Chemometrics assisted methodologies for solving real analytical problems

Chemometrics assisted methodologies for solving real analytical problems 1 Maryam Vosough Department of Green Technologies Chemistry and Chemical Engineering Research Center of Iran Website: ecgt. ccerci. ac. ir Email: vosough@ccerci. ac. ir

Introduction Due to rapid development of modern analytical instruments capable of producing multidimensional arrays, various chemometric methods have gained widespread acceptance over the past decades. There is intensive research devoted to the development and the testing of multivariate algorithms applied to progressively more difficult chemical scenarios.
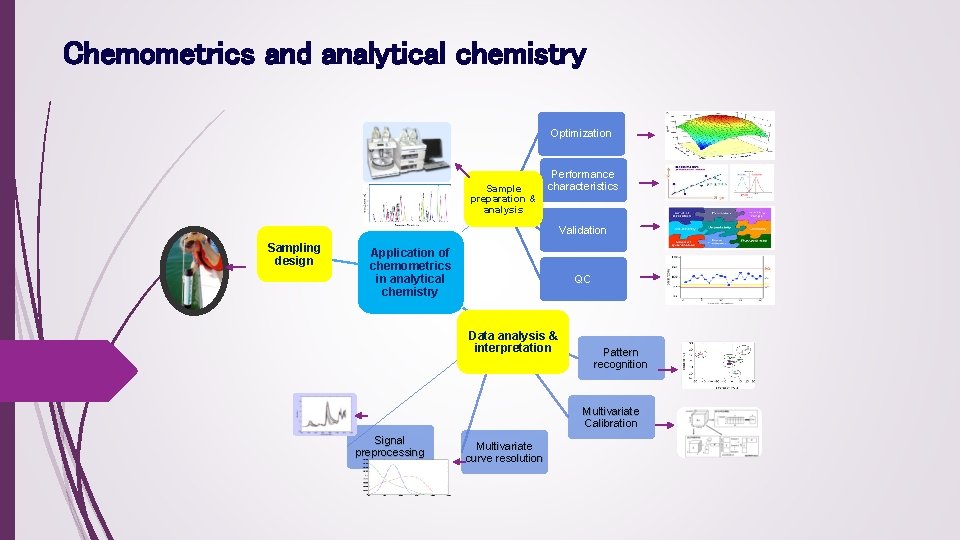
Chemometrics and analytical chemistry Optimization Performance characteristics Sample preparation & analysis Validation Sampling design Application of chemometrics in analytical chemistry QC Data analysis & interpretation Pattern recognition Multivariate Calibration Signal preprocessing Multivariate curve resolution

Main research activities Ø Ø Ø Multivariate analysis and resolution Design of experiment and optimization Multi-set and multiway analysis Classification Signal pre-processing Ø Environmental analysis (River water, drinking water, wastewater, sediments, sludge, . . . ) Ø Drugs of use and abuse impurity monitoring Ø Bioanalysis (In TDM, Urine, plasma, blood sample) Ø Food analysis( Oils, nuts, . . . ) Ø Metabolomics
5 Drugs of abuse profiling Ice profiling Slimming products
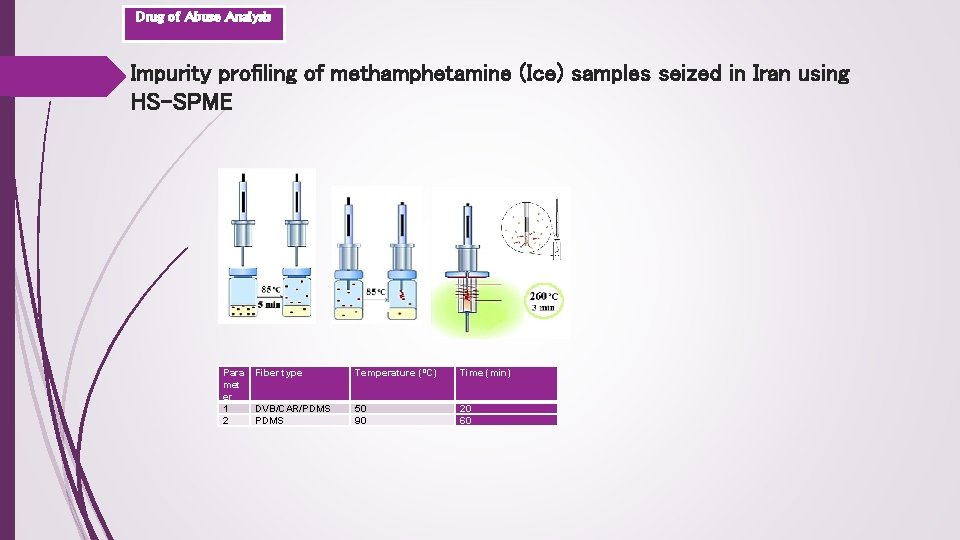
Drug of Abuse Analysis Impurity profiling of methamphetamine (Ice) samples seized in Iran using HS-SPME Para met er 1 2 Fiber type Temperature (ºC) Time (min) DVB/CAR/PDMS 50 90 20 60
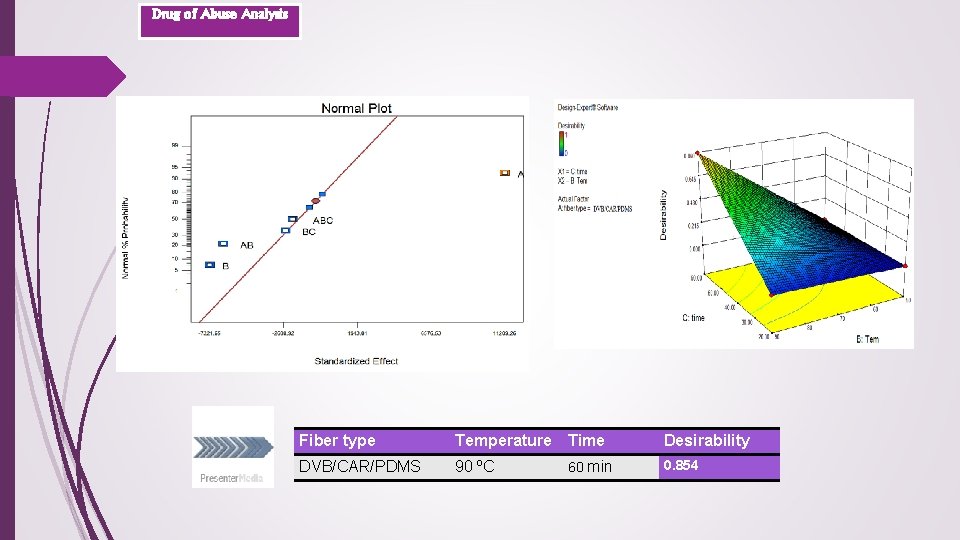
Drug of Abuse Analysis Fiber type Temperature Time Desirability DVB/CAR/PDMS 90 ºC 60 min 0. 854

Drug of Abuse Analysis GC/MS analysis of Ice HS-SPME Solvent extraction MCR/ALS modeling

9 Multiway assisted chromatographic data analysis Hyphenated chromatographic techniques such as GC-MS, LC-MS and HPLC-DAD are intensively used for analysis of environmental, drugs, biological, food products and etc. samples. Application of different high-order calibration methods for handling multi-dimensional chromatographic data is considered as an efficient complementary strategy to extract qualitative and quantitative information from complex analytical data.

10 Second- and high-order algorithms DTLD (direct trilinear decomposition) ( e pl m z) ATLD (alternating trilinear decomposition) sa PARAFAC(parallel factor analysis) APTLD (alternating penalty trilinear decomposition Wavelength ( ) SWATLD (self-weighted alternating trilinear decomposition) PARAFAC 2 MCR-ALS (multivariate curve resolution-alternating least square) BLLS( blinear least square) U-PLS/RBL(unfolded-partial least square-residual bilnearization) N-PLS/RBL(multiway partial least square-residual bilnearization) Trilinear least-squares (TLLS) with residual trilinearization (RTL) Alternating penalty quadrilinear decomposition (APQLD) x Retention time (y) GRAM (generalized rank annihilation method = N Ù Ù Ù å n=1 X Y Z+ E ijk ni nj nk

11 Calibration using multi-way data: Advantages Minimization or elimination of suffering sample preparation steps Multi-target quantifications Calibration in the presence of unknown interferences Simple or clean up-free sample preparation strategies Rapid chromatographic trends (cost and environmental impact) Increased selectivity and sensitivity
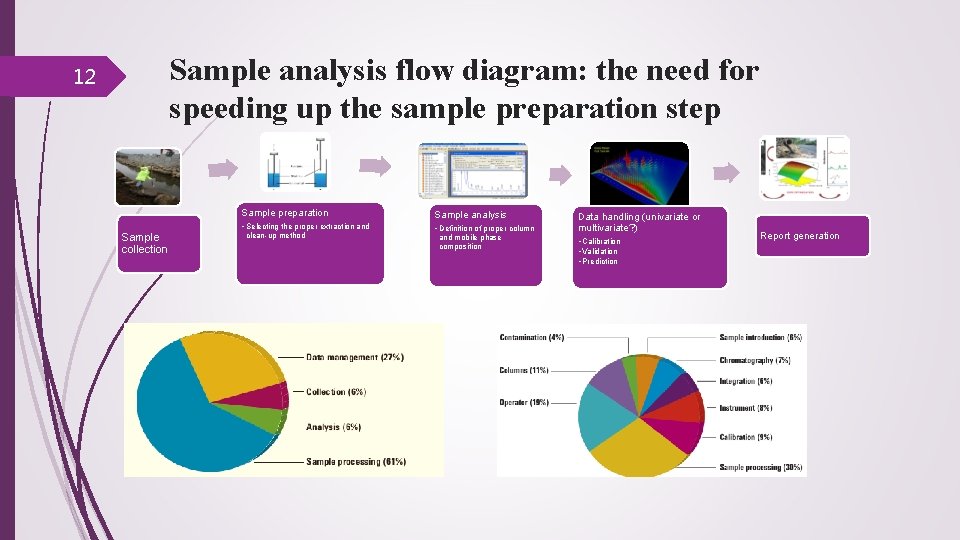
Sample analysis flow diagram: the need for speeding up the sample preparation step 12 Sample collection Sample preparation Sample analysis • Selecting the proper extraction and • Definition of proper column clean-up method and mobile phase composition Data handling (univariate or multivariate? ) • Calibration • Validation • Prediction Report generation

13 Calibration using multi-way data: Pitfalls Challenging backgrounds (A non-stable or uniform background offset) Retention time shifts of the chromatograms (non-linear, low -signal relative to matrix constituents) Complex co-eluting patterns Two or three dimensional data dividing before multi-way modeling Low intensity peaks Peak splitting Different matrix patterns of the sample class Matrix effect (first how to check we have matrix effect? Then, the calibration strategy) Similarity of spectral profiles Evaluating the recovery of extraction (mixing up with recovery of the algorithm) Maintenance of second-order advantage

14 Background drift correction (two-way) Curve fitting estimation and subtraction of background signal (local or global) Eluent background spectrum (EBS) Multidimensional extension of the spline-based approach, asymmetric least squares (ALS) Orthogonal spectral signal projection (OSSP) PCA and SIMPLISMA And many others Modeling the background variations by excessive (one or more factors) Estimating background contribution using PARAFAC and ATLD, when no baseline shape change among all samples Estimating background contribution using MCR/ALS and PARAFAC 2, when there is baseline shape change among all samples

