Chapter 9 NUCLEIC ACIDS Acknowledgement Addisa Ababa University

Chapter 9 -NUCLEIC ACIDS

Acknowledgement • • Addisa Ababa University Haramaya University Hawassa University Jimma University of Gondar American Society for Clinical Pathology Center for Disease Control and Prevention. Ethiopia

Chapter learning objective • At the end of this chapter the student will be able to : • Explain classification and chemistry of nucleic acids. RNA and DNA • Discuss metabolism of nucleotides • Explain clinical significance (Hyperuricemia and gout and Orotic aciduria • Explain DNA replication • Explain transcription • Explain translation

• 8. 1. classification and biochemistry of nucleic acid • Nucleic acids are required for the storage and expression of genetic information • Two types of nucleic acids are present in all mammalian cells including humans. • They are DNA-deoxy ribonucleic acid and RNAribonucleic acid. • DNA is present not only in chromosomes in the nucleus of eukaryotic organisms but also in mitochondria and the chloroplasts of plants

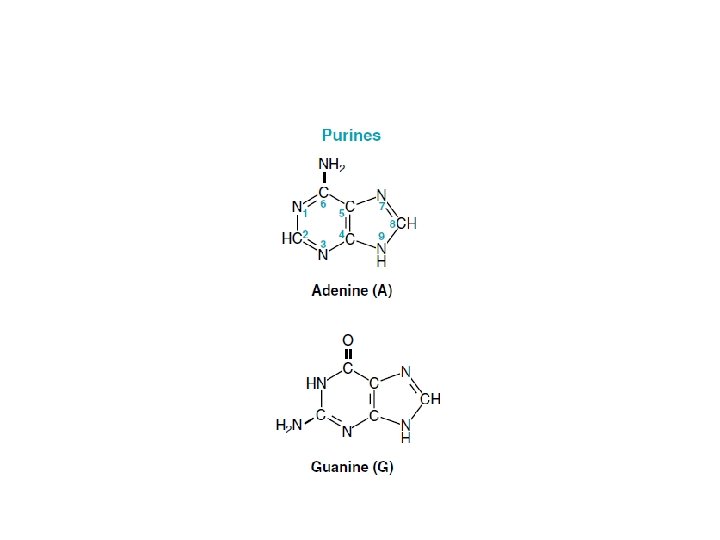
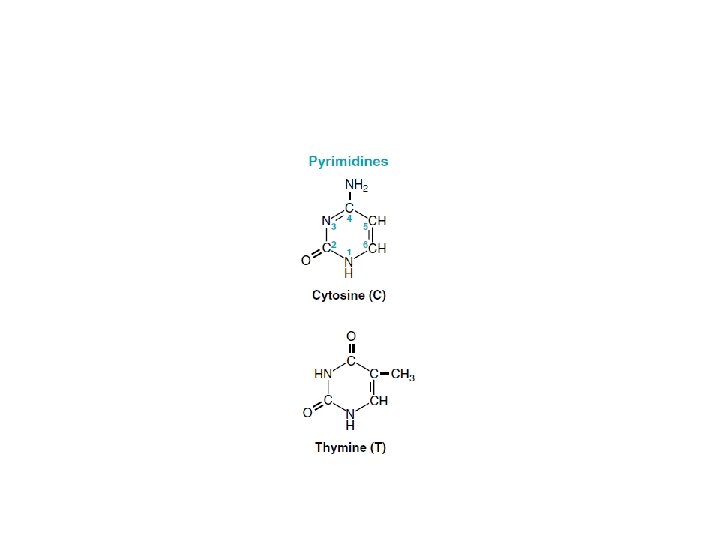
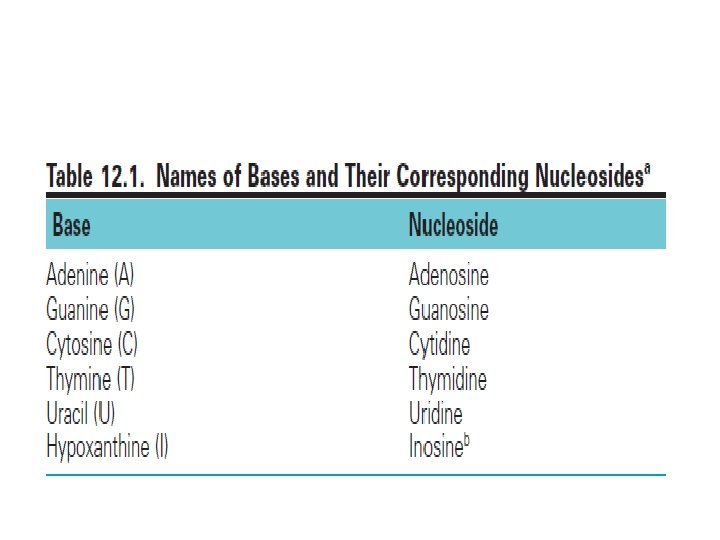
• Nucleotides are the monomeric units of the nucleic acids, DNA (deoxyribonucleic acid) and RNA (ribonucleic acid). • Each nucleotide consists of a heterocyclic nitrogenous base, a sugar, and phosphate. • DNA contains the purine bases adenine (A) and guanine (G) and the pyrimidine bases cytosine (C) and thymine (T). RNA contains A, G, and C, but it has uracil (U) instead of thymine. • In DNA, the sugar is deoxyribose, whereas in RNA it is ribose.

