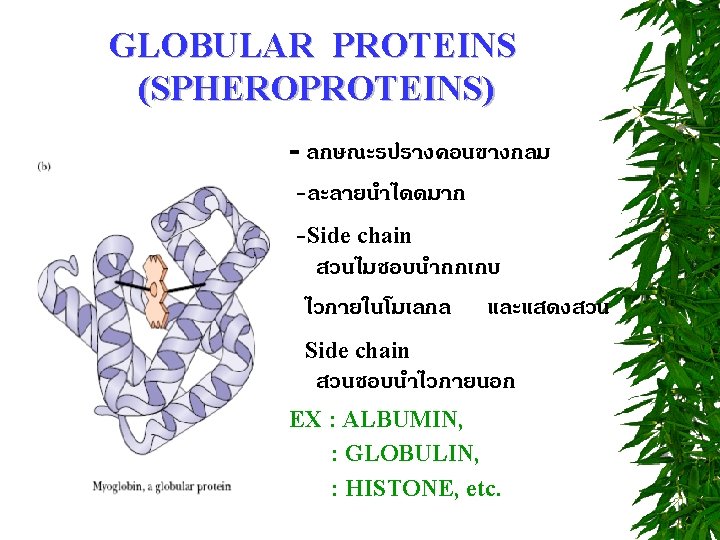
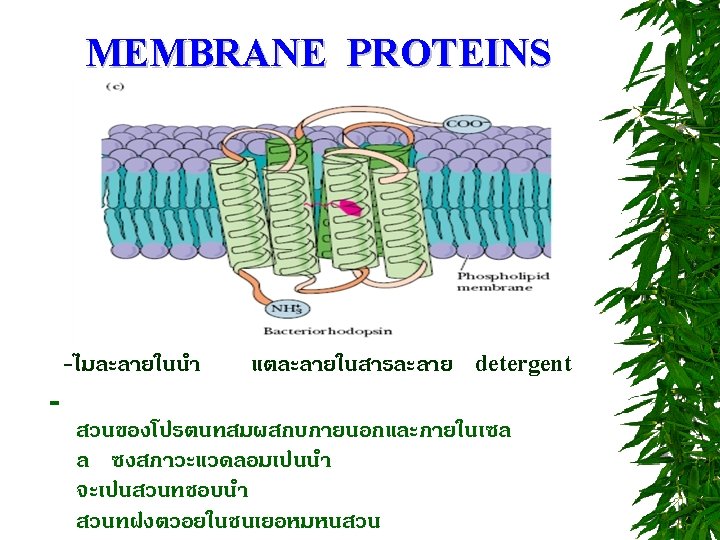
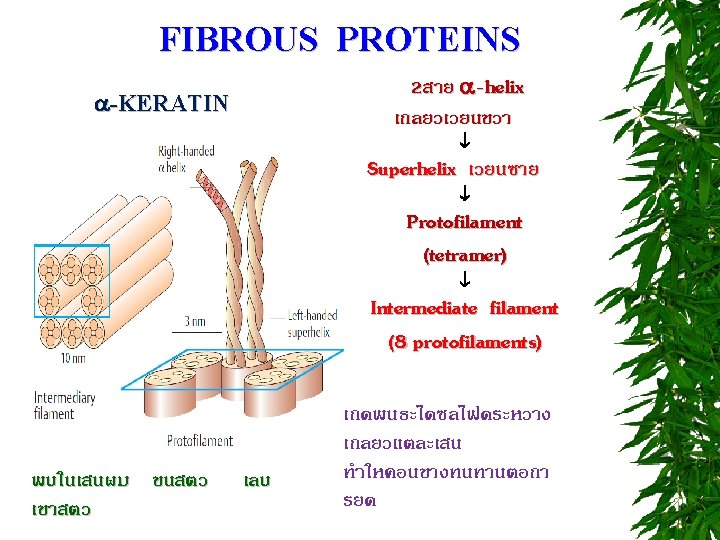
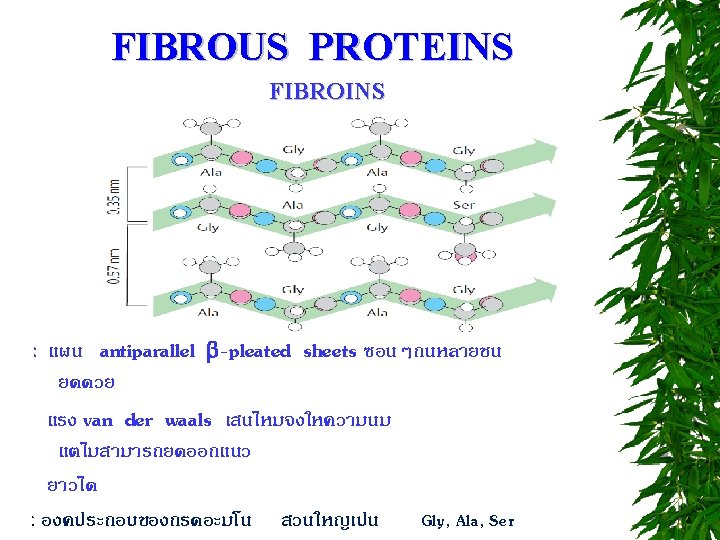
CLASSIFICATION OF PROTEINS BASED ON SHAPE FIBROUS PROTEINS (SCLEROPROTEINS) GLOBULAR PROTEINS (SPHEROPROTEINS) MEMBRANE PROTEINS




