Chromosome Biology How is gene expression regulated How
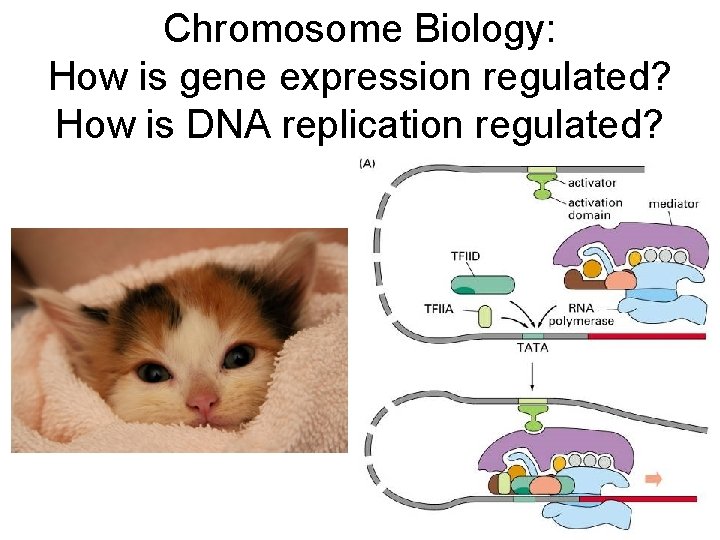
Chromosome Biology: How is gene expression regulated? How is DNA replication regulated?
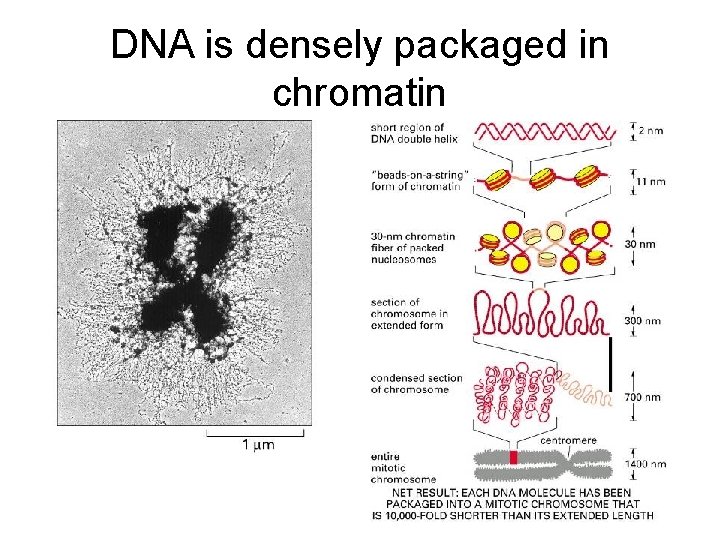
DNA is densely packaged in chromatin
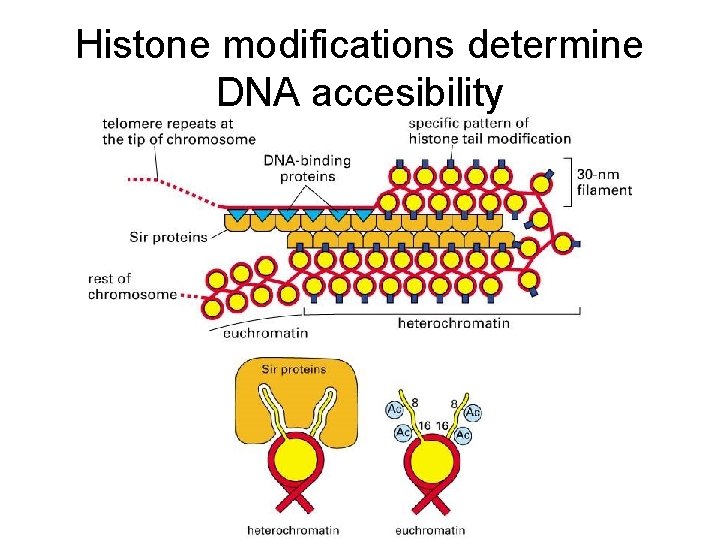
Histone modifications determine DNA accesibility
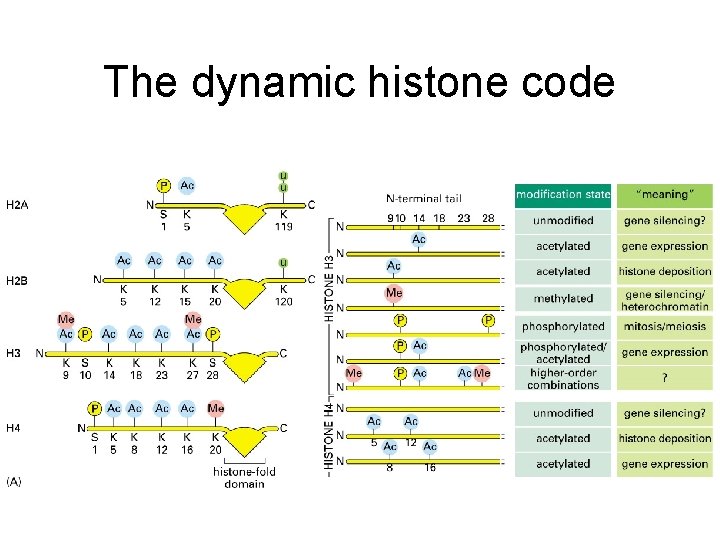
The dynamic histone code
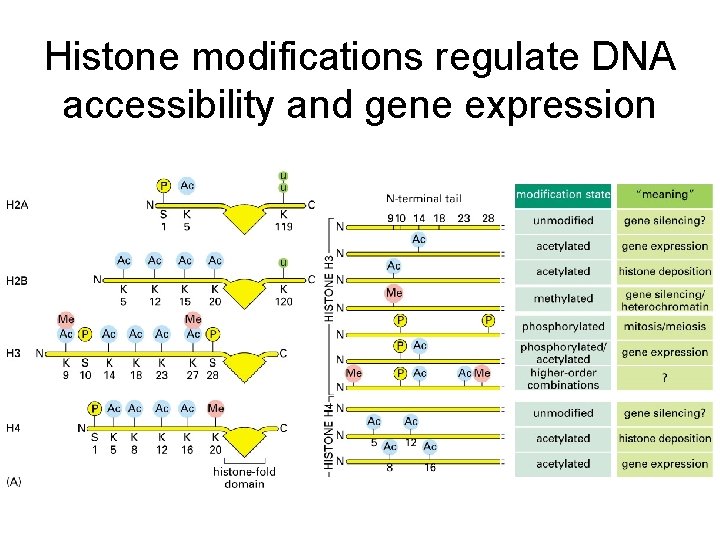
Histone modifications regulate DNA accessibility and gene expression