Outline 8. 1. Classification and chemistry of nucleic acids 8. 2. Metabolism of nucleotides 8. 3. Clinical significance (Hyperuricemia and gout and Orotic aciduria 8. 4. DNA replication 8. 5. Transcription 8. 6. Translation

• Prokaryotes have a single chromosome, but may also contain nonchromosomal DNA in the form of plasmids • RNA is present in nucleus and cytoplasm • Nucleic acids are acidic substances containing nitrogenous bases, sugar and phosphorus.

• Both DNA and RNA are polymers of nucleotides (polynucleotides) joined together by phosphodiester linkage. • Diester linkage of phosphate joins 3' OH and 5' OH belonging to two separate sugars


• Nucleotide sequence of a polynucleotide is known as primary structure of nucleic acid and it confers individuality to polynucleotide chain. • Polynucleotide chains are represented in 5' → 3' direction only. • However, the phosphodiester linkage runs in 3' → 5' direction.

• Each poly nucleotide chain has two ends. The 5' end carrying phosphate is on the left hand side and 3' end carrying unreacted hydroxyl is on the right hand side. • Primary structures of DNA and RNA exist in single stranded DNA and RNA organisms.

• Therefore, in DNA and RNA, letters A, G, C, T stands for nitrogen bases and sugar is deoxy ribose if the polynucleotide is a segment of DNA and sugar is ribose if it is RNA segment. • Hydrolysis of nucleotides produce nitrogen bases, sugars and phosphate.

• Nucleotides contain two types of nitrogenous bases: -purine -pyrimidine • In the case of purine nucleosides, the sugar is attached to N-9 of purine ring where as in pyrimidine nucleosides the sugar is attached to N-1 of pyrimidine ring. • Nucleotides are phosphorylated nucleosides.
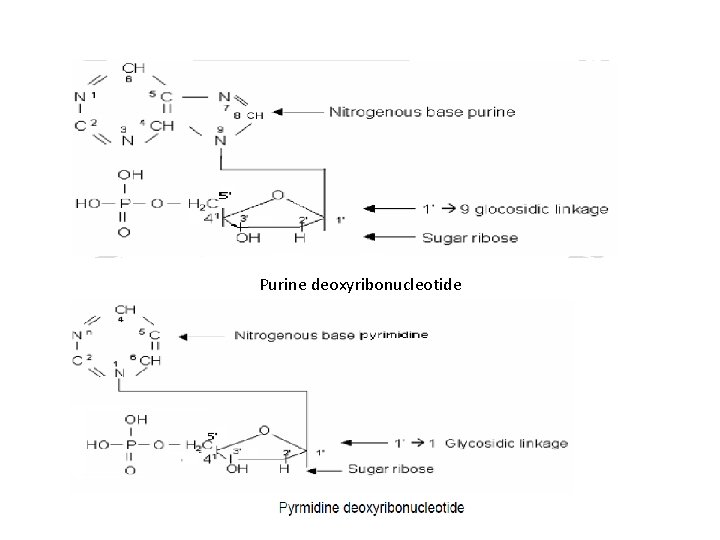
Purine deoxyribonucleotide

Structure of DNA • Deoxyribonucleic acid (DNA) is polymer of deoxyribonucleotides attached to each other by phosphodiester linkages. • Each deoxyribonucleoside is composed of nitrogen bases and a sugar deoxyribose. • The nitrogenous bases are purines and pyrimidines. • The two purine bases are adenine and guanine. • The three pyrimidine bases are cytosine, thymine and uracil.

Primary Structure of DNA • The deoxyribonucleotides are linked together by phosphodiester bonds between the 3'–hydroxyl of the sugar of one nucleotide through a phosphate molecule to the 5'–hydroxyl on the sugar of another nucleotide. • The sugar–phosphate linkages form the backbone of the polymer to which the variable bases are attached.

• The nucleotide polymer has a free phosphate group attached to 5'–position of sugar and a free 3’–hydroxyl group. • The sequence of the polymer is written in the 5’ to 3’ direction with abbreviations to different bases
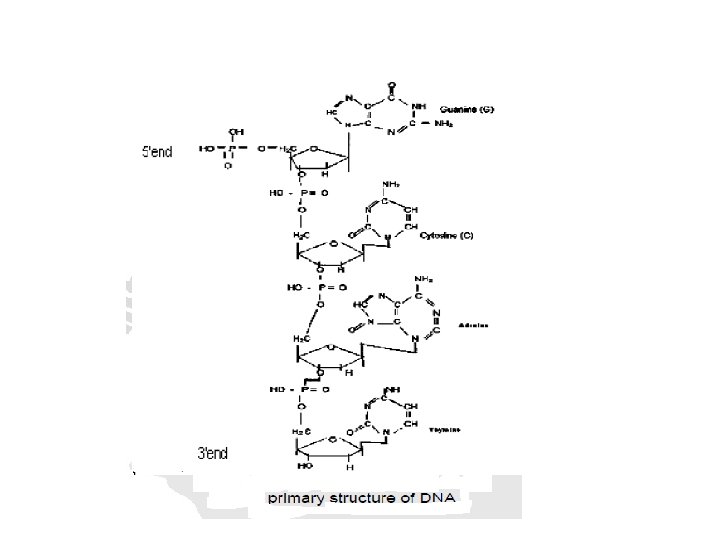