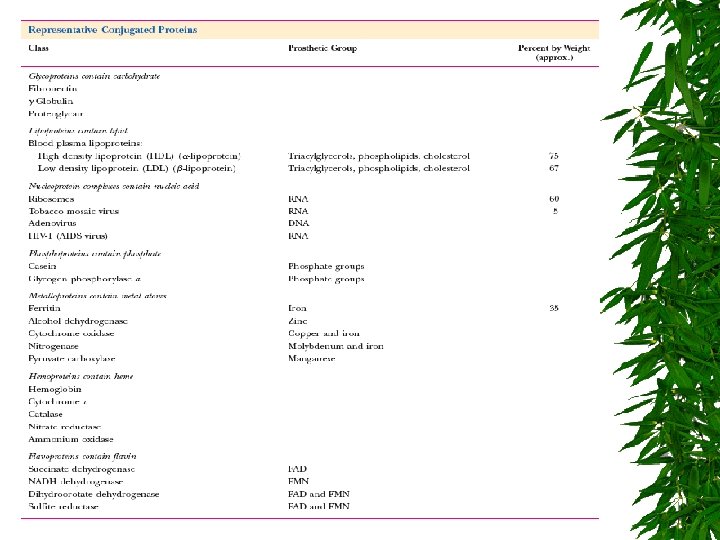
CLASSIFICATION OF PROTEINS BASED ON COMPOSITION SIMPLE PROTEINS CONSIST OF ONLY AMINO ACIDS AND CONTAIN NO OTHER CHEMICAL GROUPS EX. RIBONUCLEASE, ACTIN CONJUGATED PROTEINS ATTACHED TO NON-PROTEIN GROUP : PROSTHETIC GROUP NUCLEOPROTEINS LIPOPROTEINS CHROMOPROTEINS GLYCOPROTEINS PHOSPHOPROTEINS METALLOPROTEINS ATTACHED ATTACHED TO TO TO NUCLEIC ACID LIPID CHROMOPHORE CARBOHYDRATE PHOSPHATE METAL ION



BIOLOGICAL ROLES OF PROTEINS 1. ENERGY SOURCE 2. STRUCTURE EX : COLLAGEN, FIBOIN, ELASTIN 3. MOVEMENT EX : ACTIN, TUBULIN, CYTOSKELETON PROTEINS 4. PROTECTIVE EX : KERATIN, FIBRINOGEN AND THROMBIN, IMMUNOGLOBULIN 5. REGULATION HORMONE, TRANSCRIPTIONAL FACTOR 6. TRANSPORT EX : GLUCOSE TRANSPORTER, Na+-K+ ATPase 7. CATALYSIS ENZYME