15 Chromatographic peak alignment methods Methods based on maximum correlation (or minimum distance) Correlation optimized warping (COW) Dynamic time warping (DTW) Parametric time warping (PTW) Interval correlation optimized shifting (ICOShift) Peak alignment by means of genetic algorithm (PAGA) And many others Methods based on matrix of data per sample Rank alignment (rank minimization) Iterative target factor analysis coupled to COW(ITTFA-COW) Bi-dimensional alignment using PARAFAC MCR/ALS-COW-PARAFAC or MCR/ALS Abstract subspace difference alignment (ASSD)
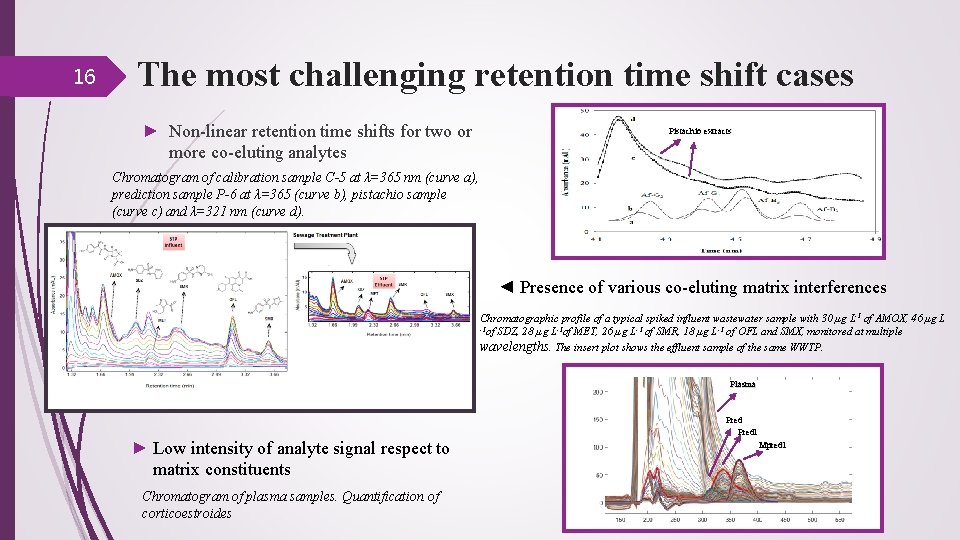
16 The most challenging retention time shift cases ► Non-linear retention time shifts for two or more co-eluting analytes Pistachio extracts Chromatogram of calibration sample C-5 at λ=365 nm (curve a), prediction sample P-6 at λ=365 (curve b), pistachio sample (curve c) and λ=321 nm (curve d). ◄ Presence of various co-eluting matrix interferences Chromatographic profile of a typical spiked influent wastewater sample with 30 µg L-1 of AMOX, 46 µg L -1 of SDZ, 28 µg L-1 of MET, 26 µg L-1 of SMR, 18 µg L-1 of OFL and SMX, monitored at multiple wavelengths. The insert plot shows the effluent sample of the same WWTP. Plasma ► Low intensity of analyte signal respect to matrix constituents Chromatogram of plasma samples. Quantification of corticoestroides Predl Mpredl
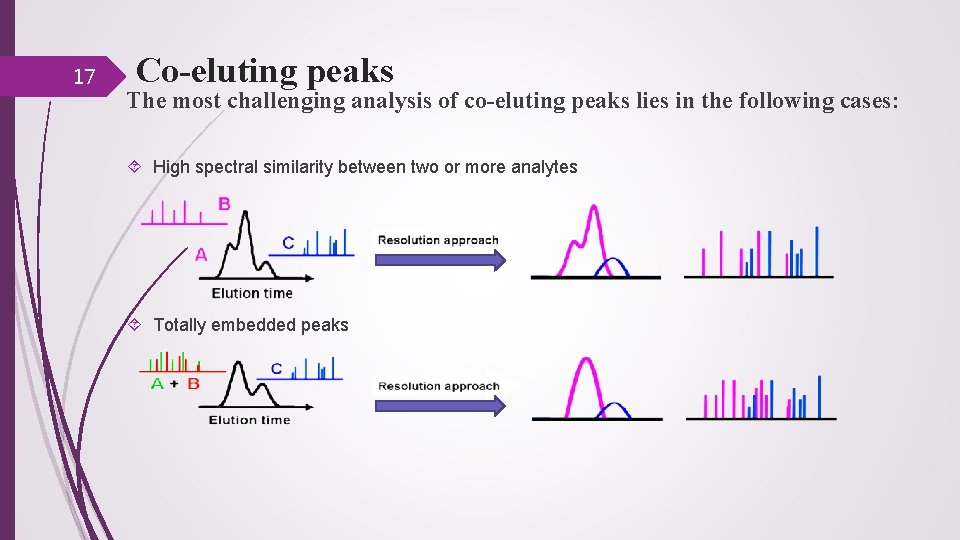
17 Co-eluting peaks The most challenging analysis of co-eluting peaks lies in the following cases: High spectral similarity between two or more analytes Totally embedded peaks

18 Handling high spectral similarity Single target modeling Multitarget modeling • Practical considerations: Developing a mobile phase method (longer chromatographic runs) which fully separated analytes. • Individually multiway modeling of chromatographic section containing the target of interest • Needs longer chromatographic runs, and thus consuming more solvent • MCR/ALS modeling. First row-wise augmentation with different detector and then column-wise augmentation for different samples. • Needs a second detector with a selective response to each analyte and it must be synchronized with the first instrument • The temporal misalignment should be corrected. • MCR/ALS modeling. The data matrices should be augmented in spectral mode R. L. Pérez, G. M. Escandar, Anal. Chim Acta 835(2014)19– 28. (S. Navea, R. Tauler, A. de Juan, Anal. Chem, 78 (2006) 4768 -4778) (Licarion Pinto, et al, Anal. Chim Acta 902 (2016) 59– 69)
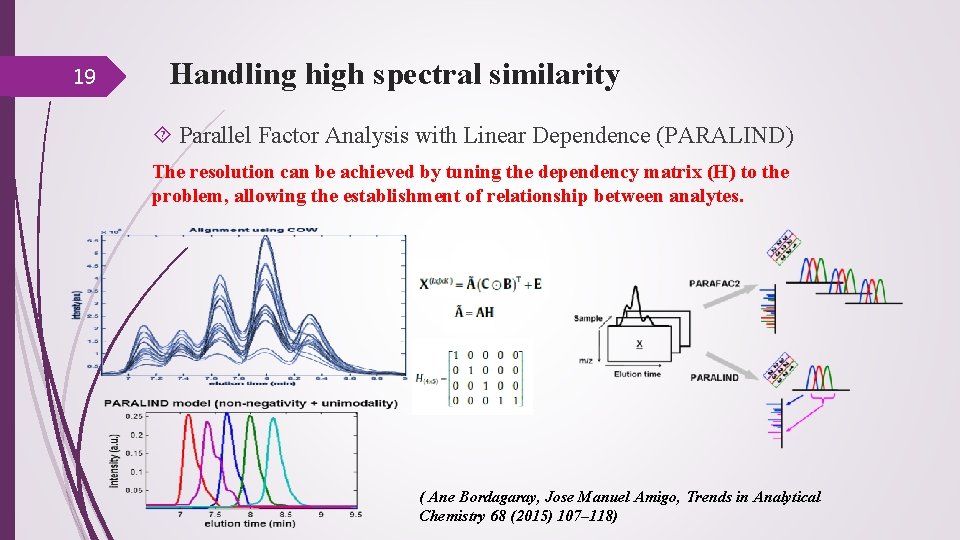
19 Handling high spectral similarity Parallel Factor Analysis with Linear Dependence (PARALIND) The resolution can be achieved by tuning the dependency matrix (H) to the problem, allowing the establishment of relationship between analytes. ( Ane Bordagaray, Jose Manuel Amigo, Trends in Analytical Chemistry 68 (2015) 107– 118)

Analysis of sunscreen agents

Quantification of sunscreen agents as emerging pollutants from suncare formulations to environmental samples and human body: Priori vs. Posteriori Analysis Chemometric tools assisted quantification of selected sunscreen agents and parabens in suncare formulations in the presence of interferences using HPLC-DAD Chro. MATHographic strategies for investigating the occurrence of UV filters as high risk pollutants in environmental water samples in Iran Determination of organic UV filter compounds in sediment samples using matrix solid phase dispersion and high performance liquid chromatography-diode array detection Chemometrics assisted dispersive liquid–liquid microextraction for quantification of seven UV filters in urine samples by HPLC-DAD
Cosmetic Analysis MBC BZ 3 OCR Analytes MP PP EDP BDM
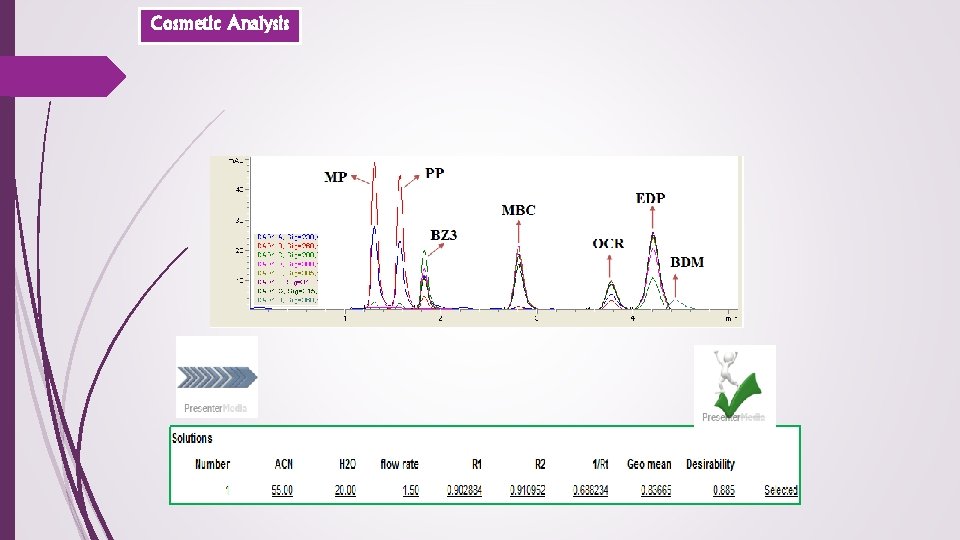
Cosmetic Analysis
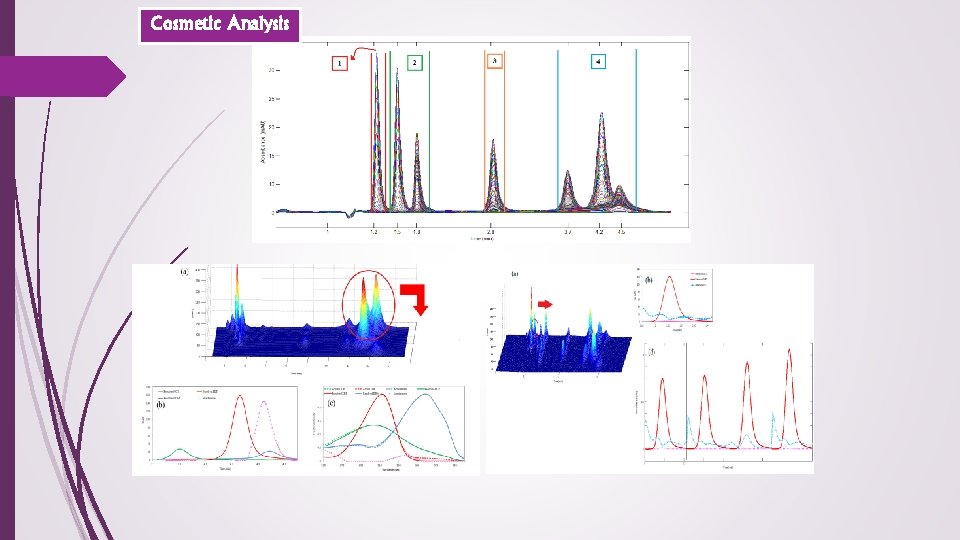
Cosmetic Analysis
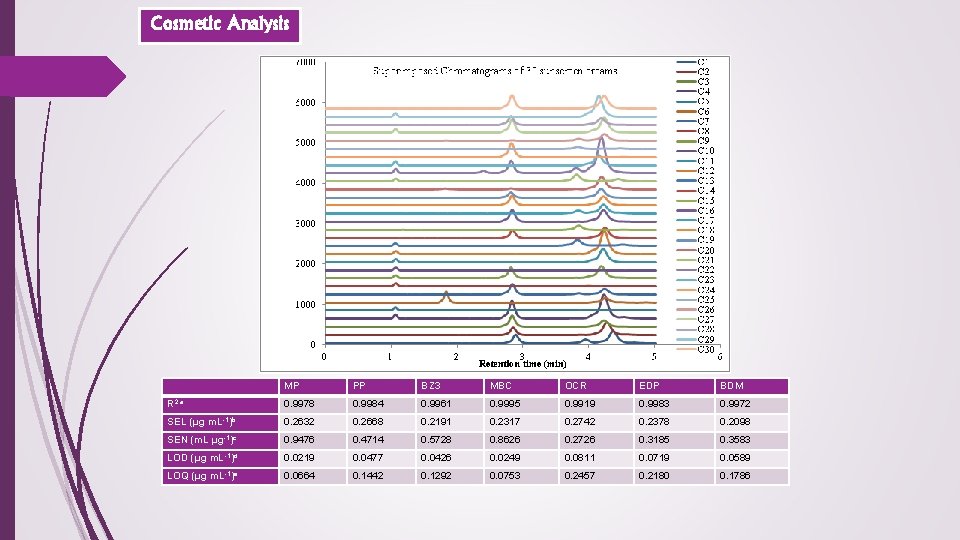
Cosmetic Analysis MP PP BZ 3 MBC OCR EDP BDM R 2 a 0. 9978 0. 9984 0. 9961 0. 9995 0. 9919 0. 9983 0. 9972 SEL (μg m. L-1)b 0. 2632 0. 2668 0. 2191 0. 2317 0. 2742 0. 2378 0. 2098 SEN (m. L μg-1)c 0. 9476 0. 4714 0. 5728 0. 8626 0. 2726 0. 3185 0. 3583 LOD (μg m. L-1)d 0. 0219 0. 0477 0. 0426 0. 0249 0. 0811 0. 0719 0. 0589 LOQ (μg m. L-1)e 0. 0664 0. 1442 0. 1292 0. 0753 0. 2457 0. 2180 0. 1786
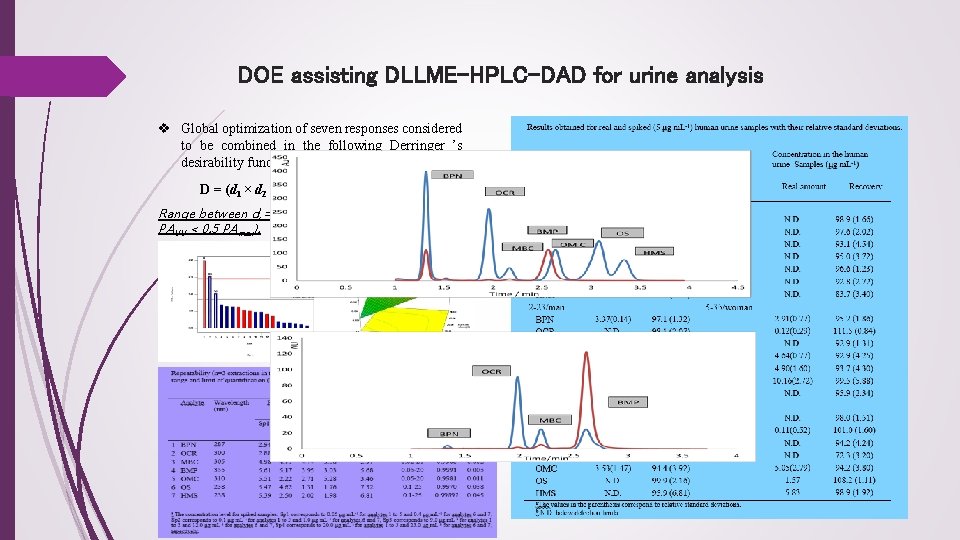
DOE assisting DLLME-HPLC-DAD for urine analysis v Global optimization of seven responses considered to be combined in the following Derringer ’s desirability function: D = (d 1 × d 2 × d 3 × d 4 × d 5 × d 6 × d 7)1/7 Range between di =1 (for PAUV=PAmax) and di =0 (for PAUV < 0. 5 PAmax).

