Chromosomal structures and transposable elements Package DNA in
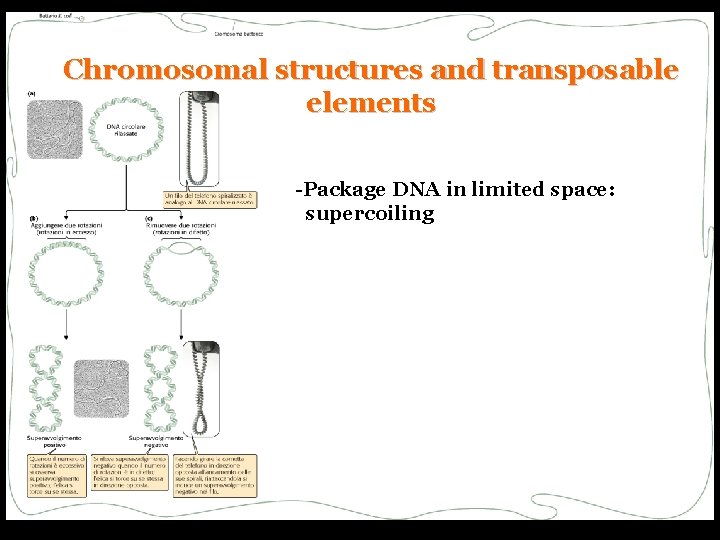
Chromosomal structures and transposable elements -Package DNA in limited space: supercoiling
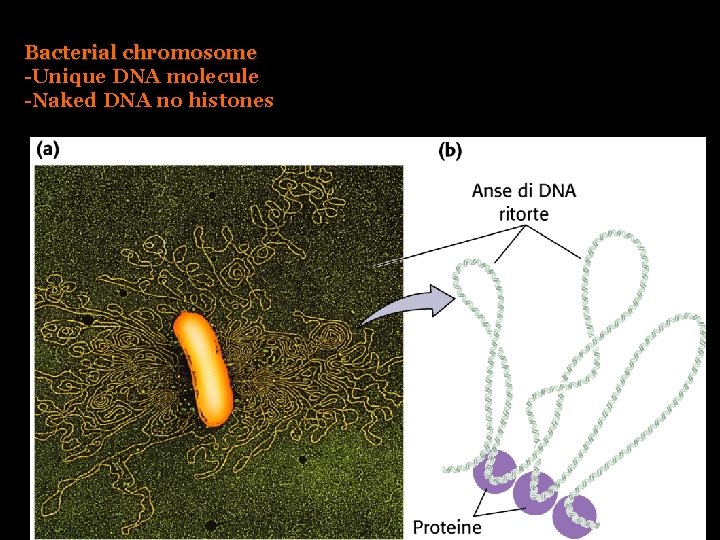
Bacterial chromosome -Unique DNA molecule -Naked DNA no histones
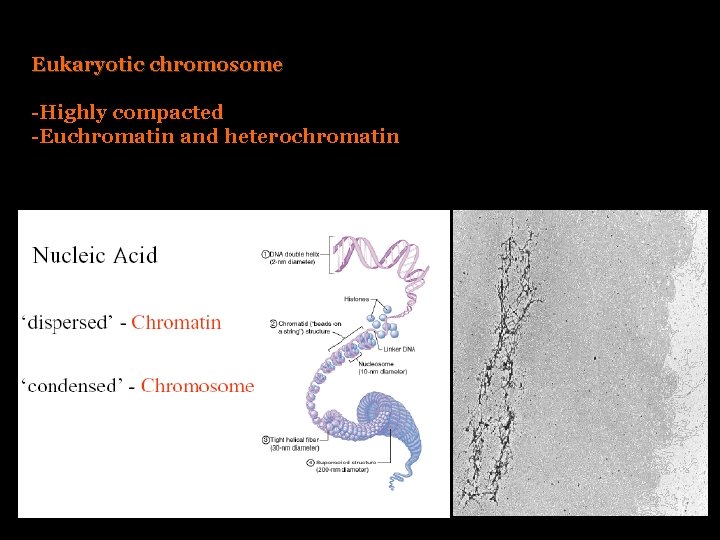
Eukaryotic chromosome -Highly compacted -Euchromatin and heterochromatin
CHROMATIN
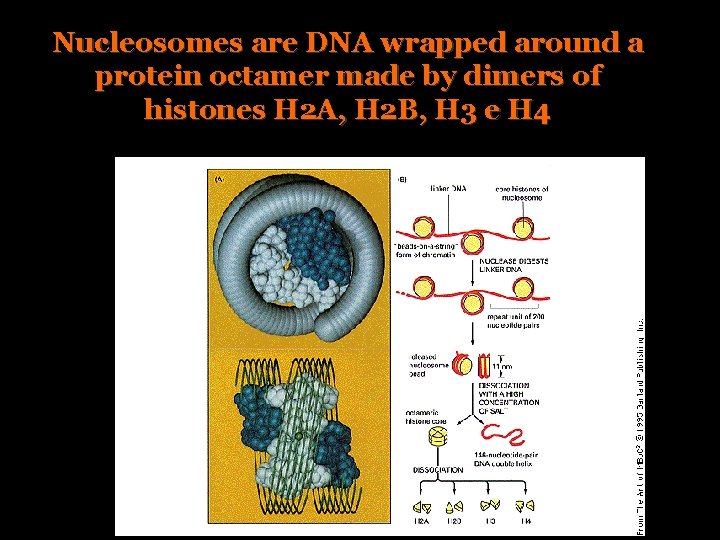
Nucleosomes are DNA wrapped around a protein octamer made by dimers of histones H 2 A, H 2 B, H 3 e H 4