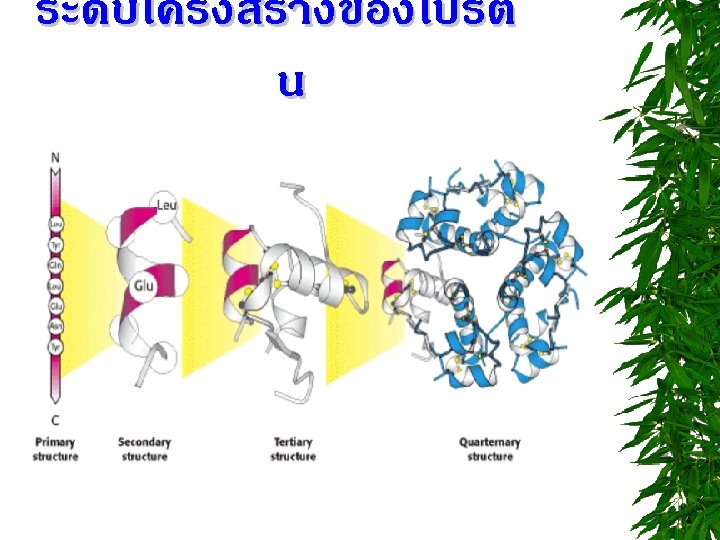
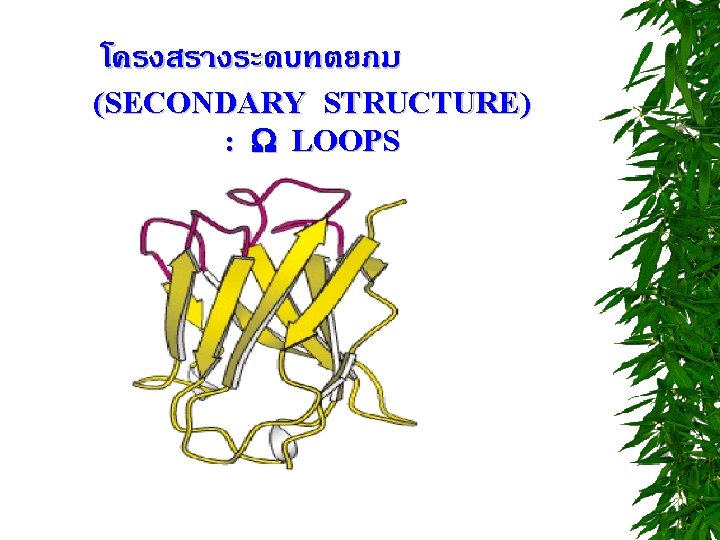
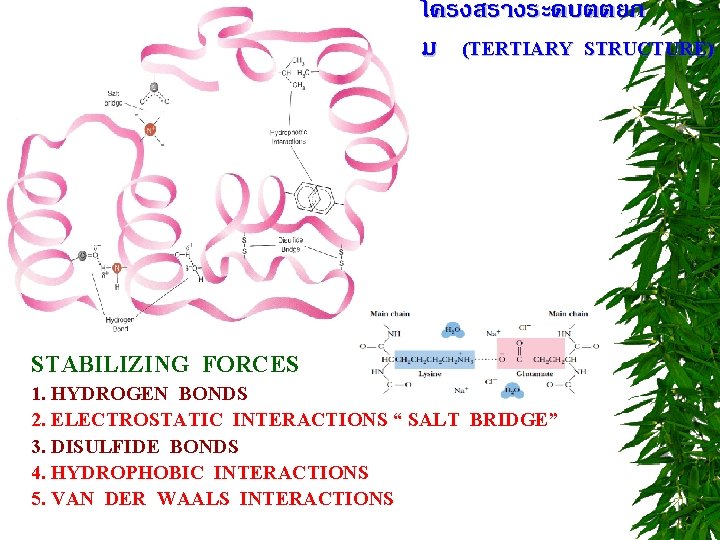
โครงสรางระดบตตยภ ม (TERTIARY STRUCTURE) STABILIZING FORCES 1. HYDROGEN BONDS 2. ELECTROSTATIC INTERACTIONS “ SALT BRIDGE” 3. DISULFIDE BONDS 4. HYDROPHOBIC INTERACTIONS 5. VAN DER WAALS INTERACTIONS



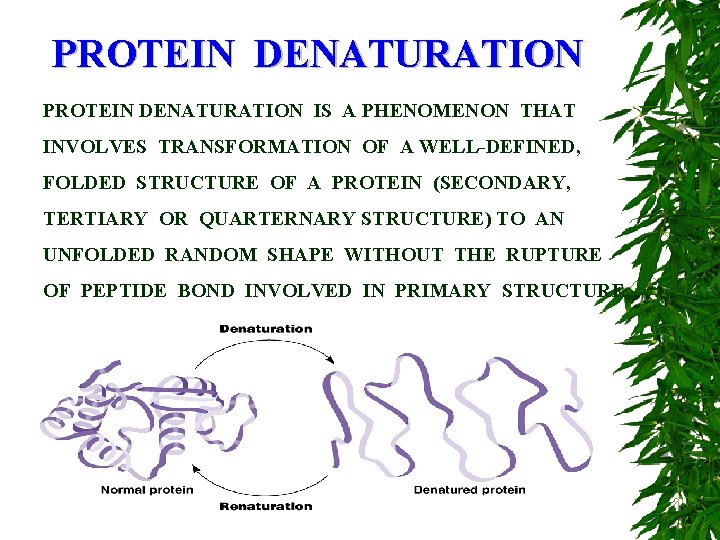
PROTEIN DENATURATION IS A PHENOMENON THAT INVOLVES TRANSFORMATION OF A WELL-DEFINED, FOLDED STRUCTURE OF A PROTEIN (SECONDARY, TERTIARY OR QUARTERNARY STRUCTURE) TO AN UNFOLDED RANDOM SHAPE WITHOUT THE RUPTURE OF PEPTIDE BOND INVOLVED IN PRIMARY STRUCTURE.

