Chapter 6 Microbial Metabolism Energy Metabolism Special Metabolism

Chapter 6 Microbial Metabolism • Energy Metabolism • Special Metabolism in Microbes • The Relationship between Catabolism and Anabolism • Regulation of Metabolism and Ferment Industry

An Overview metabolism: the sum total of all chemical reactions occurring in the cell catabolism metabolism anabolism catabolism Complex molecules Simple molecules anabolism ATP [H]

Section 1 Energy Metabolism in Microbes summarize Organic Compounds Primary Energy Sunlight Chemoheterotroph Photoautotroph Inorganic Compounds in Reduced State A T P ATP Chemoautotroph

Chemoheterotroph biological oxidation and energy Release Process——Dehydrogenation, Giving Hydrogen and Accepting Hydrogen (Electron) Function——Releasing Energy (ATP), Engendering Reducing Power [H] and Producing Intermediate Metabolites The breakdown of glucose to pyruvate biological oxidation fermentation respiration (aerobic or anaerobic respiration )


1 The breakdown of glucose to pyruvate Glocose——representative substrate of biological oxidation Embden-Meyerhof-Parnas Pathway (Glycolysis) Hexose Monophosphate Pathway Entner-Doudoroff Pathway (KDPG Pathway) PK (phosphoketolase) pathway

1)Embden-Meyerhof-Parnas Pathway (EMP) (Glycolysis, Hexose Diphosphate Pathway) glucose Ten steps pyruvate with O 2 :connecte EMP pathway with TCA pathway; without O 2:reduce some metabolism product, only energyyielding process. generates ATP by substrate-level phosphorylation: (1) glyceraldehyde 1, 3 -phosphate 3 -phoshoglyceric acid + ATP; (2) PEP pyruvate + ATP C 6 H 12 O 6+2 NAD++2 ADP+2 Pi→ 2 CH 3 COCOOH+2 NADH+2 H++2 ATP+2 H 2 O
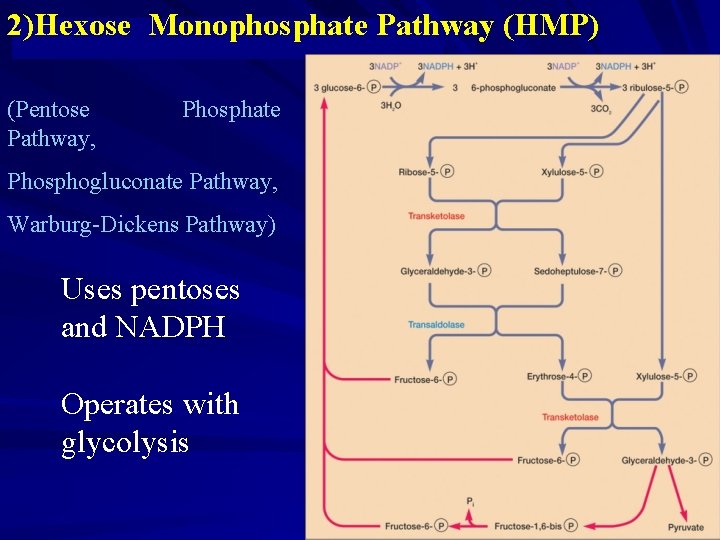
2)Hexose Monophosphate Pathway (HMP) (Pentose Pathway, Phosphate Phosphogluconate Pathway, Warburg-Dickens Pathway) Uses pentoses and NADPH Operates with glycolysis

fructose 6 -phosphates: be converted to glucose 6 -phosphates ,be returned to Pentose Phosphate Pathway glyceraldehyde 3 -phosphate : a. through EMP pthway, be converted to pyruvate,into TCA pthway b. converted to Hexose Phosphate, be returned to Pentose Phosphate Pathway The overall reaction: 6 glucose 6 -phosphates +12 NADP++3 H 2 O → 5 glucose 6 phosphates + 6 CO 2+12 NADPH+12 H++Pi

