CIMI Modelling Taskforce Report Dr Linda Bird 26

CIMI Modelling Taskforce Report Dr Linda Bird 26 th June 2013

Agenda • • • Background CIMI Modelling Approach CIMI Modelling Foundations CIMI Modelling Methodology Future Work Tomorrow: • Terminology Binding

BACKGROUND

Taskforce Members Core Members: • Linda Bird (co-chair) • Harold Solbrig (co-chair) • Tom Beale • Dave Carlson • Stephen Chu • Stan Huff • Mike Lincoln • Rahil Qamar Siddiqui • Gerard Freriks • Josh Mandel • Mark Shafarman • Michael van der Zel Secretary: • Eithne Keelaghan Technical Resources: • Peter Hendler • Galen Mulrooney • Daniel Karlsson • Cecil Lynch • Joey Coyle • Grahame Grieve • Dipak Kalra • David Moner Clinical Modelling Resource: • William Goossen • Jay Lyle • Ian Mc. Nicoll • Anneke Goossen • Heather Leslie • Sarah Ryan • Marcelo Rodrigues dos Santos

Terms of Reference This taskforce has been established to: • Develop CIMI's modelling methodology; • Create an initial set of CIMI clinical models; • Further test and develop CIMI technical models, including: – CIMI reference model – Archetype Object Model 1. 5, and – CIMI terminology.

Modelling Taskforce History May 10 th-12 th May to Sept 14 th-16 th Oct to Dec 2 nd-4 th Jan 18 th-20 th Feb – Mar 2012 Pleasanton meeting: Modelling Taskforce established Taskforce infrastructure and planning Modelling methodology Observation modelling pattern Heart rate model Rockville meeting Laboratory Results models and patterns Terminology binding methodology & reference sets Groningen: Taskforce meeting 2013 Scottsdale meeting Terminology and Modelling Tooling CIMI Modelling Style Guides (TOC) Comparative Analysis Spreadsheet template Laboratory Results Models Example Terminology bindings Lab Specialisation Models – CBC; Gas & CM Panels Demographics models CIMI Reference Model Review

Modelling Taskforce: Post-Leeds Apr 4 -6 Apr to June 2013 Leeds meetings Technical/Implementation CIMI Reference Model DSTU (link associations, instance ids) Implementation artefacts: EA, BMM, XMI, XML Schema, XML, RDF opencimi Github repository CIMI URIs Semver. org versioning rules Archetype identification rules (open. EHR) CIMI-Mindmap to ADL conversion Archetype Definition Language (ADL) 1. 5 Archetype Modelling Language (AML) Modelling Unit of Measure: modelling style and approach CIMI Model Example Instance Formats

Semver. org Versioning Rules • Major version – incremented for a breaking change • Minor version – incremented for non-breaking change • Patch version – incremented for change to the informal parts • Version modifier – e. g. “-rc” (release candidate), “-dstu” (draft standard for trial use), “+u” • Commit number – incremented every time artefact is committed • First version rule = 0. 0. 1 E. g. 1. 0. 0 -dstu

Archetype Identification Rules Knowledge Artefact Identification (open. EHR Foundation) • rm_publisher = “CIMI” • rm_closure in {“CORE”, “PARTY”, “DATA_VALUE”} • rm_class_name: name of reference model class e. g. ENTRY • concept_id: human-readable id of concept e. g. heart_rate • version: as per semver. org rules • Examples: E. g. CIMI-CORE-ENTRY. observation. v. 0. 0. 4 E. g. CIMI-CORE-CLUSTER. reference_range. v. 0. 0. 1 E. g. CIMI-PARTY-ACTOR. person. v. 0. 0. 2

Unit of Measure: Modelling Style • Where global standard exists (e. g. Systolic BP in “mm. Hg”) – CIMI Models will define specific Unit of Measure • Where no global standard exists (e. g. Body temperature) – CIMI General Model will restrict the property (e. g. mass concentration, time, volume, pressure) of each quantity units by binding to a reference set of valid alternative units for that property – Jurisdictional specialisations can specialise the CIMI general models to define the specific unit of measure used locally – CIMI ‘Canonical/Preferred Model’ will be defined, which specialises the CIMI General Model to the specific unit of measure to be used when interoperability between jurisdictions is required • CIMI Models will use SNOMED CT to define Units • Jurisdictional implementations can choose to adopt other code sets (e. g. UCUM), and map to SNOMED CT for crossjurisdictional interoperability

CIMI MODELLING APPROACH

CIMI’s Modeling Approach • • Modular for reusability Composable to meet use-cases Pattern-based for consistency Constraint-based to allow specialisation Logical for implementation in multiple formats Maximal for completeness Extensible for local requirements Bound to terminology for isosemanticity & interoperability
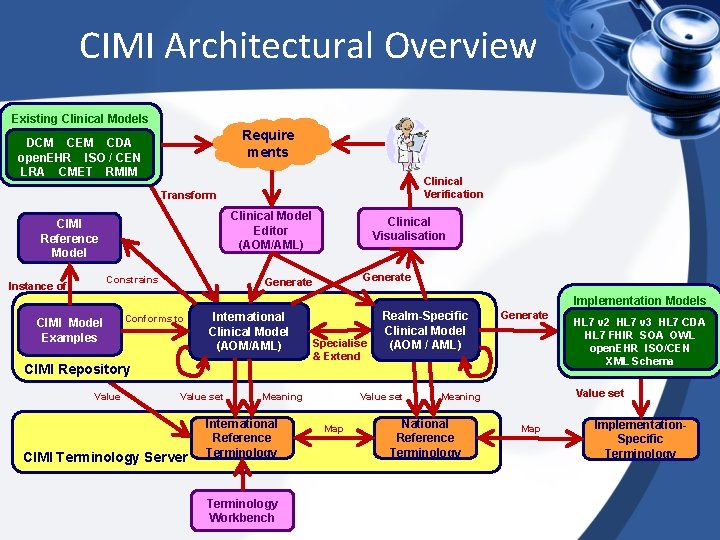
CIMI Architectural Overview Existing Clinical Models Require ments DCM CEM CDA open. EHR ISO / CEN LRA CMET RMIM Clinical Verification Transform Clinical Model Editor (AOM/AML) CIMI Reference Model Constrains Instance of M 0 M Clinical 2 Visualisation Generate Implementation Models CIMI Model Examples Conforms to International Clinical Model (AOM/AML) CIMI Repository Value set CIMI Terminology Server Specialise & Extend Meaning International Reference Terminology Workbench Realm-Specific Clinical Model (AOM / AML) Value set Map Generate Value set Meaning National Reference Terminology HL 7 v 2 HL 7 v 3 HL 7 CDA HL 7 FHIR SOA OWL open. EHR ISO/CEN XML Schema Map Implementation. Specific Terminology

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, UML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