PROTEIN DENATURATION EFFECTS : - DECREASED SOLUBILITY : UNMASKING OF HYDROPHOBIC PORTION - LOST BIOLOGICAL ACTIVITY : CATALYTIC PROPERTY : IMMUNOLOGICAL PROPERTY - INCREASE SUSCEPTIBILITY TO BREAKDOWN BY PROTEASE - INABILITY TO CRYSTALIZE

DENATURATING CONDITIONS 1. 2. 3. 4. 5. 6. 7. 8. STRONG ACID AND BASE ORGANIC SOLVENT DETERGENT REDUCING AGENTS SALT CONCENTRATION HEAVY METAL IONS TEMPERATURE CHANGE MECHANICAL STRESS

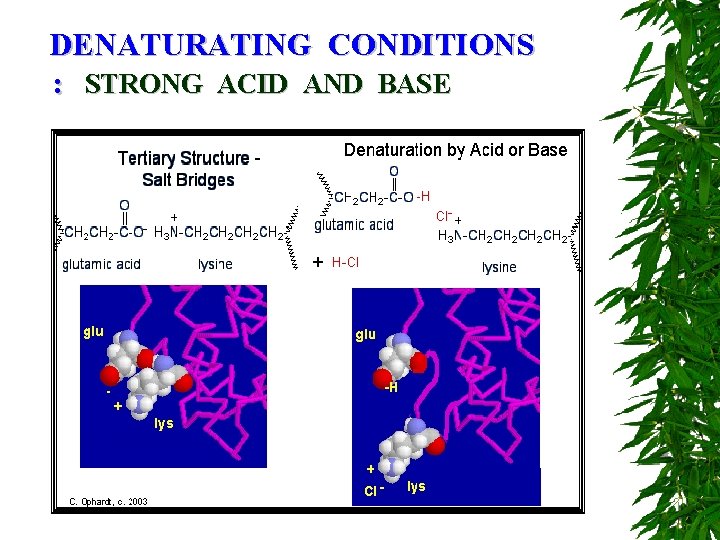
DENATURATING CONDITIONS : STRONG ACID AND BASE

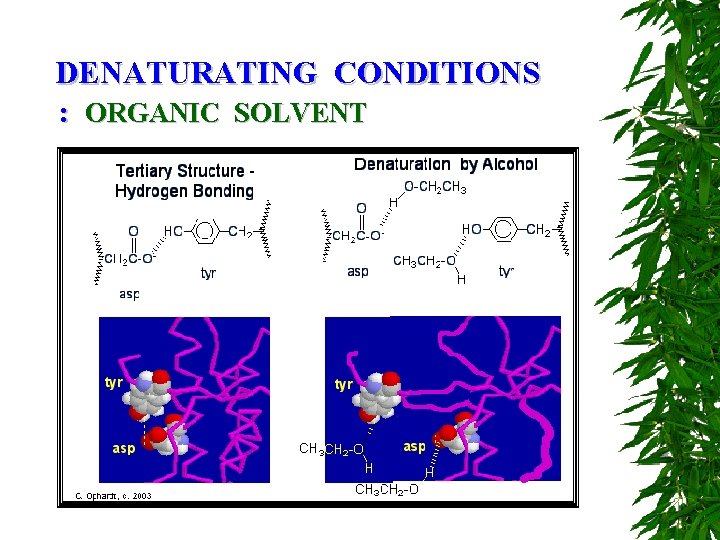
DENATURATING CONDITIONS : ORGANIC SOLVENT