What controls histone deacetylation?
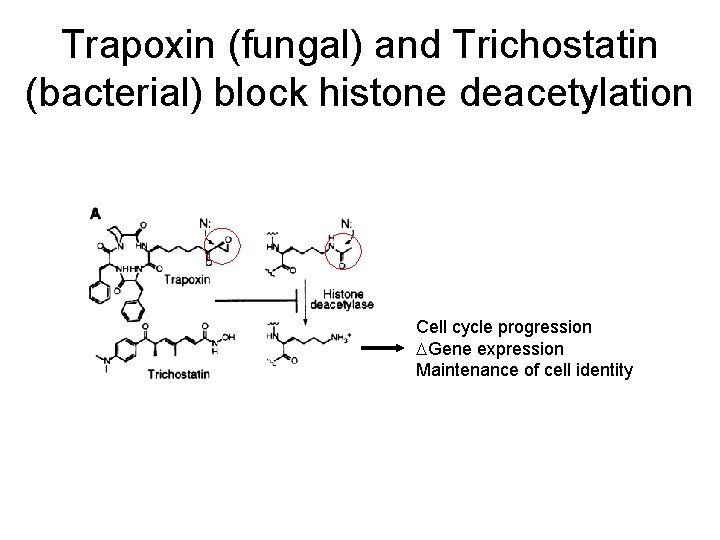
Trapoxin (fungal) and Trichostatin (bacterial) block histone deacetylation Cell cycle progression DGene expression Maintenance of cell identity
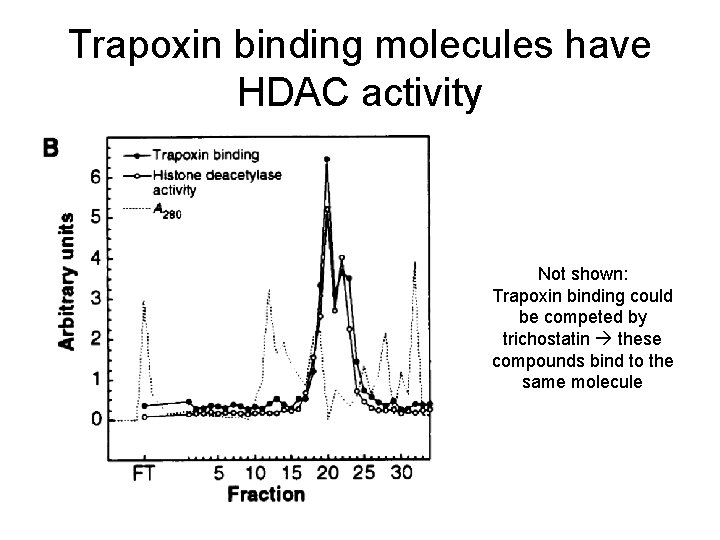
Trapoxin binding molecules have HDAC activity Not shown: Trapoxin binding could be competed by trichostatin these compounds bind to the same molecule
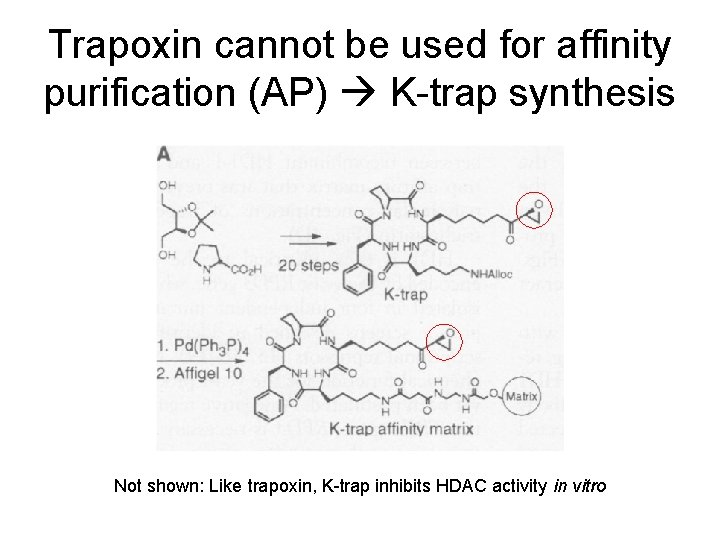
Trapoxin cannot be used for affinity purification (AP) K-trap synthesis Not shown: Like trapoxin, K-trap inhibits HDAC activity in vitro
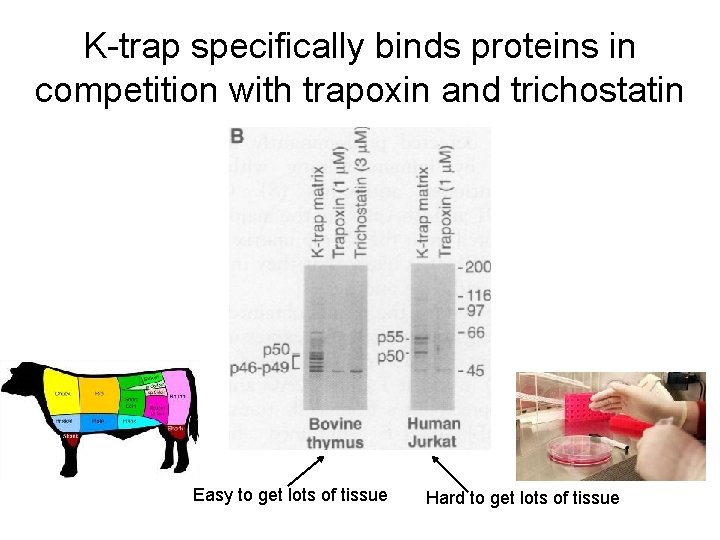
K-trap specifically binds proteins in competition with trapoxin and trichostatin Easy to get lots of tissue Hard to get lots of tissue
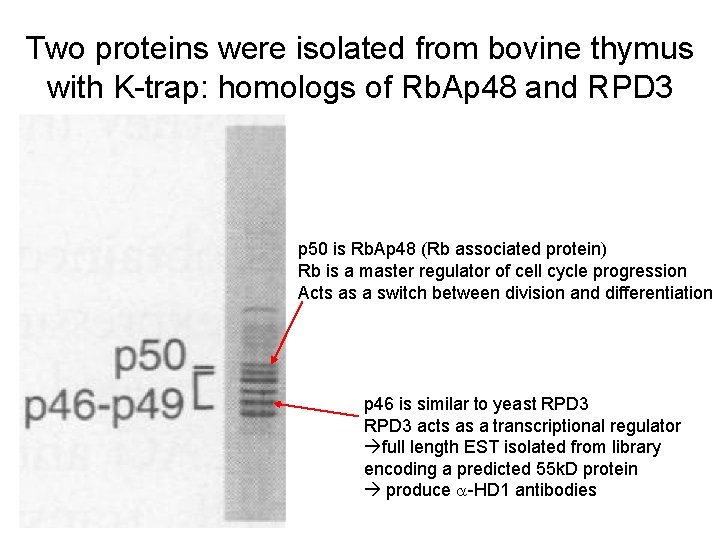
Two proteins were isolated from bovine thymus with K-trap: homologs of Rb. Ap 48 and RPD 3 p 50 is Rb. Ap 48 (Rb associated protein) Rb is a master regulator of cell cycle progression Acts as a switch between division and differentiation p 46 is similar to yeast RPD 3 acts as a transcriptional regulator full length EST isolated from library encoding a predicted 55 k. D protein produce a-HD 1 antibodies
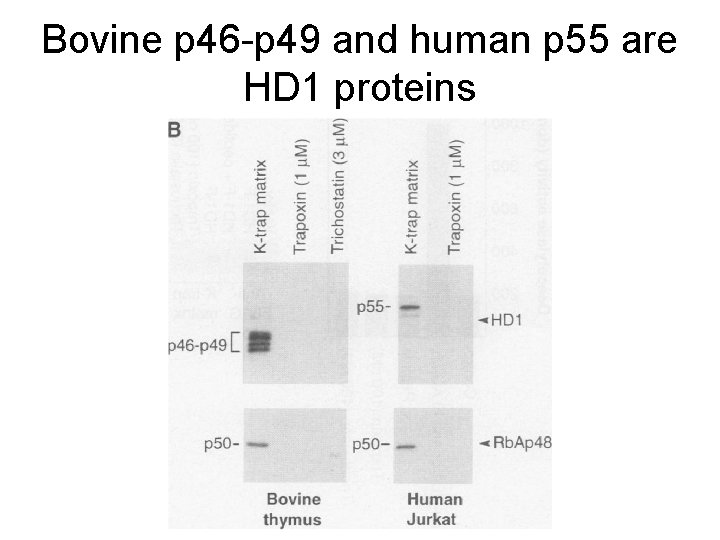
Bovine p 46 -p 49 and human p 55 are HD 1 proteins
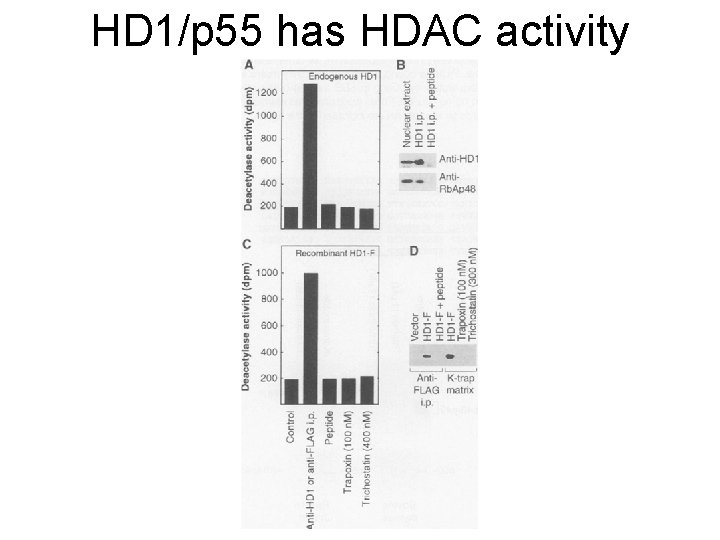
HD 1/p 55 has HDAC activity
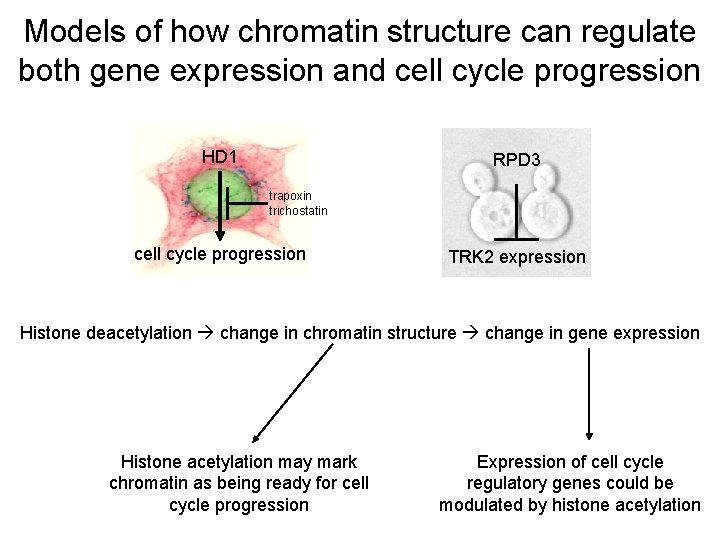
Models of how chromatin structure can regulate both gene expression and cell cycle progression HD 1 RPD 3 trapoxin trichostatin cell cycle progression TRK 2 expression Histone deacetylation change in chromatin structure change in gene expression Histone acetylation may mark chromatin as being ready for cell cycle progression Expression of cell cycle regulatory genes could be modulated by histone acetylation