3) Entner-Doudoroff Pathway (KDPG Pathway) 1952, Entner-Doudoroff :Pseudomonas saccharophila process: (4 septs) 1 Glucose glucose 6 -phosphates 6 -phosphogluconate dehydratase KDPG 2 -oxo-3 -deoxy-6 -phosphogluconate aldolase glyceraldehyde 3 -phosphate + pyruvate Produces NADPH and 1 ATP Does not involve glycolysis Pseudomonas, Rhizobium, Agrobacterium
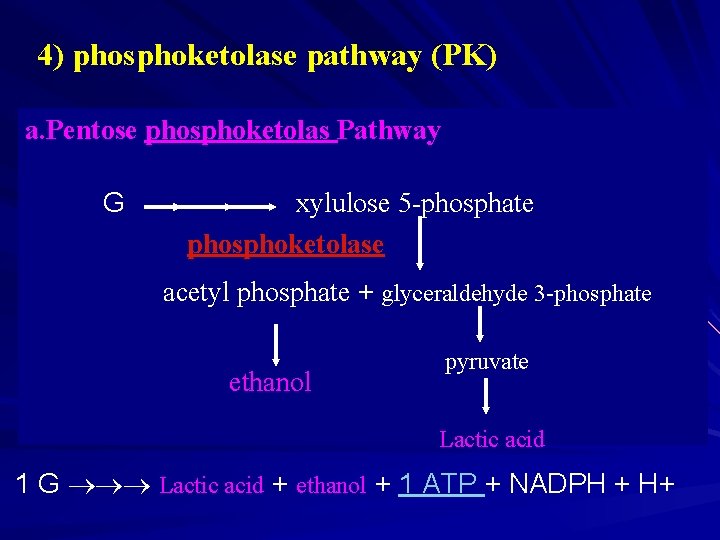
4) phosphoketolase pathway (PK) a. Pentose phosphoketolas Pathway a. G xylulose 5 -phosphate phosphoketolase acetyl phosphate + glyceraldehyde 3 -phosphate ethanol pyruvate Lactic acid 1 G Lactic acid + ethanol + 1 ATP + NADPH + H+
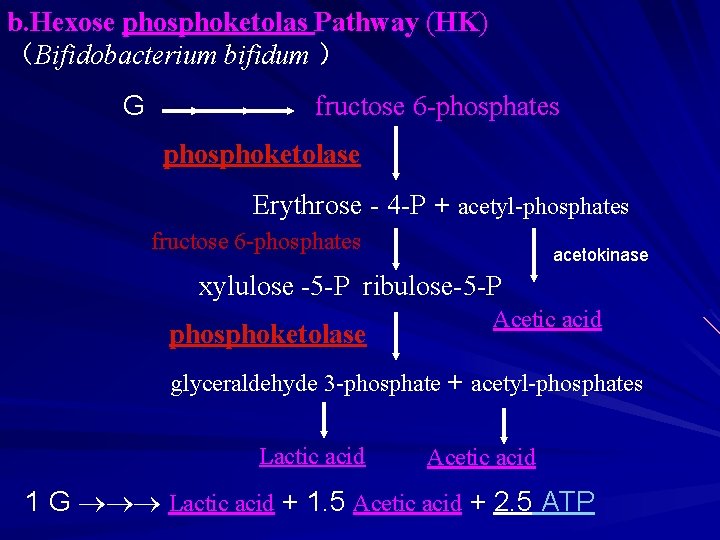
b. Hexose phosphoketolas Pathway (HK) (Bifidobacterium bifidum ) G fructose 6 -phosphates phosphoketolase Erythrose - 4 -P + acetyl-phosphates fructose 6 -phosphates acetokinase xylulose -5 -P ribulose-5 -P phosphoketolase Acetic acid glyceraldehyde 3 -phosphate + acetyl-phosphates Lactic acid Acetic acid 1 G Lactic acid + 1. 5 Acetic acid + 2. 5 ATP

2 fermentantion 1. definitions broader :use microorganism to produce useful metabolic product narrower:under anaerobic conditions, be defined as an energyyielding process using itself metabolic intermediates as the final hydrogen (electron) accepter organic molecules serve as both electron donors and acceptors. trait: 1) generates ATP by substrate-level phosphorylation; 2)the glucose is partially oxidized ,mostly energy in fermention products ; 3)lower energy; 4) generates many kinds of fermention products