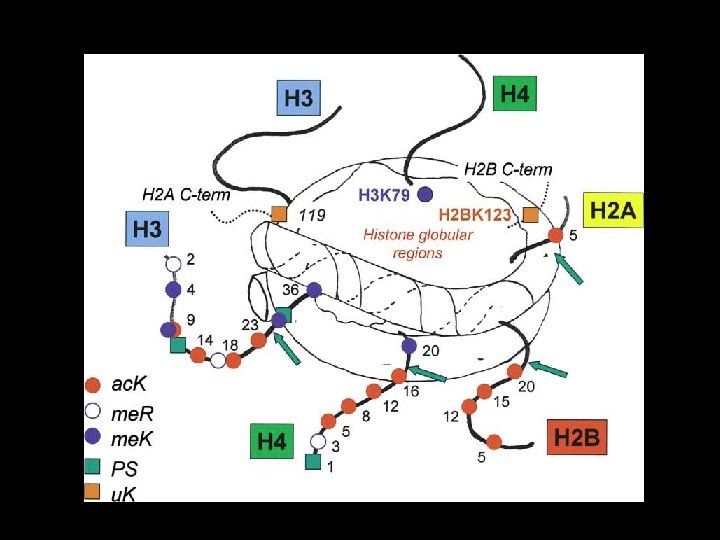
HISTONES H 1 are highly conserved, small, basic proteins Linker histone H 2 A H 2 B helix Core histones variable H 3 H 4 conserved N Histone acetylation is a reversible modification of lysines in the N-termini of the core histones. Result: • reduced binding to DNA • destabilization of chromatin
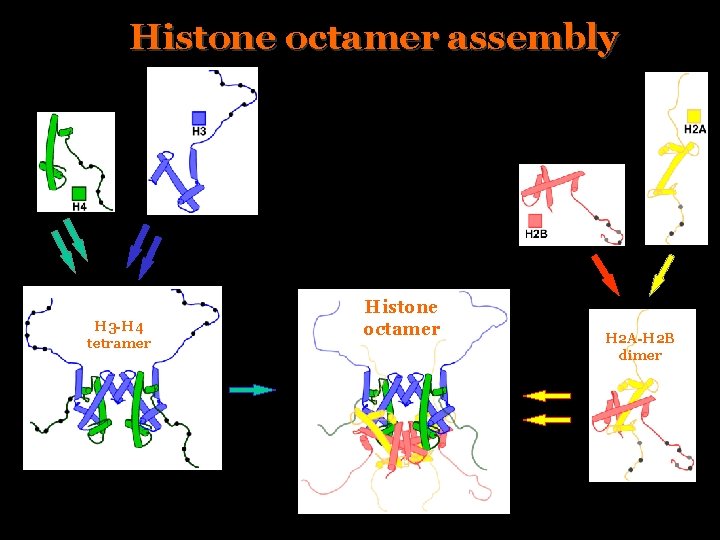
Histone octamer assembly H 3 -H 4 tetramer Histone octamer H 2 A-H 2 B dimer
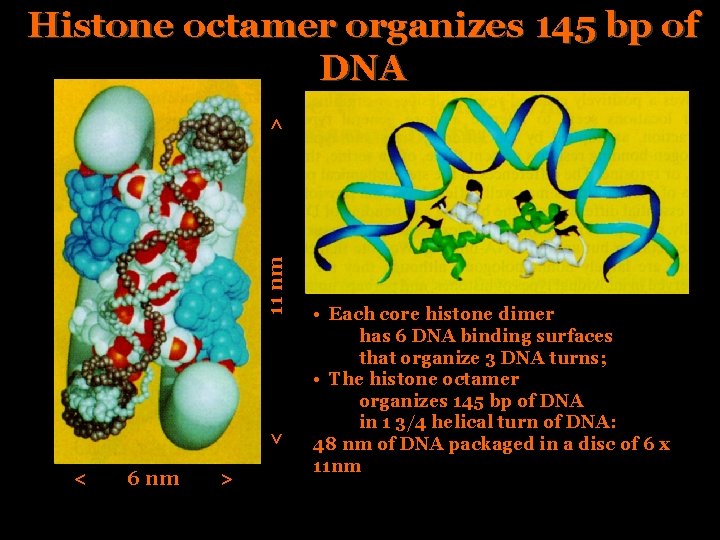
