CHAPTER SIX Nucleic acid hybridization principles and applications

CHAPTER SIX Nucleic acid hybridization: principles and applications 생물정보학협동과정 강민호

Nucleic Acid Hybridization • Nucleic acid hybridization is a fundamental tool in molecular genetics which takes advantage of the ability of individual single -stranded nucleic acid molecules to form double stranded molecules (that is, to hybridize to each other)

Standard nucleic acid hybridization assays - A labeled nucleic acid - a probe - to identify related DNA or RNA molecules - Complex mixture of unlabeled nucleic acid molecules- the target -Base complementarity with a high degree of similarity between the probe and the target.
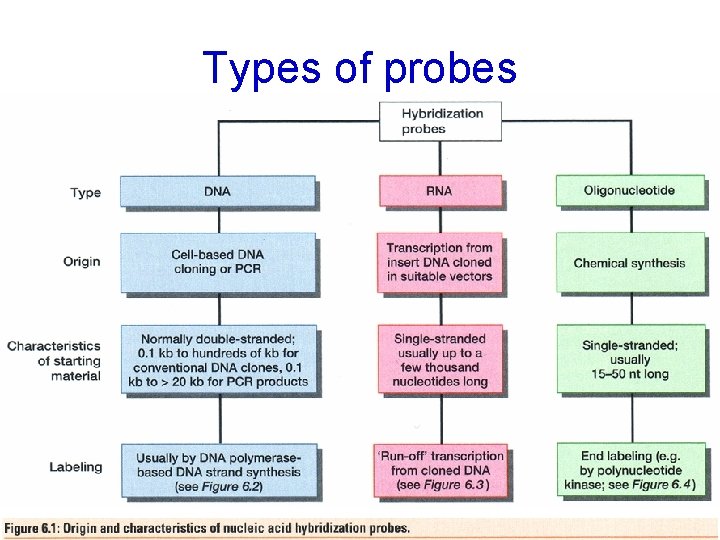
Types of probes