27

Environmental Analysis by multiway mathods: With no/the simplest method of sample preparation
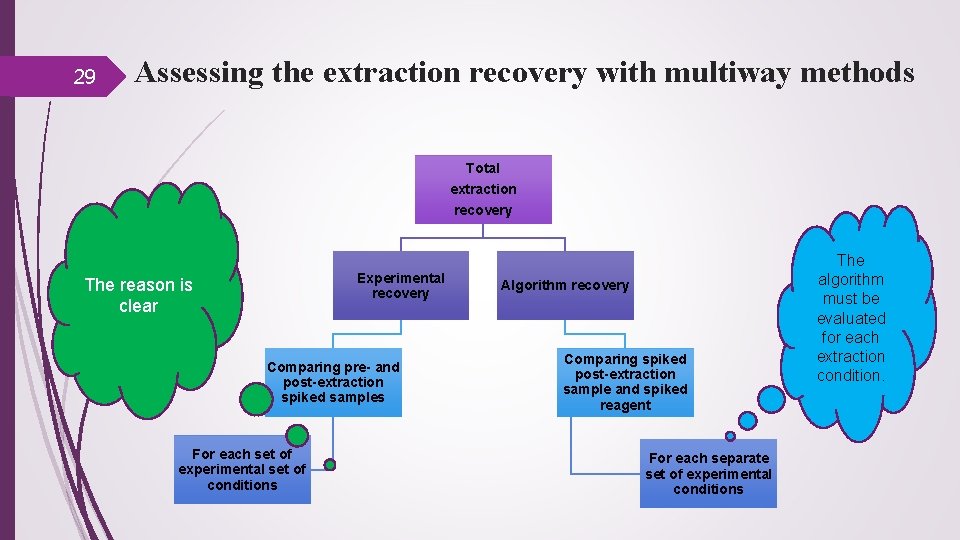
29 Assessing the extraction recovery with multiway methods Total extraction recovery Experimental recovery The reason is clear Comparing pre- and post-extraction spiked samples For each set of experimental set of conditions Algorithm recovery Comparing spiked post-extraction sample and spiked reagent For each separate set of experimental conditions The algorithm must be evaluated for each extraction condition.

30 The effect of co-extracted components The model (algorithm) recovery must be evaluated for each individual (different) extraction step The algorithm recovery may be highly dependent on the extraction procedure i. e. on the co-extracted matrix constituents ( not necessary co-eluted peaks) Example: Extraction A extracts: ANALYTE (X%), and B, C and D (as interferents) Extraction B extracts: ANALYTE (Y%), and C, D and E (as interferents) Co-extracted components may change Instrumental signal Algorithm efficiency

31 So! Although exploiting the second-order advantage can be very useful for recovery studies, but we have to consider all previously mentioned problematic effects of co-extracted matrix constituents such as: high intensity matrix peaks, retention time shift, matrix effect and so on. This is specially true for complex samples with a wide variety of matrix constituents !
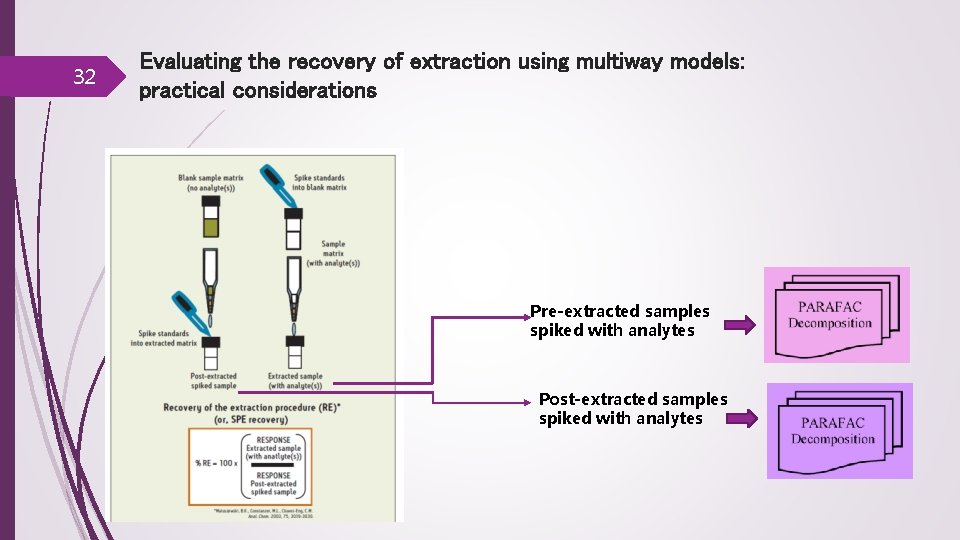
32 Evaluating the recovery of extraction using multiway models: practical considerations Pre-extracted samples spiked with analytes Post-extracted samples spiked with analytes
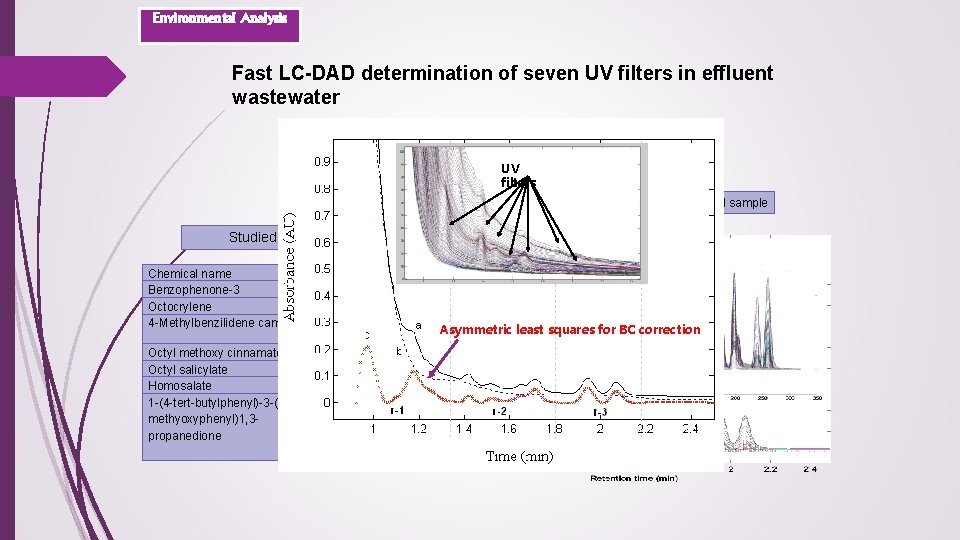
Environmental Analysis Fast LC-DAD determination of seven UV filters in effluent wastewater UV filters Effeluent wastewater sample Standard sample Studied UV filters Chemical name Benzophenone-3 Octocrylene 4 -Methylbenzilidene camphor Trade name Eusolex® 4360 Eusolex® OCR Eusolex® 6300 Octyl methoxy cinnamate Octyl salicylate Homosalate 1 -(4 -tert-butylphenyl)-3 -(4 methyoxyphenyl)1, 3 propanedione Eusolex® 2292 Eusolex® OS Eusolex® HMS Eusolex® 9020 Asymmetric least squares for BC correction
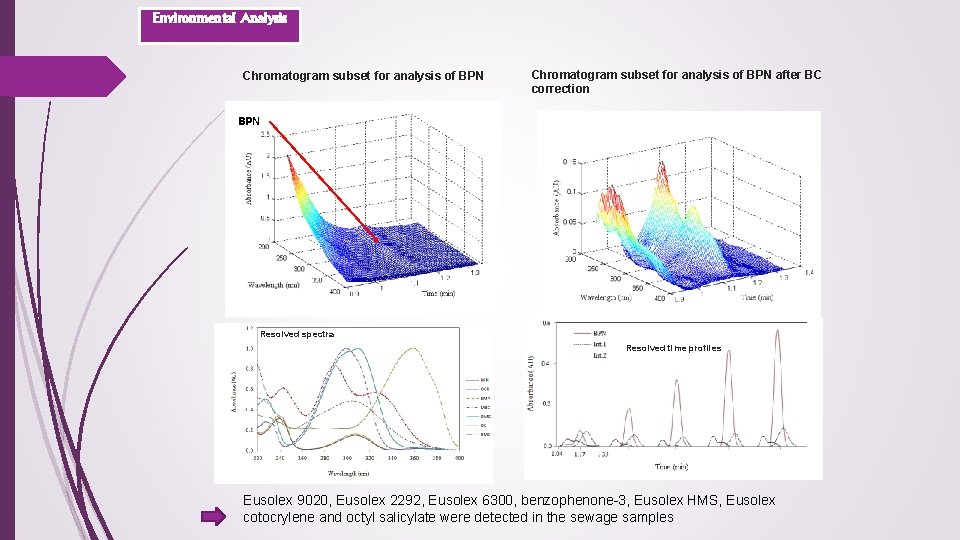
Environmental Analysis Chromatogram subset for analysis of BPN after BC correction BPN Resolved spectra Resolved time profiles Eusolex 9020, Eusolex 2292, Eusolex 6300, benzophenone-3, Eusolex HMS, Eusolex cotocrylene and octyl salicylate were detected in the sewage samples