2. fermentation sorts 1) alcohol fermentation a. yeasts 1 G (EMP) 2 pyruvate 2 aldehyde + CO 2 2 ethanol + 2 ATP condiction:p. H 3. 5~4. 5 , without O 2 Strain:Saccharomyces cerevisiae, few bacteria(Erwinia amylovora, Sarcine vintriculi) I. Add Na. HSO 4 + aldehyde sulfonic hydroxy aldehyde ii. weak basic(p. H 7. 5) 2 aldehyde 1 acetate + 1 ethanol glycerol fermentation : dihydroxyacetone as hydrogen accepter, hydrolyzed to glycerol

b. bacteria (Zymomonasmobili, Pseudomonas saccharophila ) homoalcohol fermentation homo 1 G 2 pyruvate ethanol + 1 ATP (ED) heteroalcohol fermentation (Thermoanaerobacter ethanolicu) 1 G 2 pyruvate ( Pyruvate formate lyase ) formic acids + acetyl-Co. A Without Pyruvatedecarboxylate With aldehyde dehydrogenase aldehyde ethanol

2)Lactic acid fermentation Homolactate fermentation For example, Lactobacillus delbruckii, Streptococcus faecalis —— EMP pathway(pyruvate lactate ) Heterolactate fermentation (PK pathway) Leuconostoc mesenteroides (PK) generates energy: 1 ATP Bifidobacterium bifidum (PK、HK) generates energy: 2 G 5 ATP, 1 G 2. 5 ATP
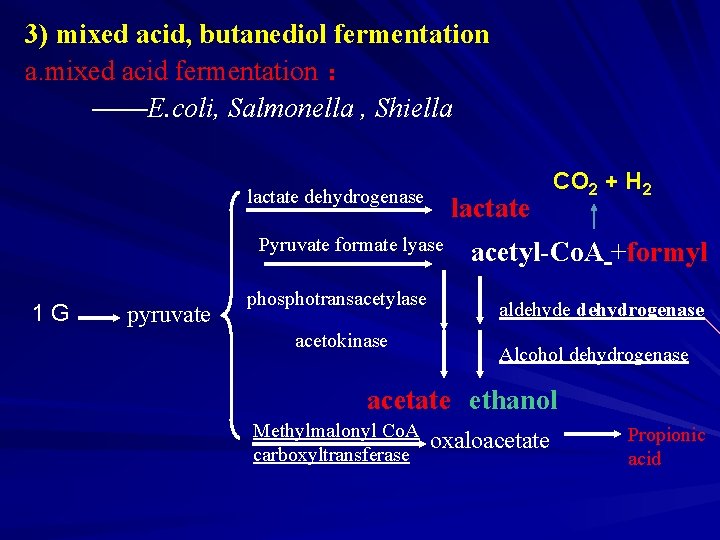
3) mixed acid, butanediol fermentation a. mixed acid fermentation : ——E. coli, Salmonella , Shiella lactate dehydrogenase lactate Pyruvate formate lyase 1 G pyruvate phosphotransacetylase acetokinase CO 2 + H 2 acetyl-Co. A +formyl aldehyde dehydrogenase Alcohol dehydrogenase acetate ethanol Methylmalonyl Co. A carboxyltransferase oxaloacetate Propionic acid

b. butanediol fermentation —— Enterobacter, Serratia pyruvate (acetolactate V. P. test: Red substance dehydrogenase) 3 -hydroxy butanone (OH-、O 2) diacetyl butanediol Two important reaction: 1. V. P. test 2. methylene red (M. R) test Enterobacter aerogenes: methylene red (-) E. coli: V. P. (-), methylene red (+) neutral
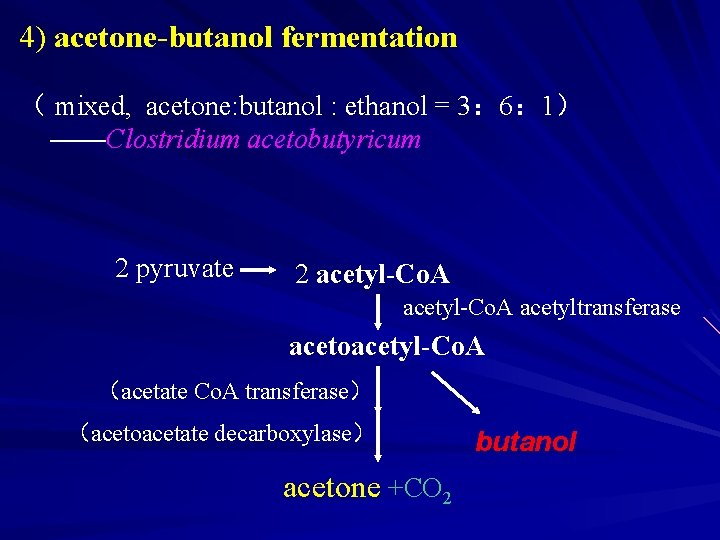
4) acetone-butanol fermentation acetone( mixed, acetone: butanol : ethanol = 3: 6: 1) acetone: ——Clostridium acetobutyricum 2 pyruvate 2 acetyl-Co. A acetyltransferase acetoacetyl-Co. A (acetate Co. A transferase) (acetoacetate decarboxylase) acetone +CO 2 butanol