Chromosome Biology: How is gene expression regulated? How is DNA replication regulated?
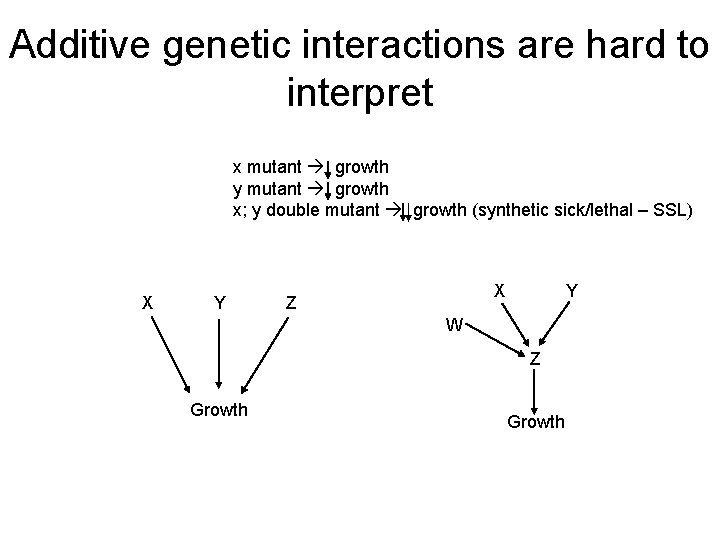
Additive genetic interactions are hard to interpret x mutant growth y mutant growth x; y double mutant growth (synthetic sick/lethal – SSL) X Y X Z Y W Z Growth
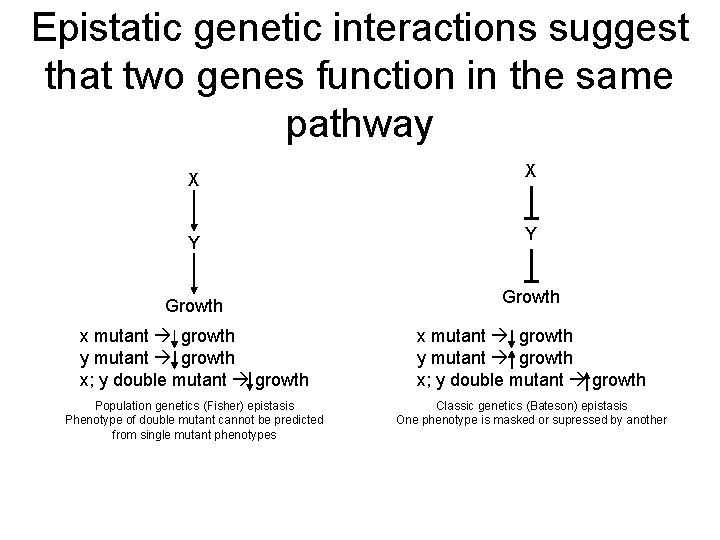
Epistatic genetic interactions suggest that two genes function in the same pathway X X Y Y Growth x mutant growth y mutant growth x; y double mutant growth Population genetics (Fisher) epistasis Phenotype of double mutant cannot be predicted from single mutant phenotypes Classic genetics (Bateson) epistasis One phenotype is masked or supressed by another
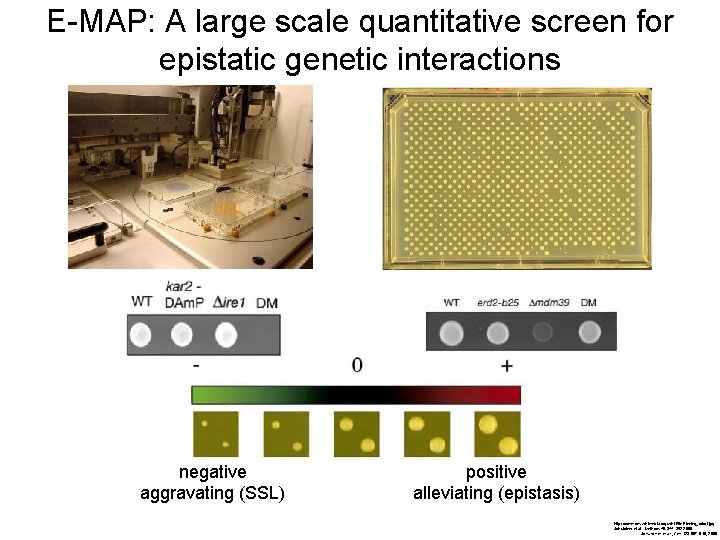
E-MAP: A large scale quantitative screen for epistatic genetic interactions negative aggravating (SSL) positive alleviating (epistasis) http: //commons. wikimedia. org/wiki/File: Pinning_robot. jpg Schuldiner et al. , Methods 40: 344– 352 2006 Schuldiner et al. , Cell 123: 507– 519, 2005
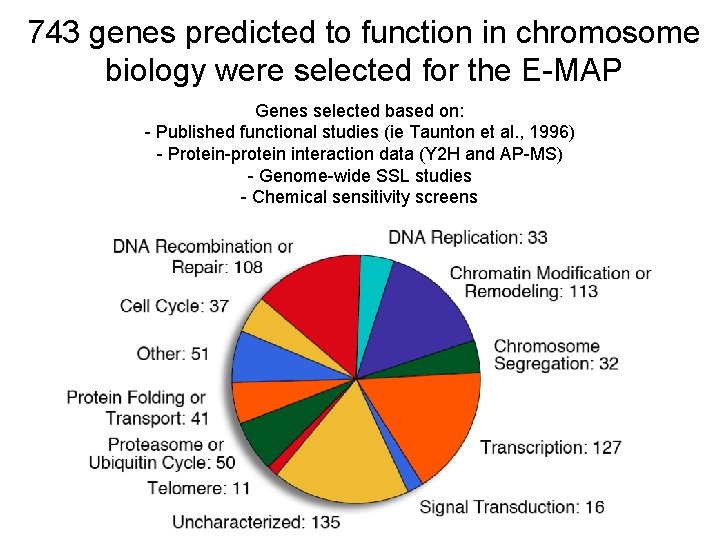
743 genes predicted to function in chromosome biology were selected for the E-MAP Genes selected based on: - Published functional studies (ie Taunton et al. , 1996) - Protein-protein interaction data (Y 2 H and AP-MS) - Genome-wide SSL studies - Chemical sensitivity screens
Network diagrams do not allow the identification of discrete biological functional modules
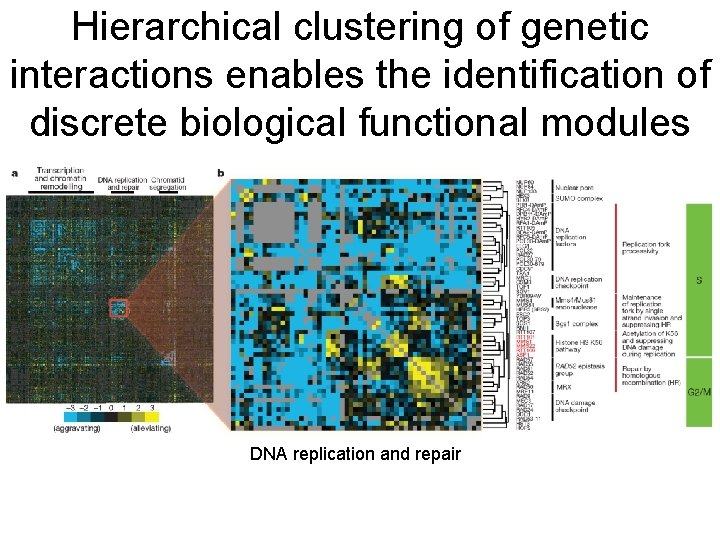
Hierarchical clustering of genetic interactions enables the identification of discrete biological functional modules DNA replication and repair