Probes • DNA labelling – 5’ – 3’ – Uniform labeling • Nick translation • Random primer • PCR-mediated labeling • RNA labelling – In vitro transcription of a cloned DNA insert • Different probes – Radioactive labeling or isotopic labeling – Nonradioactive labeling or nonisotopic labeling
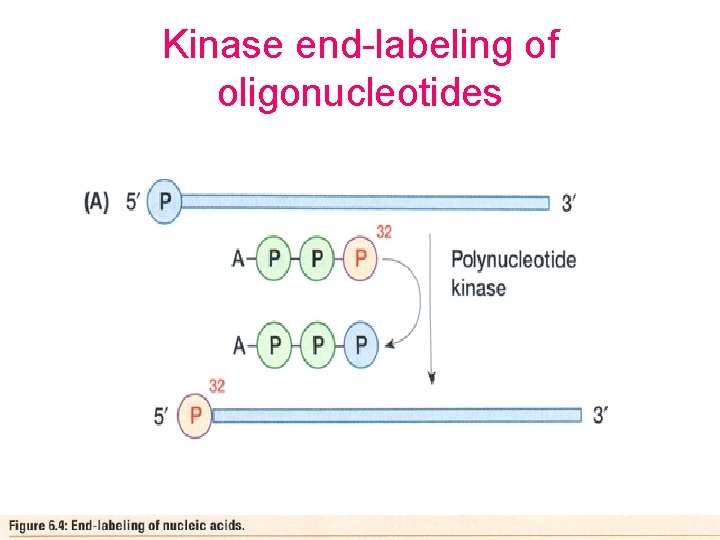
Kinase end-labeling of oligonucleotides
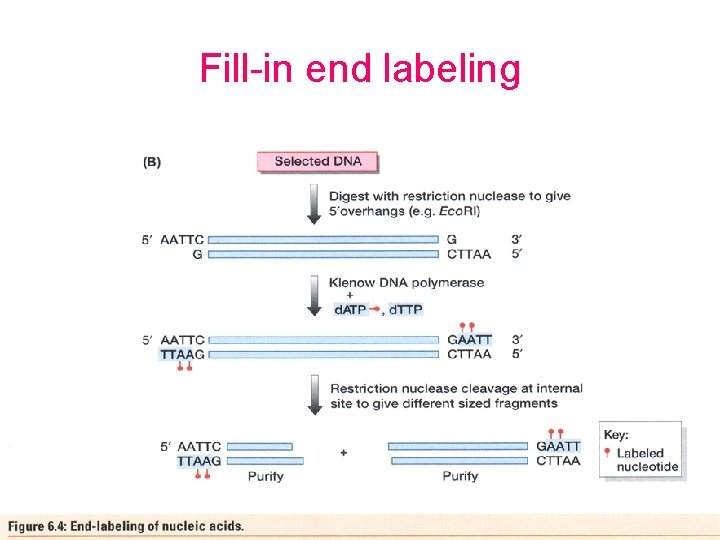
Fill-in end labeling
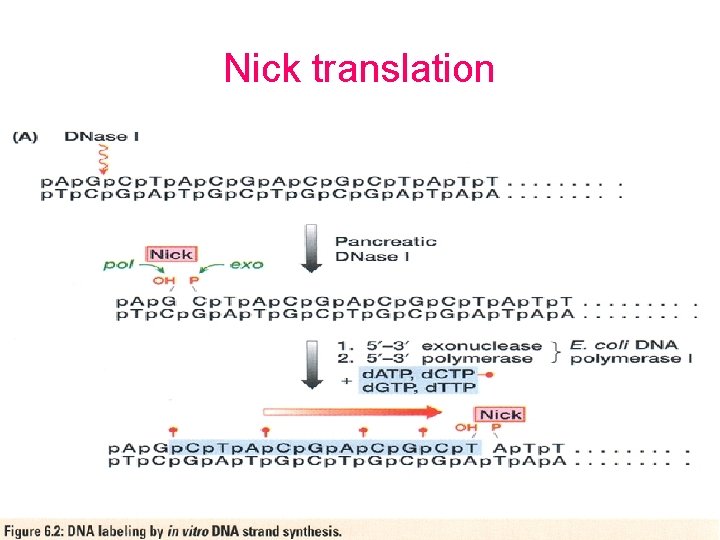
Nick translation
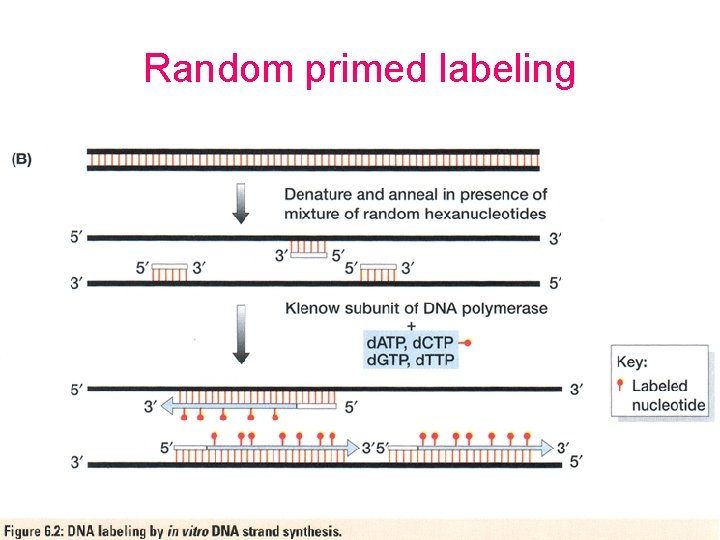
Random primed labeling
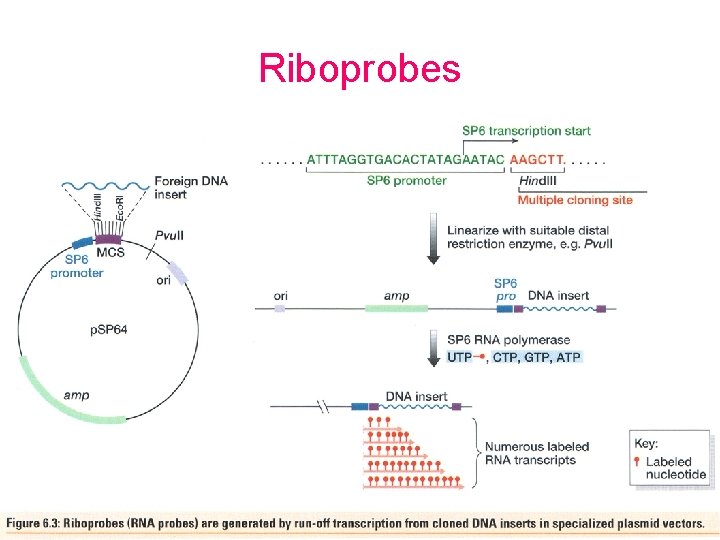
Riboprobes
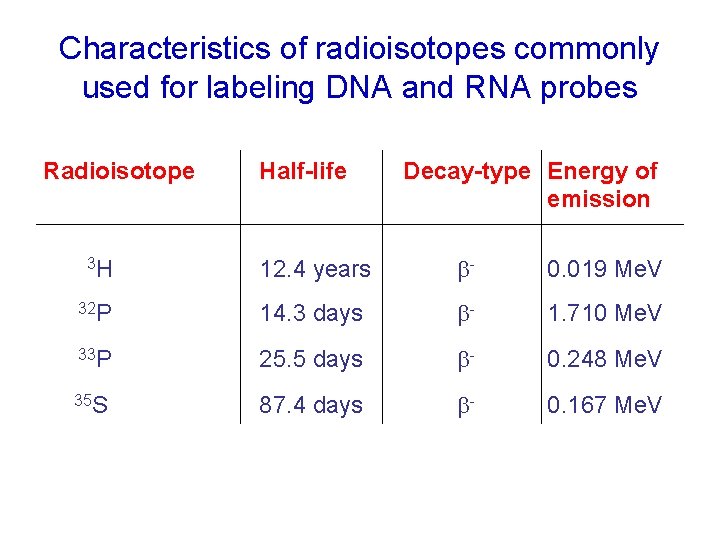
Characteristics of radioisotopes commonly used for labeling DNA and RNA probes Radioisotope Half-life Decay-type Energy of emission 12. 4 years b- 0. 019 Me. V 32 P 14. 3 days b- 1. 710 Me. V 33 P 25. 5 days b- 0. 248 Me. V 35 S 87. 4 days b- 0. 167 Me. V 3 H