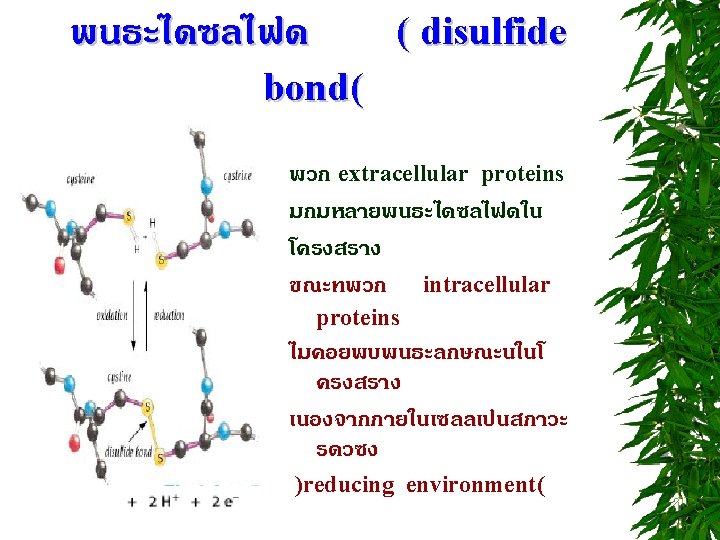
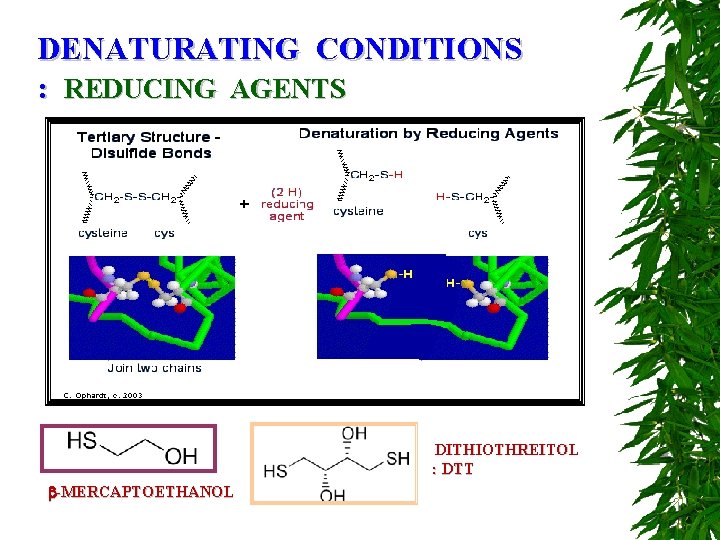
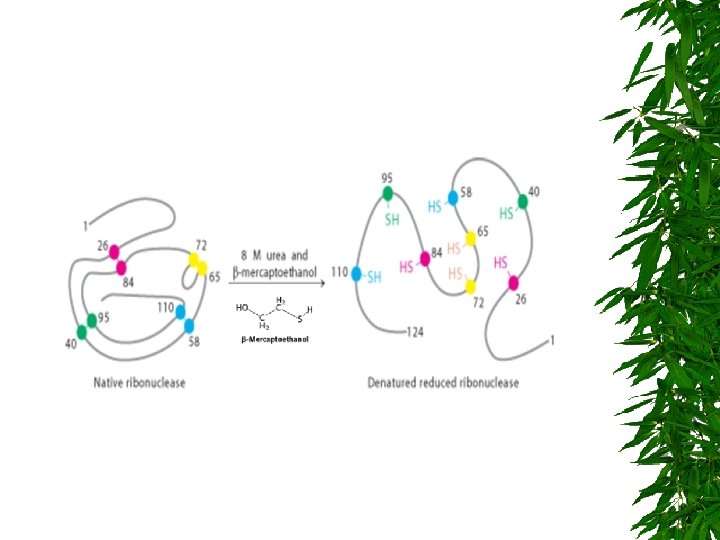
DENATURATING CONDITIONS : REDUCING AGENTS DITHIOTHREITOL : DTT -MERCAPTOETHANOL

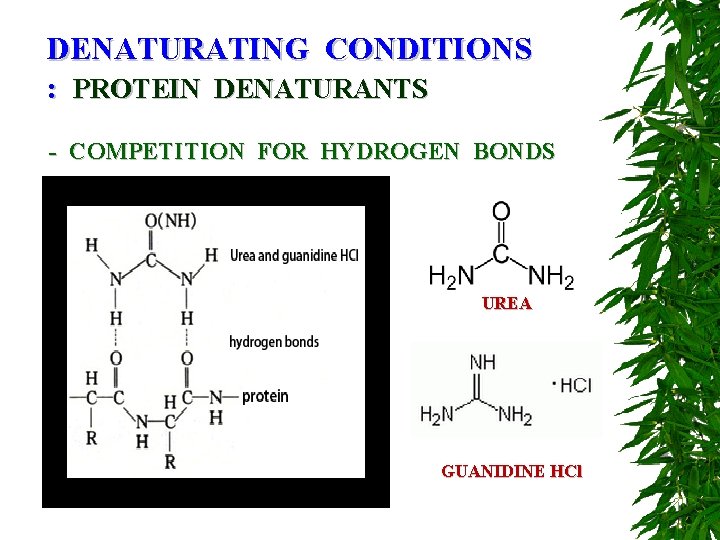
DENATURATING CONDITIONS : PROTEIN DENATURANTS - COMPETITION FOR HYDROGEN BONDS UREA GUANIDINE HCl

DENATURATING CONDITIONS : SALT CONCENTRATION

DENATURATING CONDITIONS : TEMPERATURE CHANGE HEAT : DISRUPTS HYDROGEN BONDS AND WEAK INTERACTION IN PROTEIN STRUCTURE DUE TO INCREASING OF TRANSLATIONAL AND VIBRATIONAL ENERGY COLD : FREEZING TEMPERATURE CAN DENATURE SOME PROTEINS

DENATURATING CONDITIONS : MECHANICAL STRESS AGITATION : SHEARING OF HYDROGEN BONDS

DENATURATING CONDITIONS : IRRADIATION - THE EFFECT DEPENDS ON THE WAVELENGHT AND ENERGY INVOLVED - RADIATION CAUSE : OXIDATION OF AMINO ACID RESIDUES : RUPTURE OF COVALENT BOND : IONIZATION : FORMATION OF PROTEIN FREE RADICAL