< 11 nm > Histone octamer organizes 145 bp of DNA < 6 nm > • Each core histone dimer has 6 DNA binding surfaces that organize 3 DNA turns; • The histone octamer organizes 145 bp of DNA in 1 3/4 helical turn of DNA: 48 nm of DNA packaged in a disc of 6 x 11 nm
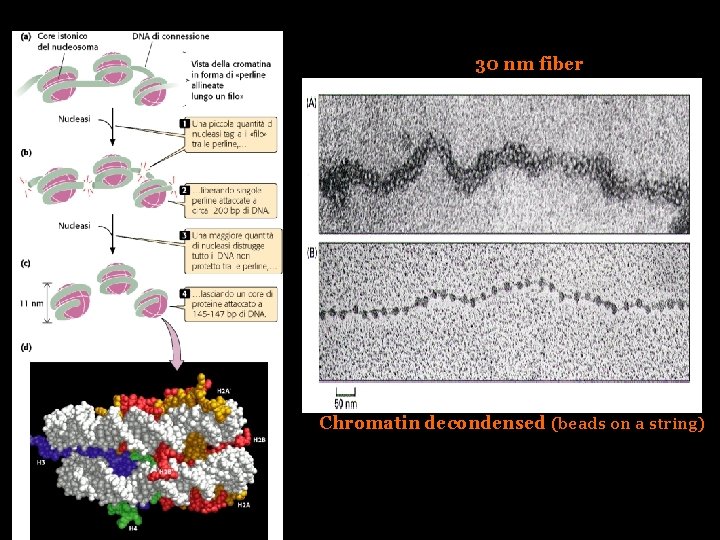
30 nm fiber Chromatin decondensed (beads on a string)
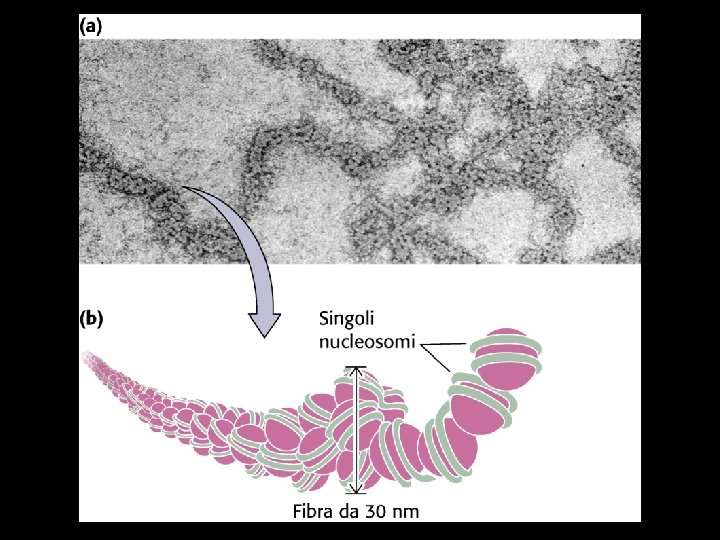
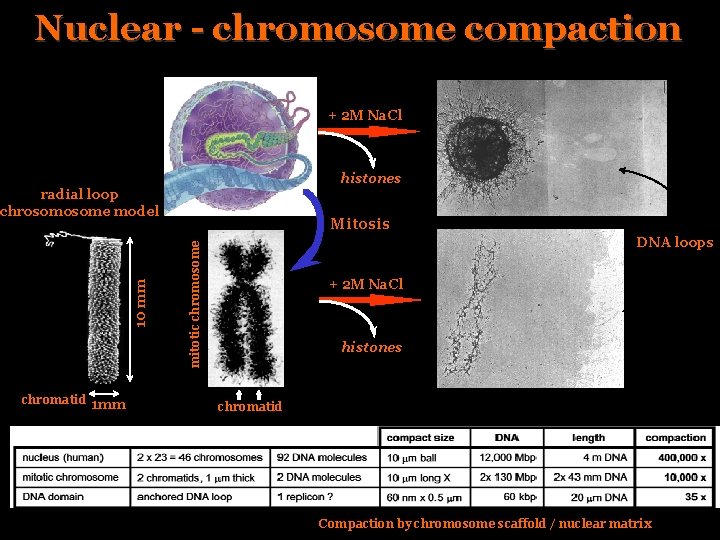
Nuclear - chromosome compaction + 2 M Na. Cl histones chromatid 1 mm Mitosis DNA loops mitotic