Nonisotopic labeling and detection • The use of nonradioactive labels has several advantages: – – – safety higher stability of a probe efficiency of the labeling reaction detection in situ less time taken to detect signal • Major types – Direct nonisotopic labeling (ex. nt labeled with a fluorophore) – Indirect nonisotopic labeling (ex. biotin. -streptavidin system)
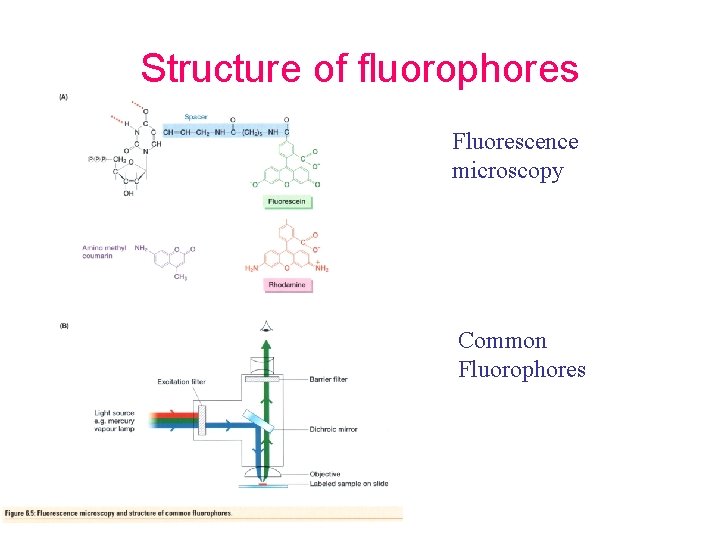
Structure of fluorophores Fluorescence microscopy Common Fluorophores
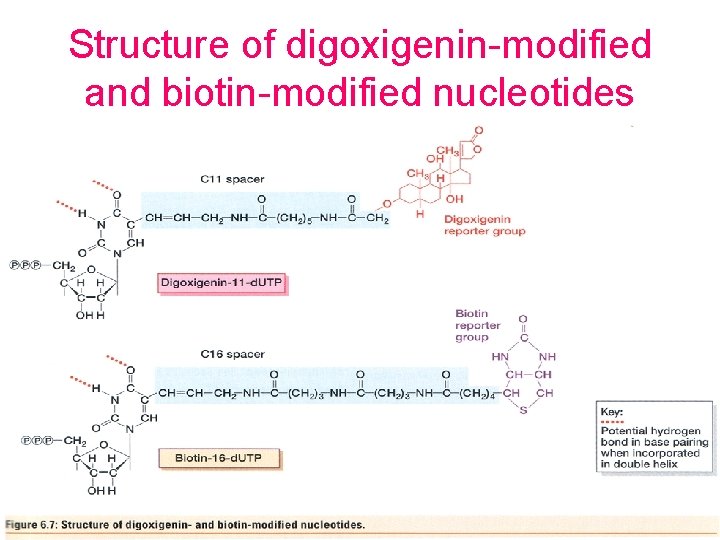
Structure of digoxigenin-modified and biotin-modified nucleotides
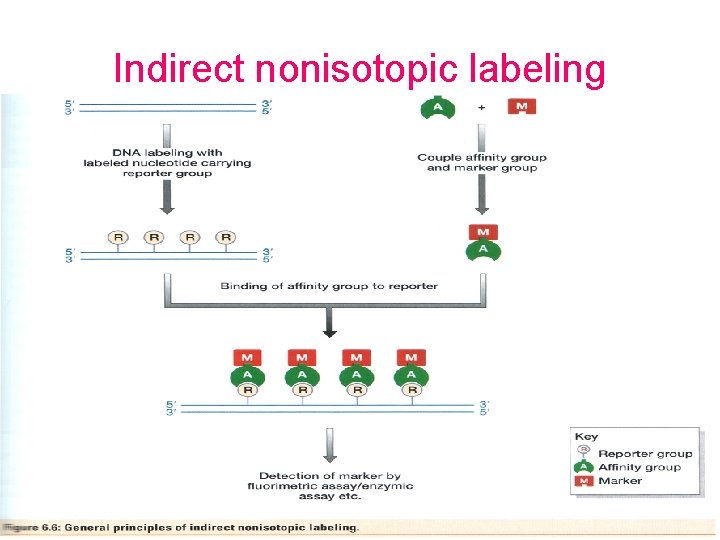
Indirect nonisotopic labeling
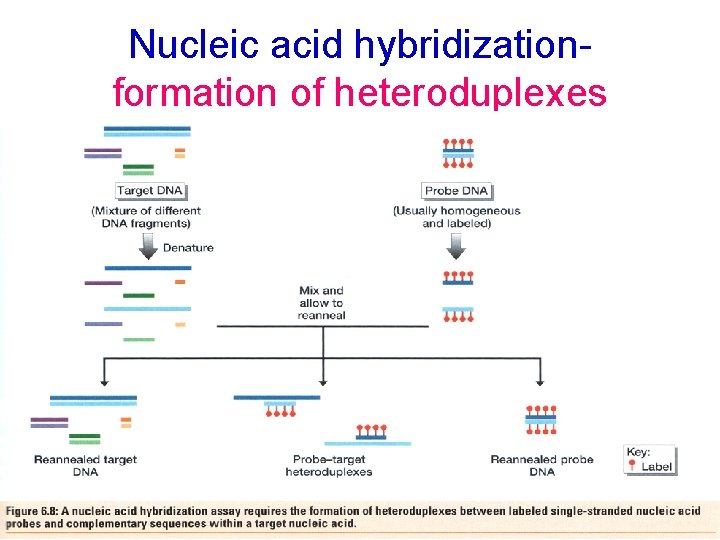
Nucleic acid hybridizationformation of heteroduplexes
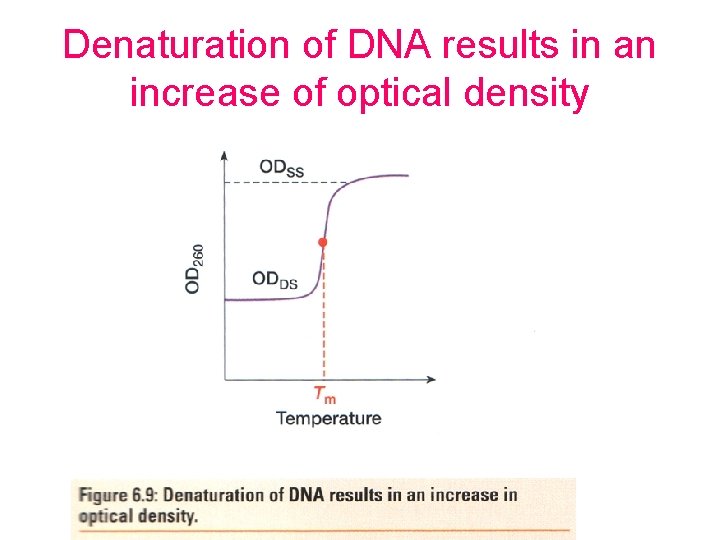
Denaturation of DNA results in an increase of optical density