Environmental Analysis Investigating the occurrence of UV filters as high risk pollutants in environmental water samples in Zarjub river (Rasht) Eusolex 9020, Eusolex 2292, Eusolex 6300 and benzophenone-3 were determined in Zarjub River water. Results of the analyses of the Ionian Sea showed the presence of benzophenone-3 as the only pollutant in common.
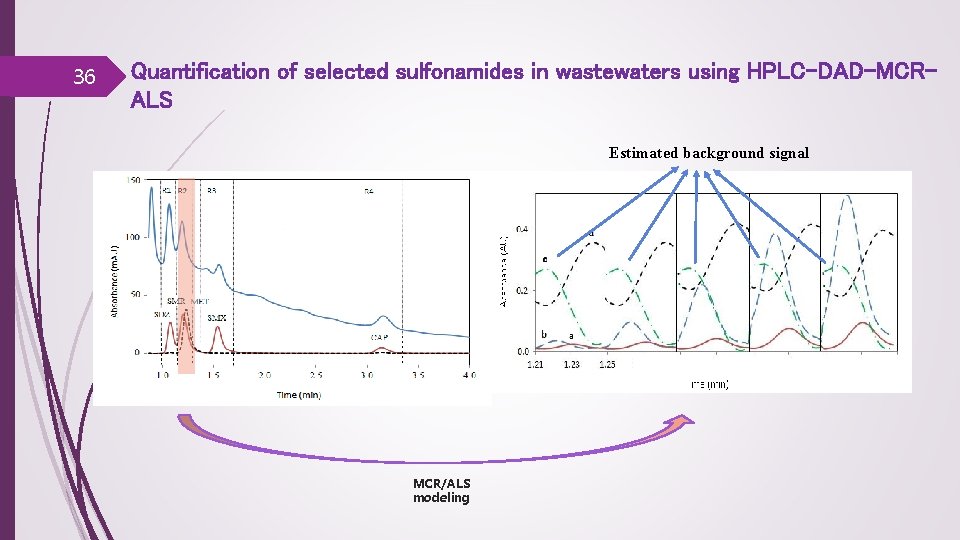
36 Quantification of selected sulfonamides in wastewaters using HPLC-DAD-MCRALS Estimated background signal MCR/ALS modeling
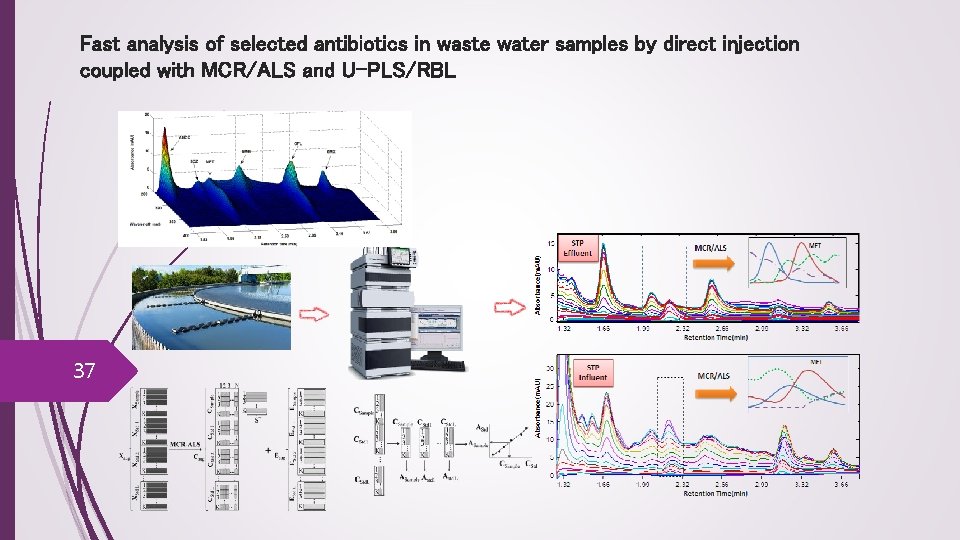
Fast analysis of selected antibiotics in waste water samples by direct injection coupled with MCR/ALS and U-PLS/RBL 37
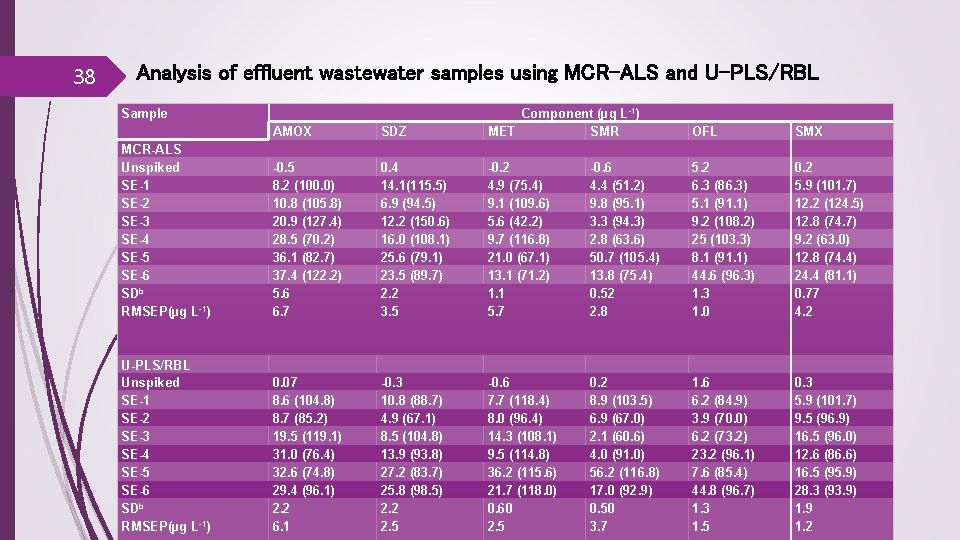
38 Analysis of effluent wastewater samples using MCR-ALS and U-PLS/RBL Sample MCR-ALS Unspiked SE-1 SE-2 SE-3 SE-4 SE-5 SE-6 SDb RMSEP(µg L-1) U-PLS/RBL Unspiked SE-1 SE-2 SE-3 SE-4 SE-5 SE-6 SDb RMSEP(µg L-1) AMOX -0. 5 8. 2 (100. 0) 10. 8 (105. 8) 20. 9 (127. 4) 28. 5 (70. 2) 36. 1 (82. 7) 37. 4 (122. 2) 5. 6 6. 7 0. 07 8. 6 (104. 8) 8. 7 (85. 2) 19. 5 (119. 1) 31. 0 (76. 4) 32. 6 (74. 8) 29. 4 (96. 1) 2. 2 6. 1 SDZ 0. 4 14. 1(115. 5) 6. 9 (94. 5) 12. 2 (150. 6) 16. 0 (108. 1) 25. 6 (79. 1) 23. 5 (89. 7) 2. 2 3. 5 -0. 3 10. 8 (88. 7) 4. 9 (67. 1) 8. 5 (104. 8) 13. 9 (93. 8) 27. 2 (83. 7) 25. 8 (98. 5) 2. 2 2. 5 Component (µg L-1) MET SMR -0. 2 -0. 6 4. 9 (75. 4) 4. 4 (51. 2) 9. 1 (109. 6) 9. 8 (95. 1) 5. 6 (42. 2) 3. 3 (94. 3) 9. 7 (116. 8) 2. 8 (63. 6) 21. 0 (67. 1) 50. 7 (105. 4) 13. 1 (71. 2) 13. 8 (75. 4) 1. 1 0. 52 5. 7 2. 8 -0. 6 7. 7 (118. 4) 8. 0 (96. 4) 14. 3 (108. 1) 9. 5 (114. 8) 36. 2 (115. 6) 21. 7 (118. 0) 0. 60 2. 5 0. 2 8. 9 (103. 5) 6. 9 (67. 0) 2. 1 (60. 6) 4. 0 (91. 0) 56. 2 (116. 8) 17. 0 (92. 9) 0. 50 3. 7 OFL 5. 2 6. 3 (86. 3) 5. 1 (91. 1) 9. 2 (108. 2) 25 (103. 3) 8. 1 (91. 1) 44. 6 (96. 3) 1. 3 1. 0 SMX 0. 2 5. 9 (101. 7) 12. 2 (124. 5) 12. 8 (74. 7) 9. 2 (63. 0) 12. 8 (74. 4) 24. 4 (81. 1) 0. 77 4. 2 1. 6 6. 2 (84. 9) 3. 9 (70. 0) 6. 2 (73. 2) 23. 2 (96. 1) 7. 6 (85. 4) 44. 8 (96. 7) 1. 3 1. 5 0. 3 5. 9 (101. 7) 9. 5 (96. 9) 16. 5 (96. 0) 12. 6 (86. 6) 16. 5 (95. 9) 28. 3 (93. 9) 1. 9 1. 2
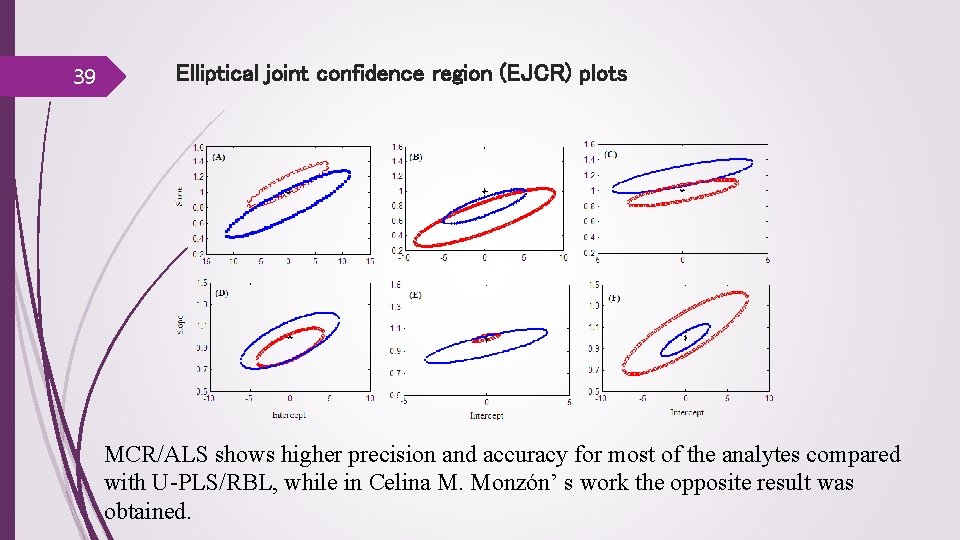
39 Elliptical joint confidence region (EJCR) plots MCR/ALS shows higher precision and accuracy for most of the analytes compared with U-PLS/RBL, while in Celina M. Monzón’ s work the opposite result was obtained.

40 Performances of U-PLS/RBL compared with MCR/ALS The observations indicated that both methods provide comparable results MCR/ALS. Good initial estimates of chromatographs and spectra are needed U-PLS/RBL benefits from some important advantages. In fact, it is a rather straight forward technique, has simpler and faster program operations, it is independent from initial estimate (guess) of the component spectra. U-PLS/RBL. It is not possible to give a clear physical interpretation of the system unless there is only one unexpected component. The success of these modeling methods depends on availability of a sufficiently large and representative set of calibration samples. Although U-PLS/RBL, as a latent based structure modeling, did not require a data set with perfect trilinear structure, we obtained more satisfactory results by performing retention time alignment

