Class 2 Basic Sequence Alignment Sequence Comparison Much

Class 2: Basic Sequence Alignment .

Sequence Comparison Much of bioinformatics involves sequences u DNA sequences u RNA sequences u Protein sequences We can think of these sequences as strings of letters u DNA & RNA: alphabet of 4 letters u Protein: alphabet of 20 letters

Sequence Comparison (cont) u Finding similarity between sequences is important for many biological questions For example: u Find genes/proteins with common origin · Allows to predict function & structure u Locate common subsequences in genes/proteins · Identify common “motifs” u Locate sequences that might overlap · Help in sequence assembly

Sequence Alignment Input: two sequences over the same alphabet Output: an alignment of the two sequences Example: u GCGCATGGATTGAGCGA u TGCGCCATTGATGACCA A possible alignment: -GCGC-ATGGATTGAGCGA TGCGCCATTGAT-GACC-A
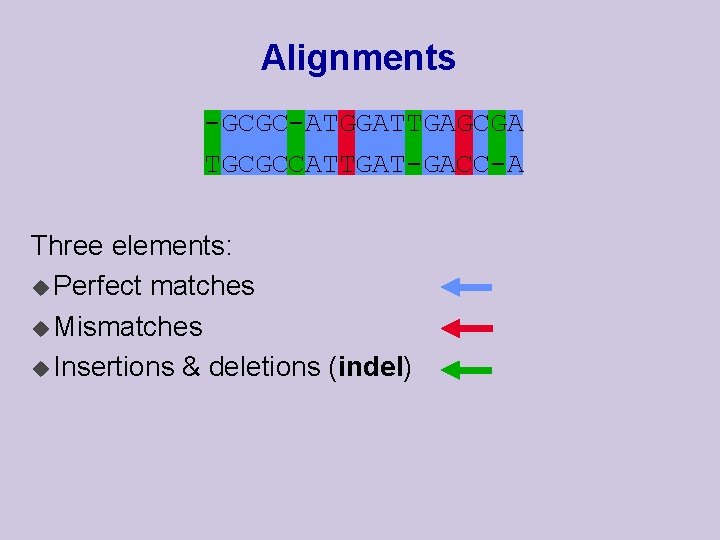
Alignments -GCGC-ATGGATTGAGCGA TGCGCCATTGAT-GACC-A Three elements: u Perfect matches u Mismatches u Insertions & deletions (indel)

Choosing Alignments There are many possible alignments For example, compare: -GCGC-ATGGATTGAGCGA TGCGCCATTGAT-GACC-A to ------GCGCATGGATTGAGCGA TGCGCC----ATTGATGACCA-Which one is better?

Scoring Alignments Rough intuition: u Similar sequences evolved from a common ancestor u Evolution changed the sequences from this ancestral sequence by mutations: · Replacements: one letter replaced by another · Deletion: deletion of a letter · Insertion: insertion of a letter u Scoring of sequence similarity should examine how many operations took place

Simple Scoring Rule Score each position independently: u Match: +1 u Mismatch: -1 u Indel -2 Score of an alignment is sum of positional scores

Example: -GCGC-ATGGATTGAGCGA TGCGCCATTGAT-GACC-A Score: (+1 x 13) + (-1 x 2) + (-2 x 4) = 3 ------GCGCATGGATTGAGCGA TGCGCC----ATTGATGACCA-Score: (+1 x 5) + (-1 x 6) + (-2 x 11) = -23

More General Scores u The choice of +1, -1, and -2 scores was quite arbitrary u Depending on the context, some changes are more plausible than others · Exchange of an amino-acid by one with similar properties (size, charge, etc. ) vs. · Exchange of an amino-acid by one with opposite properties

Additive Scoring Rules u We define a scoring function by specifying a function · (x, y) is the score of replacing x by y · (x, -) is the score of deleting x · (-, x) is the score of inserting x u The score of an alignment is the sum of position scores

Edit Distance u The edit distance between two sequences is the “cost” of the “cheapest” set of edit operations needed to transform one sequence into the other u Computing edit distance between two sequences almost equivalent to finding the alignment that minimizes the distance