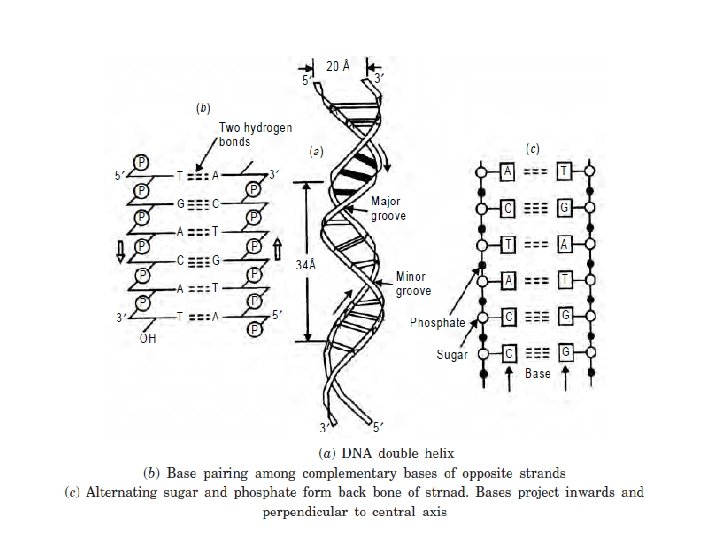
Secondary Structure of DNA • The secondary structure of DNA is formed when the two strands of DNA are paired together • In the secondary structure of DNA, the two strands are anti-parallel. That means, the 5’ ---3’ of one strand is in opposite direction to the other strand. • The bases are stacked in the inside of the two strands.

• The bases of one strand pairs with the bases of the other strand of the same plane such that adenine always pairs with thymine with two bonds. Guanine always pairs with cytosine with three bonds. • The negatively charged phosphate group and the sugar units expose themselves to the outside of the chain. • The two strands of DNA coil around a single axis forming right handed double helix.

• The planes of the sugars are at right angles to that of the bases. • The helical structure repeats after 10 residues on each chain. • Watson - Crick Model of DNA is also referred as B-DNA, which is the most stable under physiological conditions. • The mitochondrial DNA is circular and there can be formation of Z-DNA and C-DNA which can be performed during either replication or transcription.

The Structure of RNA • The building unit of RNA is ribonucleotide. • Ribonucleotide contains “O” in the carbon 2’ sugar ribose. • Uracil is found in RNA while Thymine is found in DNA. • The nuclear DNA is in secondary structure, but RNA is in primary structure. • Only t. RNA after post transcriptional process can be changed to tertiary structure.
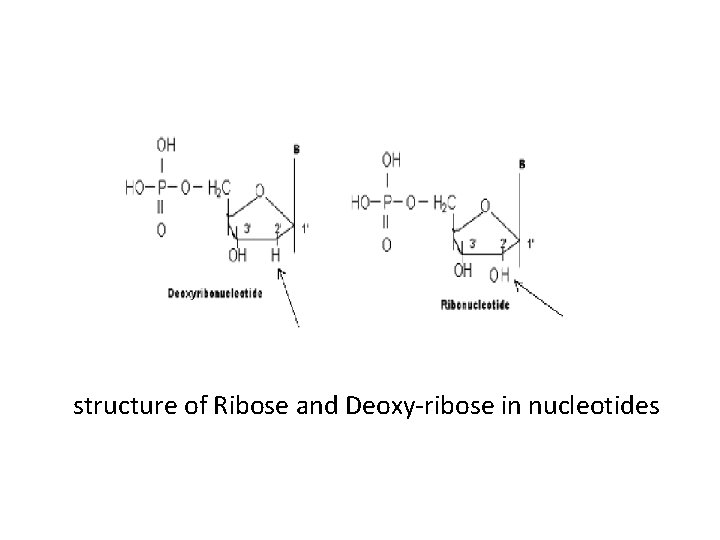
structure of Ribose and Deoxy-ribose in nucleotides

• The three RNAs that have important role in protein synthesis are: 1. Messenger RNA (m. RNA) 2. Transfer RNA (t. RNA) 3. Ribosomal RNA (r. RNA) • They differ from each other by size, function and stability.

Messenger RNA (m. RNA) • It accounts for 5% of cellular RNA. • m. RNA contains cap at the 5’ end of the chain and poly-A tail at 3’- end. • Cap characterizes 7 -methylated guanosine tri phosphate (m 7 GTP). -Cap protects from exonuclease attack. -also used for recognition during protein synthesis

• Poly- A (a polymer of adenylate) characterizes about 200 successive adenylate residues. -serves to protect the m. RNA form exonuclease attack. -serves for the transport of the m. RNA form nucleus to cytosol.