FOUNDATION 1: CIMI REFERENCE MODEL

CIMI Reference Model - Core

CIMI Reference Model – Data Values

CIMI Reference Model – Party Model

FOUNDATION 2: ARCHETYPE OBJECT MODEL / ARCHETYPE MODELLING LANGUAGE
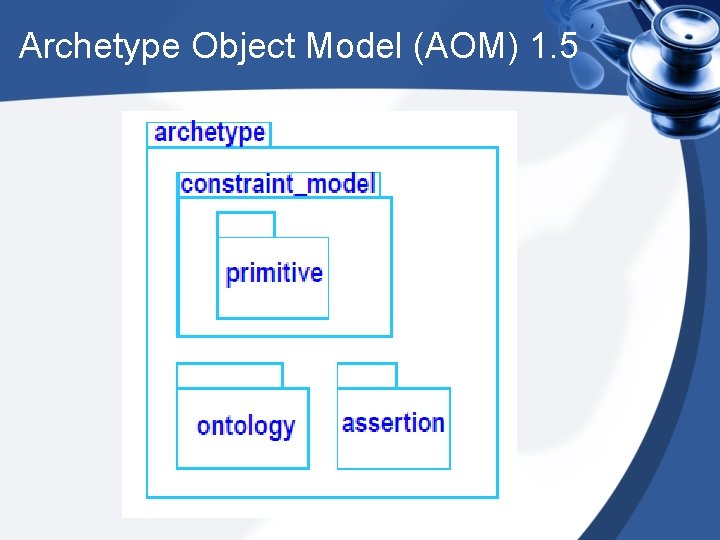
Archetype Object Model (AOM) 1. 5
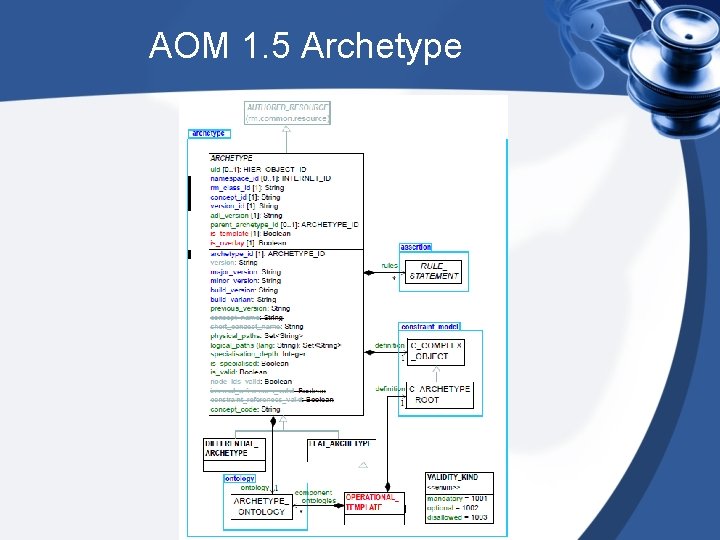
AOM 1. 5 Archetype
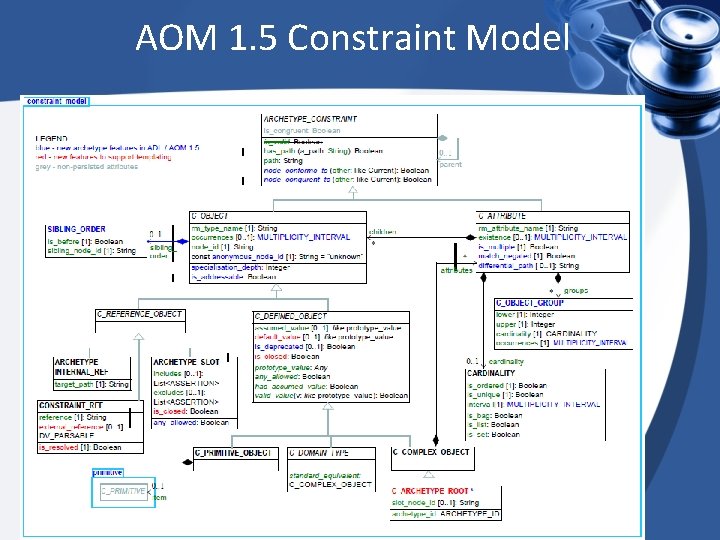
AOM 1. 5 Constraint Model
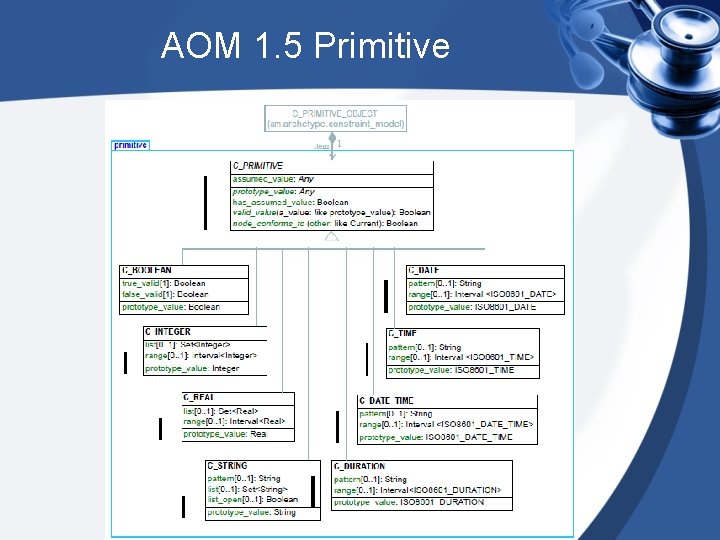
AOM 1. 5 Primitive
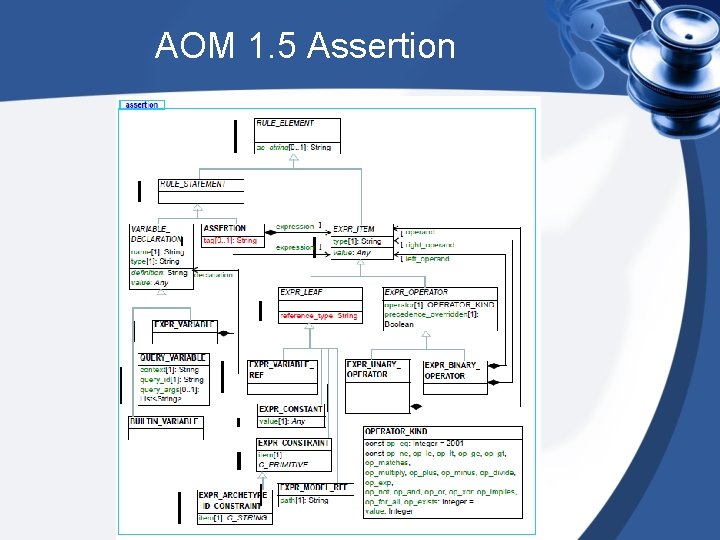
AOM 1. 5 Assertion