5)stickland reaction Main point:amino acid oxidation couples with other amino acids reduction, generates 1 ATP hydrogen donors (oxidation ) amino acid: Ala, Leu, Ile, Val, His, Ser, Phe, Tyr, Try hydrogen acceptors (reduction) amino acid: Gly, Pro, Arg, Met, Leo, and so on.

The Stickland reaction is used to oxidize several amino acids: alanine, leucine, isoleucine, valine, phenylalanine, tryptophan, and histidine. Bacteria also ferment amino acids (e. g. , alanine, glycine, glutamate, threonine, and arginine) by other mechanisms.
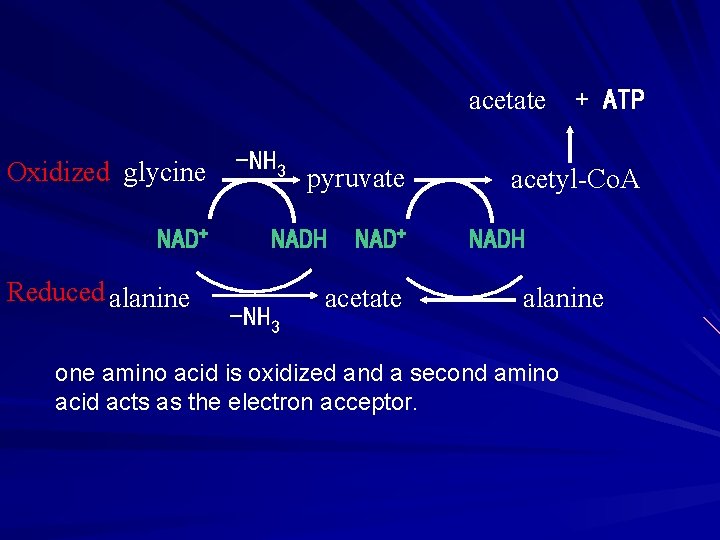
acetate Oxidized glycine NAD+ Reduced alanine -NH 3 pyruvate NADH -NH 3 NAD+ acetate + ATP acetyl-Co. A NADH alanine one amino acid is oxidized and a second amino acid acts as the electron acceptor.

3 Respiration aerobic respiration anaerobic respiration 1. Aerobic Respiration: The final electron acceptor in the electron transport chain is molecular oxygen (O 2).
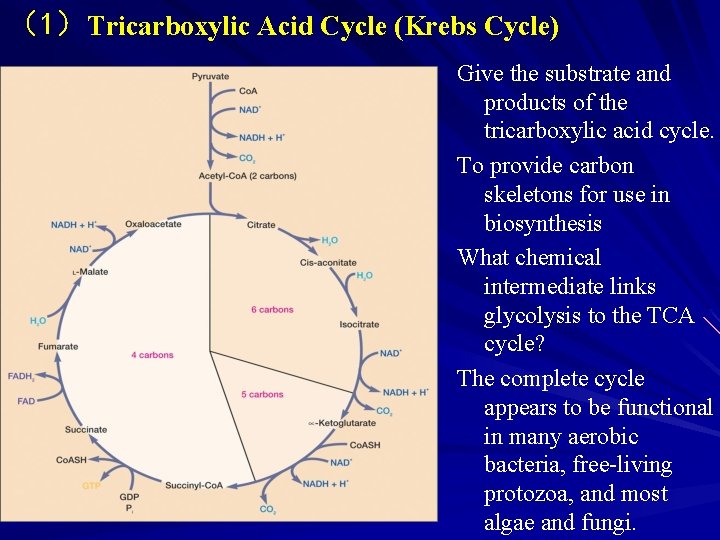
(1)Tricarboxylic Acid Cycle (Krebs Cycle) Give the substrate and products of the tricarboxylic acid cycle. To provide carbon skeletons for use in biosynthesis What chemical intermediate links glycolysis to the TCA cycle? The complete cycle appears to be functional in many aerobic bacteria, free-living protozoa, and most algae and fungi.