chromosome 10 mm radial loop chrosome model + 2 M Na. Cl histones chromatid Compaction by chromosome scaffold / nuclear matrix

Variation in the chromatin structure -Polytenic chromosomes in Drosophila, DNA replication non associated to cell division. -Sensitivity to DNAse I correlates with gene activity
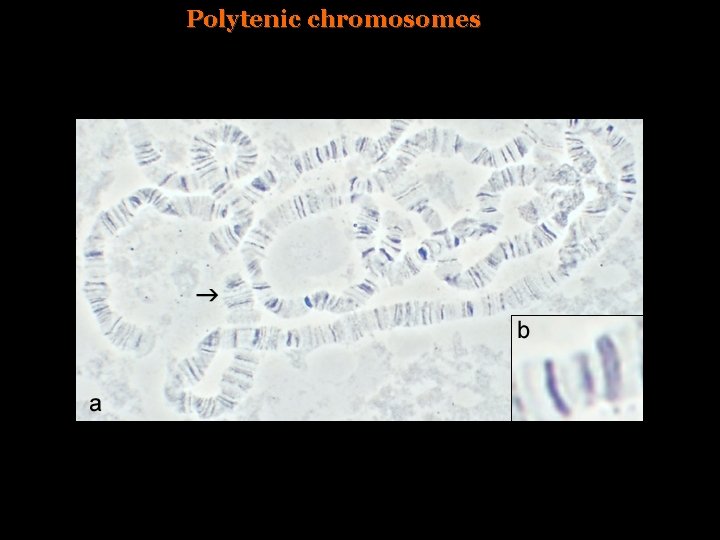
Polytenic chromosomes
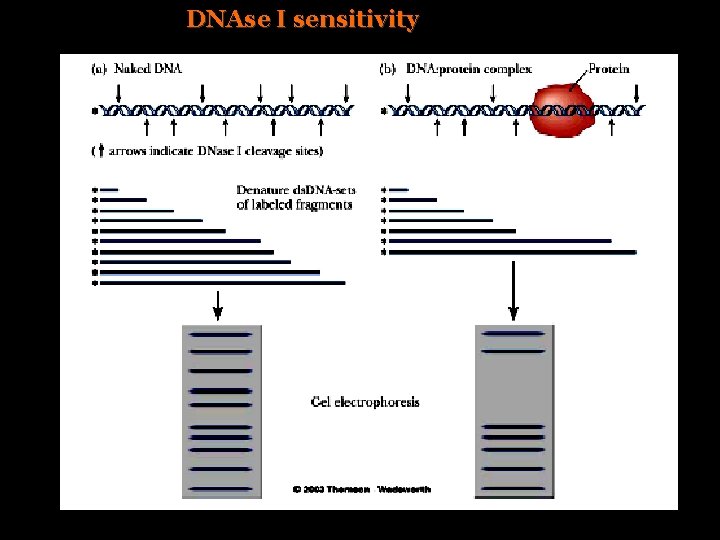
DNAse I sensitivity
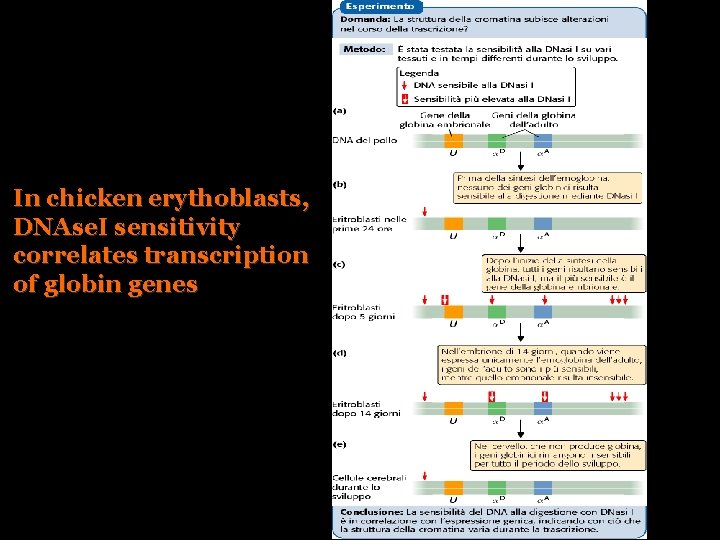
In chicken erythoblasts, DNAse. I sensitivity correlates transcription of globin genes