• They consist of 1000 -10, 000 nucleotides • m. RNA molecules have different life spans that ranges from few minutes to days. • In prokaryotes 5' end of m. RNA contains a sequence rich in A and G. Such sequence is known as Shine-Dalgarno sequence. It helps attachment of m. RNA with ribosome during protein synthesis. • Some prokaryotic m. RNA has secondary structure. Intrastrand base pairing among complementary bases allows folding of linear molecule. As a result hairpin, or loop like secondary structure is formed.

Transfer RNA (t. RNA) • t. RNA accounts for 15% of total cell RNA. • They are the smallest of all the RNAs. Usually consist of 50 -100 nucleotides. • All t. RNA have 4 arms. 1. Amino acid arm: the one that carries amino acid 2. DHU (dihydrouridine) arm: the one that binds with active center of the enzyme aminoacyl t. RNA synthetase.

3. TϕC (ribothymidine-pseudouridine-cytidine) arm: the one that binds to ribosome during protein synthesis. Greek alphabet ϕ (Psi) stands for pseudo uridine. 4. Anticodon arm: which pairs with the codon of m. RNA during protein synthesis. • Secondary structure of all the t. RNAs is in the form of clover leaf

• An amino acid arm is where amino acid is attached to 3'-OH of adenosine moiety of t. RNA. • ACC is the common base sequence at the 3'-end. • An anti-codon arm recognizes codon on m. RNA. • The 5' end of t. RNA is phosphorylated and residue is guanosine.
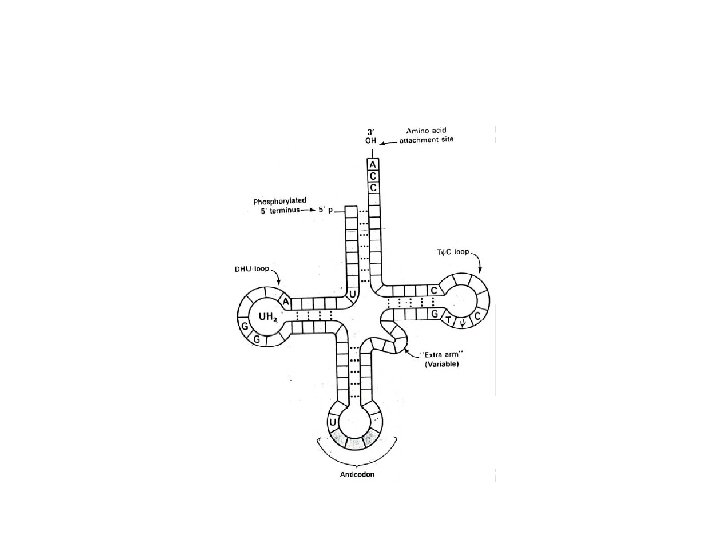

Ribosomal RNA (r. RNA) • r. RNA is highly methylated as compared to the other RNAs. • r. RNA, ribosomal proteins and Mg++ constitute ribosome. • Each ribosome consists of two big and small subunits. • r. RNA accounts for 80% of total cellular RNA.

• In ribosomes, r. RNA is found in combination with protein. It is known as ribonucleoprotein. • The length of r-RNA ranges form 100 -600 nucleotides. • r. RNAs differ in sedimentation coefficients (S). 1. In prokaryotic cells as 23 s, 16 s, and 5 s 2. In eukaryotic cells as 28 s, 18 s, 5. 8 s and 5 s. • S is for Svedberg unit which is related to molecular weight and shape
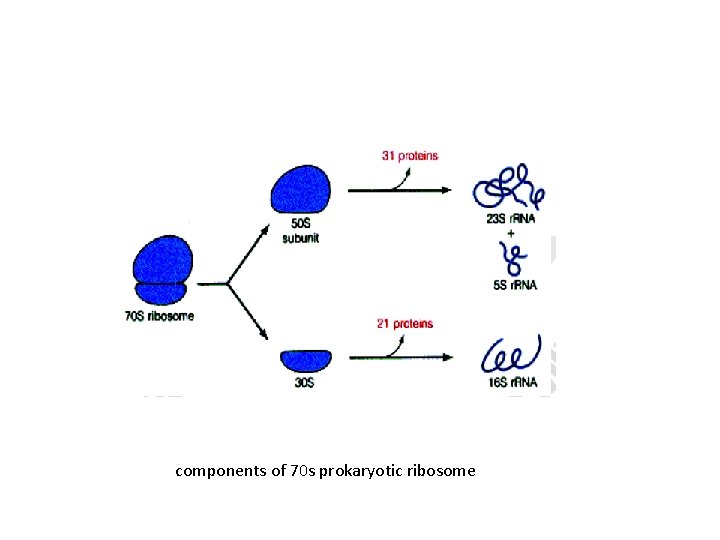
components of 70 s prokaryotic ribosome
Secondary structure of 16 S r. RNA

8. 2. Catabolism of Nucleic acids • Pancreatic enzymes called nucleases hydrolyze both DNA and RNA to nucleotides and then to nucleoside and phosphoric acid in the intestinal tract. • Nucleic acid is also hydrolyzed by lysosomal enzymes inside tissues. • The nucleoside is absorbed in to blood and transported to peripheral tissues. Excess nitrogen bases are further degraded. • Finally adenine and guanine are converted to uric acid in our body which is excreted through urine.