Epistasis groups correlate with known physical complexes
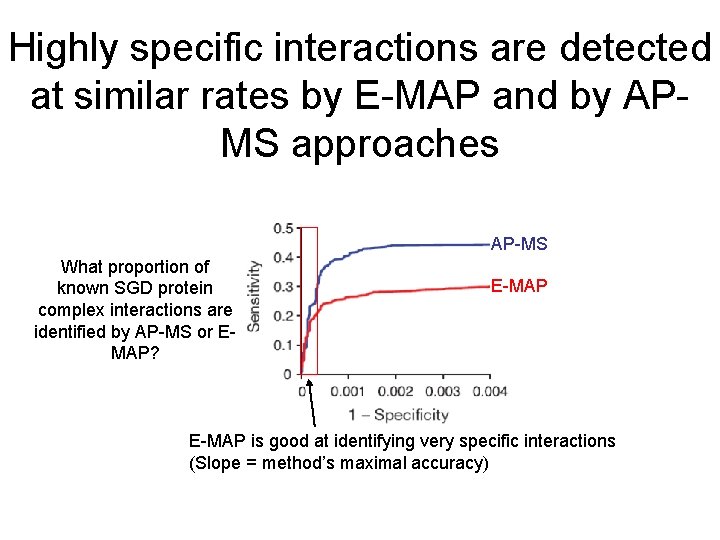
Highly specific interactions are detected at similar rates by E-MAP and by APMS approaches AP-MS What proportion of known SGD protein complex interactions are identified by AP-MS or EMAP? E-MAP is good at identifying very specific interactions (Slope = method’s maximal accuracy)
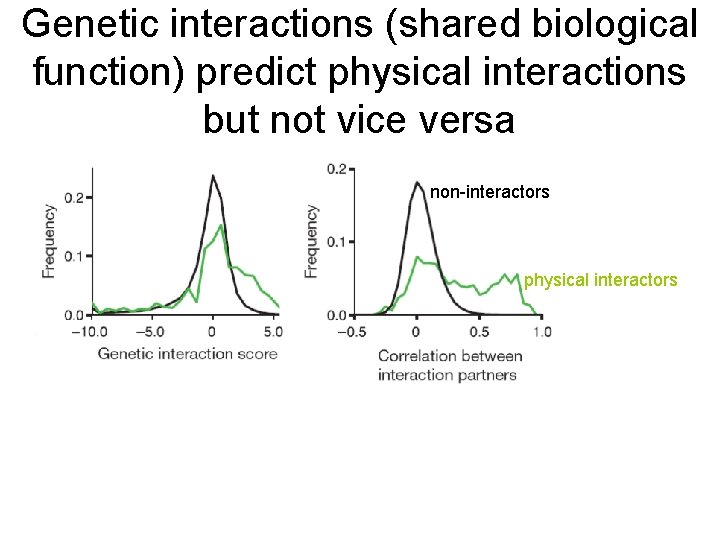
Genetic interactions (shared biological function) predict physical interactions but not vice versa non-interactors physical interactors

An example of dissecting multiprotein complexes into functional modules using genetic interaction mapping
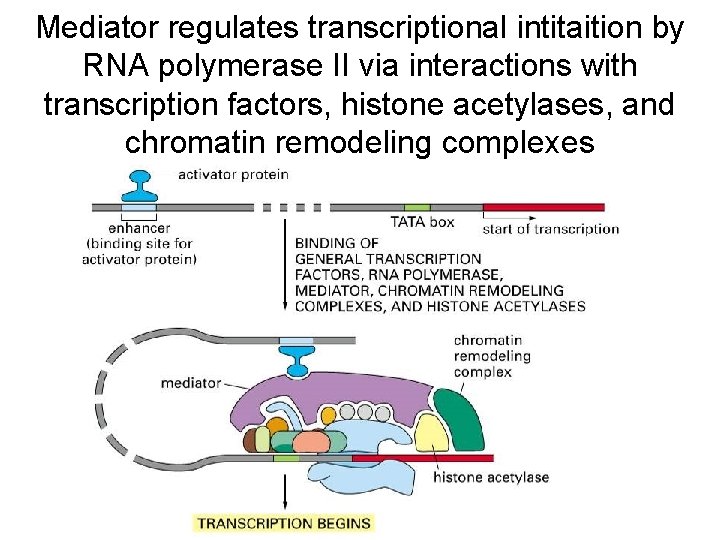
Mediator regulates transcriptional intitaition by RNA polymerase II via interactions with transcription factors, histone acetylases, and chromatin remodeling complexes
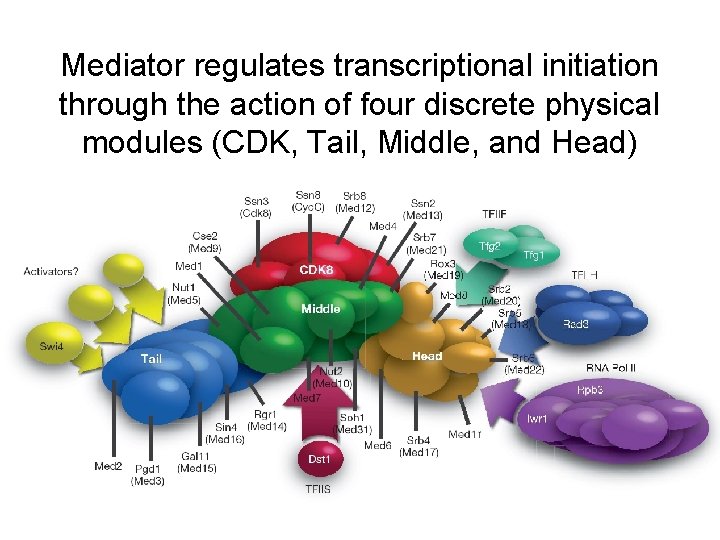
Mediator regulates transcriptional initiation through the action of four discrete physical modules (CDK, Tail, Middle, and Head)
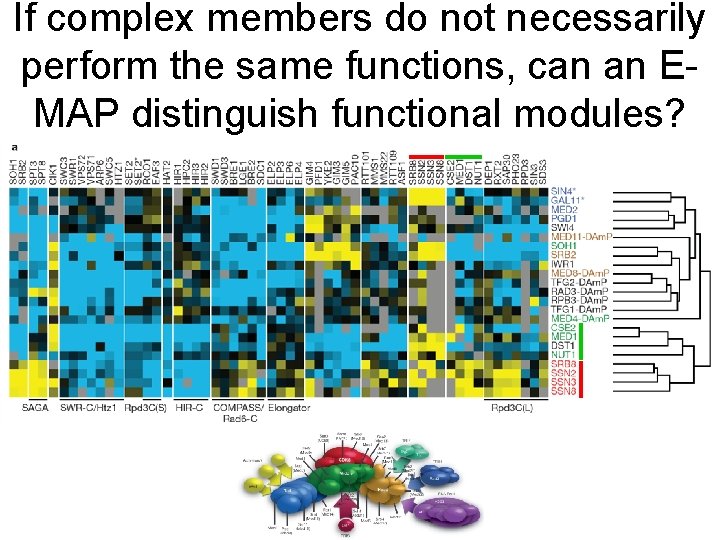
If complex members do not necessarily perform the same functions, can an EMAP distinguish functional modules?
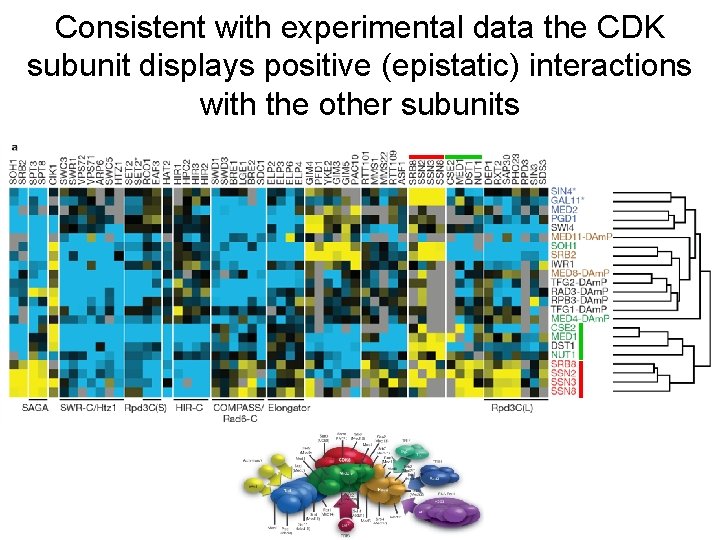
Consistent with experimental data the CDK subunit displays positive (epistatic) interactions with the other subunits
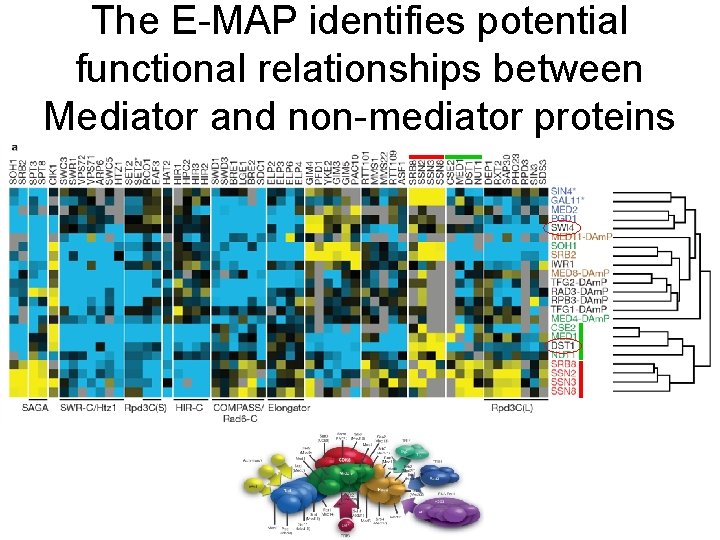
The E-MAP identifies potential functional relationships between Mediator and non-mediator proteins

