Chapter 9 Molecular Geometry Bonding Theories Molecular Shapes

Chapter 9 Molecular Geometry & Bonding Theories

Molecular Shapes Determined by bond angles; made by joining nuclei of atoms in a molecule Bond angles are generally symmetrical within a molecule (CH 4 = 109. 5 o) Basic Regents shapes: linear (180 o), trigonal planar (120 o), tetrahedral (109. 5 o), trigonal pyramidal (107 o), bent (104. 5 o) All shapes are “AB” molecules (central & peripheral atoms)
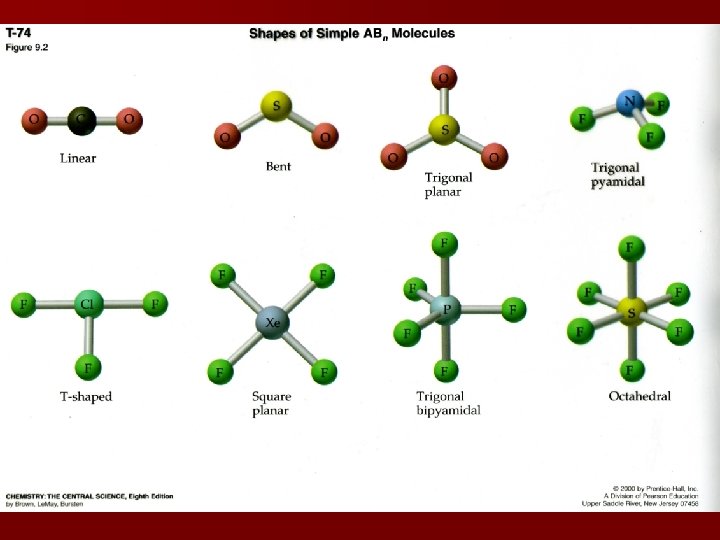

VSEPR Model Valence Shell Electron Pair Repulsion theory creates shapes and bond angles based on the premise that electrons will repel each other within the limited space they are allowed to occupy inside an atom Bonding pair: area most likely to find electrons Lone pair: electron domain located solely on one atom Electron domains will be arranged to minimize repulsive forces

Electron Domain Geometry The geometry presented by both bonded and lone pairs inside an atom Water has a tetrahedral electron domain geometry as it contains 2 bonds and 2 lone pairs


Molecular Geometry The geometry presented by only the atoms and the bonded pairs of a molecule Water has a bent molecular geometry because it has two bonded pairs and two lone pairs

Predicting Molecular Geometry 1. 2. 3. 4. 5. Sketch the Lewis structure Count electron domains on central atom Arrange them to minimize repulsions Describe molecular geometry in terms of angular arrangement of bonded atoms Multiple bonds are considered one domain


Predict both electron domain & molecular geometries: Sn. Cl 31 - O 3

Other Effects: Lone Pairs Lone pairs force remaining bonds closer together More repulsive force because electrons are closer to the nucleus than electrons attracted to two nuclei Bond angle decreases as lone pairs are added CH 4>NH 3>H 2 O (109. 5>107>104. 5)

Other Effects: Multiple Bonds Multiple bonds are more repulsive than single bonds Higher electron density causes other bonds to move closer together; decreases bond angle COCl 2 shows 111 o angle between Cl-C-Cl bond, not the expected 120 o