Computing Edit Distance u How can we compute the edit distance? ? · If |s| = n and |t| = m, there are more than alignments u The additive form of the score allows to perform dynamic programming to compute edit distance efficiently
![Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. . Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. .](http://slidetodoc.com/presentation_image/0ed7760dd694142241e02bbb6ce07399/image-14.jpg)
Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. . m+1] The best alignment must be in one of three cases: 1. Last position is (s[n+1], t[m +1] ) 2. Last position is (s[n +1], -) 3. Last position is (-, t[m +1] )
![Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. . Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. .](http://slidetodoc.com/presentation_image/0ed7760dd694142241e02bbb6ce07399/image-15.jpg)
Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. . m+1] The best alignment must be in one of three cases: 1. Last position is (s[n+1], t[m +1] ) 2. Last position is (s[n +1], -) 3. Last position is (-, t[m +1] )
![Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. . Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. .](http://slidetodoc.com/presentation_image/0ed7760dd694142241e02bbb6ce07399/image-16.jpg)
Recursive Argument u Suppose we have two sequences: s[1. . n+1] and t[1. . m+1] The best alignment must be in one of three cases: 1. Last position is (s[n+1], t[m +1] ) 2. Last position is (s[n +1], -) 3. Last position is (-, t[m +1] )

Recursive Argument Define the notation: u Using the recursive argument, we get the following recurrence for V:

Recursive Argument u Of course, we also need to handle the base cases in the recursion:
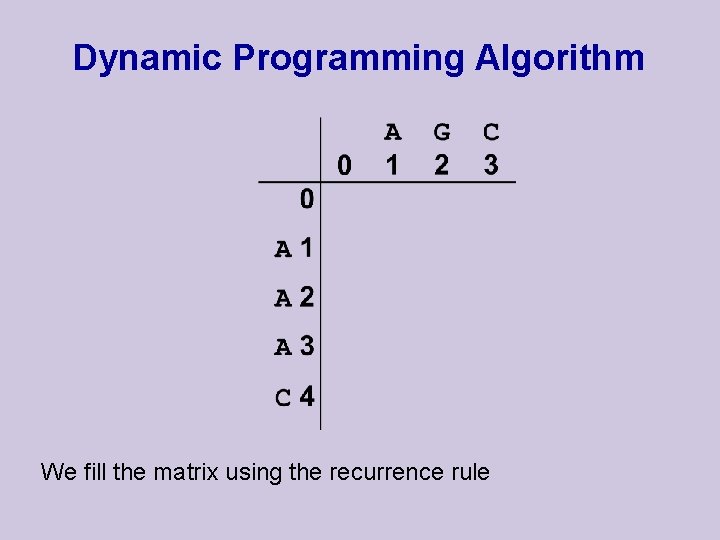
Dynamic Programming Algorithm We fill the matrix using the recurrence rule

Dynamic Programming Algorithm Conclusion: d(AAAC, AGC) = -1

Reconstructing the Best Alignment u To reconstruct the best alignment, we record which case in the recursive rule maximized the score

Reconstructing the Best Alignment u We now trace back the path the corresponds to the best alignment AAAC AG-C

Reconstructing the Best Alignment u Sometimes, score AAAC A-GC more than one alignment has the best

Complexity Space: O(mn) Time: O(mn) u Filling the matrix O(mn) u Backtrace O(m+n)

Space Complexity real-life applications, n and m can be very large u The space requirements of O(mn) can be too demanding · If m = n = 1000 we need 1 MB space · If m = n = 10000, we need 100 MB space u We can afford to perform extra computation to save space · Looping over million operations takes less than seconds on modern workstations u In u Can we trade off space with time?

Why Do We Need So Much Space? To compute d(s, t), we only need O(n) space u Need to compute V[n, m] u Can fill in V, column by column, only storing the last two columns in memory Note however u This “trick” fails when we need to reconstruct the sequence u Trace back information “eats up” all the memory