(2)The Electron Transport Chain Some important electron transport chain carriers of the respiration chain in microbes Nicotianamide adenine dinucleotide (NAD) and nicotianamide adenine dinucleotide phosophate (NADP) Flavin adenine dinucleotide (FAD) and flavin mononucleotide (FMN) Iron-sulphur protein Ubiquinone (Coenzyme Q) Cytochrome system
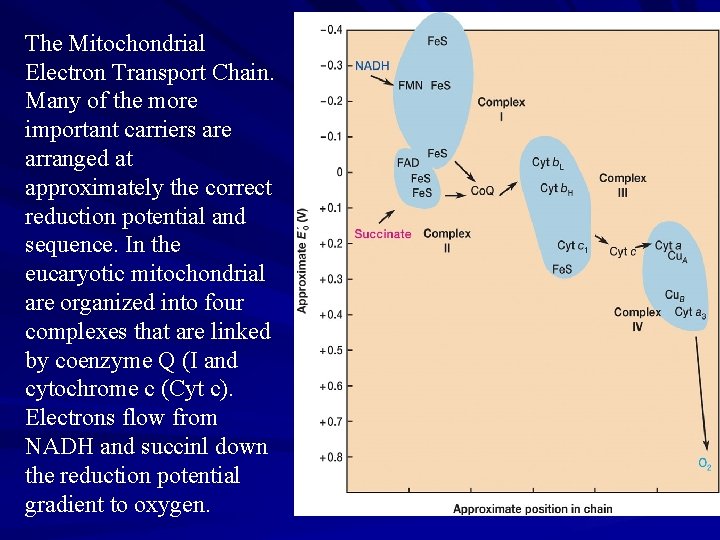
The Mitochondrial Electron Transport Chain. Many of the more important carriers are arranged at approximately the correct reduction potential and sequence. In the eucaryotic mitochondrial are organized into four complexes that are linked by coenzyme Q (I and cytochrome c (Cyt c). Electrons flow from NADH and succinl down the reduction potential gradient to oxygen.

The electron transport chain in Prokaryote Main outline: electron accepter multiplicity:O 2, NO 3 -, NO 2 -, NO-, SO 42 -, S 2 -, CO 32 - et al; Electron donors: H 2, S, Fe 2+, NH 4+, NO 2 -, G, other orgnisim et al; various cytochrome: a, a 1, a 2, a 4, b, b 1, c, c 1, c 4, c 5, d, o; Terminal oxidase: cyt a 1, a 2, a 3, d, o,catalase, peroxid enzyme; Respiration Chain component variable, being branched respiration chain : Bacterial chains also may be shorter and have lower P/O ratios than mitochondrial transport chains, from the several position of electron transport chain into and off by Terminal oxidase in several position. E. coli (absent O 2) Co. Q cyt. b 556 cyt. o cyt. b 558 cyt. d The cytochromes 0 branch has moderately high affinity for oxygen,is a proton pump,and operates at higher oxygen concentrations; The cytochromes d branch has very high affinity for oxygen and functions at low oxygen levels。 electron transport multiplicity.

(3)Oxidative phosphorylqtion Chemiosmotic hypothesis-first formulated in 1961 by the British biochemist Peter.Mitchell. According to the Chemiosmotic hypothesis, the electron transport chain is organized so that protons move outward from the mitochondrial matrix and electrons are transported inward. Protons movement may result either from carrier loops, as shown in figure 9. 14, or from the action of special proton pumps that derive their energy from electron transport. The result is proton motive force(PMF), composed of a gradient of protons and a membrane porential due to the unequal distribution of charges. When protons return to the mitochondrial matrix driven by the proton motive force, ATP is synthesized in a reversal of the ATP hydrolysis reaction.

2. Anaerobic Respiration The final electron acceptor in the electron transport chain is not O 2. Yields less energy than aerobic respiration because only part of the Krebs cycles operations under anaerobic conditions. Nitrate Respiration( Denitrification) Nitrate reduction outline: a. have completely Respiration system; b. in the absence of O 2 only, nitrate reductase A and nitrous acid reductase needing for Denitrification are induced. c. facultative anaerobic bacteria: Bacillus licheniformis, Paracoccus denitrificans, Pseudomonas aeruginosa and so on.