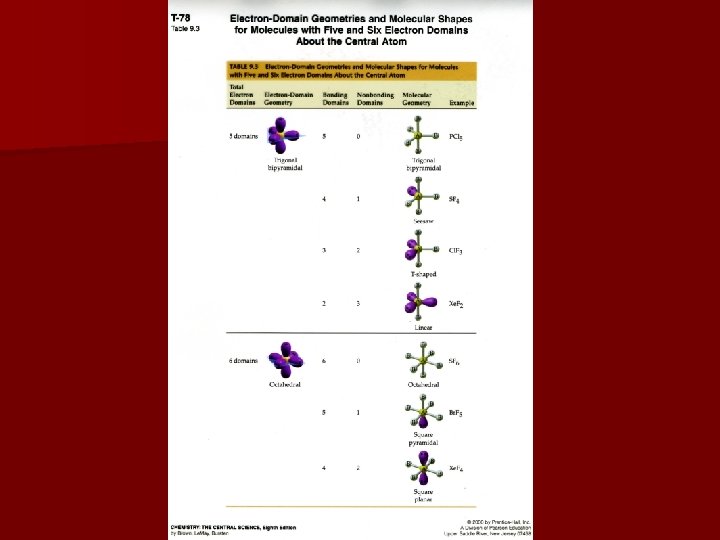
Shapes & Expanded Valence Greater than 4 electron domains (typically 5 or 6); noted by axial or equatorial position 5 domains = trigonal bipyramidal 6 domains = octahedral When placing lone pairs, minimize repulsions equatorial is better in bipyramidal geometry; octahedral all positions are symmetrical

Variations of Expanded Shapes Bipyramidal: seesaw – 1 equatorial lone pair t-shaped - 2 equatorial lone pairs linear - 3 equatorial lone pairs Octahedral: square pyramidal – one axial lone pair square planar – two axial lone pairs

Complex Compounds When you have more than one central atom, determine geometry for each “central atom” Example: amino acid

Polyatomic Molecules & Polarity Bond dipole: dipole moment of only 2 atoms in a bond Overall dipole moment: sum of the magnitude and directions of individual bond dipoles (vector-like) CO 2 vs H 2 O When symmetrically surrounded by identical atoms, the overall dipole moment is zero

Valence Bond Theory Combines electron pair bonds and atomic orbitals Atomic orbitals overlap; 2 orbitals share a high electron density in the same region in space Occurs when electrons occupy space between 2 atoms by spinning in opposite directions (Pauli – ms + ½ or – ½ ) Optimum overlap distance can be graphically shown – too far = no attraction; too close = nuclei repel; optimum distance =lowest energy state, most stable

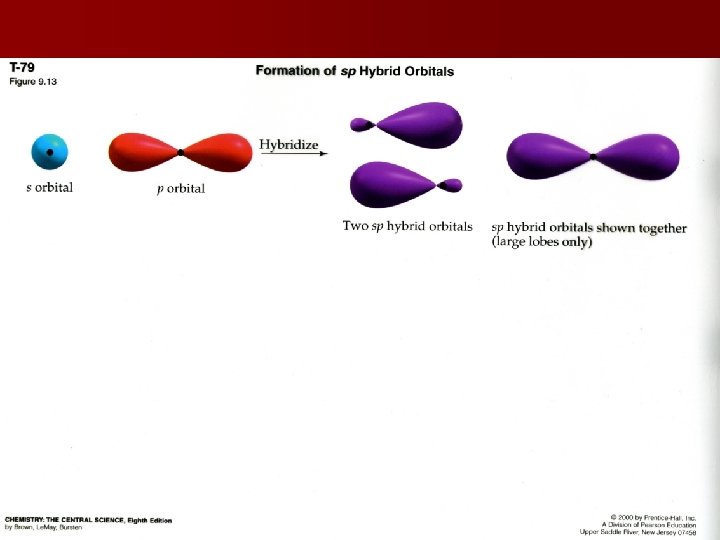
Hybridization Mixing 2 orbitals to result in equal energies Ex: Be. F 2 is covalent in liquid or gaseous state, but orbital notation shows no available bonding electrons One electron is promoted from the 2 s to the 2 p in order to create 2 sp hybrid orbitals with equal energy Energy used in promotion of electrons is recouped in the formation of covalent bonds