Why Do We Need So Much Space? To find d(s, t), need O(n) space u Need to compute V[n, m] u Can fill in V, column by 0 column, storing only two columns in memory A 1 Note however A 2 u This “trick” fails when we A 3 need to reconstruct the sequence C 4 u Trace back information “eats up” all the memory 0 0 A G C 1 2 3 -2 -4 -6 -2 1 -4 -1 -1 -3 0 -2 -6 -3 -2 -1 -8 -5 -4 -1
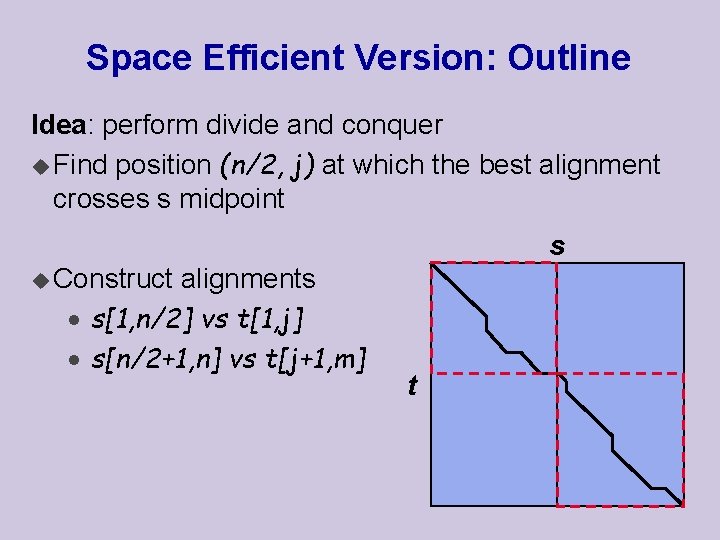
Space Efficient Version: Outline Idea: perform divide and conquer u Find position (n/2, j) at which the best alignment crosses s midpoint s u Construct alignments · s[1, n/2] vs t[1, j] · s[n/2+1, n] vs t[j+1, m] t
![Finding the Midpoint Suppose s[1, n] and t[1, m] are given u We can Finding the Midpoint Suppose s[1, n] and t[1, m] are given u We can](http://slidetodoc.com/presentation_image/0ed7760dd694142241e02bbb6ce07399/image-29.jpg)
Finding the Midpoint Suppose s[1, n] and t[1, m] are given u We can write the score of the best alignment that goes through j as: d(s[1, n/2], t[1, j]) + d(s[n/2+1, n], t[j+1, m]) u Thus, we need to compute these two quantities for all values of j
![Finding the Midpoint (cont) Define u F[i, j] = d(s[1, i], t[1, j]) u Finding the Midpoint (cont) Define u F[i, j] = d(s[1, i], t[1, j]) u](http://slidetodoc.com/presentation_image/0ed7760dd694142241e02bbb6ce07399/image-30.jpg)
Finding the Midpoint (cont) Define u F[i, j] = d(s[1, i], t[1, j]) u B[i, j] = d(s[I+1, n], t[j+1, m]) u F[i, j] + B[i, j] = score of best alignment through (i, j) compute F[i, j] as we did before u We compute B[i, j] in exactly the same manner, going “backward” from B[n, m] u We

Time Complexity Analysis mid-point: cmn (c - a constant) u Recursive sub-problems of sizes (n/2, j) and (n/2, m-1 -1) u Finding T(m, n) = cmn + T(j, n/2) + T(m-j-1, n/2) Lemma: T(m, n) 2 cmn Time complexity is linear in size of the problem u At worse, twice the cost of regular solution.

Local Alignment Consider now a different question: u Can we find similar substring of s and t u Formally, given s[1. . n] and t[1. . m] find i, j, k, and l such that d(s[i. . j], t[k. . l]) is maximal

Local Alignment u As before, we use dynamic programming u We now want to set. V[i, j] to record the best alignment of a suffix of s[1. . i] and a suffix of t[1. . j] u How should we change the recurrence rule?

Local Alignment New option: u We can start a new match instead of extend previous alignment Alignment of empty suffixes

Local Alignment Example s = TAATA t = ATCTAA

Local Alignment Example s = TAATA t = TACTAA
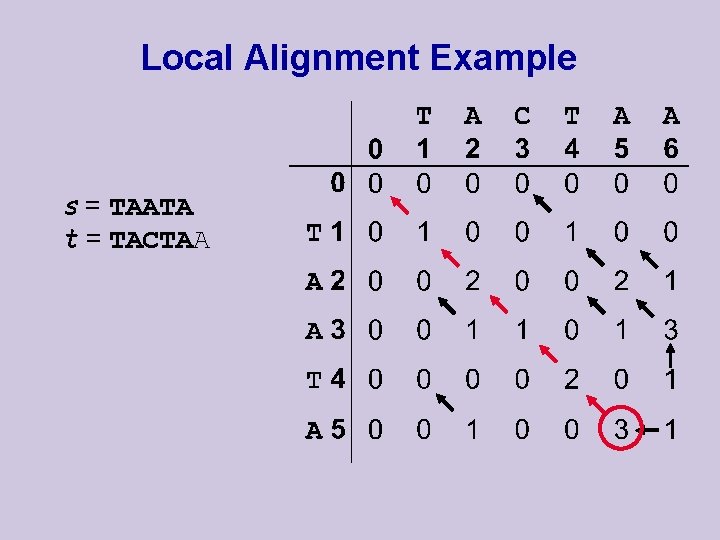
Local Alignment Example s = TAATA t = TACTAA

Local Alignment Example s= TAATA t = TACTAA

Sequence Alignment We seen two variants of sequence alignment: u Global alignment u Local alignment Other variants: u Finding best overlap (exercise) All are based on the same basic idea of dynamic programming
- Slides: 39