9. 3. clinical correlates • Since uric acid precipitates, excess uric acid causes kidney stone in kidney and gout in joints • On degradation of pyrimidine bases: 1. Cytosine and uracil are converted to ammonia, carbondioxide and beta-alanine 2. Thymine to NH 3, CO 2, H 2 O and α-methyl β alanine.

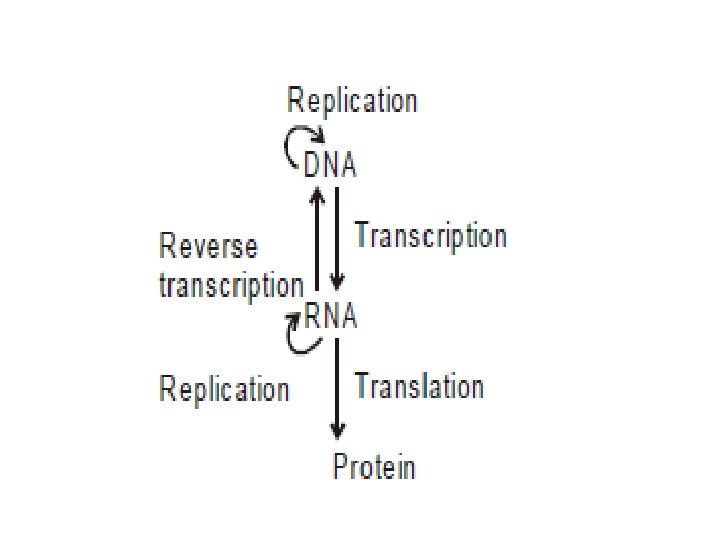
BIOSYNTHESIS OF NUCLEIC ACIDS • Genetic information stored in DNA in the form of nucleotide sequence flows from DNA to DNA, DNA to RNA then from RNA to protein. • This flow of genetic information is the central dogma of molecular genetics. • Usually involves three different processes. 1. Replication • Is synthesis of new DNA or information copying • In this process information is transmitted from parent to daughter. The new DNA is identical to parent DNA

2. Transcription • Is synthesis of RNA from DNA or information transfer • In this process, information is transferred from DNA to RNA 3. Translation • Synthesis of proteins using information present in RNAs or information decoding. • In this process, information present in RNA in the form of nucleotide sequence, is converted into sequence of amino acids

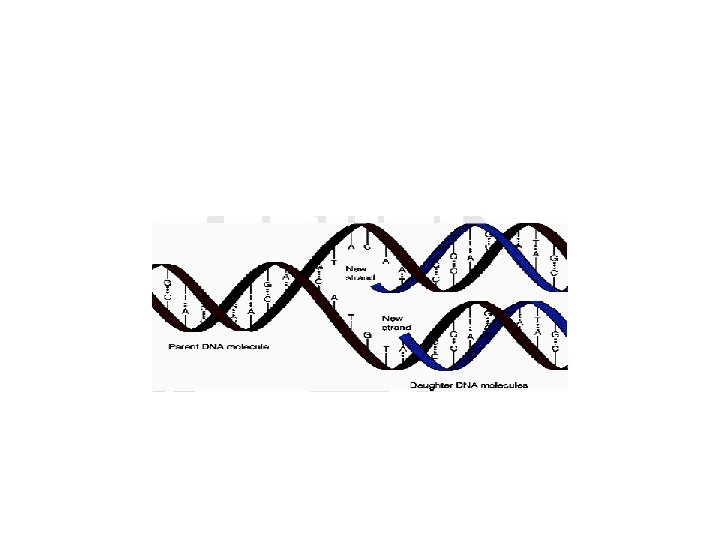
9. 4. DNA REPLICATION • Is synthesis of DNA • It is the way in which the genetic information can pass from parental cell to daughter cell. • As stated before, the double helical structure of DNA depends on the base complementarity.

• Also this complementarity represents the fundamental basis for the formation of new DNA strands from the parent DNA strand in a semi conservative manner. • In this process, two daughter DNA’s are produced, each has one parent strand (conserved) and newly synthesized strand.

• The enzymes involved are template-directed polymerases that can synthesize the complementary sequence of each strand with extraordinary fidelity. • Steps in prokaryotic DNA synthesis -separation of the two complementary DNA strands and formation of the replication fork -RNA primer synthesis -chain elongation -excision of RNA primers and their replacement by DNA

Separation of the two complementary DNA strands and formation of the replication fork • The parental double helical DNA must separate (melt) in order to replicate • DNA replication begins at unique nucleotide sequence, origin of replication

• In prokaryotic cells origin is at one site. In eukaryotic cells origin is at many sites. • These sites include a short sequence composed almost exclusively of AT base pairs • As the two strands unwind and separate they form a “V” where active synthesis occurs, replication fork

Proteins required for DNA strand separation ( prepriming complex) • Dna. A protein: bind to specific nucleotide sequences at the origin of replication. -It separates the DNA strand forming localized regions of single-stranded DNA • DNA helicases: bind to single-stranded DNA near the replication fork and then move into the neighboring double stranded region, forcing the strand apart in effect.