41

Environmental analysis: Chemometrics assisting nanomaterials based sample preparation methodologies
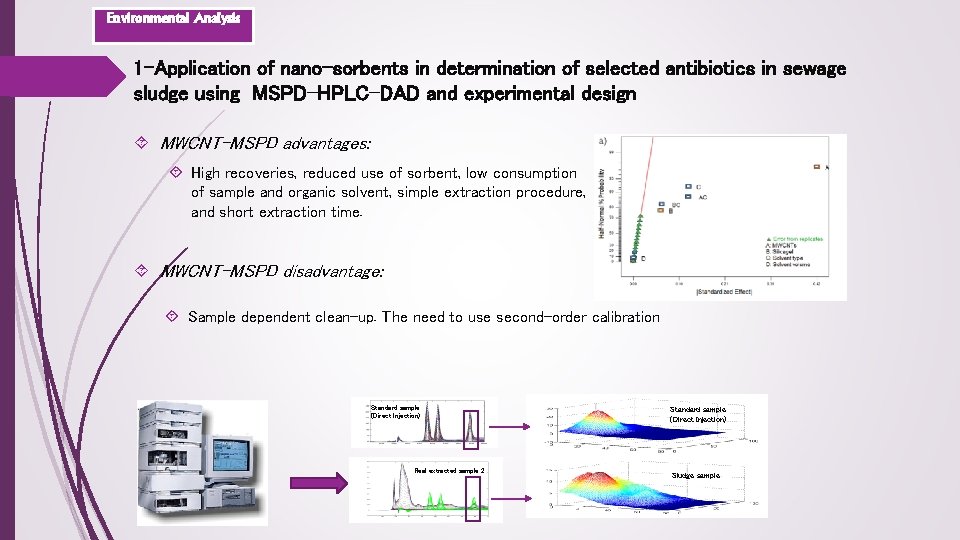
Environmental Analysis 1 -Application of nano-sorbents in determination of selected antibiotics in sewage sludge using MSPD-HPLC-DAD and experimental design MWCNT-MSPD advantages: High recoveries, reduced use of sorbent, low consumption of sample and organic solvent, simple extraction procedure, and short extraction time. MWCNT-MSPD disadvantage: Sample dependent clean-up. The need to use second-order calibration Standard sample (Direct Injection) Real extracted sample 2 Standard sample (Direct Injection) Sludge sample
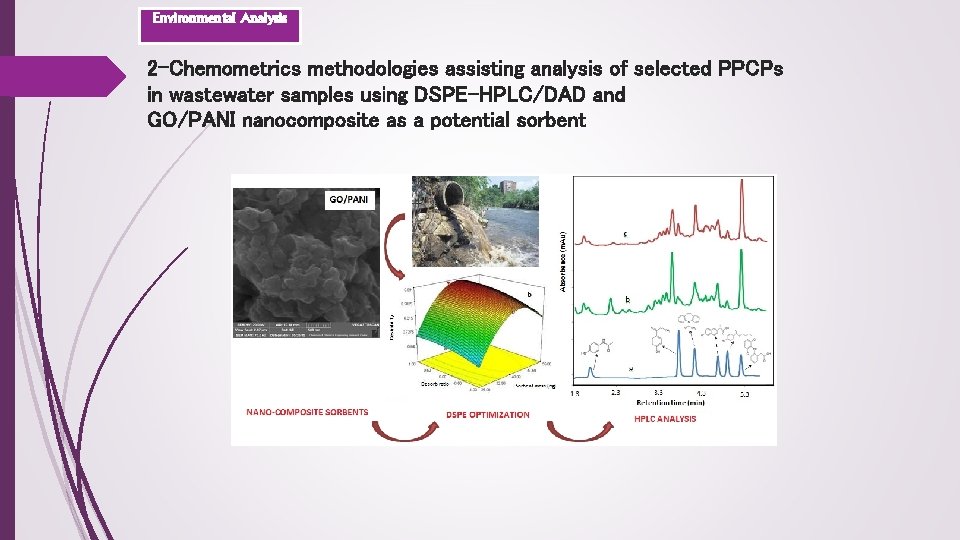
Environmental Analysis 2 -Chemometrics methodologies assisting analysis of selected PPCPs in wastewater samples using DSPE-HPLC/DAD and GO/PANI nanocomposite as a potential sorbent
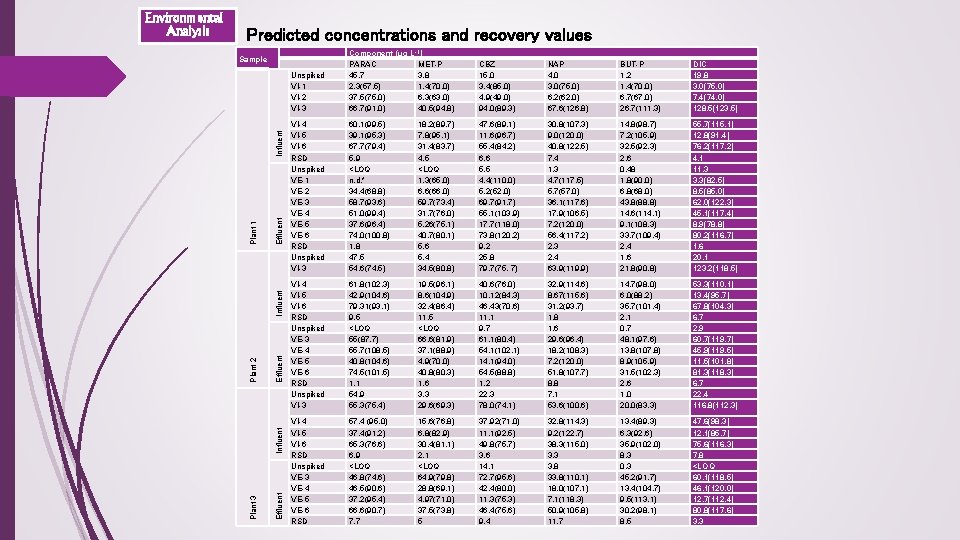
Predicted concentrations and recovery values Effluent Influent Plant 2 Influent Plant 1 Influent Sample Plant 3 Environmental Analysis Unspiked VI-1 VI-2 VI-3 Component (µg L-1) PARAC MET-P 45. 7 3. 8 2. 3(57. 5) 1. 4(70. 0) 37. 5(75. 0) 6. 3(63. 0) 66. 7(91. 0) 40. 5(94. 8) CBZ 15. 0 3. 4(85. 0) 4. 9(49. 0) 94. 0(89. 3) NAP 4. 0 3. 0(75. 0) 6. 2(62. 0) 67. 6(126. 8) BUT-P 1. 2 1. 4(70. 0) 6. 7(67. 0) 26. 7(111. 3) DIC 19. 8 3. 0(75. 0) 7. 4(74. 0) 128. 5(123. 5) VI-4 VI-5 VI-6 RSD Unspiked VE-1 VE-2 VE-3 VE-4 VE-5 VE-6 RSD Unspiked VI-3 60. 1(99. 5) 39. 1(95. 3) 67. 7(79. 4) 5. 9 <LOQ n. d. * 34. 4(68. 8) 58. 7(93. 6) 51. 0(99. 4) 37. 6(96. 4) 74. 0(100. 8) 1. 8 47. 5 54. 6(74. 5) 18. 2(89. 7) 7. 8(95. 1) 31. 4(83. 7) 4. 5 <LOQ 1. 3(65. 0) 6. 6(66. 0) 59. 7(73. 4) 31. 7(76. 0) 5. 26(75. 1) 40. 7(80. 1) 5. 6 5. 4 34. 5(80. 8) 47. 6(89. 1) 11. 6(96. 7) 55. 4(84. 2) 6. 6 5. 5 4. 4(110. 0) 5. 2(52. 0) 69. 7(91. 7) 55. 1(103. 9) 17. 7(118. 0) 73. 8(120. 2) 9. 2 25. 8 79. 7(75. 7) 30. 8(107. 3) 9. 0(120. 0) 40. 8(122. 5) 7. 4 1. 3 4. 7(117. 5) 5. 7(57. 0) 36. 1(117. 6) 17. 9(106. 5) 7. 2(120. 0) 56. 4(117. 2) 2. 3 2. 4 63. 9(119. 9) 14. 8(98. 7) 7. 2(105. 9) 32. 5(92. 3) 2. 6 0. 48 1. 8(90. 0) 6. 8(68. 0) 43. 8(88. 8) 14. 6(114. 1) 9. 1(108. 3) 33. 7(109. 4) 2. 4 1. 6 21. 8(90. 8) 55. 7(115. 1) 12. 8(91. 4) 76. 2(117. 2) 4. 1 11. 3 3. 3(82. 5) 8. 5(85. 0) 62. 0(122. 3) 45. 1(117. 4) 8. 9(78. 8) 80. 2(116. 7) 1. 6 20. 1 123. 2(118. 5) VI-4 VI-5 VI-6 RSD Unspiked VE-3 VE-4 VE-5 VE-6 RSD Unspiked VI-3 61. 8(102. 3) 42. 9(104. 6) 79. 31(93. 1) 9. 5 <LOQ 55(87. 7) 55. 7(108. 5) 40. 8(104. 6) 74. 5(101. 5) 1. 1 54. 9 55. 3(75. 4) 19. 5(96. 1) 8. 6(104. 9) 32. 4(86. 4) 11. 5 <LOQ 66. 6(81. 9) 37. 1(88. 9) 4. 9(70. 0) 40. 8(80. 3) 1. 6 3. 3 29. 6(69. 3) 40. 6(76. 0) 10. 12(84. 3) 46. 43(70. 6) 11. 1 9. 7 61. 1(80. 4) 54. 1(102. 1) 14. 1(94. 0) 54. 5(88. 8) 1. 2 22. 3 78. 0(74. 1) 32. 9(114. 6) 8. 67(115. 6) 31. 2(93. 7) 1. 8 1. 6 29. 6(96. 4) 18. 2(108. 3) 7. 2(120. 0) 51. 8(107. 7) 8. 8 7. 1 53. 6(100. 6) 14. 7(98. 0) 6. 0(88. 2) 35. 7(101. 4) 2. 1 0. 7 48. 1(97. 6) 13. 8(107. 8) 8. 9(105. 9) 31. 5(102. 3) 2. 6 1. 0 20. 0(83. 3) 53. 3(110. 1) 13. 4(95. 7) 67. 8(104. 3) 6. 7 2. 9 60. 7(119. 7) 45. 9(119. 5) 11. 5(101. 8) 81. 3(118. 3) 6. 7 22. 4 116. 8(112. 3) VI-4 VI-5 VI-6 RSD Unspiked VE-3 VE-4 VE-5 VE-6 RSD 57. 4 (95. 0) 37. 4(91. 2) 65. 3(76. 6) 6. 9 <LOQ 46. 8(74. 6) 46. 5(90. 6) 37. 2(95. 4) 66. 6(90. 7) 7. 7 15. 6(76. 8) 6. 8(82. 9) 30. 4(81. 1) 2. 1 <LOQ 64. 9(79. 8) 28. 8(69. 1) 4. 97(71. 0) 37. 5(73. 8) 5 37. 92(71. 0) 11. 1(92. 5) 49. 8(75. 7) 3. 6 14. 1 72. 7(95. 6) 42. 4(80. 0) 11. 3(75. 3) 46. 4(75. 6) 9. 4 32. 8(114. 3) 9. 2(122. 7) 38. 3(115. 0) 3. 3 3. 8 33. 8(110. 1) 18. 0(107. 1) 7. 1(118. 3) 50. 9(105. 8) 11. 7 13. 4(89. 3) 6. 3(92. 6) 35. 9(102. 0) 8. 3 0. 3 45. 2(91. 7) 13. 4(104. 7) 9. 5(113. 1) 30. 2(98. 1) 8. 5 47. 6(98. 3) 12. 1(85. 7) 75. 6(116. 3) 7. 8 <LOQ 60. 1(118. 5) 46. 1(120. 0) 12. 7(112. 4) 80. 8(117. 6) 3. 3
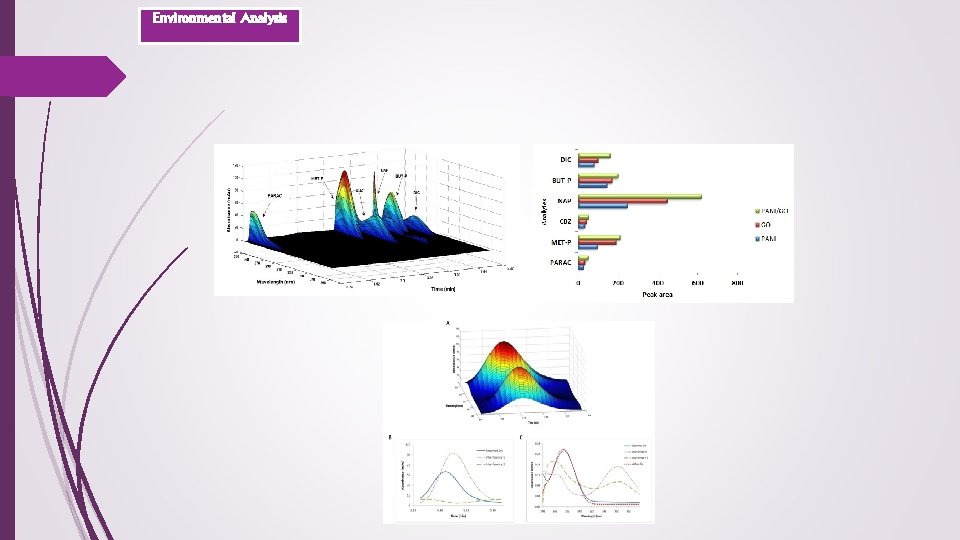
Environmental Analysis
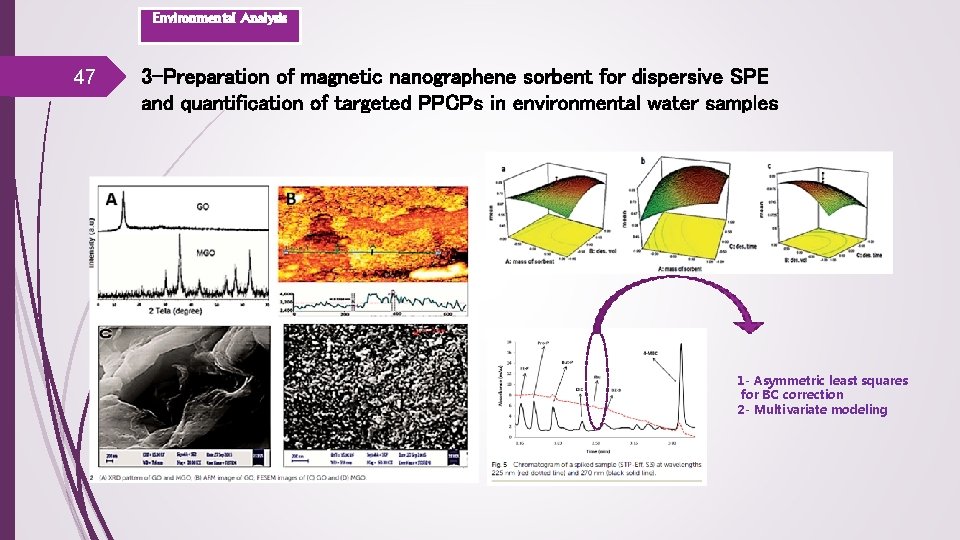
Environmental Analysis 47 3 -Preparation of magnetic nanographene sorbent for dispersive SPE and quantification of targeted PPCPs in environmental water samples 1 - Asymmetric least squares for BC correction 2 - Multivariate modeling

48

49

Solving Real Quantitative Bioanalysis Problems with Multiway Enhanced HPLC and Fluorescence Spectroscopy