TECHNIQUES IN PROTEIN STUDY

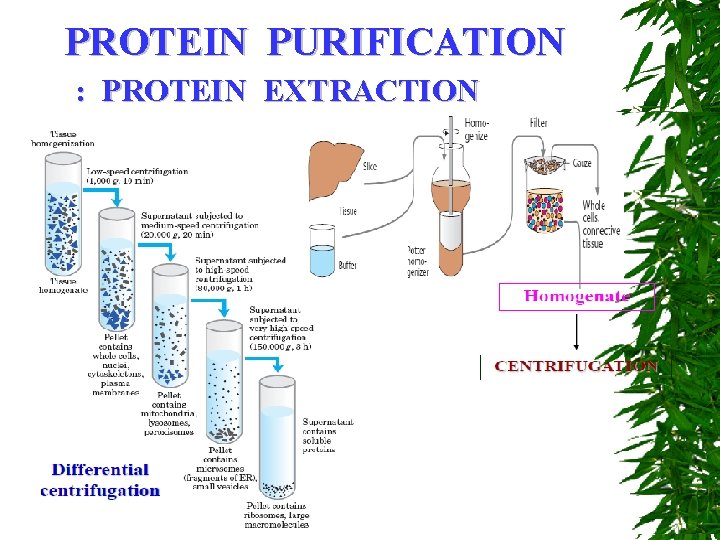
PROTEIN PURIFICATION : PROTEIN EXTRACTION

PROTEIN PURIFICATION : SALT FRACTIONATION

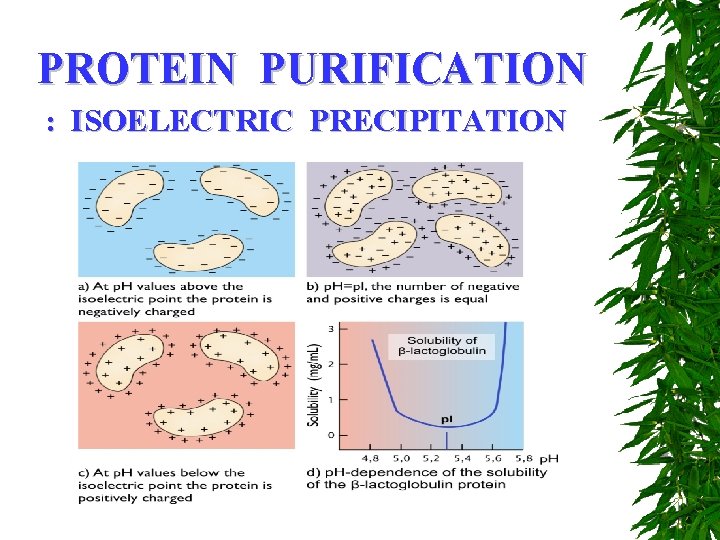
PROTEIN PURIFICATION : ISOELECTRIC PRECIPITATION

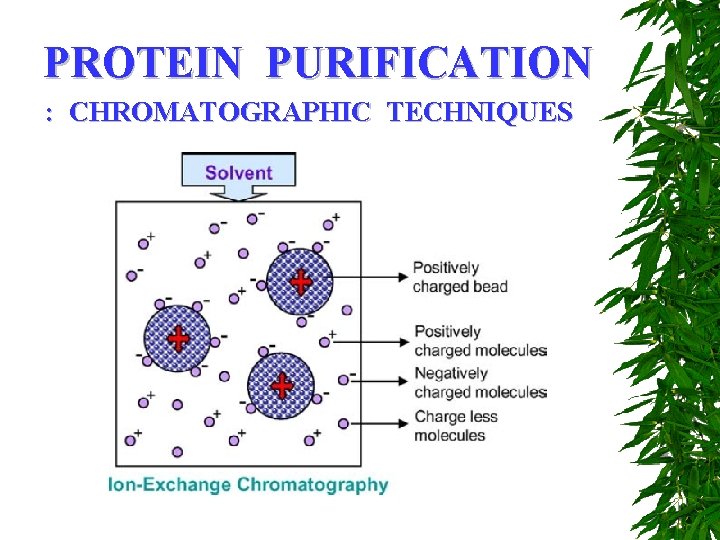
PROTEIN PURIFICATION : CHROMATOGRAPHIC TECHNIQUES