Hybridization & Shape Valence bond model equates: sp hybrids with a linear shape sp 2 hybrids with a trigonal planar shape sp 3 hybrids with tetrahedral geometries sp 3 d hybrids with bipyramidal geometries sp 3 d 2 hybrids with octahedral geometries Last two hybridizations occur in the 3 rd period elements or higher
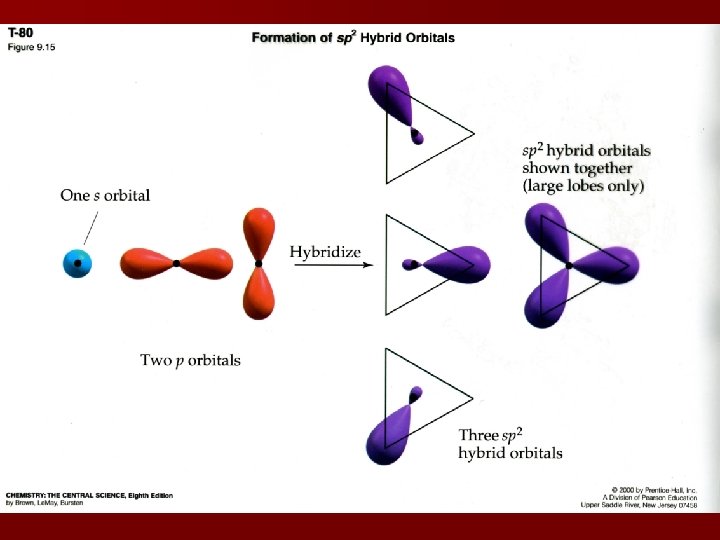
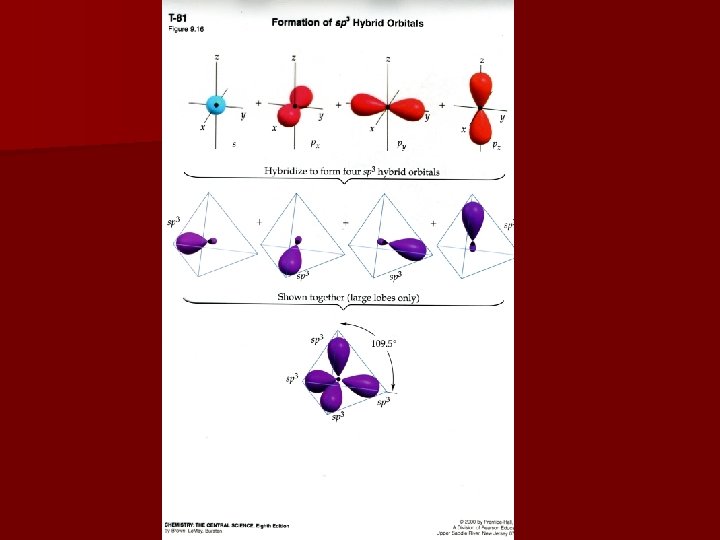
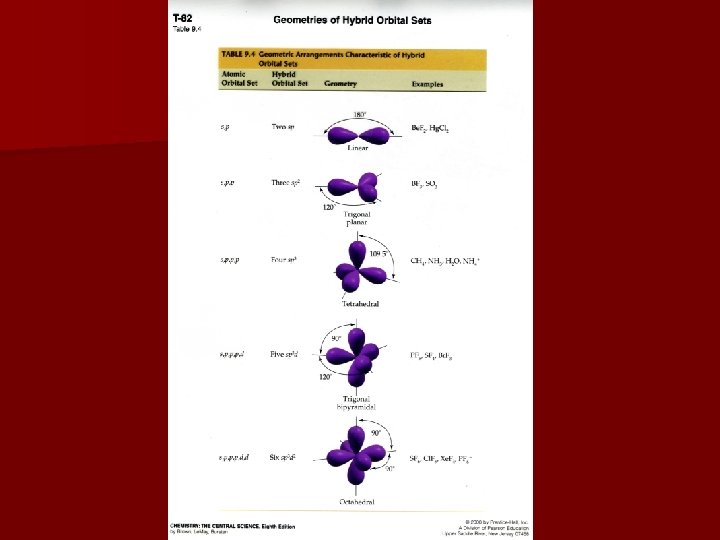

Predicting Hybridization Draw Lewis structure 2. Determine electron domain geometry with the VSEPR model 3. Specify hybrid orbitals needed to accommodate the electron domain geometry 1.