Factors affecting Tm of nucleic acid hybrids • • • Destabilizing agents (ex. formamide, urea) Ionic strenght Base composition (G/C%, repetitive DNA) Mismatched base pairs Duplex lenght Different equations for calculating Tm for: • DNA-DNA hybrids • DNA-RNA hybrids • RNA-RNA hybrids • Oligonucleotide probes
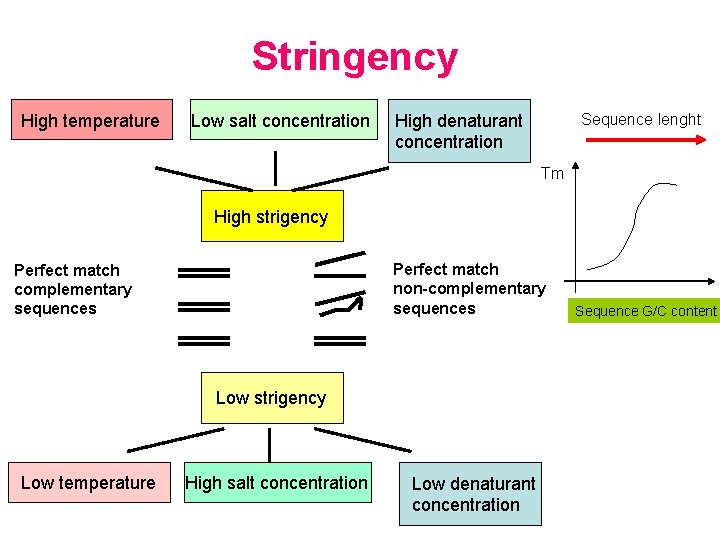
Stringency High temperature Low salt concentration Sequence lenght High denaturant concentration Tm High strigency Perfect match non-complementary sequences Perfect match complementary sequences Low strigency Low temperature High salt concentration Low denaturant concentration Sequence G/C content