AOM 1. 5 Ontology

Archetype Definition Language 1. 5 archetype (adl_version=1. 5) CIMI-RM-CLUSTER. anatomical_location. v 1 concept [at 0000] -- Anatomical location language original_language = <[ISO_639 -1: : en]> description original_author = < ["name"] = <"Ian Mc. Nicoll"> ["date"] = <"29/11/2012“> > details = < ["en"] = < language = <[ISO_639 -1: : en]> purpose = <" Identification of a single anatomical structure, or area, of the human body"> use = <""> misuse = <""> copyright = <"“> > > lifecycle_state = <"Author. Draft"> other_contributors = <> other_details = <> definition CLUSTER[at 0000] matches { -- Anatomical location items matches { ELEMENT[at 0001] occurrences matches {0. . 1} matches { -- Body site name value matches { TEXT matches {*} } } ELEMENT[at 0002] occurrences matches {0. . 1} matches { -- Body site description value matches { TEXT matches {*} } } ELEMENT[at 0003] occurrences matches {0. . 1} matches { -- Body side value matches { TEXT matches {*} } }}}} ontology term_definitions = < ["en"] = < items = < ["at 0000"] = < text = <"Anatomical location"> description = <" Identification of a single anatomical structure, or area, of the human body“>> ["at 0001"] = < text = <"Body site name"> description = <"Name of anatomical location, as specific as is possible. “> > ["at 0002"] = < text = <"Body site description"> description = <"Description of anatomical location. “> > ["at 0003"] = < text = <"Body side">

FOUNDATION 3: CIMI MODELLING PATTERNS
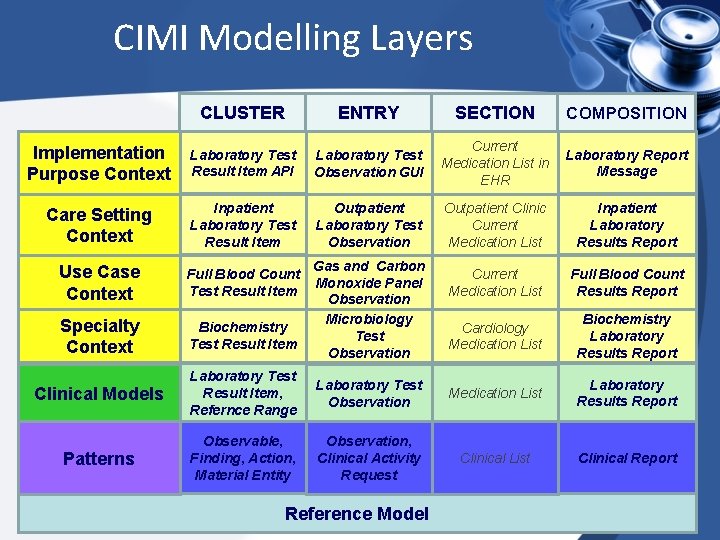
CIMI Modelling Layers CLUSTER ENTRY SECTION COMPOSITION Implementation Purpose Context Laboratory Test Result Item API Laboratory Test Observation GUI Current Medication List in EHR Laboratory Report Message Care Setting Context Inpatient Laboratory Test Result Item Outpatient Laboratory Test Observation Outpatient Clinic Current Medication List Inpatient Laboratory Results Report Use Case Context Full Blood Count Test Result Item Current Medication List Full Blood Count Results Report Specialty Context Biochemistry Test Result Item Cardiology Medication List Biochemistry Laboratory Results Report Clinical Models Laboratory Test Result Item, Refernce Range Laboratory Test Observation Medication List Laboratory Results Report Patterns Observable, Finding, Action, Material Entity Observation, Clinical Activity Request Clinical List Clinical Report Gas and Carbon Monoxide Panel Observation Microbiology Test Observation Reference Model
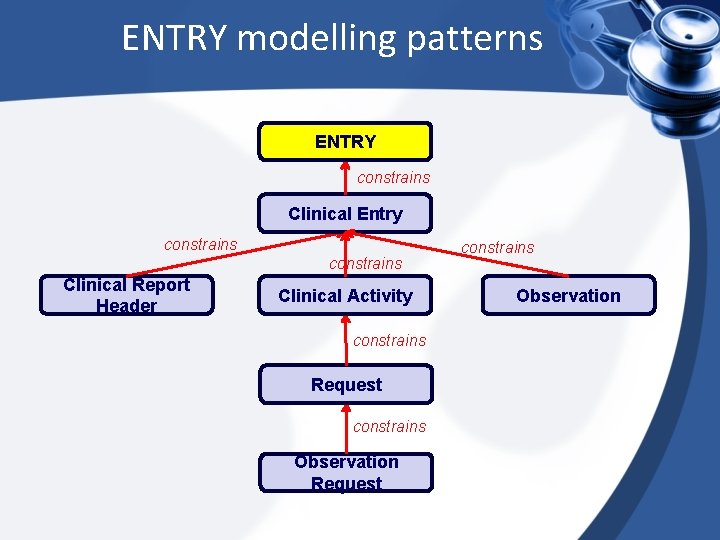
ENTRY modelling patterns ENTRY constrains Clinical Entry constrains Clinical Report Header Clinical Activity constrains Request constrains Observation
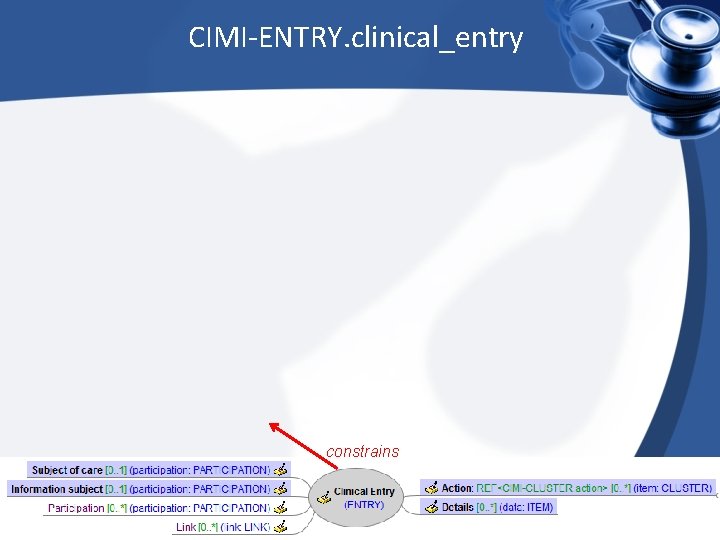
CIMI-ENTRY. clinical_entry constrains
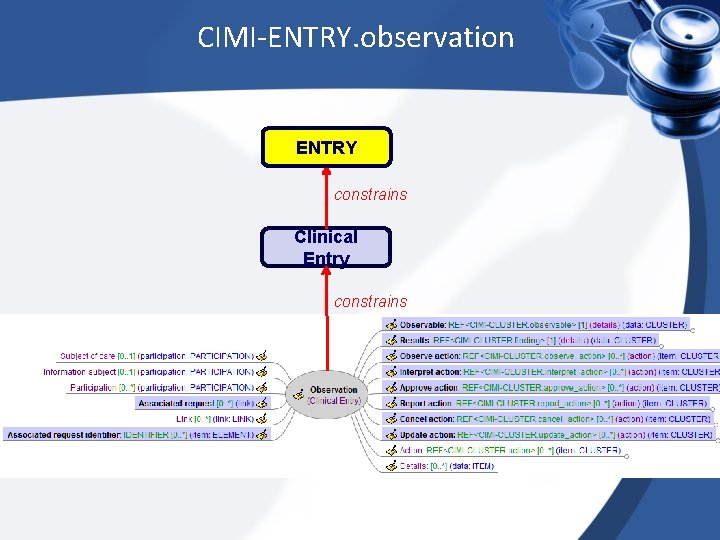
CIMI-ENTRY. observation ENTRY constrains Clinical Entry constrains
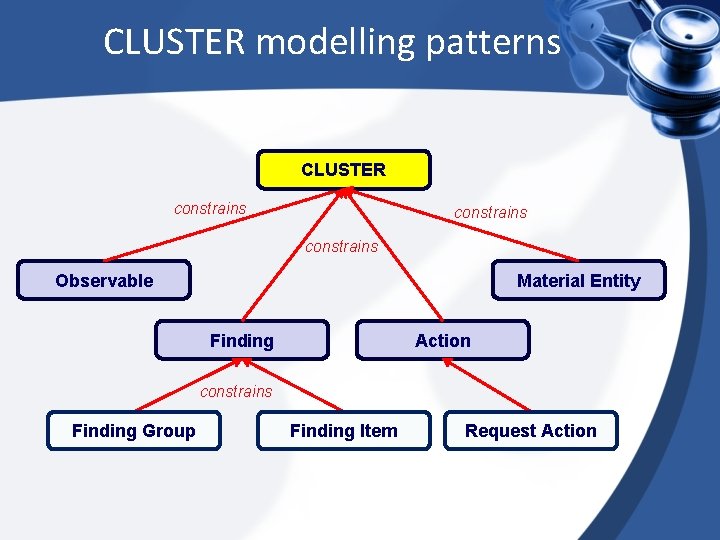
CLUSTER modelling patterns CLUSTER constrains Observable Material Entity Finding Action constrains Finding Group Finding Item Request Action