OK – so the E-MAP is good at confirming what we already know about functional and physical interactions. Can it be used as a tool for identifying novel functions related to chromosome biology?

What does this uncharacterized epistasis group in the DNA damage cluster do? ?

What is known about the proteins in this novel epistasis cluster? • RTT 101 • MMS 22 Physically associated with each other and other proteins Complex has characteristic features of a ubiquitin ligase • RTT 109 - poorly characterized, regulates Ty 1 transposition • ASF 1 - histone chaperone required for histone assembly, supression of spurious transcriptional initiation, and H 3 K 56 acetylation
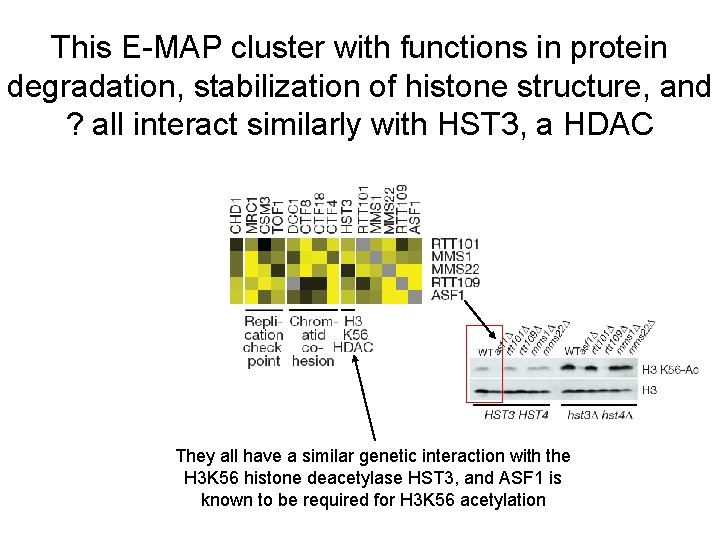
This E-MAP cluster with functions in protein degradation, stabilization of histone structure, and ? all interact similarly with HST 3, a HDAC They all have a similar genetic interaction with the H 3 K 56 histone deacetylase HST 3, and ASF 1 is known to be required for H 3 K 56 acetylation
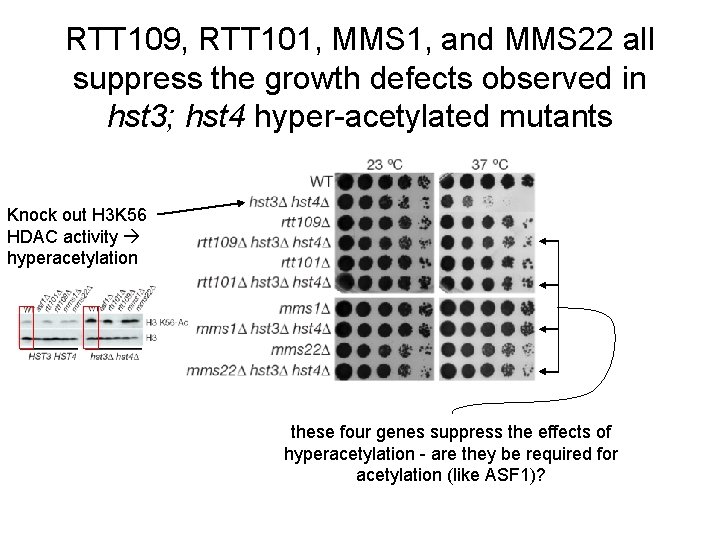
RTT 109, RTT 101, MMS 1, and MMS 22 all suppress the growth defects observed in hst 3; hst 4 hyper-acetylated mutants Knock out H 3 K 56 HDAC activity hyperacetylation these four genes suppress the effects of hyperacetylation - are they be required for acetylation (like ASF 1)?
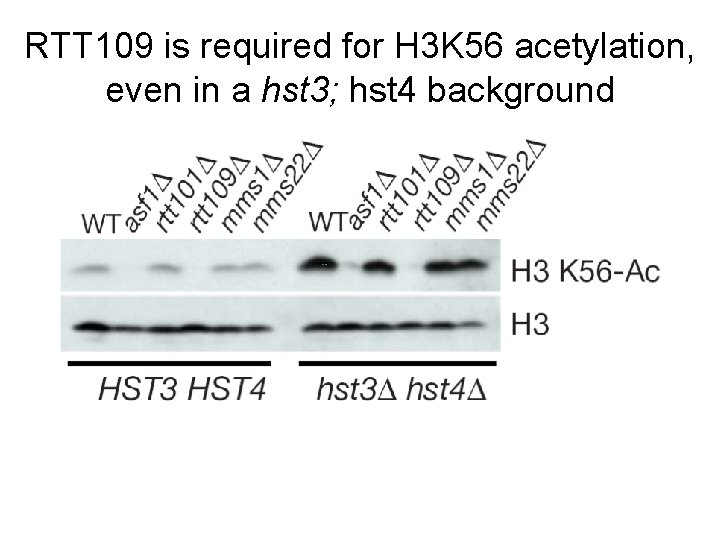
RTT 109 is required for H 3 K 56 acetylation, even in a hst 3; hst 4 background
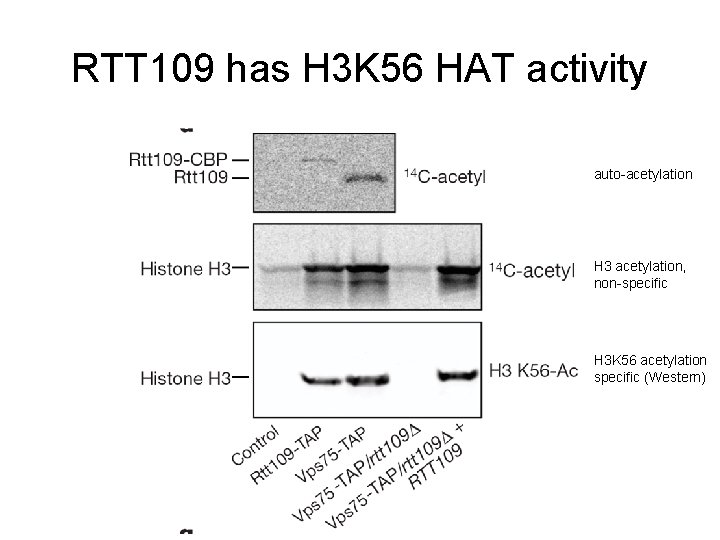
RTT 109 has H 3 K 56 HAT activity auto-acetylation H 3 acetylation, non-specific H 3 K 56 acetylation specific (Western)
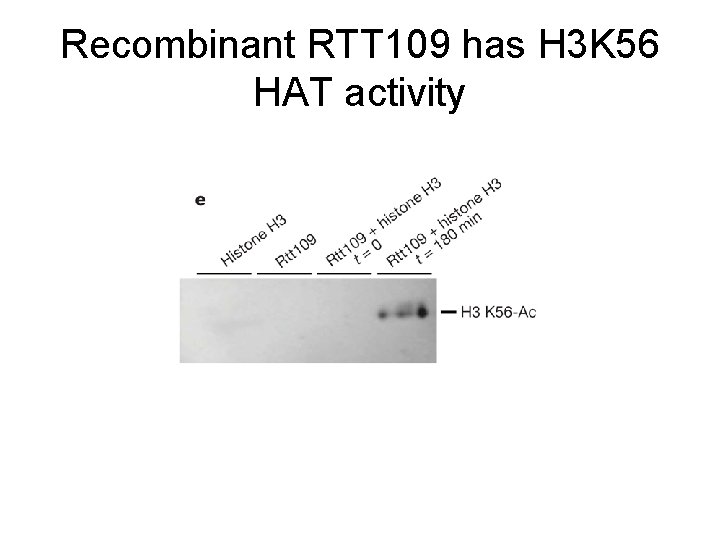
Recombinant RTT 109 has H 3 K 56 HAT activity