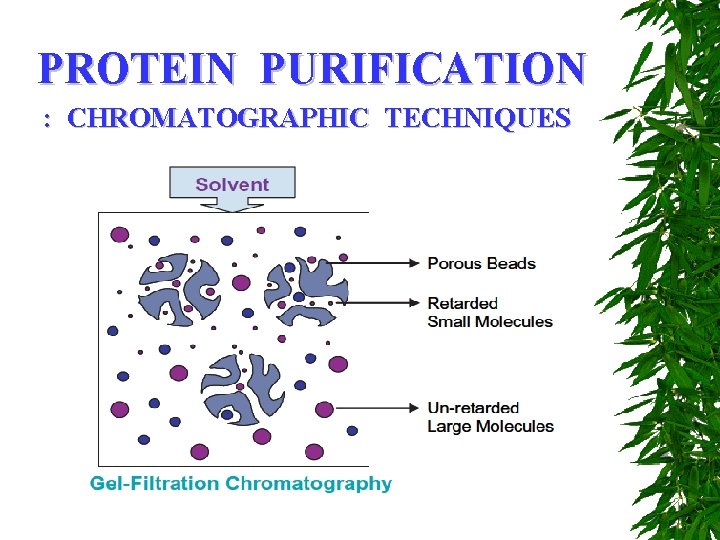
PROTEIN PURIFICATION : CHROMATOGRAPHIC TECHNIQUES

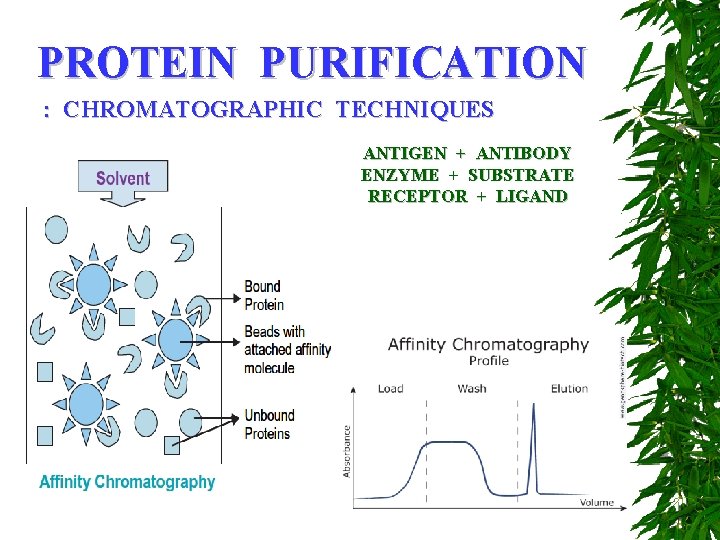
PROTEIN PURIFICATION : CHROMATOGRAPHIC TECHNIQUES ANTIGEN + ANTIBODY ENZYME + SUBSTRATE RECEPTOR + LIGAND