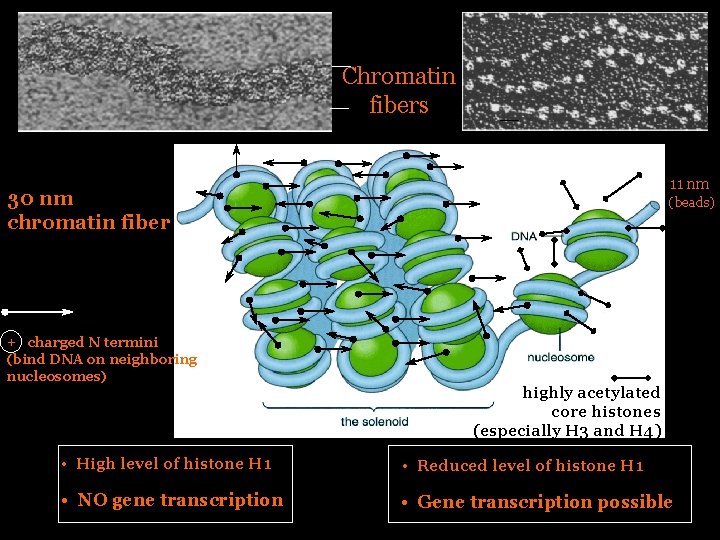
Chromatin fibers 11 nm 30 nm chromatin fiber + charged N termini (bind DNA on neighboring nucleosomes) (beads) highly acetylated core histones (especially H 3 and H 4) • High level of histone H 1 • Reduced level of histone H 1 • NO gene transcription • Gene transcription possible
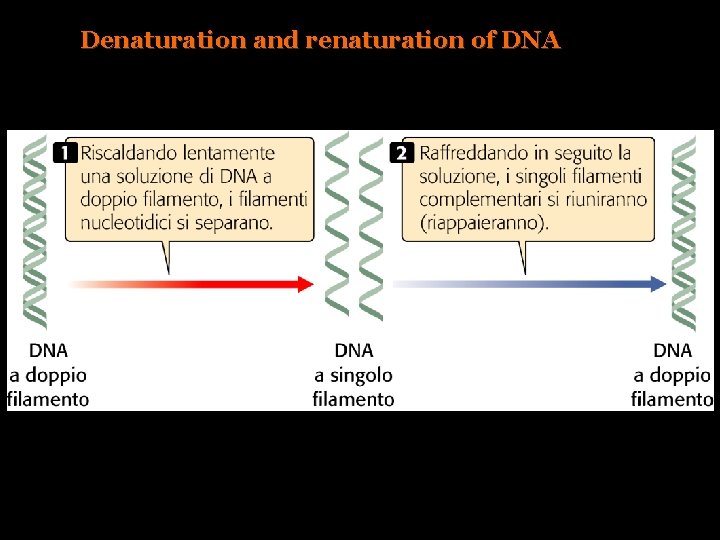
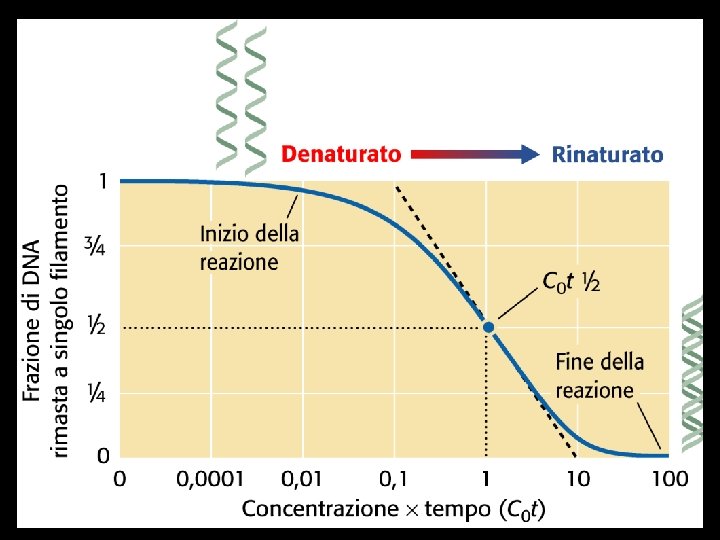
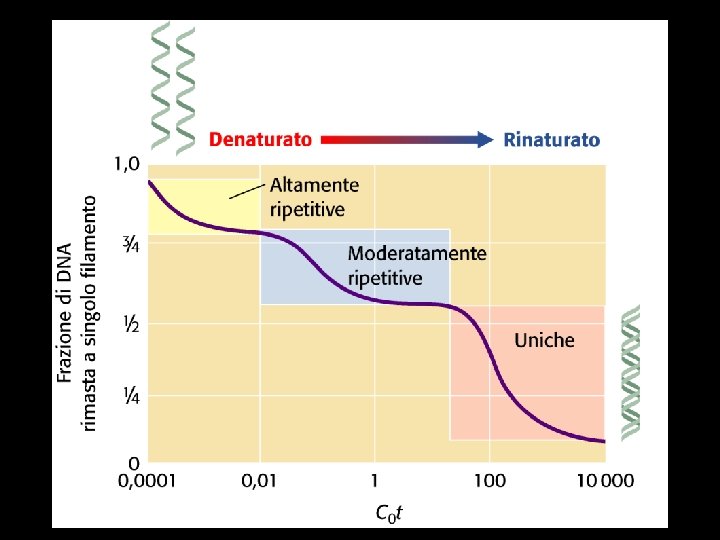
Denaturation and renaturation of DNA
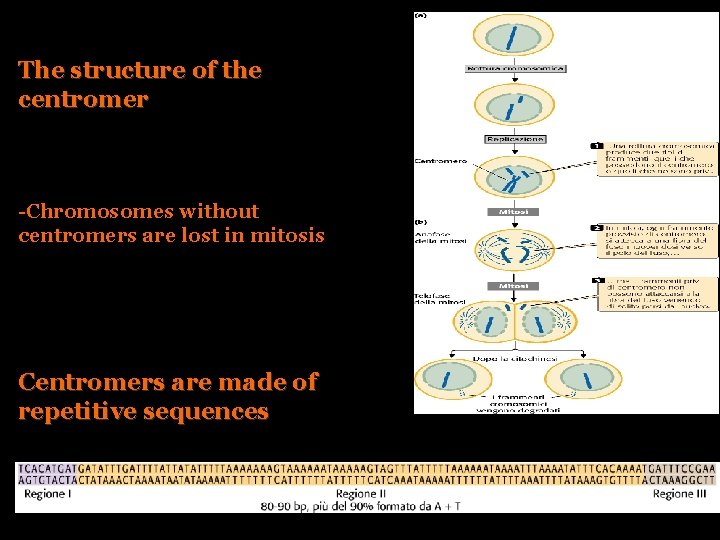