FOUNDATION 4: STYLE GUIDES

CIMI Modelling Guides • User Guide – Modelling Framework – Modellng Methodology – Modelling Examples and Use Cases • Editorial Guide – Modelling Principles – Modelling Patterns • Terminology Binding Guide – – Types of Terminology Bindings Terminology Binding Rules Terminology Binding Patterns Terminology Binding Examples and Use Cases • Technical Guide – – – CIMI Reference Model Archetype Object Model Archetype Definition Language Archetype Modelling Language Model Instance Representation Model Transformation and Implementation

MODELLING METHODOLOGY

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, AML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

Comparative Analysis Template • Purpose: To compare a set of existing clinical models, and identify the maximal set of data items to be included in the international CIMI models. In so doing, it also documents the high-level mapping from the CIMI data items to the source data models.
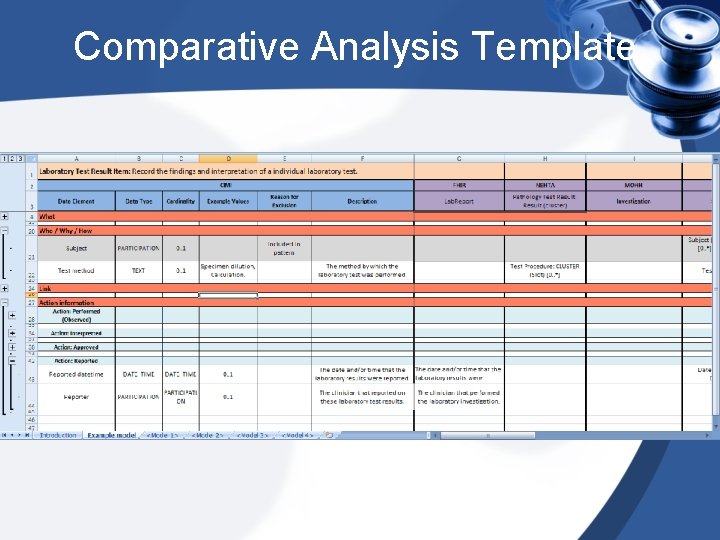
Comparative Analysis Template