Nitrate Respiration Homotype: NO 3 - NH 3 - N R - NH 2 Hetertype: in the absence of O 2 ,use NO 3 -as the final hydrongen accepter Facultative anaerobe (viz denitrify bacteriua) NO 3 - Na. R denitrification: NO 2 NO N 2 O N 2 Ni. R NOR N 2 OR 1) make N(NO 3 -) in soil reduced into N 2,and disappear, reduce edaphic fertility; 2) Denitrification has the importance action in nitrogen cycle.

Sulfate Respiration(sulfate reduction) ——in the absent of O 2, SO 42 -, SO 32 -, S 2 O 32 - as the final electron accepter outline: a. obligate anaerobic bacteria ; b. mostly ancient bacteria c. mostil obligate chemoheterotroph,few mixed; d. end product: H 2 S; SO 42 - SO 32 - SO 2 S H 2 S e. use organic nutrients (organic acid, fatty acid, 醇类)as hydrogen donors or electron donors ; f. environment: contain SO 42 - , anaerobic environment(soil, seawater) For example, Desulfovibrio desulfuricans, D. Gigas, Desulfotomaculum nigrificans and so on.

Sulfur Respiration ( Sulfur reduction) —— use element Sulfur as the final electron accepter only. electron donors:aceticm acid, small peptide, glucose, carbohydrate polymers; For example, Desulfuromonas acetoxidans Carbonate Respiration( Carbonate reduction ) —— use CO 2、HCO 3 - as the final electron accepter methane-producing bacteria: —— use H 2 as electron donors (energy resources), CO 2 as accepter,produce CH 4; Producing acetic acid bacteria — H 2 / CO 2 carry out Anaerobic Respiration ,product acetic acid

other anaerobic respiration —— use Fe 3+, Mn 2+ , many organic oxide as the final electron accepter fumaric acid succinate + 1 ATP For example, Escherichia, Proteus, Salmonella, Klebsiella in some facultative anaerobic bacteria, Bacteroides, Propionibacterium, Vibrio succinogenes in some anaerobic bacteria. Use Desulfotomaculum auripigmentum reduces As. O 43 into As 2 S 3

Section 2 special metabolism in microbes 1 Bacterial Photosynthesis 1. Cyclic photophosphorylation 2. Noncyclic photophosphorylation 3. Photosynthesis of purple membrane in halophilic bacteria
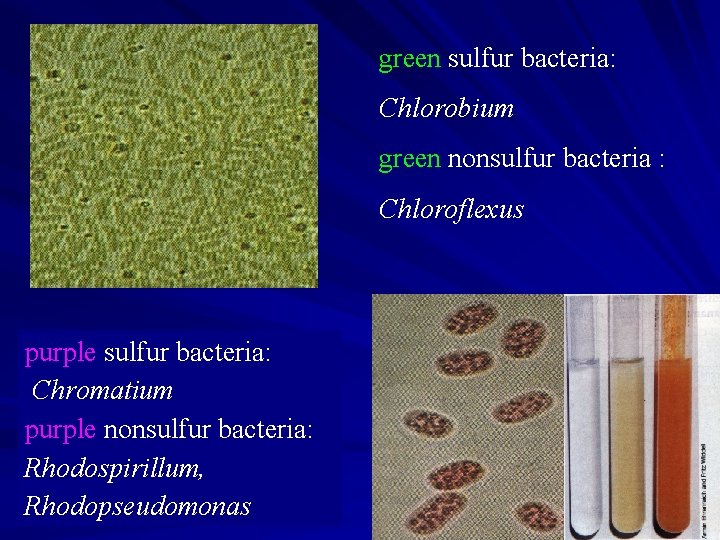
green sulfur bacteria: Chlorobium green nonsulfur bacteria : Chloroflexus purple sulfur bacteria: Chromatium purple nonsulfur bacteria: Rhodospirillum, Rhodopseudomonas

Purple Nonsulfur Bacterial Photosynthesis. Cyclic photophosphorylation The photosynthetic electron transport system in the purple nonsulfur bacterium, Rhodobacter sphaeroides. This scheme is incomplete and tentative. Ubiquinone (Q) is very similar to coenzyme Q. BPh stands bacteriopheophytin. NAD+ and the electron source succinate are in color.