The structure of the centromer -Chromosomes without centromers are lost in mitosis Centromers are made of repetitive sequences
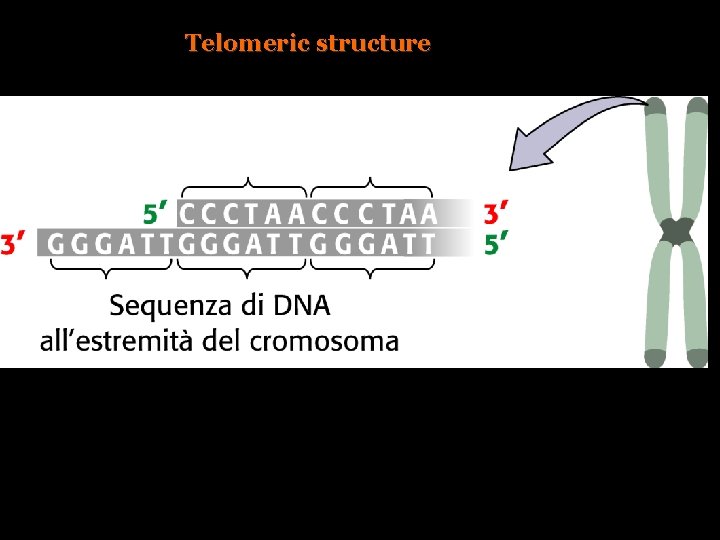
Telomeric structure
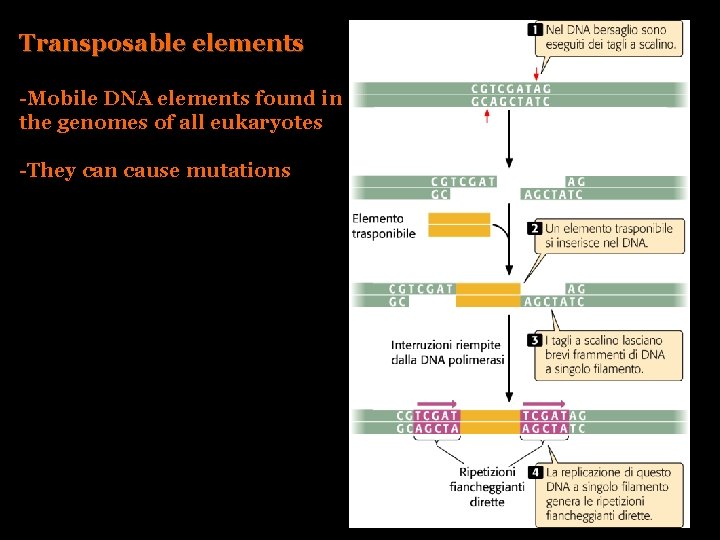
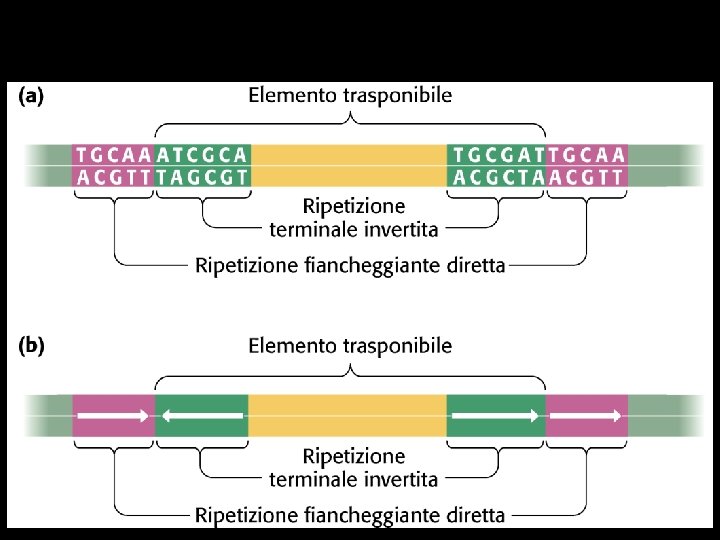
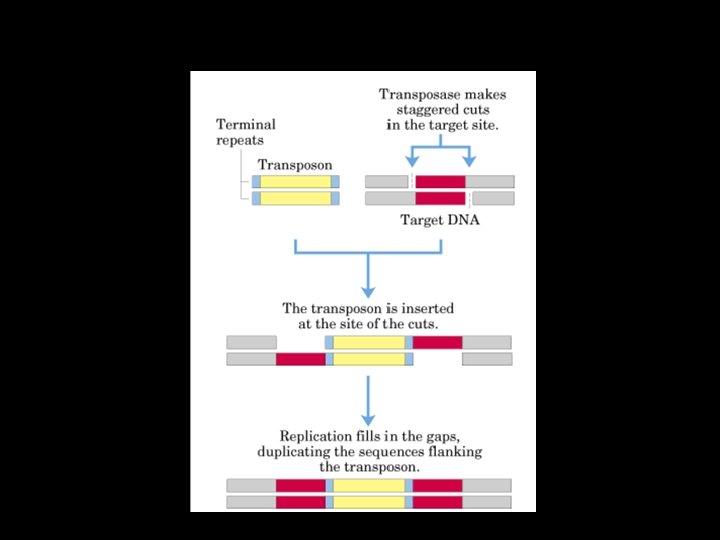
Transposable elements -Mobile DNA elements found in the genomes of all eukaryotes -They can cause mutations
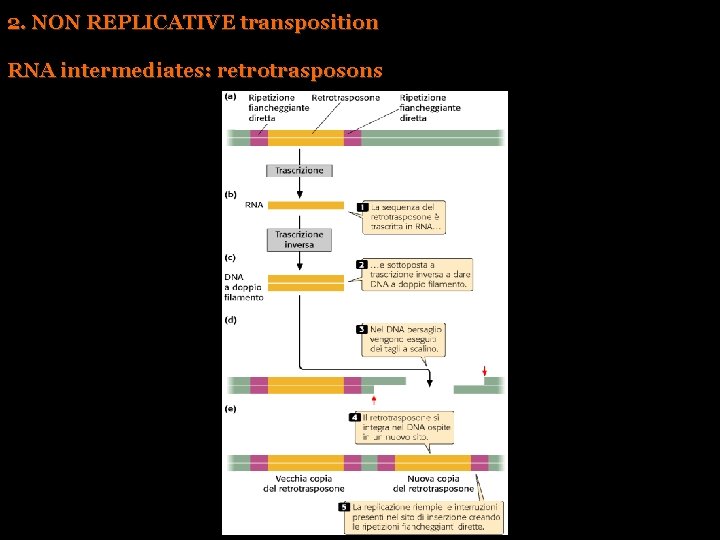
2. NON REPLICATIVE transposition RNA intermediates: retrotrasposons