51 Matrix effect and calibration strategy Instrumental matrix effect Signal enhancement or suppression Large signal shifts (shape and position) Extraction matrix effect The effect of matrix components in extraction recovery In case of any of the mentioned matrix effect standard addition strategy is the suitable calibration strategy. Can second-order advantage be exploited to simultaneously analyze the standard added samples with different interfering patterns?
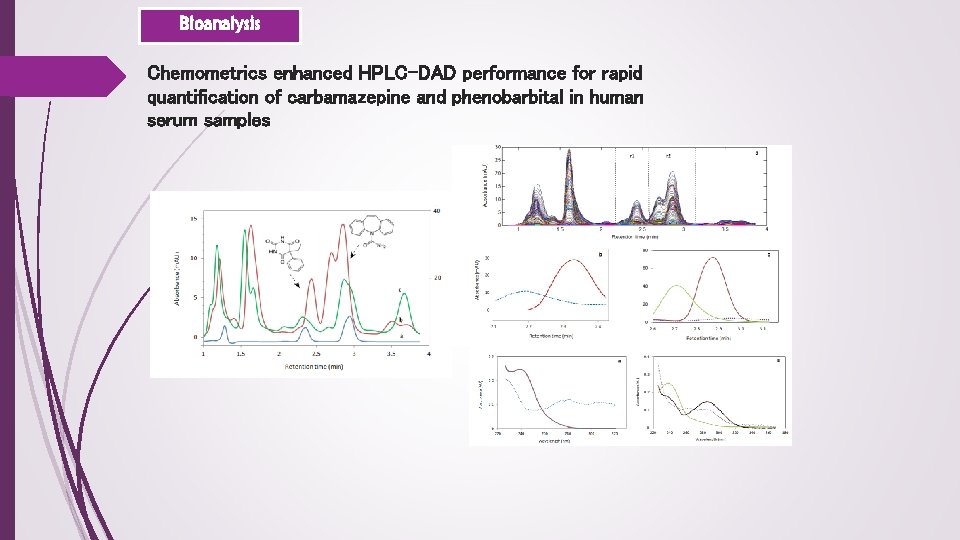
Bioanalysis Chemometrics enhanced HPLC-DAD performance for rapid quantification of carbamazepine and phenobarbital in human serum samples
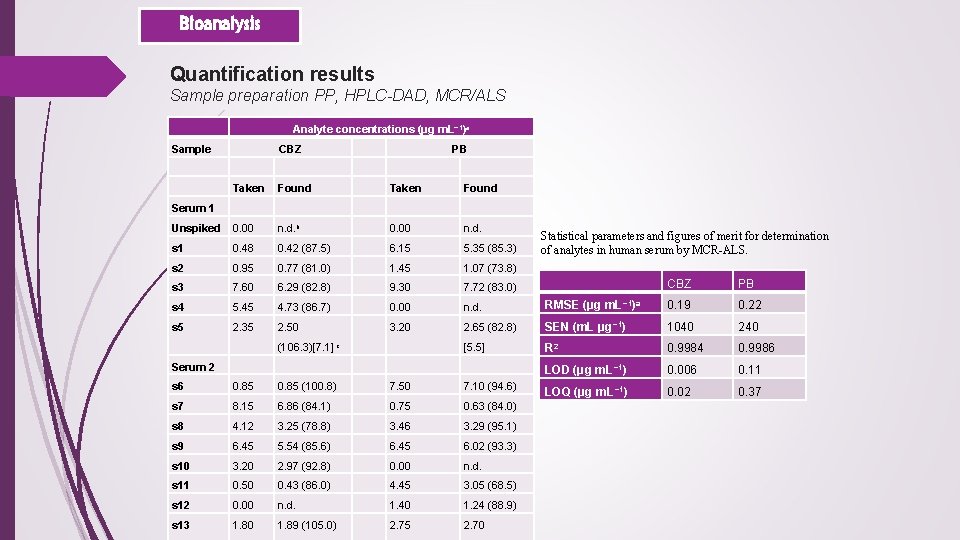
. Bioanalysis Quantification results Sample preparation PP, HPLC-DAD, MCR/ALS Analyte concentrations (μg m. L− 1)a Sample CBZ Taken Found Serum 1 Unspiked 0. 00 n. d. b s 1 0. 48 s 2 PB Taken Found 0. 00 n. d. 0. 42 (87. 5) 6. 15 5. 35 (85. 3) 0. 95 0. 77 (81. 0) 1. 45 1. 07 (73. 8) s 3 7. 60 6. 29 (82. 8) 9. 30 7. 72 (83. 0) s 4 5. 45 4. 73 (86. 7) 0. 00 s 5 2. 35 2. 50 3. 20 (106. 3)[7. 1] c Statistical parameters and figures of merit for determination of analytes in human serum by MCR-ALS. CBZ PB n. d. RMSE (μg m. L− 1)a 0. 19 0. 22 2. 65 (82. 8) SEN (m. L μg− 1) 1040 240 [5. 5] R 2 0. 9984 0. 9986 Serum 2 LOD (μg m. L− 1) 0. 006 0. 11 s 6 0. 85 (100. 8) 7. 50 7. 10 (94. 6) LOQ (μg m. L − 1) 0. 02 0. 37 s 7 8. 15 6. 86 (84. 1) 0. 75 0. 63 (84. 0) s 8 4. 12 3. 25 (78. 8) 3. 46 3. 29 (95. 1) s 9 6. 45 5. 54 (85. 6) 6. 45 6. 02 (93. 3) s 10 3. 20 2. 97 (92. 8) 0. 00 n. d. s 11 0. 50 0. 43 (86. 0) 4. 45 3. 05 (68. 5) s 12 0. 00 n. d. 1. 40 1. 24 (88. 9) s 13 1. 80 1. 89 (105. 0) 2. 75 2. 70
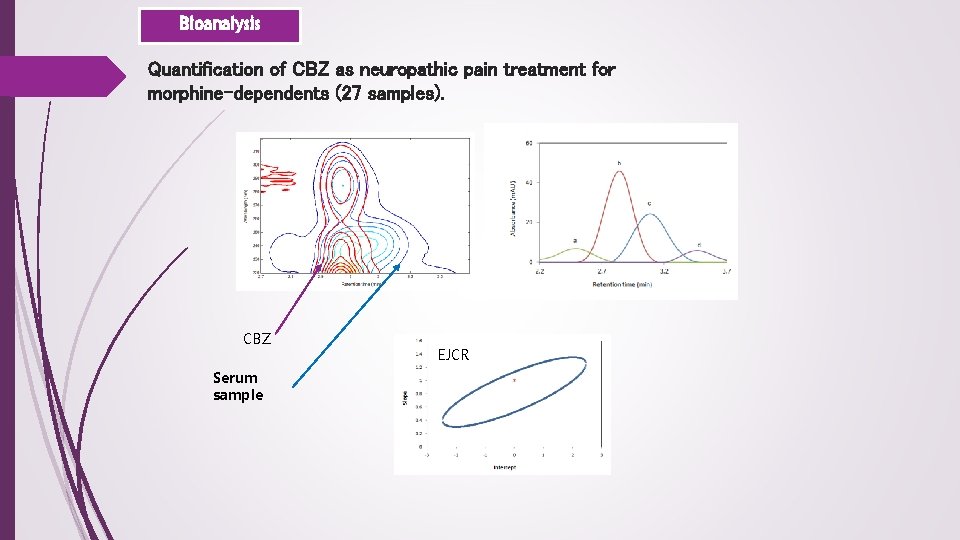
Bioanalysis Quantification of CBZ as neuropathic pain treatment for morphine-dependents (27 samples). CBZ Serum sample EJCR
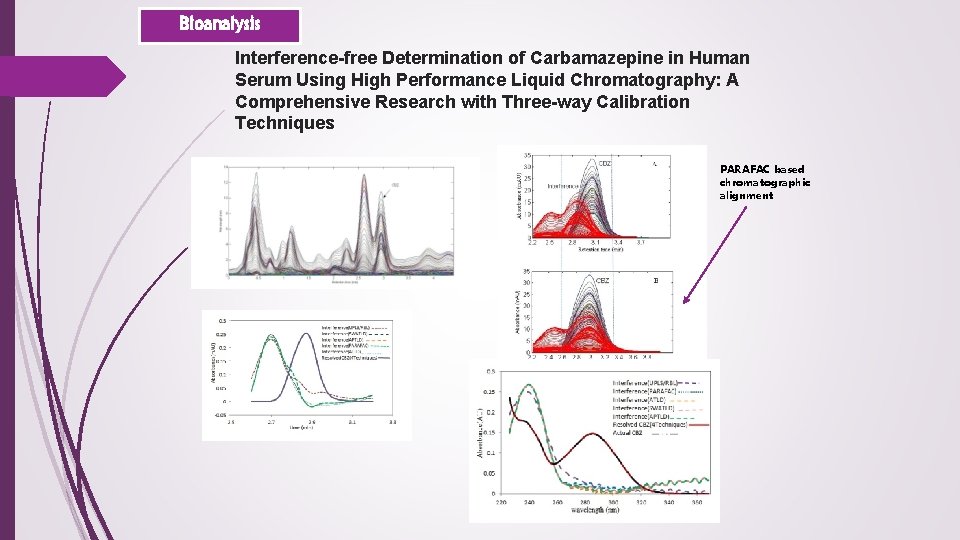
Bioanalysis Interference-free Determination of Carbamazepine in Human Serum Using High Performance Liquid Chromatography: A Comprehensive Research with Three-way Calibration Techniques PARAFAC based chromatographic alignment
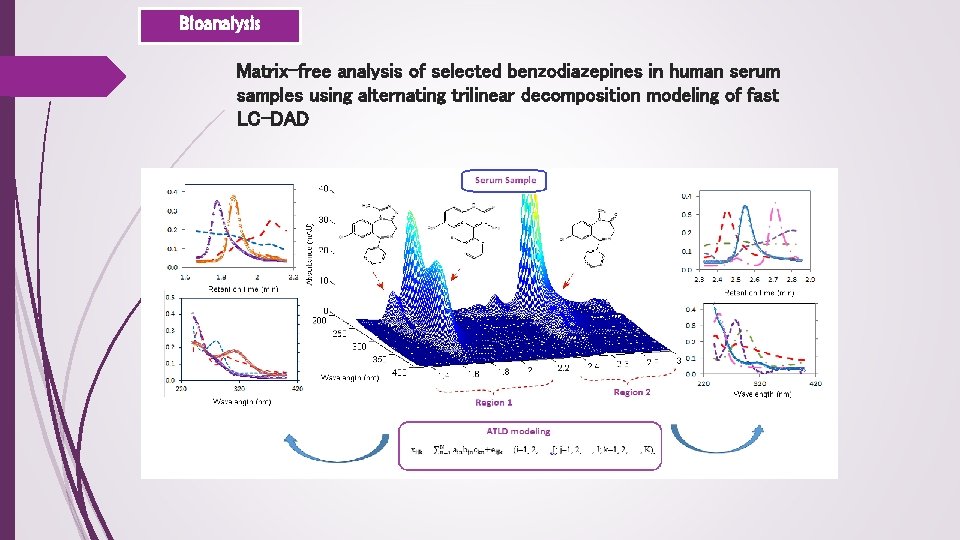
Bioanalysis Matrix-free analysis of selected benzodiazepines in human serum samples using alternating trilinear decomposition modeling of fast LC-DAD
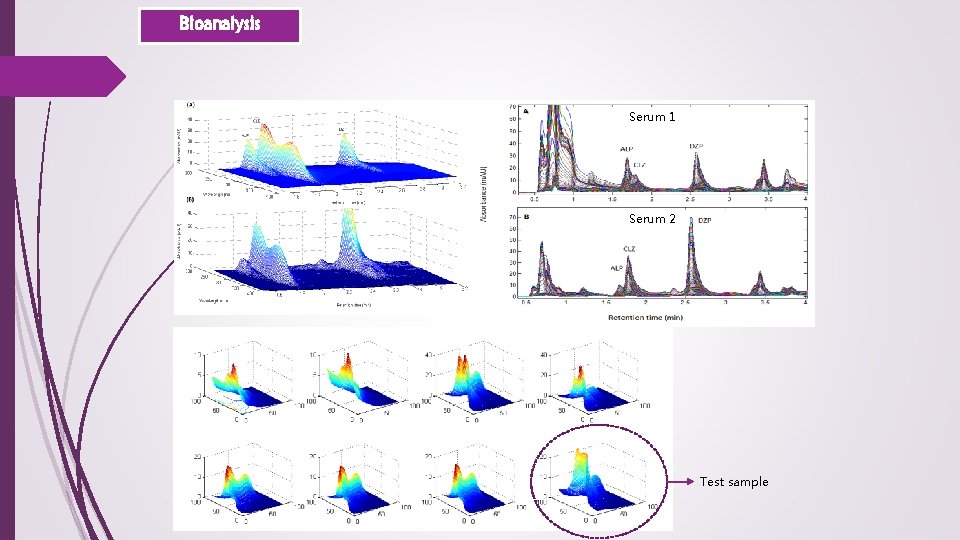
Bioanalysis Serum 1 Serum 2 Test sample
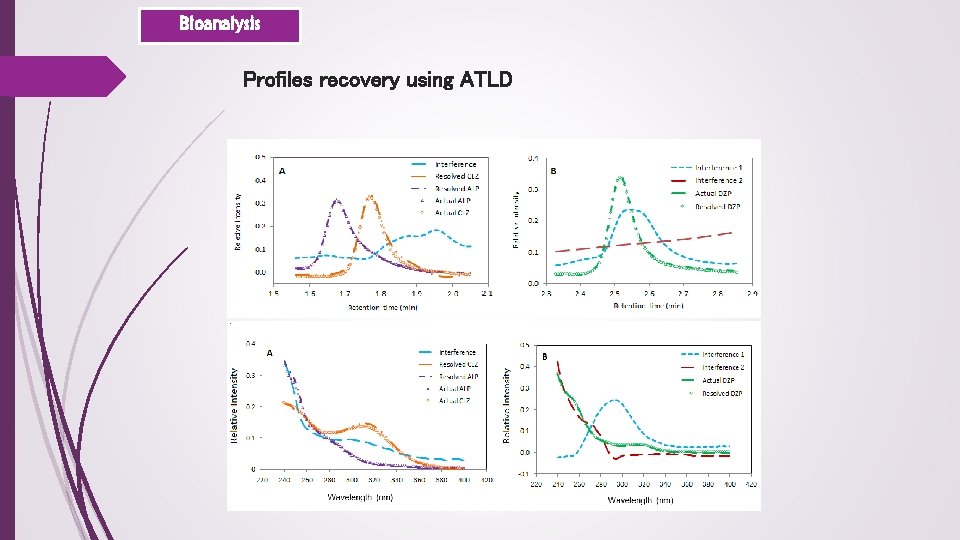
Bioanalysis Profiles recovery using ATLD
Bioanalysis Quantitative results Predicted concentration (µg m. L-1)a ALP CLZ DZP Prediction samples P 1 0. 445 (106. 9) 0. 153 (92. 1) 0. 218 (103. 3) P 2 3. 327 (99. 8) 2. 529 (94. 8) 0. 290 (92. 9) P 3 0. 172 (103. 4) 3. 550 (106. 5) 0. 515 (103. 3) P 4 0. 048 (107. 3) 1. 470 (107. 6) 2. 070 (103. 9) P 5 1. 012 (102. 4)[1. 8]b 0. 857 (102. 9)[2. 6] 3. 120 (89. 4)[2. 4] Spiked Samples Un-spiked n. dc n. d S 1 1. 809 (78. 9) 0. 235 (78. 5) 3. 943 (118. 3) S 2 0. 756 (95. 5) 1. 733 (75. 6) 0. 929 (92. 9) S 3 0. 819 (81. 9) 0. 018 (112. 9) 1. 282 (102. 5) S 4 2. 746 (85. 6) [4. 6] 1. 305 (92. 2) [3. 5] 1. 903 (99. 6) [3. 2] S 5*d 0. 017 (68. 1) 0. 302 (91. 7) 0. 019 (76. 8) S 6* 1. 152 (106. 3) 0. 090 (115. 1) 0. 391 (81. 1) S 7* 1. 325 (99. 4) 0. 882 (117. 6) 0. 280 (93. 3) S 8* 0. 768 (96. 9) [5. 7] 1. 987 (86. 7) [6. 8] 3. 107 (93. 2) [8. 1]
Bioanalysis Multivariate figures of merit ALP CLZ DZP RMSEP (μg m. L− 1) 0. 2627 0. 2228 0. 2351 SEN (m. L μg− 1) 39. 55 72. 54 21. 33 R 2 0. 9989 0. 9957 0. 9994 LOD (μg m. L− 1) 0. 015 0. 008 0. 027 LOQ (μg m. L − 1) 0. 044 0. 024 0. 081