The identification of specific sequences in a complex mixture.
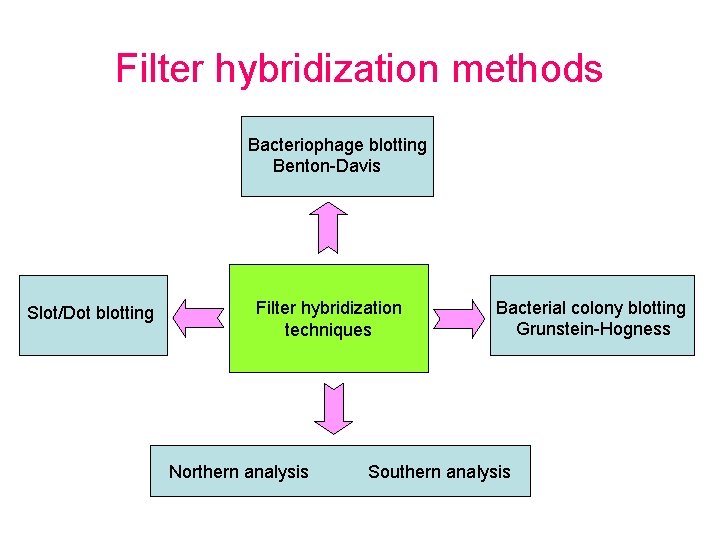
Filter hybridization methods Bacteriophage blotting Benton-Davis Slot/Dot blotting Filter hybridization techniques Northern analysis Bacterial colony blotting Grunstein-Hogness Southern analysis