OK – so the E-MAP can be used to generate testable hypotheses about the function of uncharacterized genes. Isn’t this a lot of work to identify the H 3 K 56 HAT? What does the E-MAP tell you that conventional approaches can’t?
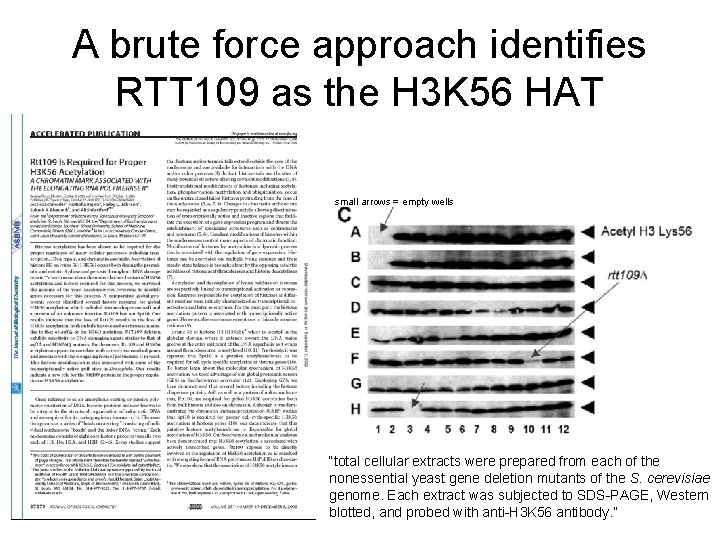
A brute force approach identifies RTT 109 as the H 3 K 56 HAT small arrows = empty wells “total cellular extracts were prepared from each of the nonessential yeast gene deletion mutants of the S. cerevisiae genome. Each extract was subjected to SDS-PAGE, Western blotted, and probed with anti-H 3 K 56 antibody. ”
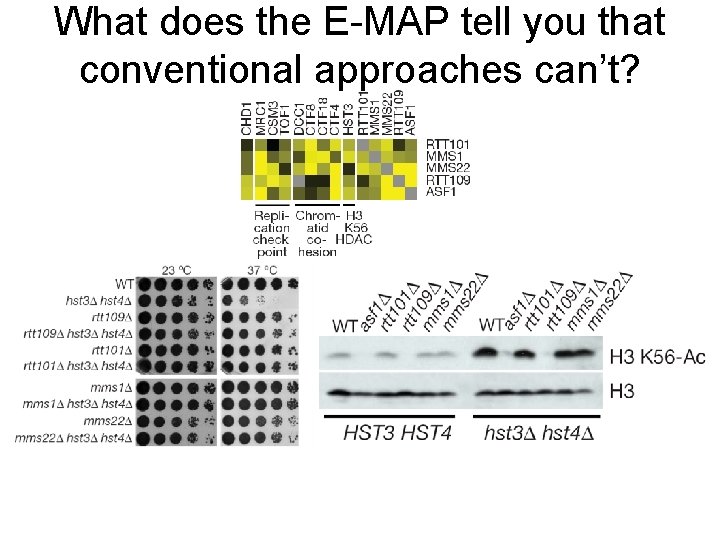
What does the E-MAP tell you that conventional approaches can’t?
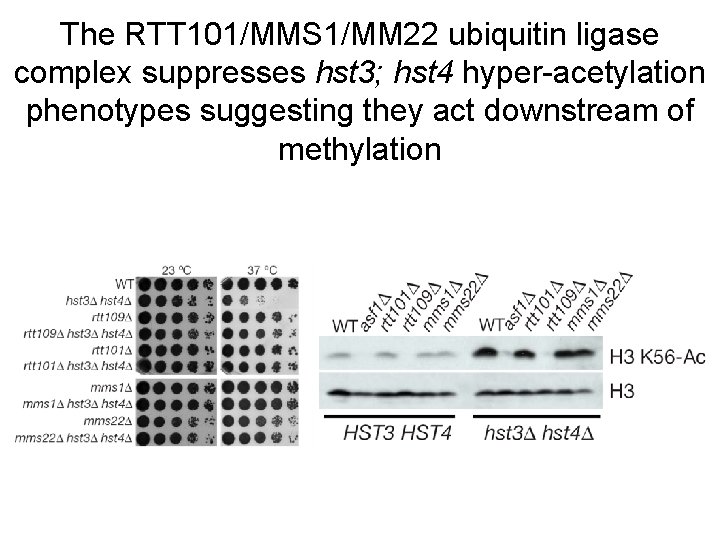
The RTT 101/MMS 1/MM 22 ubiquitin ligase complex suppresses hst 3; hst 4 hyper-acetylation phenotypes suggesting they act downstream of methylation
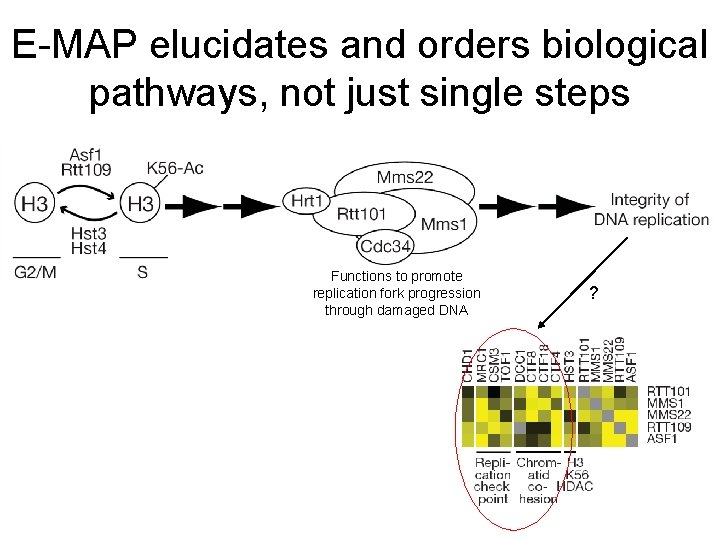
E-MAP elucidates and orders biological pathways, not just single steps Functions to promote replication fork progression through damaged DNA ?