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, AML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, AML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models
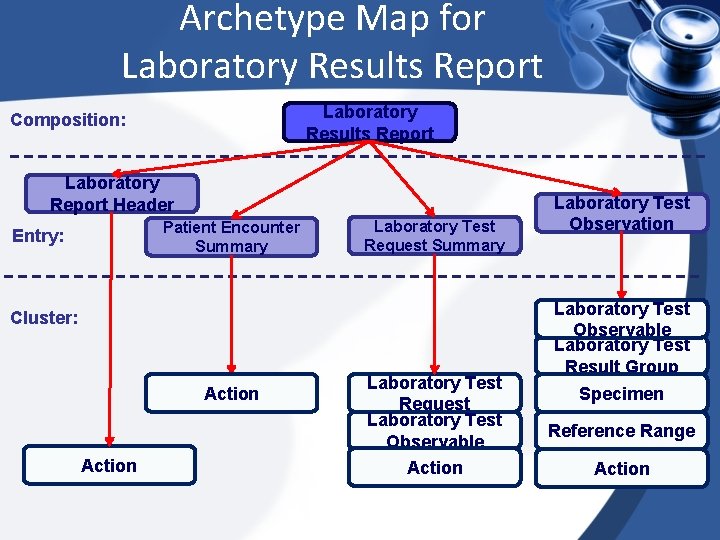
Archetype Map for Laboratory Results Report Composition: Laboratory Report Header Patient Encounter Summary Entry: Laboratory Test Request Summary Cluster: Action Laboratory Test Request Laboratory Test Observable Action Laboratory Test Observable Laboratory Test Result Group Specimen Reference Range Action
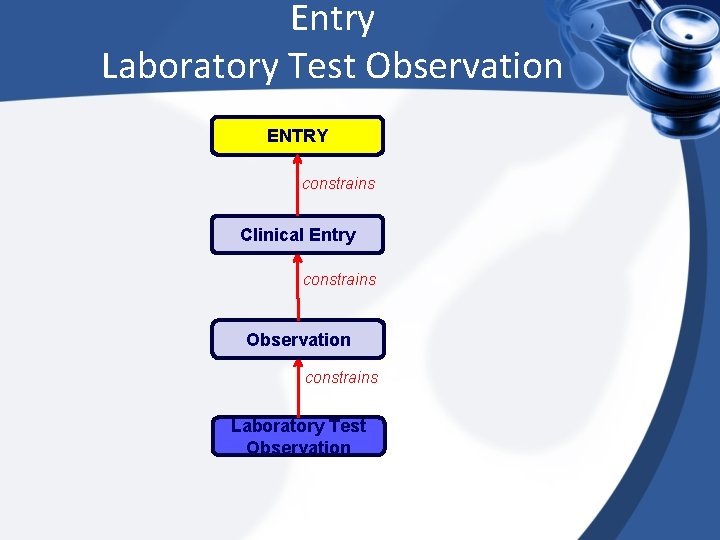
Entry Laboratory Test Observation ENTRY constrains Clinical Entry constrains Observation constrains Laboratory Test Observation
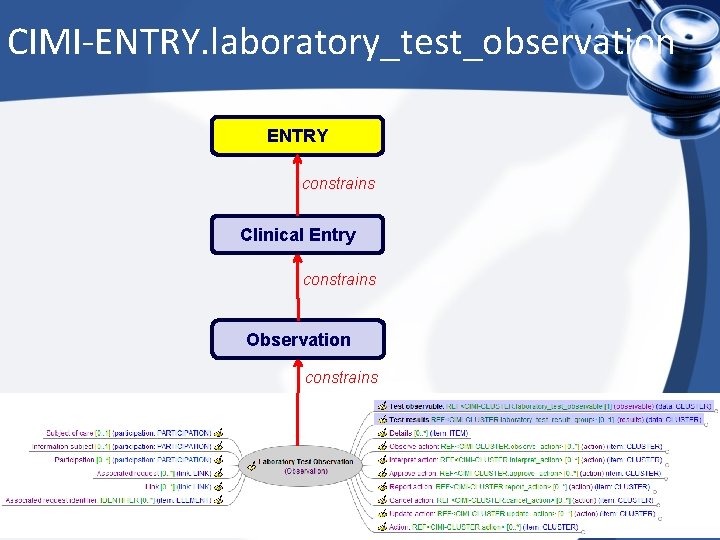
CIMI-ENTRY. laboratory_test_observation ENTRY constrains Clinical Entry constrains Observation constrains
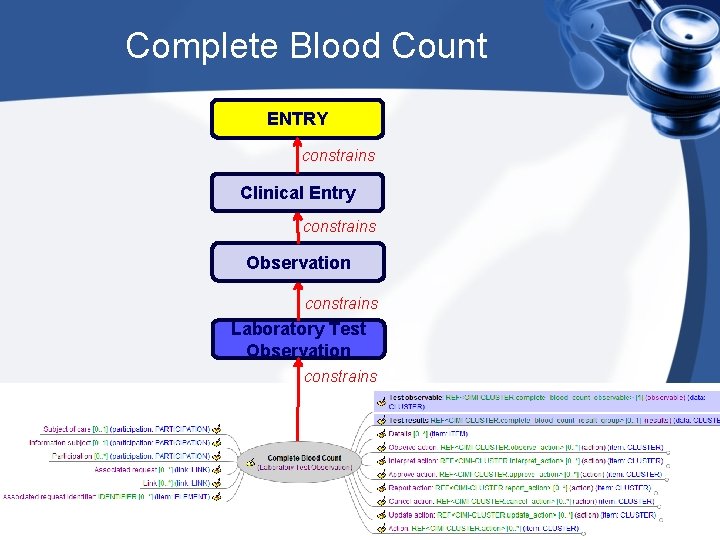
Complete Blood Count ENTRY constrains Clinical Entry constrains Observation constrains Laboratory Test Observation constrains

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, AML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, AML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

Types of Example Instances • Different example representation formats, e. g. : – Clinical user interface form – Mindmap example instance – XML instance • Reference Model vs Model Specific Instances • Full vs Lite/Green Instances Goal: Automated bi-directional transforms • XML Instance Mindmap instance Clinical form • Reference Model Instance Model-Specific Instance • Full instance Green instance + Archetype

Mindmap Example Instance 1
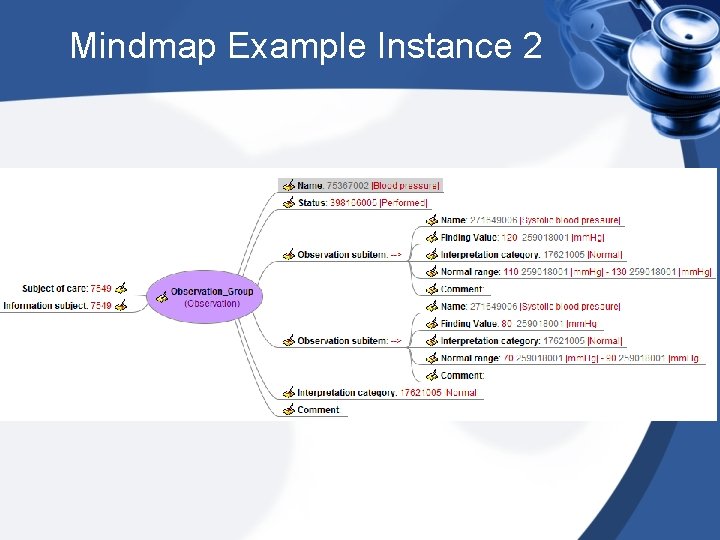
Mindmap Example Instance 2
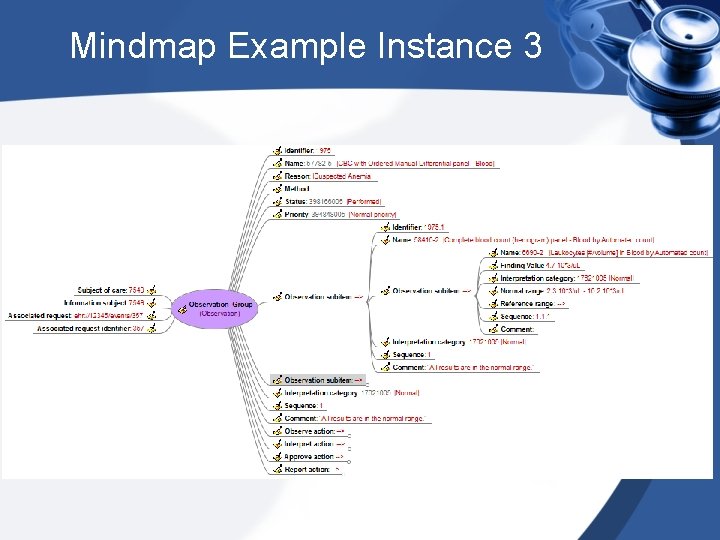
Mindmap Example Instance 3
Reference Model Instance ENTRY constrains Clinical Entry constrains Observation constrains Blood Pressure