• Single-stranded DNA-binding (SSB) proteins (helix-destablizing proteins): bind only to the single-stranded DNA and keep the two strands of DNA separated in the area of replication origin to provide the single stranded template required by polymerase. -Also protect the DNA from nucleases that cleave single-stranded DNA

Solving the problem of supercoils • As the two strands are separated there will be appearance of positive supercoils (supertwists) ahead of the replication fork • Topoisomerases are responsible for removing supercoils in the helix • Type I DNA topoisomerases: reversibly cut a single strand of the double helix. -

have both nuclease (strand-cutting) and ligase (strand-resealing) activities • Type II DNA topoisomerases: bind tightly to the DNA double helix and make transient breaks in both strands
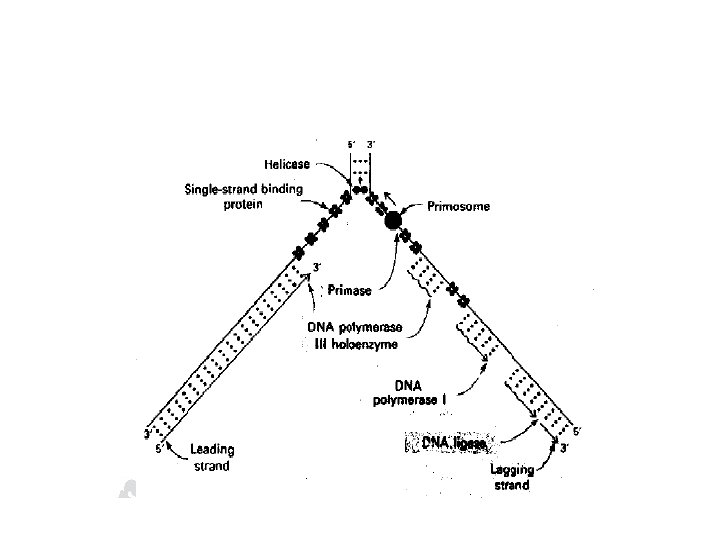

Direction of DNA replication • The DNA polymerase responsible for copying the DNA templates are only able to “read” the parental nucleotide sequences in the of 3' → 5' direction and synthesize the new DNA in the 5' → 3‘ • Leading strand: the strand that is being copied in the direction of the advancing replication fork and is synthesized almost continuously

• Lagging strand: the strand that is being copied in the direction away from the replication fork, synthesized discontinuously -these short stretches of discontinuous DNA (Okazaki fragment) are eventually joined to become a single continuous strand

RNA primer • DNA polymerase cannot initiate synthesis of a complementary strand of DNA on a totally singlestranded template, rather require an RNA primer • RNA primer: a short, double-stranded region consisting of RNA base-paired to the DNA template with a free hydroxyl group on the 3' end of the RNA strand.

Chain elongation • Prokaryotic and eukaryotic DNA polymerase elongate a new DNA strand by adding deoxyribonucleotides, one at a time, to the 3' –end of the growing chain

• DNA polymerase III: catalyze DNA chain elongation. It remains bound to the template strands as it moves along. -The new strand grows in the 5' → 3‘, antiparallel to the parental strand -the nucleotide building blocks are 5‘deoxyribonucleoside triphosphate

Proofreading: the nucleotide sequence of DNA is replicated with as few errors as possible -misreading of the template sequence could result in deleterious, perhaps lethal, mutations -the proof reading activity requires an exonuclease that move 3‘→ 5' direction , not 5' → 3‘ like the polymerase activity, because excision must be done in the reverse direction from that of synthesis.

Excision of RNA primers and their replacement by DNA • DNA polymerase III continues to synthesize DNA on the lagging strand until it is blocked by proximity to an RNA primer. • DNA polymerase I has a 5' → 3‘ exonuclease activity that is able to hydrolytically remove the RNA primer and fill the gap (5' → 3‘ polymerase activity) • RNase H catalyzes this reaction in eukaryotes

DNA ligase • Catalyzes the final phosphodiester linkage between the 5'-phosphate group on the DNA chain synthesized by DNA polymerase III and the 3‘-hydroxyl group on the chain made by DNA polymerase I
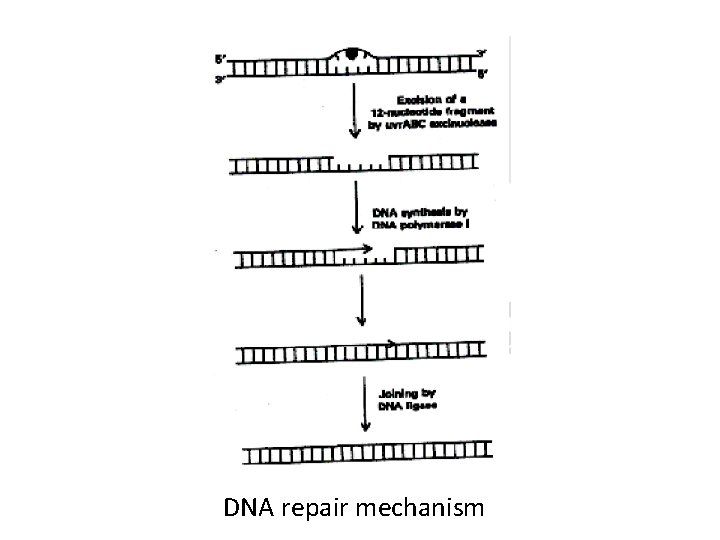
DNA repair mechanism

9. 5. Transcription • Transcription is the process of RNA Synthesis directed by a DNA template. • Following their synthesis, m. RNAs are translated into sequence of amino acids. r. RNA, t. RNA and additional small RNA molecules perform specialized structures and regulatory functions and are not translated.