Just because a data set is large does not mean it is complete

There is a lot of PPI data out there

How good is available interactome data? Uetz-array excluded because the interactions were targeted/biased Ito-core interactions were found at least 3 times, nocore were only found 1 or 2 times LC-multiple interactions were found in at least 2 articles
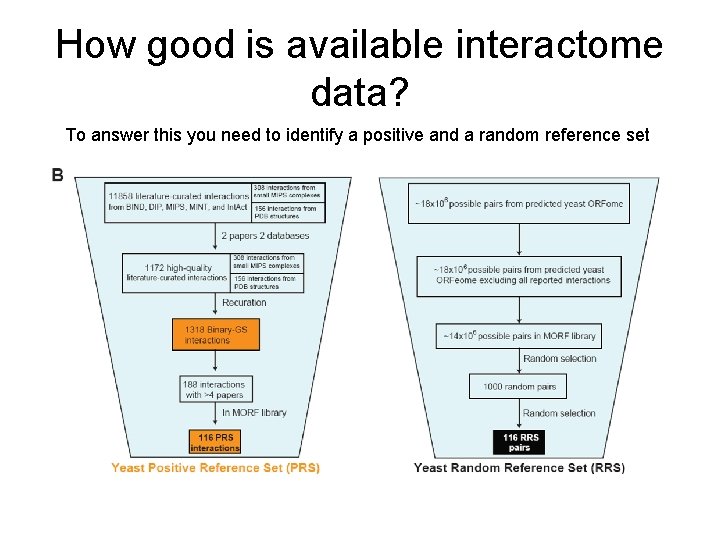
How good is available interactome data? To answer this you need to identify a positive and a random reference set
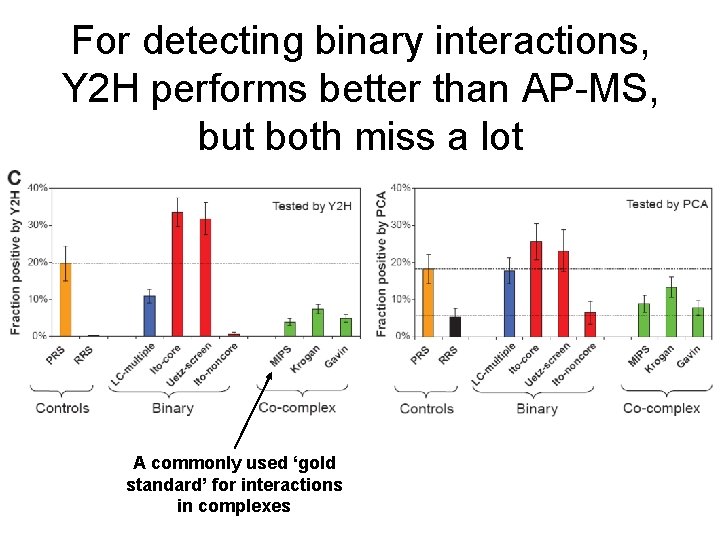
For detecting binary interactions, Y 2 H performs better than AP-MS, but both miss a lot A commonly used ‘gold standard’ for interactions in complexes
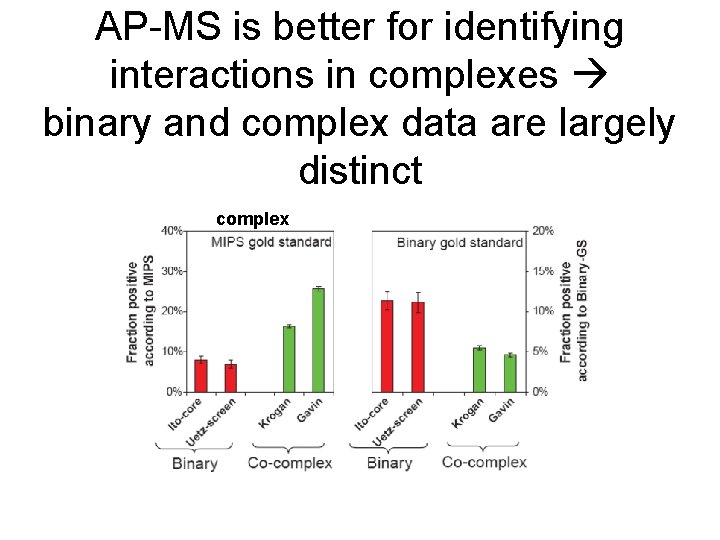
AP-MS is better for identifying interactions in complexes binary and complex data are largely distinct complex
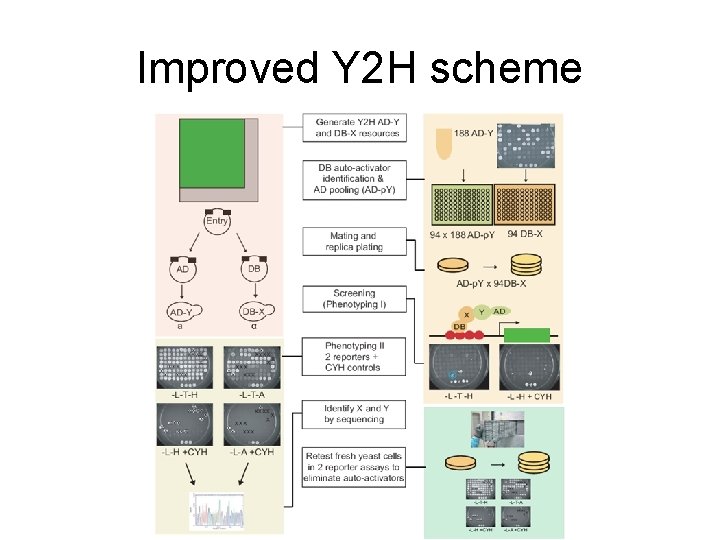
Improved Y 2 H scheme

Three factors result in incomplete Y 2 H binary datasets • Incomplete libraries (10% missing only search 66% of possible interactions) • Low sensitivity (Y 2 H identifies only 20% of PRS interactions) • Limited sampling
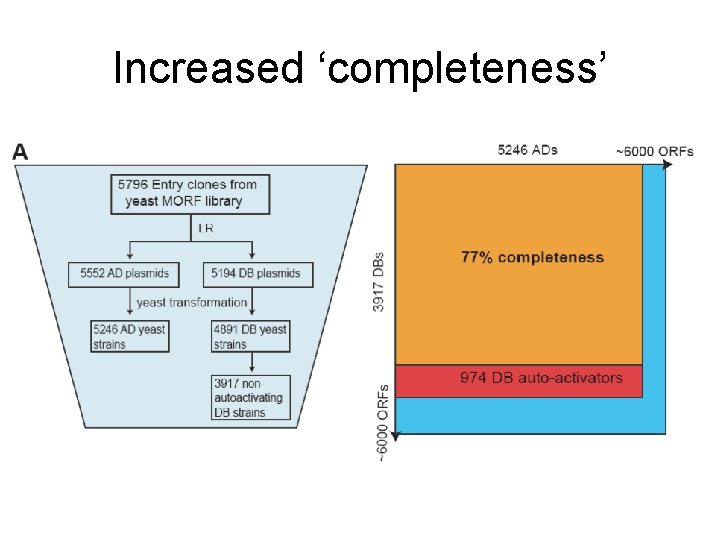
Increased ‘completeness’
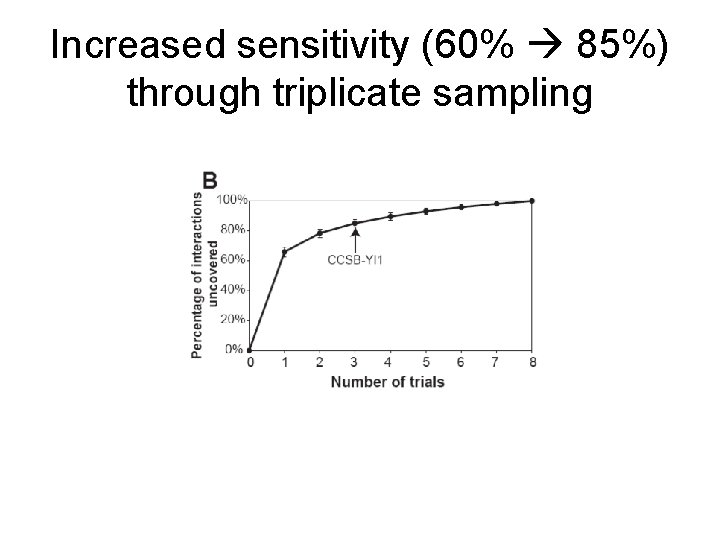
Increased sensitivity (60% 85%) through triplicate sampling

CCSB-YI 1 tested 13, 790, 256 unique pairs and identified 1809 interactions Factoring in completeness, precision, and sensitivity, the yeast binary interactome should contain ~18, 000 interactions 90% remain unidentified by this screen!