CIMI-ENTRY

Reference-Model Instance (XML) <Entry xmlns: xsi=http: //www. w 3. org/2001/XMLSchema-instance xmlns=http: //opencimi. org archetype_node_id = “at 0000. 1. 1” name = “Blood_pressure_observation” uid = “ 123456”>. . . <Data xsi: type = “CLUSTER” archetype_node_id = “at 0002. 1. 1” name = “Observable” > <Item xsi: type = “ELEMENT” archetype_node_id = “at 0003. 1. 1” name = “Name” > <Value xsi: type = “CODED_TEXT” value = “Blood pressure ” code = “ 75367002” /> </Item> </Data> <Data xsi: type = “CLUSTER” archetype_node_id = “at 0004. 1. 1” name = “Results” > <Item xsi: type = “CLUSTER” archetype_node_id = “at 0005. 1. 1” name = “Systolic_blood_pressure” > < Item xsi: type = “ELEMENT” archetype_node_id = “at 0007. 1. 1” name “Systolic_value”> <Value xsi: type = “QUANTITY” value = “ 120” > <Units xsi: type = “CODED_TEXT” value = “mm. Hg” code = “ 259018001” /> </Value> </Item> <Item xsi: type = “CLUSTER” archetype_node_id = “at 0008. 1. 1” name = “Diastolic_blood_pressure” > < Item xsi: type = “ELEMENT” archetype_node_id = “at 00010. 1. 1” name “Diastolic_value”> <Value xsi: type = “QUANTITY” value = “ 80” > <Units xsi: type = “CODED_TEXT” value = “mm. Hg” code = “ 259018001” /> </Value> </Item> </Data> </Entry> = =
Model Specific Instance ENTRY constrains Clinical Entry constrains Observation constrains Blood Pressure

Full Model Specific Instance <Blood_pressure_observation xmlns: xsi=http: //www. w 3. org/2001/XMLSchema-instance xmlns=http: //opencimi. org rm_type = “Entry” archetype_node_id = “at 0000. 1. 1” uid = “ 123456”>. . . <Observable rm_type = “CLUSTER” archetype_node_id = “at 0002. 1. 1” > <Name rm_type = “ELEMENT” archetype_node_id = “at 0003. 1. 1” > <Value xsi: type = “CODED_TEXT” value = “Blood pressure ” code = “ 75367002” /> </Name> </Observable> <Results rm_type = “CLUSTER” archetype_node_id = “at 0004. 1. 1” > <Archetype_details archetype_id = “CIMI-ENTRY. finding_group. v 1” /> <Systolic_blood_pressure rm_type = “CLUSTER” archetype_node_id = “at 0005. 1. 1” > <Name rm_type = “ELEMENT” archetype_node_id = “at 0006. 1. 1” > <Value xsi: type = “CODED_TEXT” value = “Systolic blood pressure” code = “ 271649006” /> </Name> <Systolic_value rm_type = “ELEMENT” archetype_node_id = “at 0007. 1. 1” > <Value xsi: type = “QUANTITY” value = “ 120” > <Units xsi: type = “CODED_TEXT” value = “mm. Hg” code = “ 259018001” /> </Value> </Systolic_value> </Systolic_blood_pressure> <Diastolic_blood_pressure rm_type = “CLUSTER” archetype_node_id = “at 0008. 1. 1” > <Name rm_type = “ELEMENT” archetype_node_id = “at 0009. 1. 1” > <Value xsi: type = “CODED_TEXT” value = “Diastolic blood pressure” code = “ 271650006”/> </Name> <Diastolic_value rm_type = “ELEMENT” archetype_node_id = “at 00010. 1. 1” > <Value xsi: type = “QUANTITY” value = “ 80” > <Units xsi: type = “CODED_TEXT” value = “mm. Hg” code = “ 259018001”/> </Value> </Diastolic_value> </Diastolic_blood_pressure> </Results>

Green Model Specific Instance <Blood_pressure_observation> <Subject_of_care> <Party> 456789 </Party> </Subject_of_care> <Results > <Systolic_blood_pressure> <Value value = “ 120”> <Units value = “mm. Hg” code = “ 259018001”/ </Value> </Systolic_blood_pressure> <Diastolic_blood_pressure> <Value value = “ 80”> <Units value = “mm. Hg” code = “ 259018001”/> </Value> </Diastolic_blood_pressure> </Results> </Blood_pressure_observation> Full instance Green instance + Archetype

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, UML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, AML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

FUTURE WORK

Future Work • Foundations – Reference model: Documentation and testing – Archetype object model: Finalise support for terminology binding Archetype modelling language: Finalise UML profile – Modelling patterns: Documentation, Terminology bindings and Review – Style guides: Complete content • Modelling – Laboratory Results models: Example instances, specialisations, terminology bindings – Immunization models: Gather requirements, and define models – Temperature and other priorities: As above • Implementation – – – – • Complete Mindmap-to-ADL generation for modelling patterns CIMI Model Validators (Mindmaps, ADL, AML) AML editor ADL-to-instance generation Model repository and visualisations Model instance visualisations Transformations to implementation formats Governance – Establish modelling development, review and publication processes and procedures

Online References • Taskforce Minutes – http: //informatics. mayo. edu/CIMI/index. php/Main_Page • open. CIMI Github repository – https: //github. com/opencimi • Google groups email list (cimi-modelling-taskforce) – http: //groups. google. com/group/cimi-modellingtaskforce? hl=en-GB

QUESTIONS

TERMINOLOGY BINDING

Modelling Methodology • Foundations 1. 2. 3. 4. CIMI Reference Model Archetype Object Model / Archetype Modelling Language CIMI Modelling Patterns CIMI Style Guide • Modelling Approach 1. 2. 3. 4. 5. 6. 7. 8. Analyse clinical models submitted (with value sets) Identify maximal set of data elements Remove ‘out of scope’ data elements (Style Guide) Select appropriate CIMI Modelling Patterns(Style Guide) Define CIMI model (Mindmap, ADL, UML) Add Terminology bindings o o Meaning (relationship, object, modifier) Value sets (maximal set from submitted models) Add Example Model Data Instances Technical Validation o ADL, UML 9. Clinical Validation / Review 10. Confirm mappings from submitted models

Use Cases for Terminology in Models 1. Management and quality control of model libraries a) b) c) Searching model libraries Identifying semantic overlap between models Inconsistency of model interdependencies 2. Transforming between isosemantic representations of the model: both a) b) Different levels of precoordination Different model formalisms 3. Querying data instances of models (including clinical decision support) which use different representations – for example: a) b) c) Different level of precoordiation versus structure Different modeling design choices Subsumption testing of values 4. Supporting data validation and semantic interoperability

Requirements for using Terminology in Models 1. Standard (reproducible) way of doing terminology bindings 2. The ability to represent the valid set of values for a given coded element. 3. The ability to state the association between the intended interpretation of nodes in the model and concepts in the terminology 4. Terminology bindings that are agnostic as to whether nodes are connected using a hierarchy or using links. 5. Terminology bindings that allow the values to be represented in a way that is agnostic to the degree of precoordination versus structure. 6. Terminology bindings that enable the transformation between isosemantic representations of the same model 7. Terminology bindings that allow consistency to be checked within models, and between models related by specialisation or used to fill slots (using an underlying ontology).