Multiple Bonds s bond: electron density lies on the internuclear axis p bond: electron density lies above and below the internuclear axis Single bonds = one s bond Double bonds = one s bond + one p bond Triple bonds = one s bond + two p bonds Refer to packet for visual representation
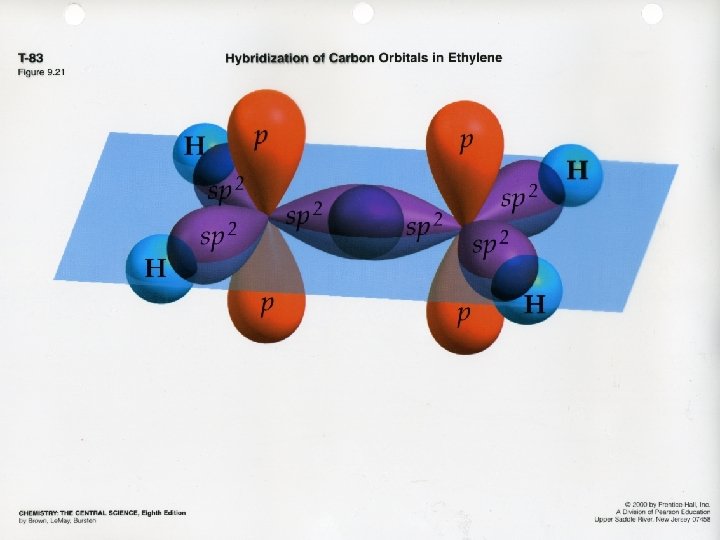
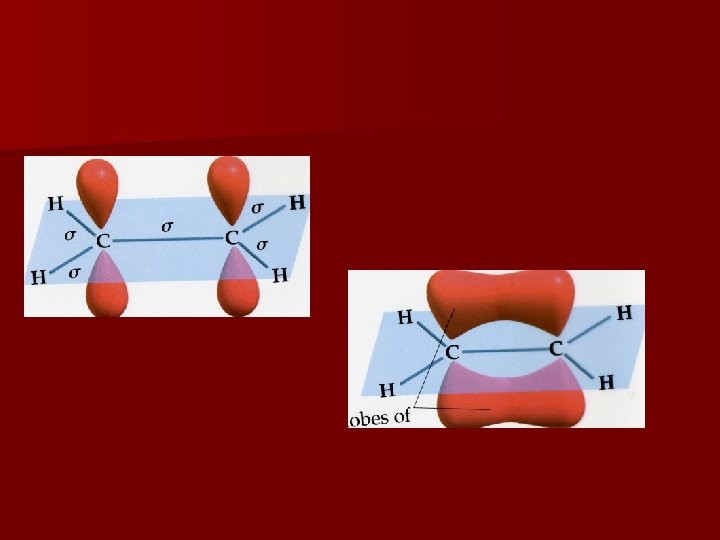

Delocalization of p Bonds Localized electrons are associated only with the two atoms that share the bond Delocalized electrons are associated with more than the 2 atoms in the bond (resonance structures) Examples: C 6 H 6 O 3 NO 3 -
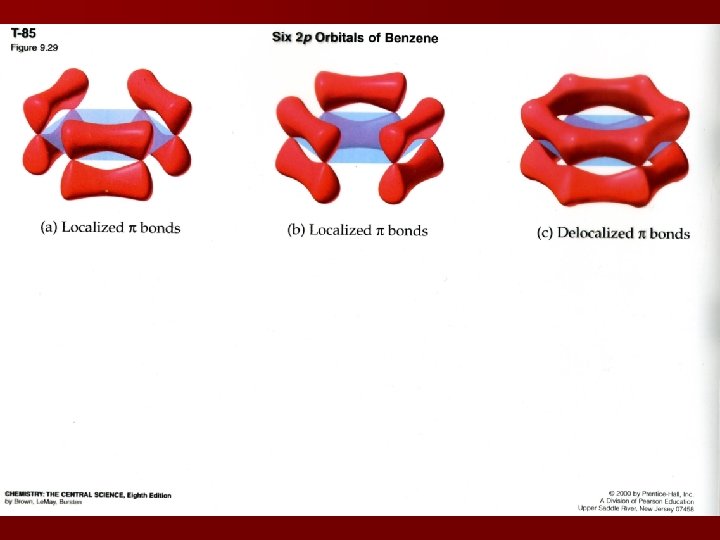