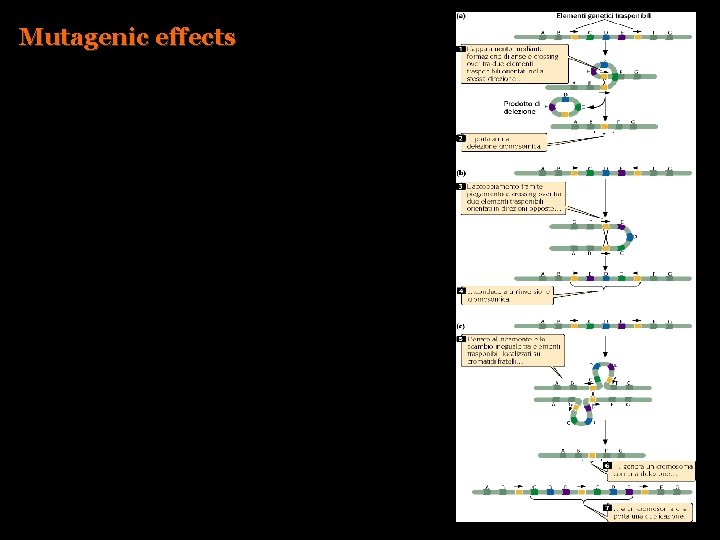
Mutagenic effects
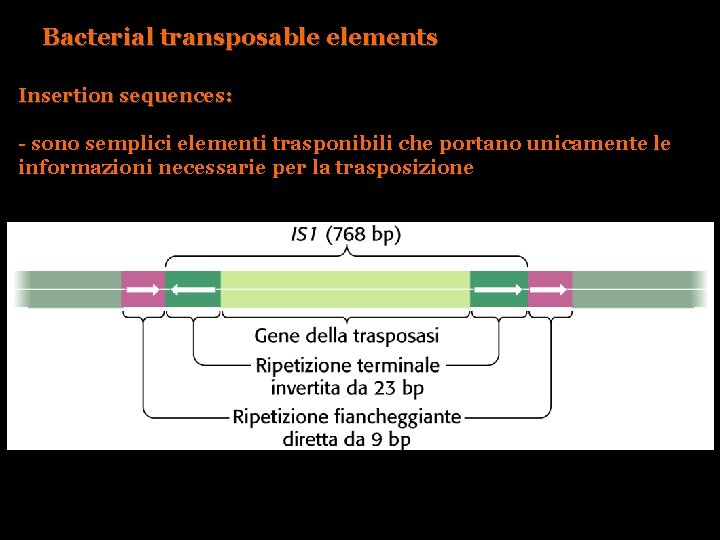
Bacterial transposable elements Insertion sequences: - sono semplici elementi trasponibili che portano unicamente le informazioni necessarie per la trasposizione
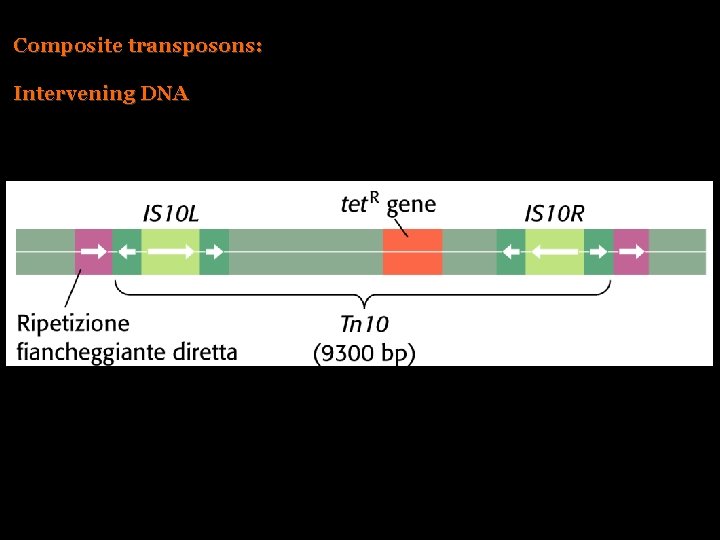
Composite transposons: Intervening DNA
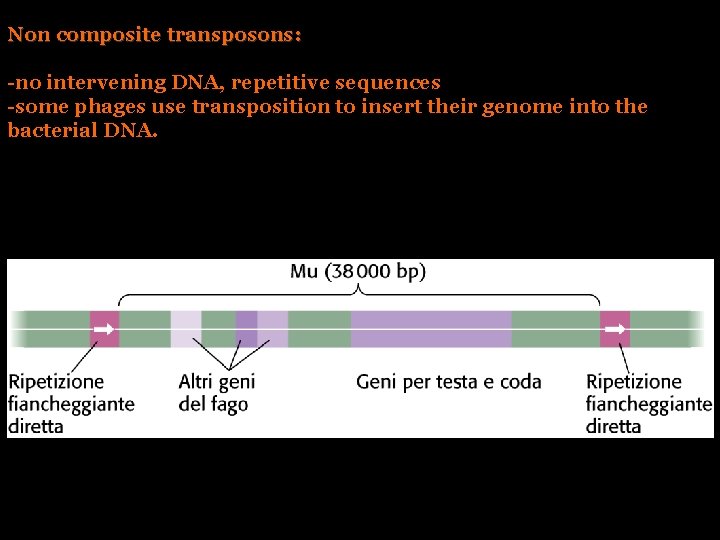
Non composite transposons: -no intervening DNA, repetitive sequences -some phages use transposition to insert their genome into the bacterial DNA.