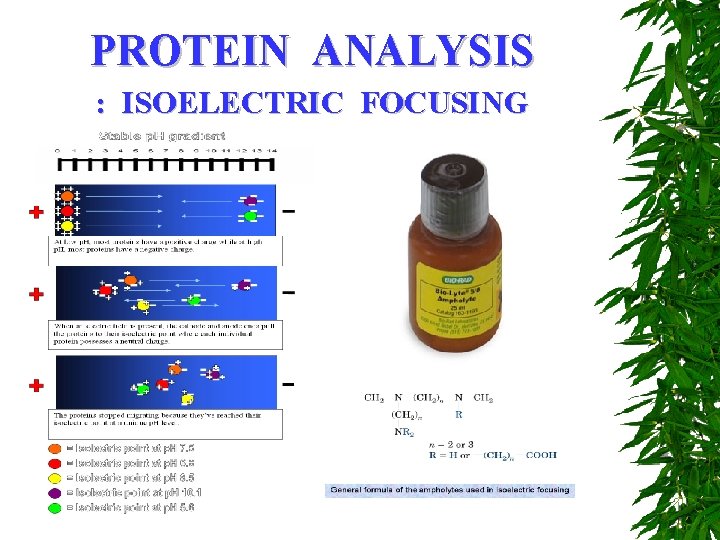
PROTEIN ANALYSIS : ISOELECTRIC FOCUSING

DETERMINATION OF AMINO ACID SEQUENCE SANGER’S FDNB METHOD DANSYL CHLORIDE EDMAN DEGRADATION

SPECIFICITY OF SOME POLYPEPTIDE CLEAVING REAGENTS

DETERMINATION OF AMINO ACID SEQUENCE

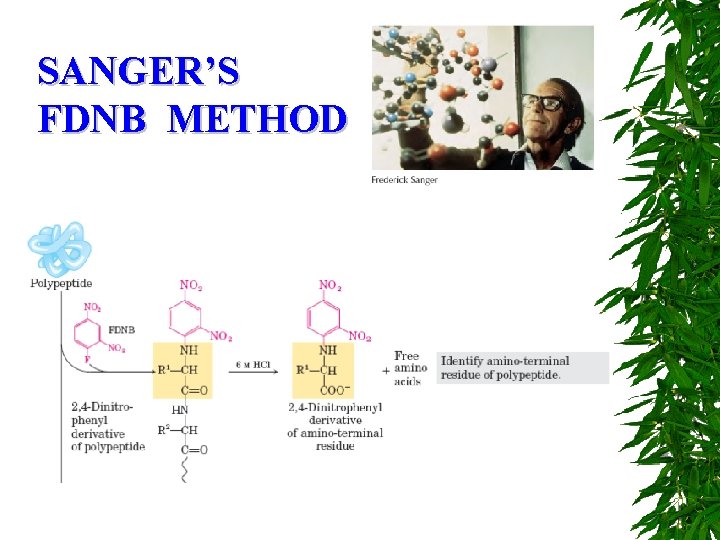
SANGER’S FDNB METHOD

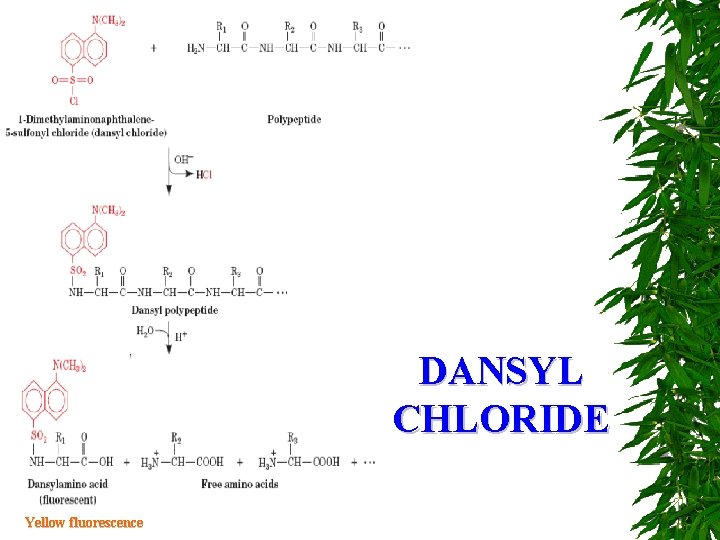
DANSYL CHLORIDE Yellow fluorescence

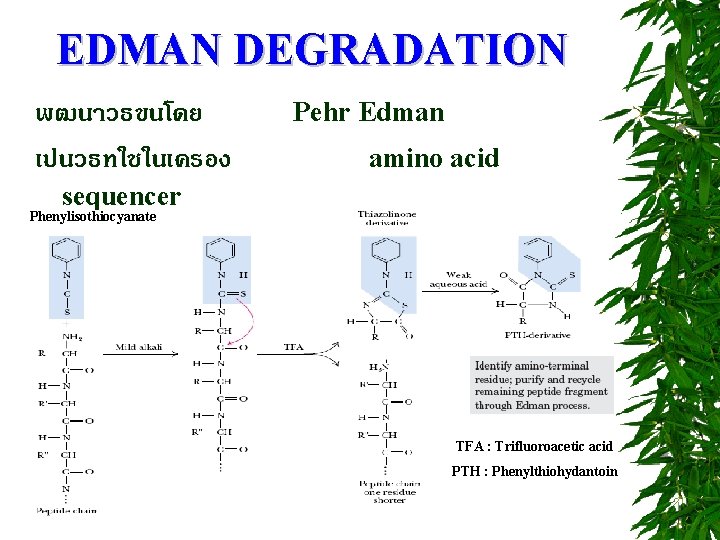
EDMAN DEGRADATION พฒนาวธขนโดย เปนวธทใชในเครอง Pehr Edman amino acid sequencer Phenylisothiocyanate TFA : Trifluoroacetic acid PTH : Phenylthiohydantoin

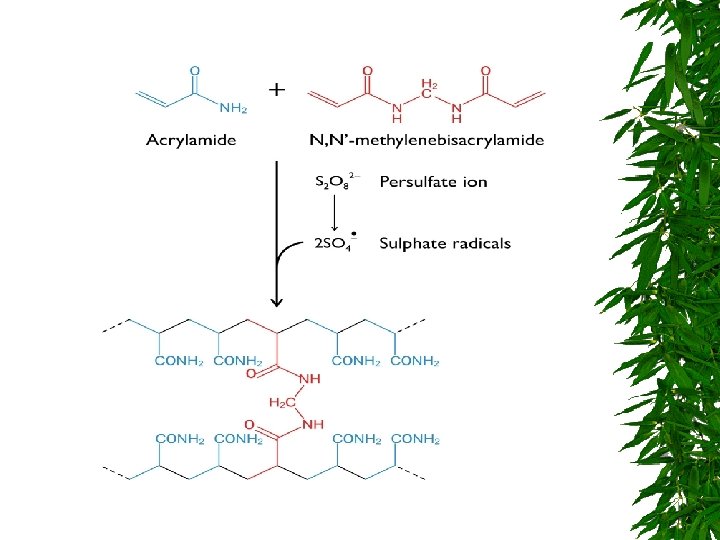
SDS GEL ELECTROPHORESIS