What do you learn from all this interaction data?
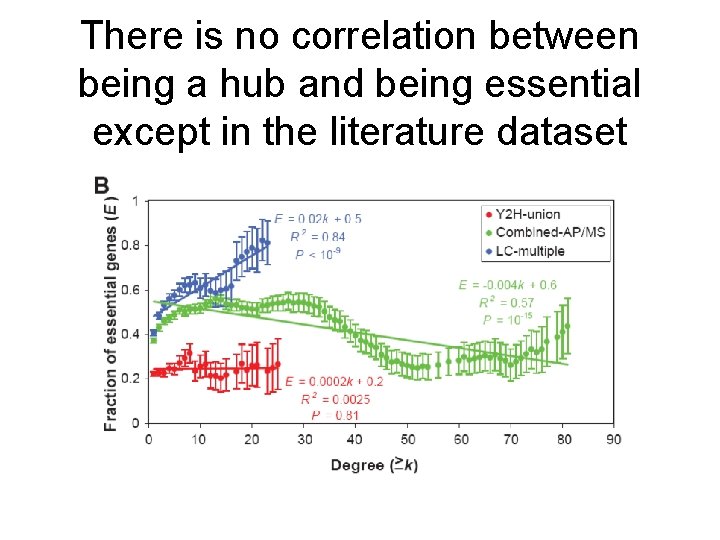
There is no correlation between being a hub and being essential except in the literature dataset
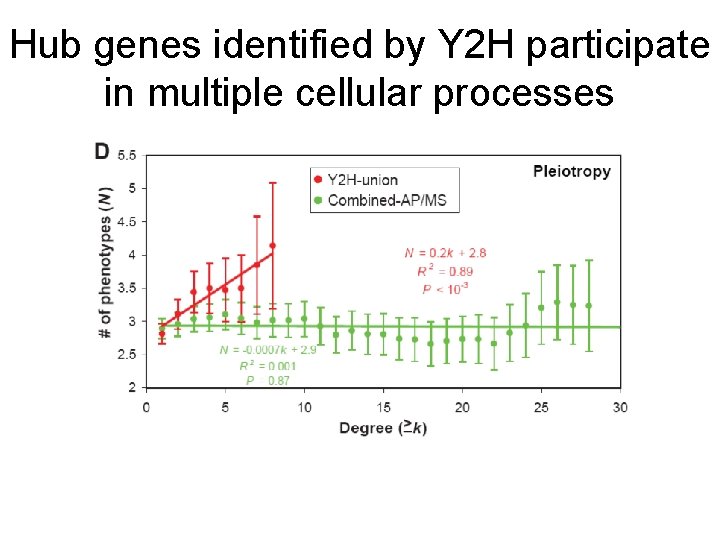
Hub genes identified by Y 2 H participate in multiple cellular processes
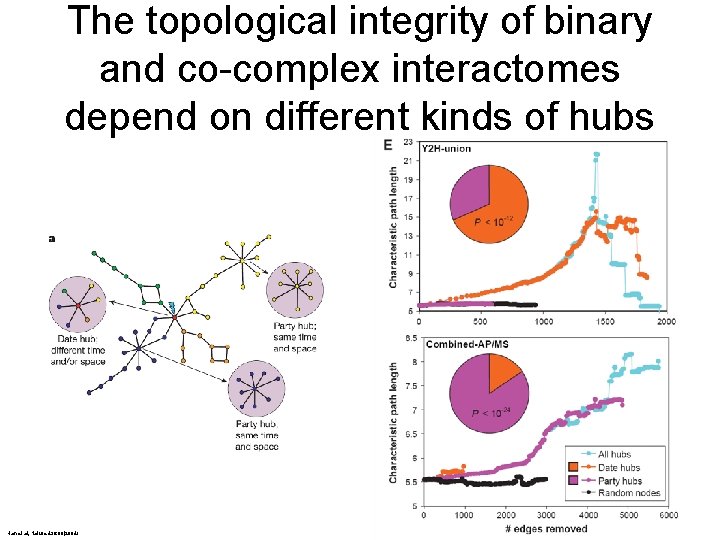
The topological integrity of binary and co-complex interactomes depend on different kinds of hubs Han et al, Nature 430: 88(2004)

Removing date and party hubs from a yeast interactome network whole Han et al, Nature 430: 88(2004) - date - party
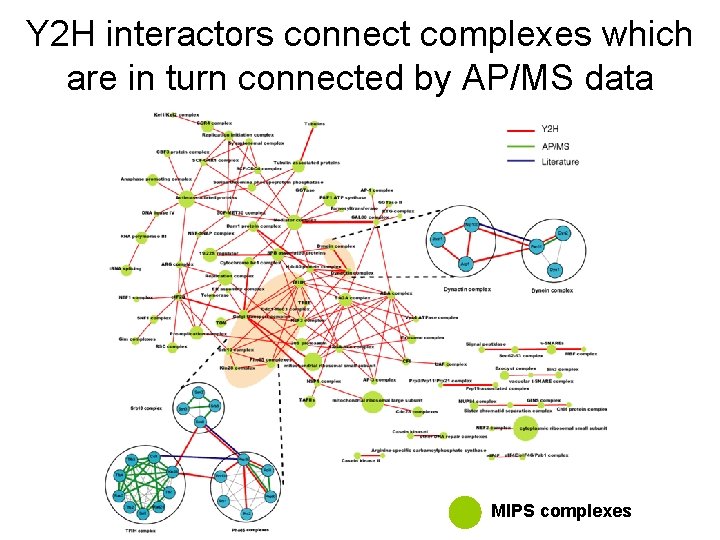
Y 2 H interactors connect complexes which are in turn connected by AP/MS data MIPS complexes
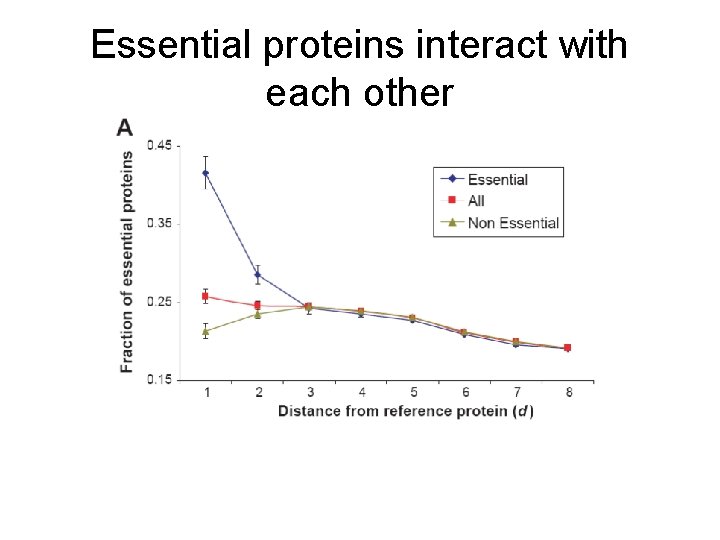
Essential proteins interact with each other
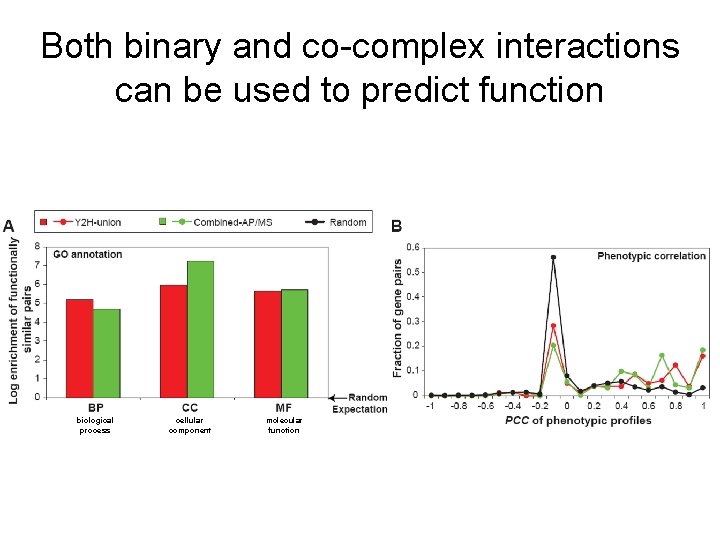
Both binary and co-complex interactions can be used to predict function biological process cellular component molecular function
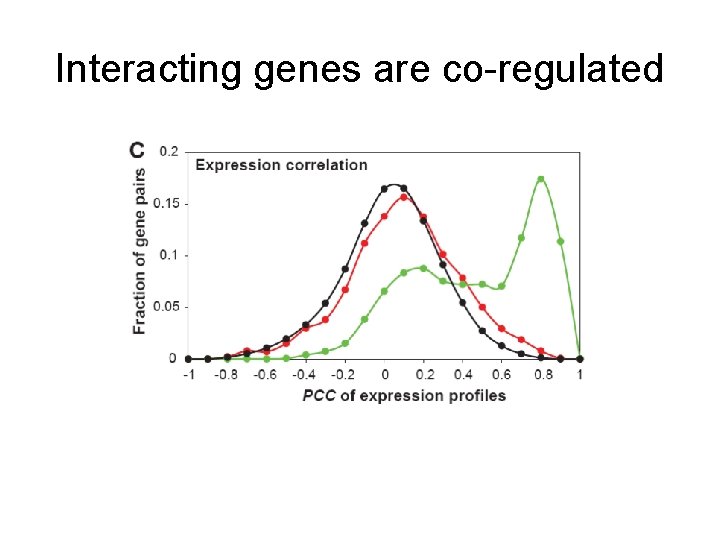
Interacting genes are co-regulated
- Slides: 63