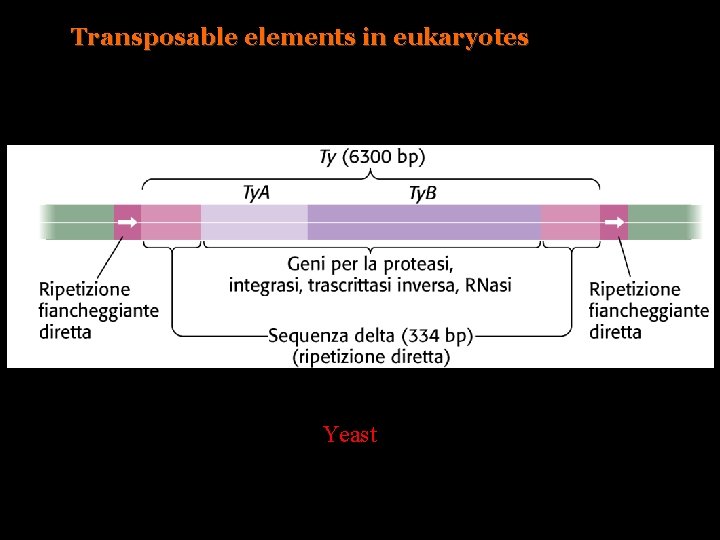
Transposable elements in eukaryotes Yeast

Barbara Mc. Clintock discovered transposable elements studying variegated maize

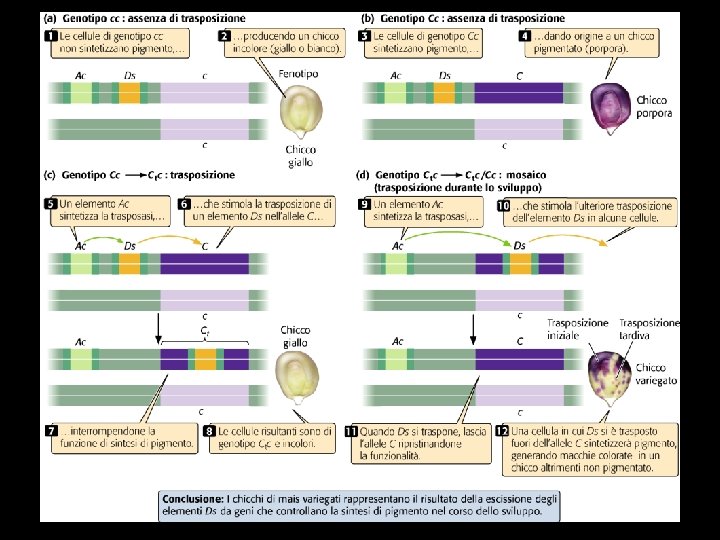
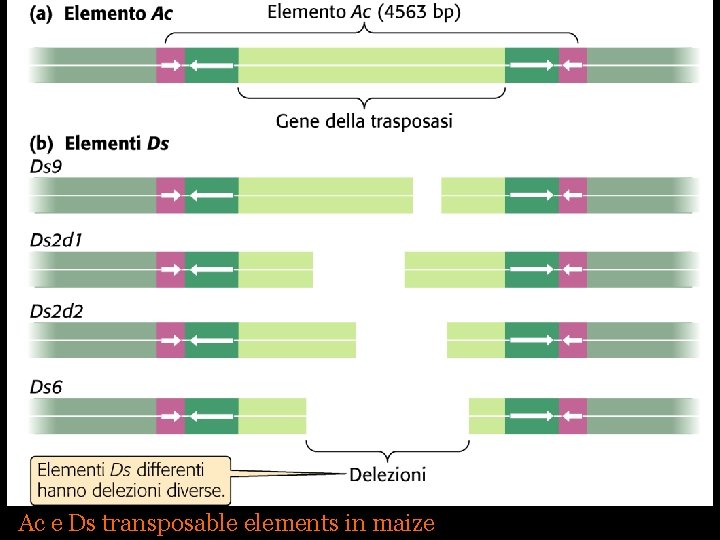
Ac e Ds transposable elements in maize
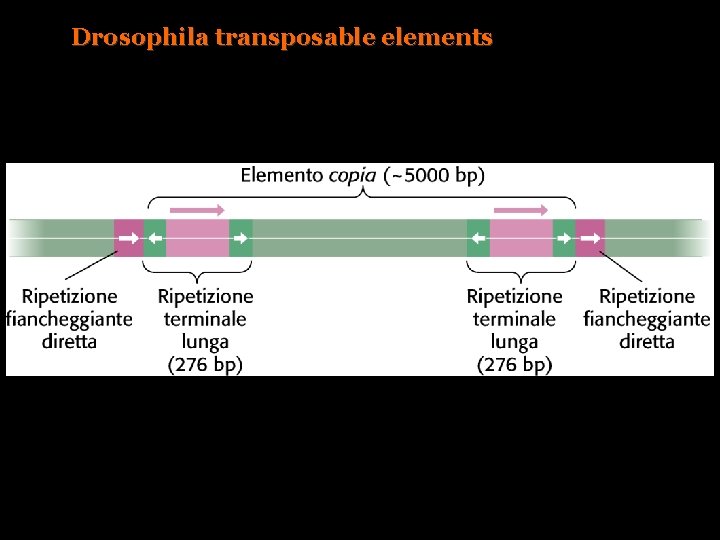
Drosophila transposable elements
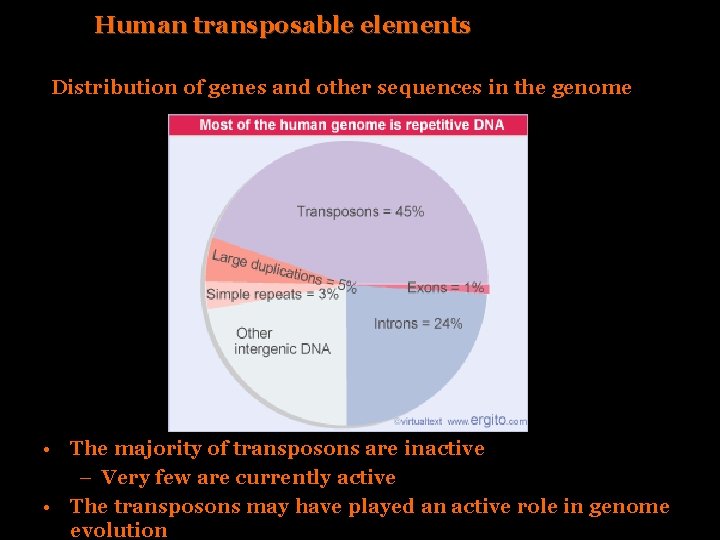
Human transposable elements Distribution of genes and other sequences in the genome • The majority of transposons are inactive – Very few are currently active • The transposons may have played an active role in genome evolution

- Slides: 38