Filters or Membranes • • Nitrocellulose Nylon Positive charged nylon (hybond) PVDF (hydrophobic polyvinylidene difloride) • Different properties: – – Binding capacity (mg nucleic acids/cm 2) Tensile strenght Mode of nucleic acid attachment Lower size limit for efficient nucleic acid retention
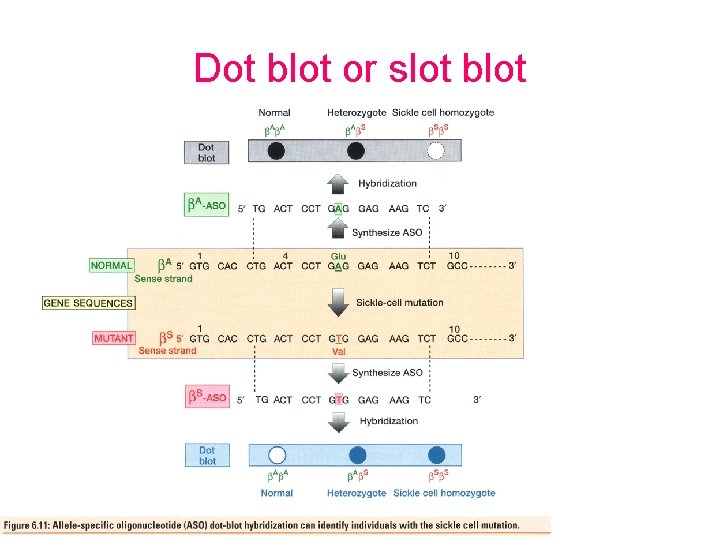
Dot blot or slot blot
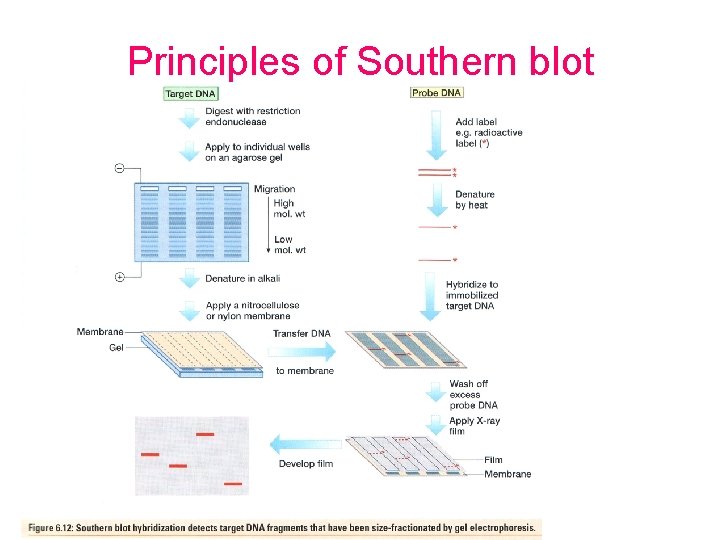
Principles of Southern blot
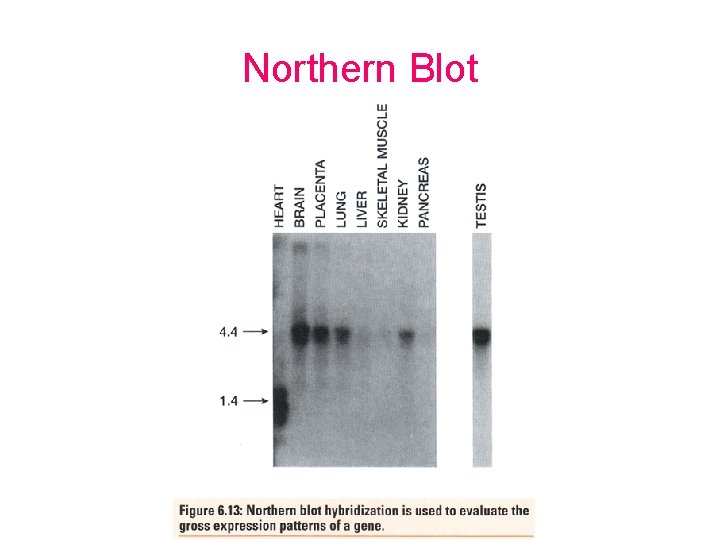
Northern Blot
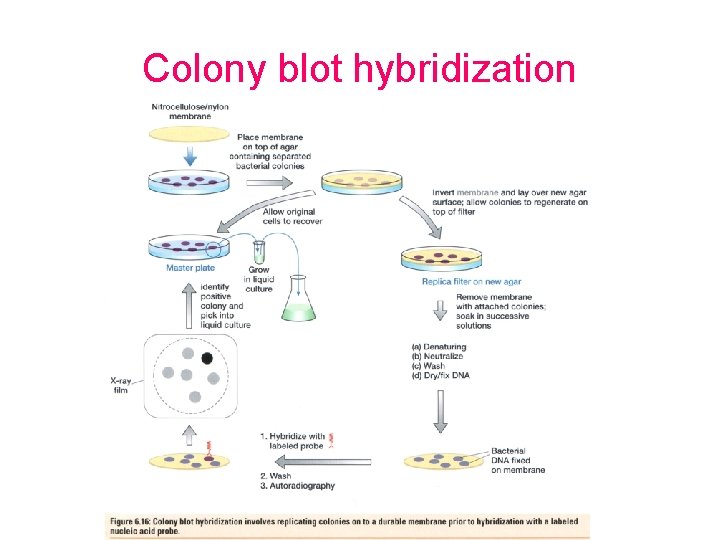
Colony blot hybridization