• It occurs in three phases: 1. Initiation 2. Elongation 3. Termination 1. Initiation • Involves the binding of the RNA polymerase holoenzyme to a region on the DNA that determines the specificity of transcription of that particular gene (Promoter region).

• The left side of that particular point is called upstream. The promoter is found at this upstream. • The RNA polymerase is made up of σ- (sigma) subunit 2 α-and 2β-subunits. • Those that are recognized by RNA polymerase σ (sigma) factors include

Pribnow box: a stretch of 6 nucleotides (5'TATAAT- 3') centered about 8 -10 nucleotides to the left of the transcription start site - a base in the promoter region is assigned a negative number if it occurs prior to (“upstream” of) the transcription start site - -35 sequence: a second consensus nucleotide sequence (5'-TTGACA- 3‘), is centered about 35 bases to the left of the transcription start site

• In between the consensus and TATA box the base sequences are highly variable and it is this sequence that can be recognized by σ (sigma) subunit or RNA polymerase in prokaryotic cell and by transcription factor in eukaryotic cell. • To initiate transcription the sigma subunit of RNA polymerase recognizes the promoter site on DNA and separates the two strands.

• After the two strands of DNA are separated the RNA Polymerase starts the pairing of purine nucleotide. • RNA polymerase does not require a primer and has no known endonuclease or exonuclease activity.

2. Elongation • Once RNA synthesis is started, there occurs the step by step addition of ribonucleoside triphosphates (ATP, GTP, CTP and UTP) at 3’ – OH end of RNA and releases pyrophosphate each time a nucleotide is added. • The forward movement of RNA polymerase continuous until a termination signal is reached. • The area of DNA transcribed by polymerase rewinds back to the double helix.
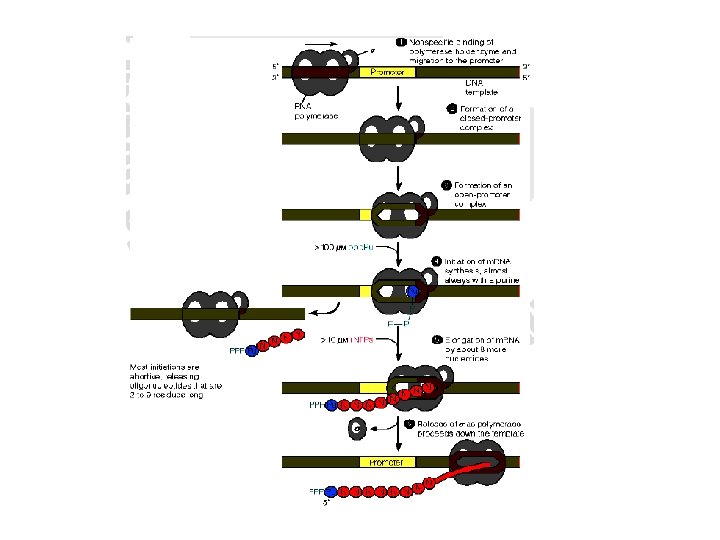

3. Termination • The process is carried out by two ways a. The RNA transcript must be able to form a stable hair pin turn that slows down the progress of RNA polymerase and causes it to pause temporarily (rho independent termination). b. An additional protein ρ (rho) factor may be required for the release of the RNA product (ρdependent termination).

• This factor binds to the C-rich region near the 3’end of the newly synthesized RNA, and migrates along behind the RNA polymerase in the 5' → 3‘ direction until the termination site is reached, then the factor displaces the DNA template strand, facilitating the dissociation of the RNA molecule.

• These two termination mechanisms are carried out in prokaryotic cells. But in eukaryotic cells termination may occur by transcription factors themselves.

9. 6. Translation • The genetic information which flows from DNA to m. RNA will be translated to the universal language called protein. • During transcription the base sequence of DNA determines the base sequence of m. RNA. • Under translation the base sequence of m. RNA determines the amino acid sequences in protein.