Isomers Bond rotation is free around a s bond Bond rotation is restricted around a p bond Rotation around a p bond is endothermic because it destroys the p bond

Bond Isomers Cis isomer: 2 groups located on the same side of a multiple bond or ring (octahedral) Trans isomer: 2 groups located on the opposite side of a multiple bond or ring (octahedral) Optical isomer: structures differing in a 3 -d relationship to each other (mirror images – like L/R hands); a form of stereoisomerism Enzymes can distinguish different optical isomers, show preference for one form or another Optical isomers also respond differently to polarized light

Molecular Orbital Theory Describes electrons in molecules through wave functions (Schrodinger & y 2) Molecular orbital (MO): specific wave function for an electron in a molecule MOs can hold 2 electrons max, have a definite amount of energy, can be represented by a contour drawing, and are associated with the entire molecule, not simply one atom

Overlapping MOs 2 atomic orbitals (ex: 1 s) overlapping (bonded) form 2 molecular orbitals: Bonding MO: electron density concentrated between 2 atomic nuclei (more stable than 1 s) Antibonding MO: electron density concentrated outside of nuclei; equal to zero between nuclei, less stable than 1 s orbital (higher energy state)
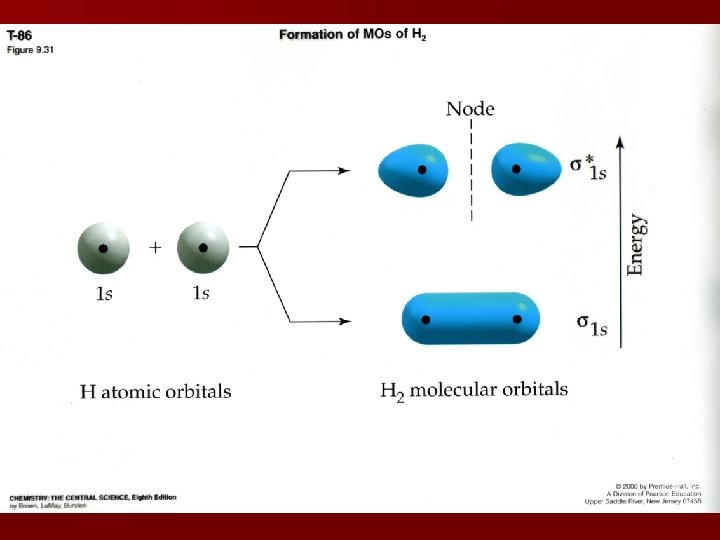
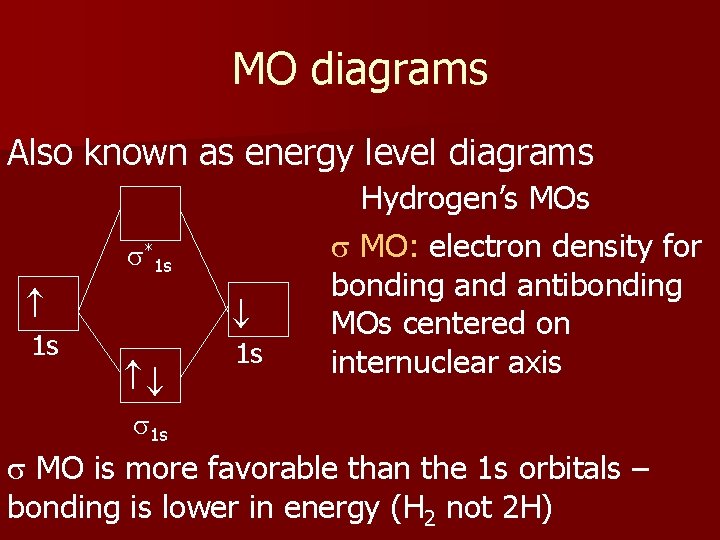
MO diagrams Also known as energy level diagrams s*1 s 1 s 1 s Hydrogen’s MOs s MO: electron density for bonding and antibonding MOs centered on internuclear axis s 1 s s MO is more favorable than the 1 s orbitals – bonding is lower in energy (H 2 not 2 H)