61
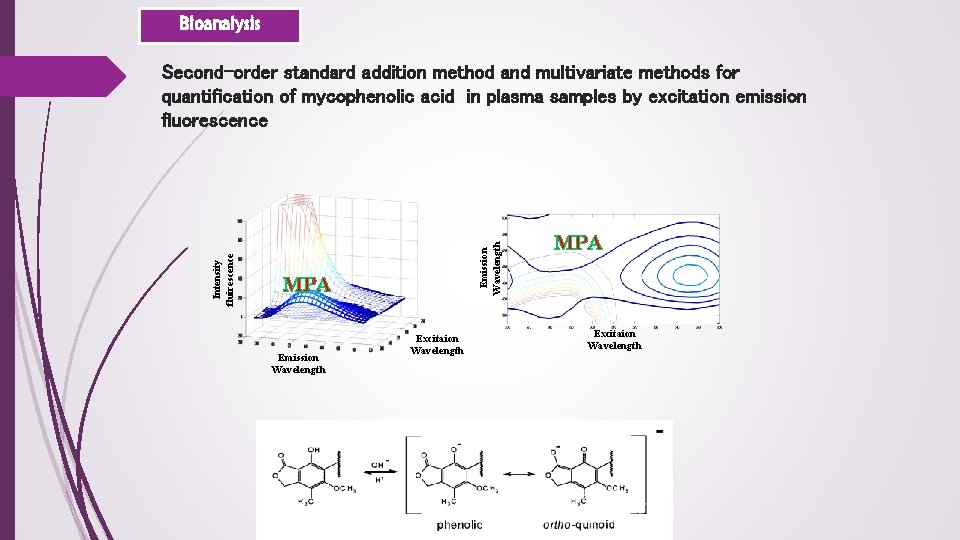
Bioanalysis Emission Wavelength Intensity fluirescence Second-order standard addition method and multivariate methods for quantification of mycophenolic acid in plasma samples by excitation emission fluorescence MPA Emission Wavelength Excitaion Wavelength MPA Excitaion Wavelength
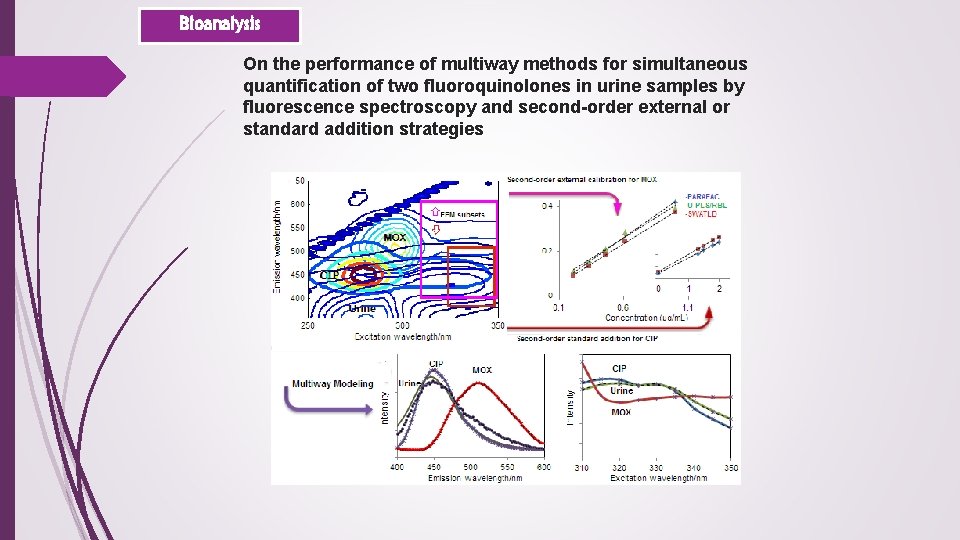
Bioanalysis On the performance of multiway methods for simultaneous quantification of two fluoroquinolones in urine samples by fluorescence spectroscopy and second-order external or standard addition strategies
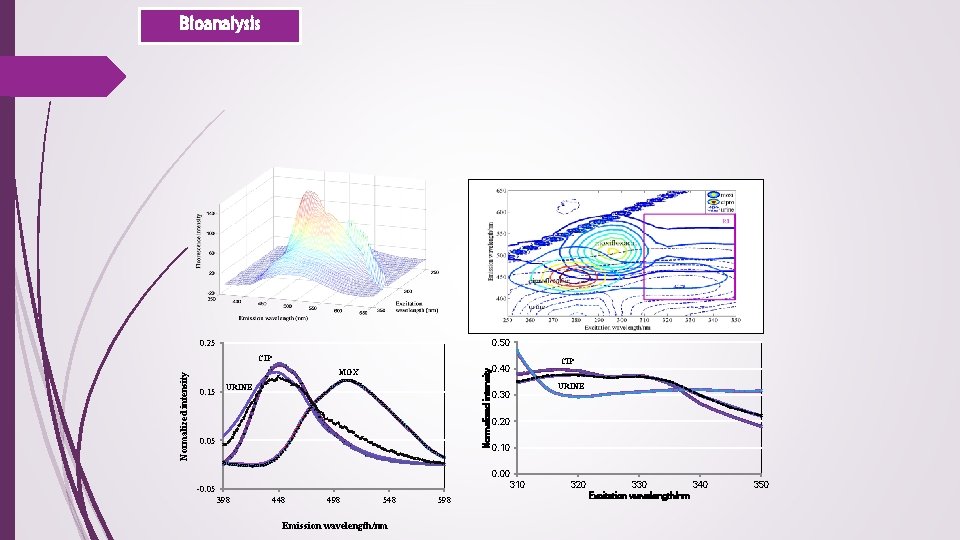
Bioanalysis 0. 50 0. 25 Normalized intensity CIP MOX 0. 15 URINE 0. 05 CIP 0. 40 URINE 0. 30 0. 20 0. 10 0. 00 310 -0. 05 398 448 498 548 Emission wavelength/nm 598 320 330 340 Excitation wavelength/nm 350
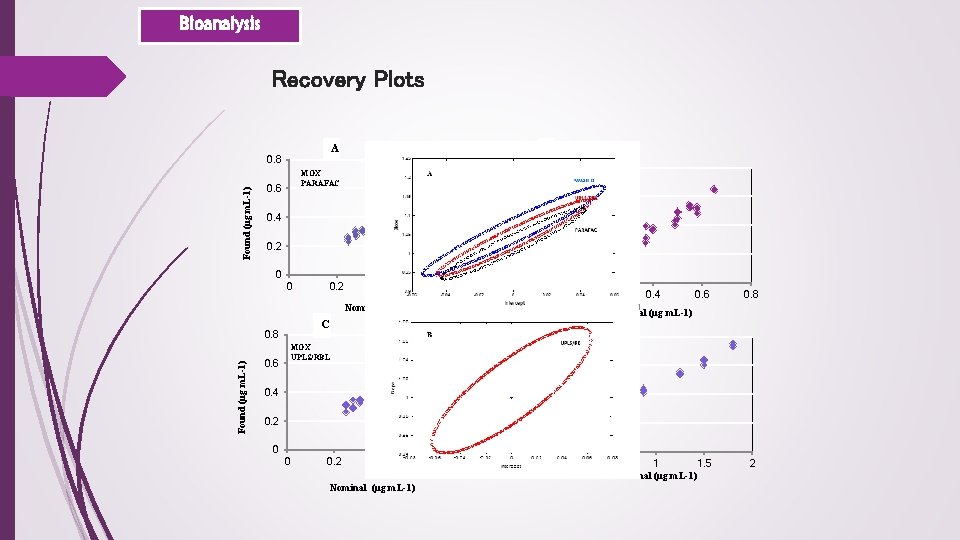
Bioanalysis Recovery Plots A B 0. 8 MOX PARAFAC 0. 6 Found (μg m. L-1) 0. 8 0. 4 0. 2 0 0 0. 2 0. 4 0. 6 MOX SWATLD 0. 6 0. 4 0. 2 0 0. 8 0 0. 2 0. 6 0. 8 0. 5 1 1. 5 Nominal (μg m. L-1) 2 MOX UPLS/RBL 0. 6 Found (μg m. L-1) 0. 8 Nominal (μg m. L-1) D C 0. 4 0. 2 0 0 0. 2 0. 4 Nominal (μg m. L-1) 0. 6 0. 8 0. 4 CIP UPLS/RBL 1. 5 1 0. 5 0 0