In situ hybridization • Chromosome in situ hybridization – Metaphase or protometaphase chromosomes are probed with labeled DNA. The DNA can be labeled with a fluorochrome (FISH). • Tissue in situ hybridization – Sliced or whole mounted preparations can be probed with RNA probes to detect m. RNA expression
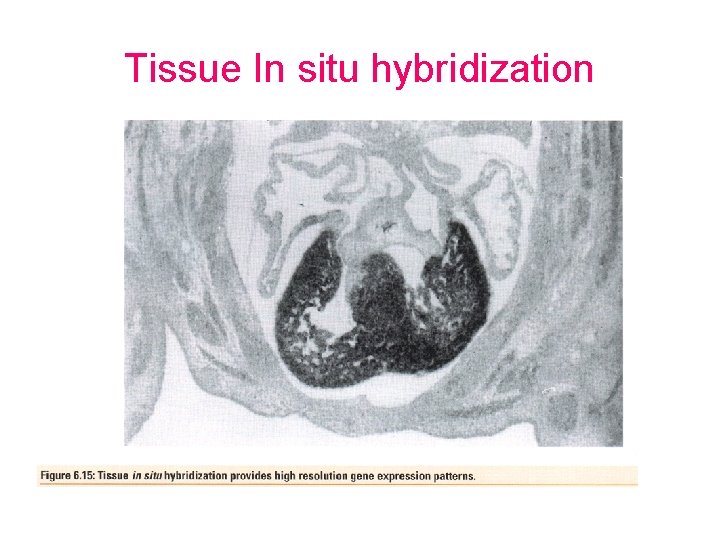
Tissue In situ hybridization

Gridded clone hybridization
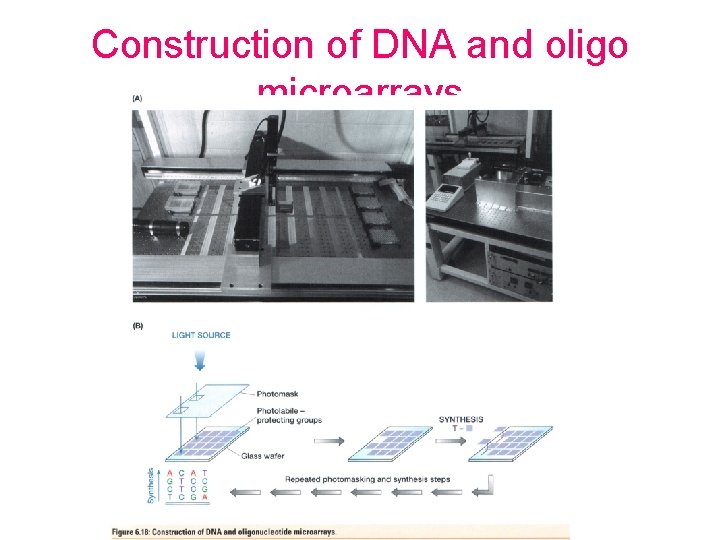
Construction of DNA and oligo microarrays

Gene expression profiling by hybridization

Summary I • Hybridization is due to complementarity of DNA strands. • DNA can be labeled various ways • Isotopic and non isotopic • Hybridization can detect identical or similar sequences.

Summary II • A variety of techniques utilize hybridization of DNA or RNA probes – ASO – Southern Blot, RFLP, VNTRs, Mutation detection, deletion detection – Northern Blot, tissue specific expression – In situ hybridization • Chromosome location and integrity • Tissue specific expression

Summary III • Colony hybridization can be used to identify specific clones. Once you have one clone you can find others that hybridize to it. • Screening of gridded clones. One can identify genomic clones homologous to a c. DNA or identify c. DNA expressed in a cell line. • Microarrays can be used in many ways to analyze gene expression in various cell types, in response to various stimuli.
- Slides: 34