Helium’s MOs s*1 s s 1 s Antibonding MOs are more unfavorable than bonding MOs are favorable – He 2 is not more stable than 2 He

Bond Order A method of determining the level of attraction between 2 atoms BO = ½ (#bonding e - #antibonding e) 0 = no bond 1 = single bond 2 = double bond 3 = triple bond 1/2, 3/2, 5/2 are possible (He 2+ has a bond order of ½ and will be stable in its formation)

Other MOs Second row diatomic molecules possess more than one type of MO Rules for MO formation: 1. #MOs formed = #AOs combined 2. AOs combine most effectively with atomic orbtials of similar energy 3. Effectiveness of combination of AOs is proportional to the degree of overlap (more overlap = bonding MO and antibonding MO) 4. Each orbital has 2 e max w/ paired spins 5. When MOs are degenerate, follow Hund’s rule

Rules for Multiple MOs in Li 2 s*1 s is lower than s 2 s s 1 s is nullified by s*1 s therefore the bond is generated by s 2 s orbitals Bond order = 1/2 (4 bonding – 2 anti) = 1 Predicts a single bond For Be 2, s 1 s, s*1 s, s 2 s, and s*2 s are filled, therefore the bond order = 0 and no bond is formed

2 p MOs p MO: the electron density is concentrated outside the internuclear axis 2 degenerate p 2 p MO and 2 degenerate p*2 p MOs There is less overlap between the p 2 p MOs than in the s 2 p MOs s 2 p < p*2 p < s*2 p Ch 9 Packet: p 20 – you should be able to sketch these representations!
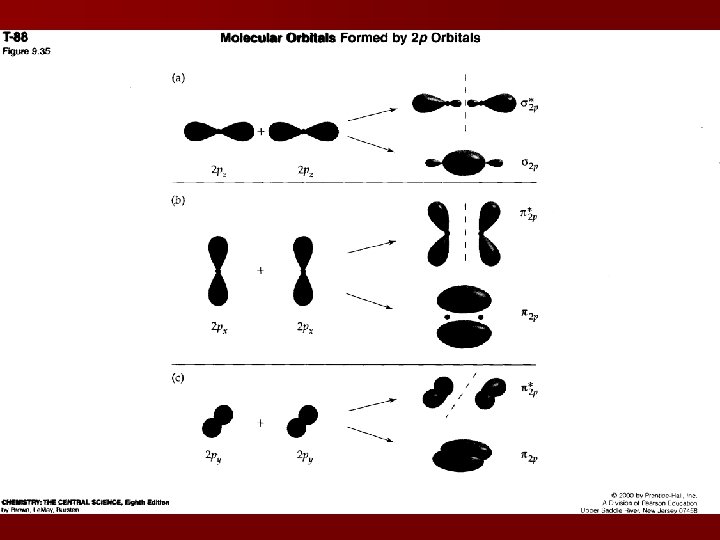

2 s-2 p interaction Interaction between the sublevels causes s 2 s MO to decrease in energy and a s 2 p MO to increase in energy New order when this occurs: s 2 s < s*2 s < p 2 p < s 2 p < p*2 p < s*2 p Occurs in B 2, C 2, N 2

Magnetic Interaction Paramagnetic: 1 or more unpaired electrons; will be attracted into a magnetic field Diamagnetic: no unpaired electrons; will be weakly repelled from a magnetic field HOMO: highest occupied molecular orbital LUMO: lowest unoccupied molecular orbital

HOMO-LUMO Gap & Color HOMO-LUMO gap is the energy difference between the HOMO and the LUMO; corresponds to the minimum energy needed to excite an electron in a molecule Colorless or white substances have a very large HOMO-LUMO gap; visible light does not have enough energy to excite the electrons to the higher level

Conjugated p bonds When p orbitals are arranged so that electrons may be delocalized over the entire molecule, bonds are said to be conjugated (alternating single and double bonds over the majority of the molecule) Increased conjugation decreases the HOMO -LUMO gap in molecules Most compounds that exhibit visible colors are large organic compounds with many conjugated double bonds
- Slides: 46