66
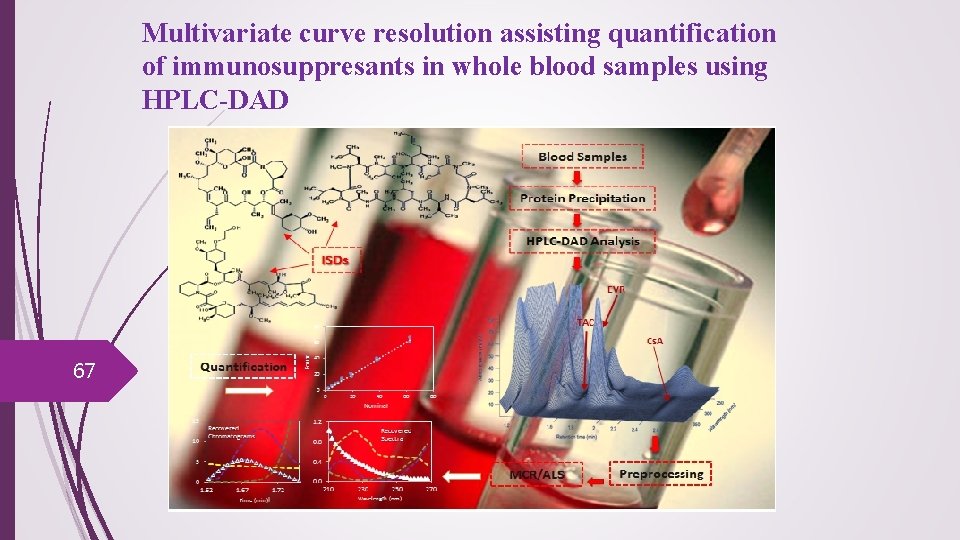
Multivariate curve resolution assisting quantification of immunosuppresants in whole blood samples using HPLC-DAD 67
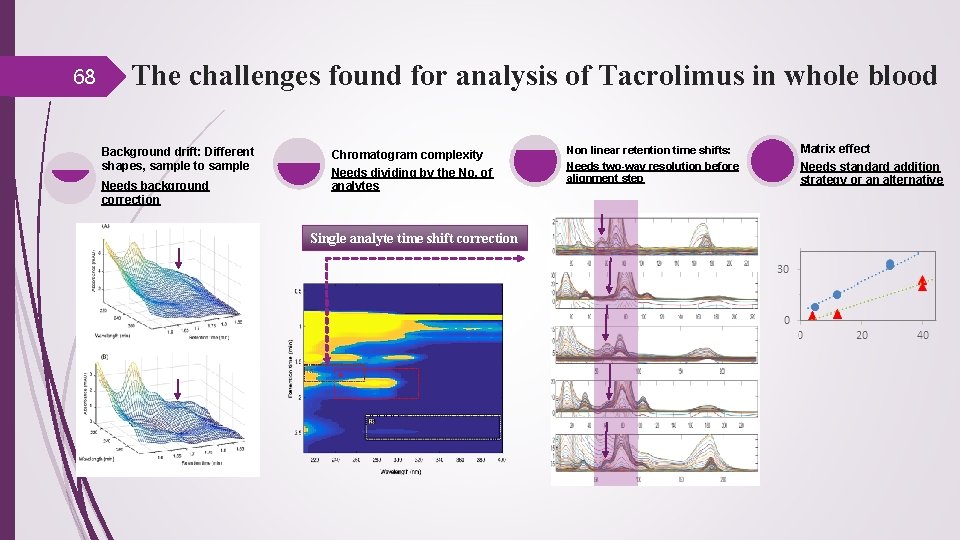
68 The challenges found for analysis of Tacrolimus in whole blood Background drift: Different shapes, sample to sample Needs background correction Chromatogram complexity Needs dividing by the No. of analytes Single analyte time shift correction Non linear retention time shifts: Needs two-way resolution before alignment step Matrix effect Needs standard addition strategy or an alternative
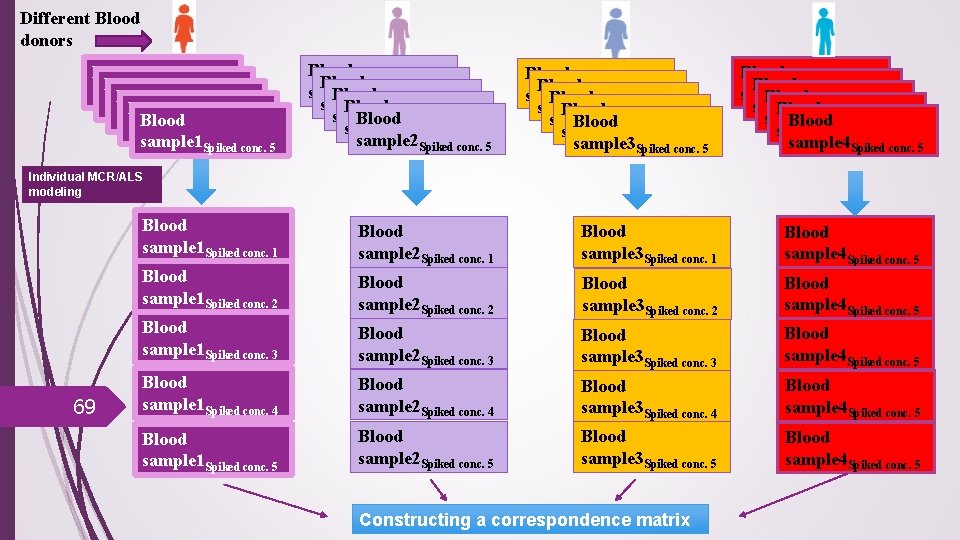
Different Blood donors Blood sample 1 Blood. Spiked conc. 1 sample 1 Blood. Spiked conc. 2 sample 1 Blood. Spiked conc. 3 sample 1 Spiked conc. 4 sample 1 Spiked conc. 5 Blood sample 2 Blood. Spiked conc. 1 sample 2 Blood. Spiked conc. 2 sample 2 Blood. Spiked conc. 3 sample 2 Spiked conc. 4 sample 2 Spiked conc. 5 Blood sample 3 Blood. Spiked conc. 1 sample 3 Blood. Spiked conc. 2 sample 3 Blood. Spiked conc. 3 sample 3 Spiked conc. 4 sample 3 Spiked conc. 5 Blood sample 4 Blood. Spiked conc. 1 sample 4 Blood. Spiked conc. 2 sample 4 Blood. Spiked conc. 3 sample 4 Spiked conc. 4 sample 4 Spiked conc. 5 Individual MCR/ALS modeling 69 Blood sample 1 Spiked conc. 1 Blood sample 2 Spiked conc. 1 Blood sample 3 Spiked conc. 1 Blood sample 1 Spiked conc. 2 Blood sample 4 Spiked conc. 5 Blood sample 2 Spiked conc. 2 Blood sample 1 Spiked conc. 3 Blood sample 3 Spiked conc. 2 Blood sample 4 Spiked conc. 5 Blood sample 2 Spiked conc. 3 Blood sample 3 Spiked conc. 3 Blood sample 4 Spiked conc. 5 Blood sample 1 Spiked conc. 4 Blood sample 2 Spiked conc. 4 Blood sample 3 Spiked conc. 4 Blood sample 4 Spiked conc. 5 Blood sample 1 Spiked conc. 5 Blood sample 2 Spiked conc. 5 Blood sample 3 Spiked conc. 5 Blood sample 4 Spiked conc. 5 Constructing a correspondence matrix
70 MCR-BANDS Unique solution
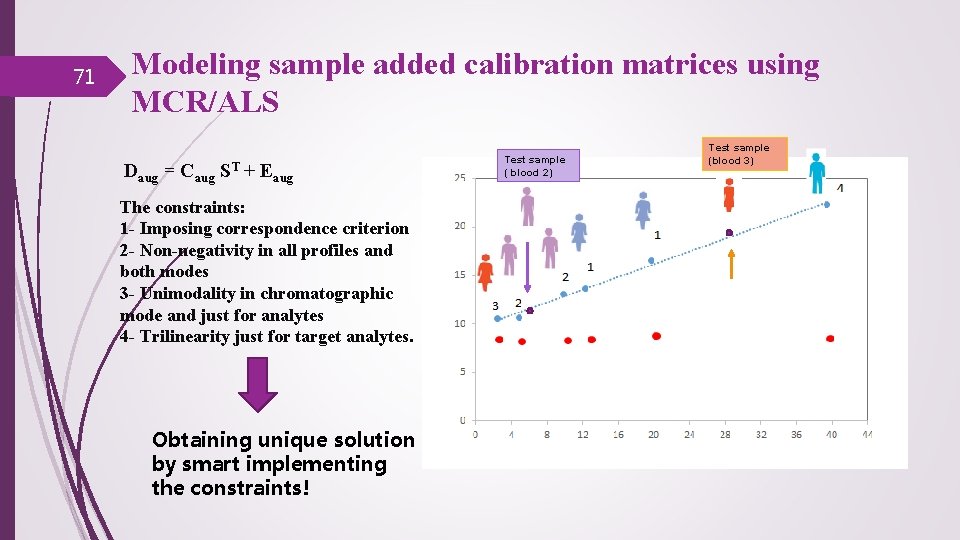
71 Modeling sample added calibration matrices using MCR/ALS Daug = Caug ST + Eaug The constraints: 1 - Imposing correspondence criterion 2 - Non-negativity in all profiles and both modes 3 - Unimodality in chromatographic mode and just for analytes 4 - Trilinearity just for target analytes. Obtaining unique solution by smart implementing the constraints! Test sample ( blood 2) Test sample (blood 3)
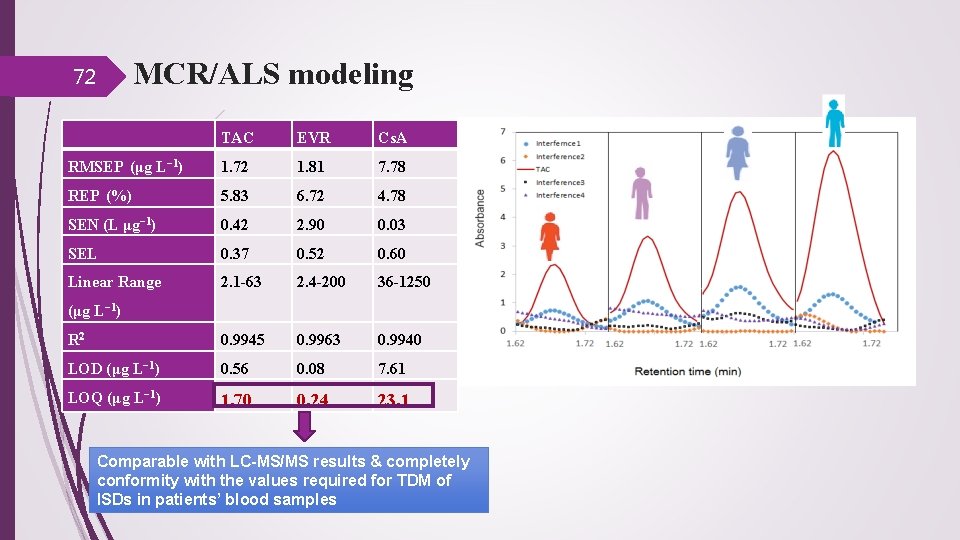
MCR/ALS modeling 72 TAC EVR Cs. A RMSEP (μg L− 1) 1. 72 1. 81 7. 78 REP (%) 5. 83 6. 72 4. 78 SEN (L μg− 1) 0. 42 2. 90 0. 03 SEL 0. 37 0. 52 0. 60 Linear Range 2. 1 -63 2. 4 -200 36 -1250 R 2 0. 9945 0. 9963 0. 9940 LOD (μg L− 1) 0. 56 0. 08 7. 61 LOQ (μg L− 1) 1. 70 0. 24 23. 1 (μg L− 1) Comparable with LC-MS/MS results & completely conformity with the values required for TDM of ISDs in patients’ blood samples

Concluding Remarks ü The basic knowledge of the chemometrics methods is necessary to find out a proper knowledge about the system under study ü Identifying where the proper chemometrics methods can be of use ü Having an estimate of instrumental noise level may facilitates the work ü Multiway calibration methods have several significant benefits for multi-target analysis in high throughput screenings. ü Exploiting chemometric data handling methods, sample preparation can be simplified. ü Because no priori knowledge of the interfering components in the sample is necessary, analysis time can be reduced. ü A target oriented preprocessing step together with a proper two-way data partitioning ü Smart implementation of constraints is necessary to resolve complex samples ü Having a good knowledge of experimental sections is necessary (Chemometrics can not do a miracle!) ü Performing more and more case studies for different algorithms can enable us to show and confirm its advantages, disadvantages, its chllenges and finally its establishment or can be a good starting point for its developments. ü. .

Special Thanks to: Chemometrics students in CCERCI • Dr Kourosh Tabar Heidar, CCERCI • Dr Amir Salemi, SBU Maesumeh Rashvand Mahin Bayat • Dr Dr S. Hamid Ahmadi, CCERCI Nafiseh Shakari Nahal Rahimdoost • Dr Kazem Kargosha, CCERCI • Reza Zadmard, CCERCI Shabnam Nasrollahi Hadi Esfahani Shiva Ghafghazi Samaneh Gorji • & Sara Nouroozi Elaheh Mazlumi • Prof. Jahanbakhsh Ghasemi Hadi Mohamedian • Prof. Marcel Maeder Farzad Torabi • Thanks to: • • • CCERCI Research Council NAJA-Research Center INSF Shaheed beheshti University Hospital Taleghani, Modarres Treatment plant Fatemi, Ekbatan, Zargande, Shush,

75 THANK YOU
- Slides: 75