• The pathway translates the three-letter alphabet of nucleotide sequence on m. RNA into the twenty letter alphabet of amino acids that constitute proteins • Takes place in 3 stages: 1. Initiation 2. Elongation 3. Termination

1. Initiation of translation • There are two mechanisms by which the ribosome recognizes the nucleotide sequence that initiate translation a. Shine-Dalgarno sequence: (5’-UAGGAGG-3’) is located 6 to 10 bases upstream of the AUG codon on the m. RNA molecule, near its 5’-end. • The 16 S r. RNA component of the 30 S ribosomal subunit has a nucleotide sequence near its 3’end that is complementary to all or part of the Shine-Dalgarno sequence

• Eukaryotes do not have Shine-Dalgarno sequence. The 40 S ribosomal sub units binds to the cap structure of m. RNA and moves until it encounters the initiator AUG codon.

b. Initiation codon: the codon AUG is recognized by a special initiator t. RNA. The recognition is facilitated by IF-2 • In eukaryotes there at least 10 IF’s are known (e. IF)
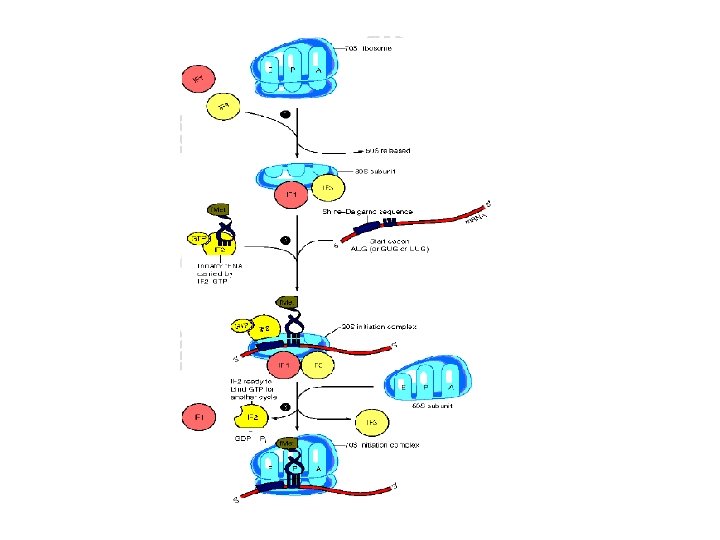

2. Elongation of protein chain • Involves the addition of amino acids to the carboxyl end of the growing chain • During elongation the ribosome moves from the 5’end to the-3’ end of the m. RNA that is being translated

• Delivery of the aminoacyl-t. RNA is facilitated by elongation factors ( EF-Tu, EF-Ts). • Elongation cycle takes place in 3 steps: a. Aminoacyl-t. RNA is delivered to A site b. Formation of peptide bond c. Translocation
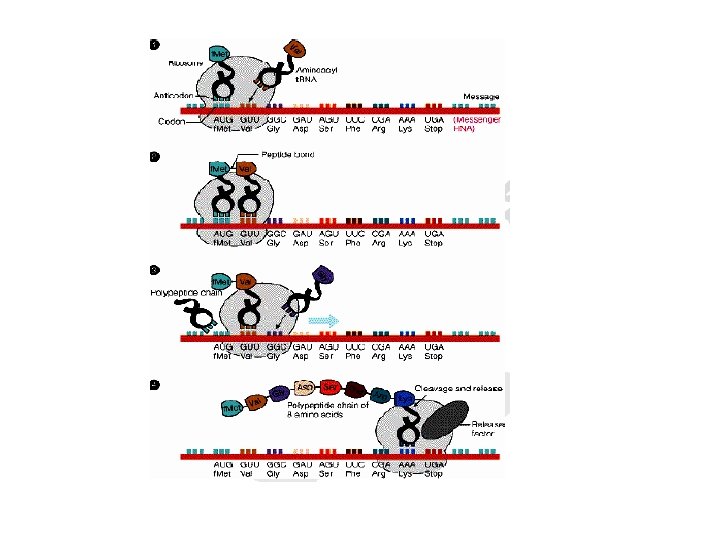

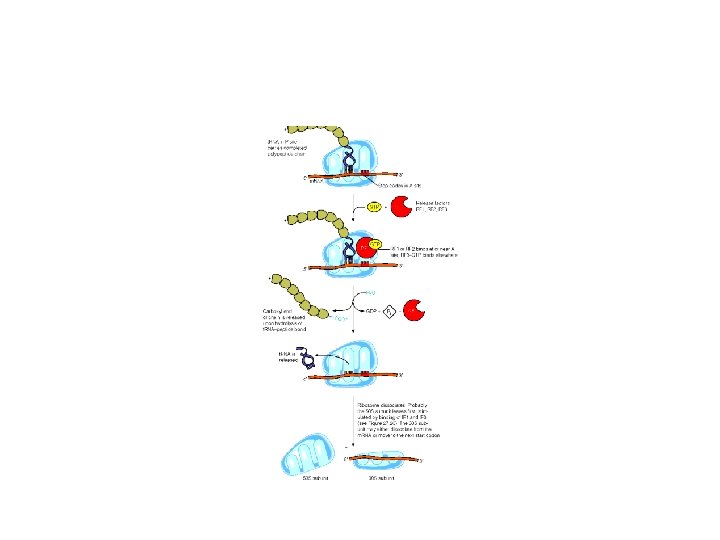
3. Termination stage • The movement of the ribosome consequently reaches to the termination site of m. RNA. • Termination occurs when one of the three stop codons moves into the A site • Release factor 1(RF-1) recognizes UAG and UAA; RF-2, UGA and UAA; RF-3, binds GTP and stimulates RF-1 and RF-2

• The factors cause the newly synthesized protein to be released and also dissociation of the ribosome from the m. RNA • No t. RNA has anticodons that pairs with the stop codons.
- Slides: 87