Terminology Binding Approach The meaning of each node has 3 parts: • Relationship: The relationship from the parent node to this node • Object: The ‘class’ of things defined by this node’s values • Modifier: The context of the node’s meaning – including subject-relationship context, temporal context, procedure/finding context, negation, state, certainty

CIMI Terminology Binding Approach STRUCTURE BINDING TERMINOLOGY Meaning Value Set Relationship Object Modifier (Linkage concept) Pharm/biol product (Context values) - Medication Name (Linkage concept) Pharm/biol product (Context values) Medication Ref_Set Element: Ingredient Has ingredient Substance (Context values) Substance Ref_Set Element: Basis of Strength Has basis of strength substance Substance (Context values) Substance Ref_Set Element: Strength Has strength Measurement Finding (Context values) - Dose form Has dose form Drug dose form (Context values) Dose_Form Ref_Set Cluster: Element: Medication
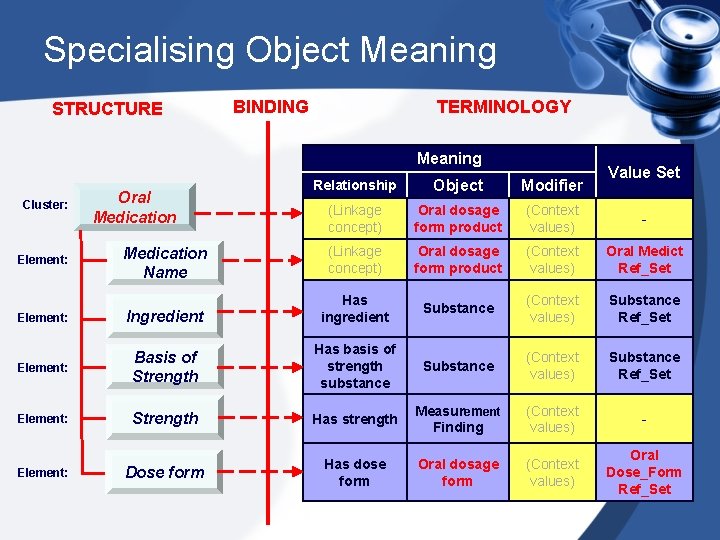
Specialising Object Meaning STRUCTURE BINDING TERMINOLOGY Meaning Value Set Relationship Object Modifier (Linkage concept) Oral dosage form product (Context values) - Medication Name (Linkage concept) Oral dosage form product (Context values) Oral Medict Ref_Set Element: Ingredient Has ingredient Substance (Context values) Substance Ref_Set Element: Basis of Strength Has basis of strength substance Substance (Context values) Substance Ref_Set Element: Strength Has strength Measurement Finding (Context values) - Dose form Has dose form Oral dosage form (Context values) Oral Dose_Form Ref_Set Cluster: Element: Oral Medication
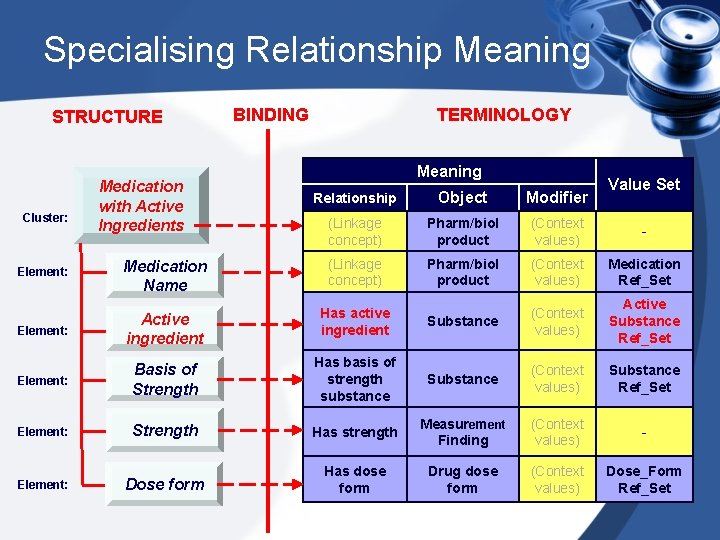
Specialising Relationship Meaning STRUCTURE Medication with Active Ingredients BINDING TERMINOLOGY Meaning Value Set Relationship Object Modifier (Linkage concept) Pharm/biol product (Context values) - Medication Name (Linkage concept) Pharm/biol product (Context values) Medication Ref_Set Element: Active ingredient Has active ingredient Substance (Context values) Active Substance Ref_Set Element: Basis of Strength Has basis of strength substance Substance (Context values) Substance Ref_Set Element: Strength Has strength Measurement Finding (Context values) - Element: Dose form Has dose form Drug dose form (Context values) Dose_Form Ref_Set Cluster: Element:
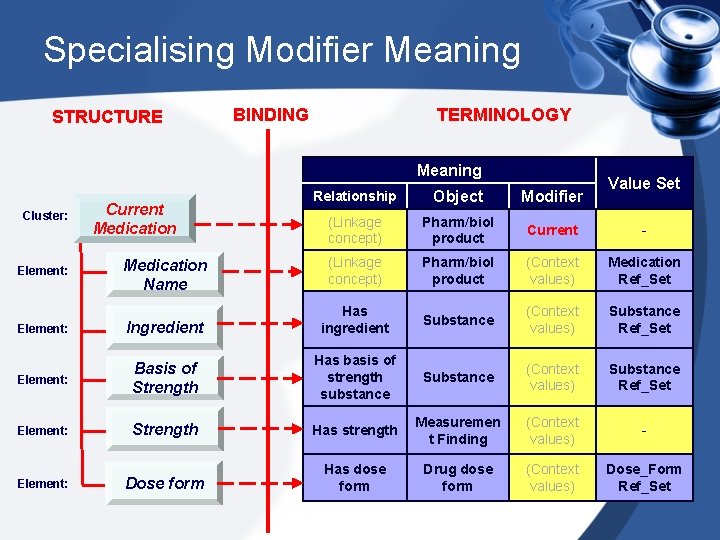
Specialising Modifier Meaning STRUCTURE BINDING TERMINOLOGY Meaning Value Set Relationship Object Modifier (Linkage concept) Pharm/biol product Current - Medication Name (Linkage concept) Pharm/biol product (Context values) Medication Ref_Set Element: Ingredient Has ingredient Substance (Context values) Substance Ref_Set Element: Basis of Strength Has basis of strength substance Substance (Context values) Substance Ref_Set Element: Strength Has strength Measuremen t Finding (Context values) - Dose form Has dose form Drug dose form (Context values) Dose_Form Ref_Set Cluster: Element: Current Medication
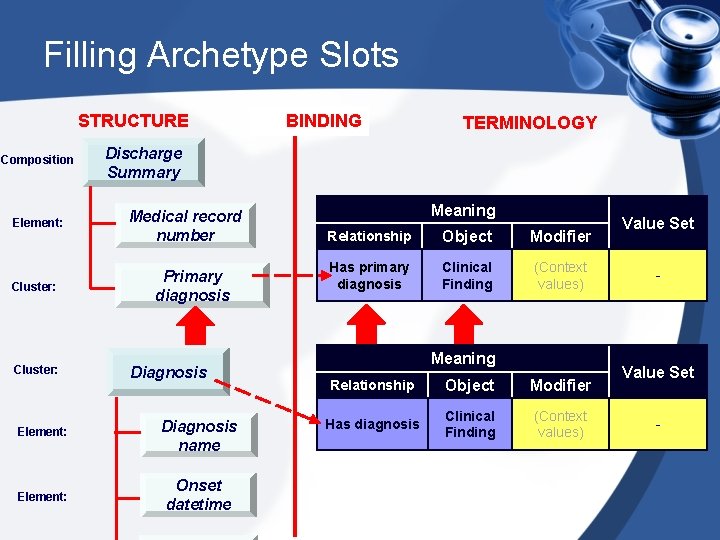
Filling Archetype Slots STRUCTURE Composition Element: Cluster: BINDING TERMINOLOGY Discharge Summary Medical record number Primary diagnosis Diagnosis Element: Diagnosis name Element: Onset datetime Meaning Relationship Object Modifier Has primary diagnosis Clinical Finding (Context values) Meaning Relationship Object Modifier Has diagnosis Clinical Finding (Context values) Value Set -
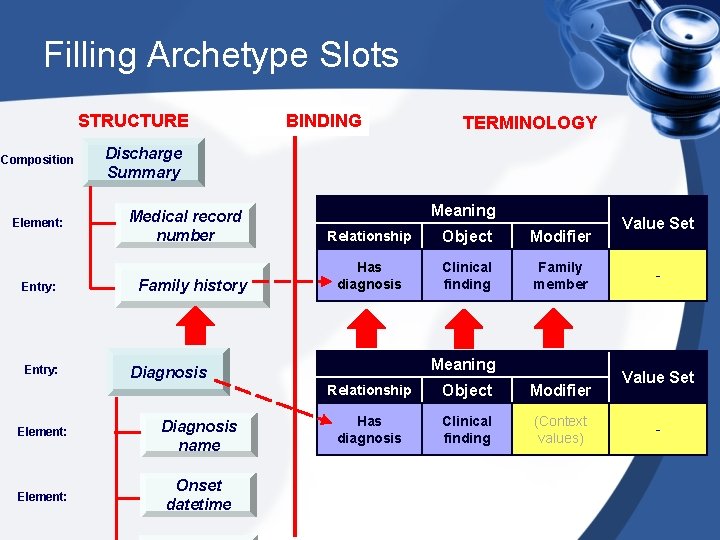
Filling Archetype Slots STRUCTURE Composition Element: Entry: BINDING TERMINOLOGY Discharge Summary Medical record number Family history Meaning Relationship Object Modifier Has diagnosis Clinical finding Family member Meaning Diagnosis Element: Diagnosis name Element: Onset datetime Relationship Object Modifier Has diagnosis Clinical finding (Context values) Value Set -
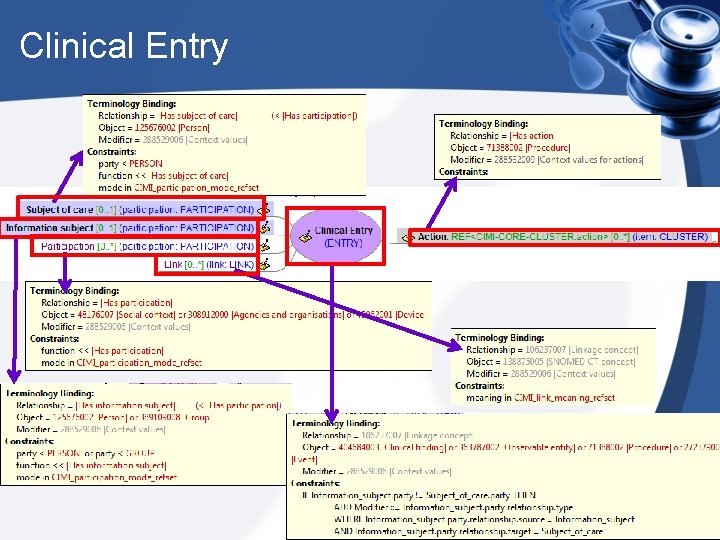
Clinical Entry
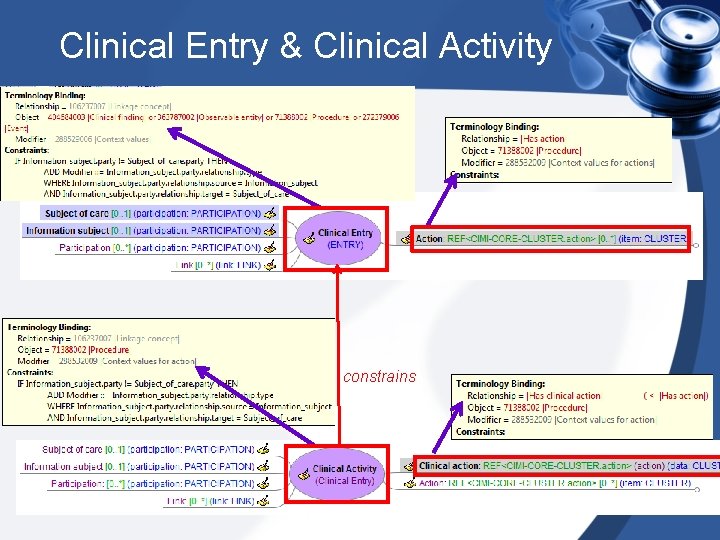
Clinical Entry & Clinical Activity constrains

Clinical Activity & Request constrains

Request & Observation Request constrains

Observation Request & Laboratory Test Request Summary constrains

Proposed Observation Bindings
- Slides: 78