Green Sulfur Bacterial Photosynthesis. Light energy is used to make ATP by cyclic photophosphorylation move electrons from sulfur donors (green and blue) to NAD+ (red). The electron transport chain has a quinone called menaquinone (MK). The photosynthetic electron transport system in the green sulfur bacterium, Chlorobium limicola.

2)noncyelic photophosphorylation Electrons also can travel in a noncyclic pathway involving both photosystems. P 700 is excited and donates electrons to ferredoxin as before. In the noncyclic route, however, reduced ferredoxin reduces NADP+ to NADPH. Because the electrons contributed to NADP+ cannot be used to reduce oxidized P 700, photosystem II participation is required. It donates electrons to oxidized P 700 and generates ATP in the process. The photosystem II antenna absorbs light energy and excites P 680, which then reduces pheophytin a. Pheophytin a is chlorophyll a in which two hydrogen atoms have replaced the central magnesium. Electrons subsequently travel to Q (probably a plastoquinone) and down the electron transport chain to P 700. Oxidized P 680 then obtains an electron from the oxidation of water to O 2.
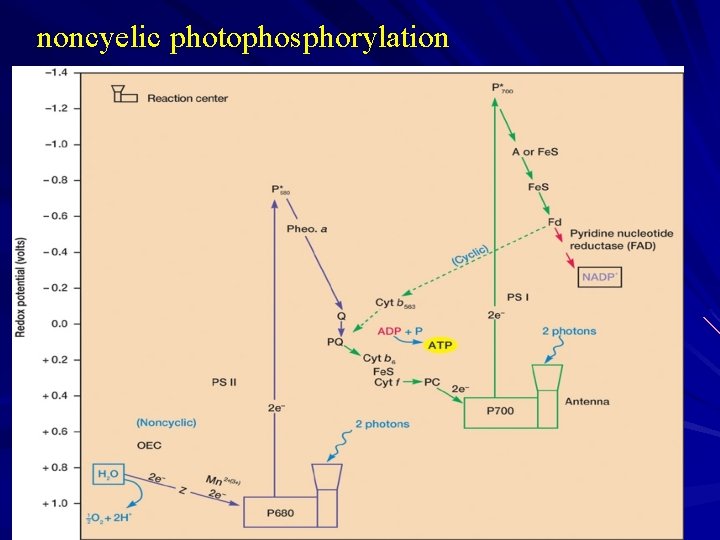
noncyelic photophosphorylation

Outline : electrons flow from water all the way to NADP with the aid of energy from two photosystems, ATP is synthesized by noncyelic photophosphorylation. one ATP and one NADPH are formed when two electrons travel through noncyclic pathway.


3) Photosynthesis of purple membrane in halophilic bacteria Halobacterium uses bacteriorhodopsin, not chlorophyll, to generate electrons for a chemiosmotic proton pump.
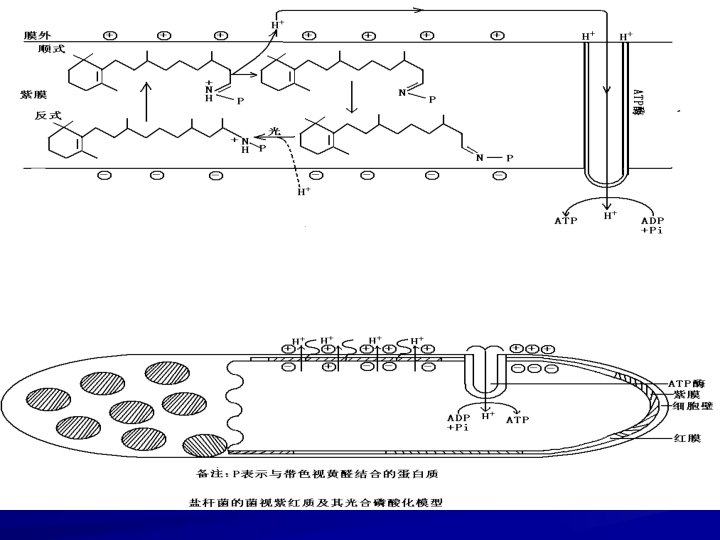

2 Chemolithotroph Biological Oxidation, Energy Release and CO 2 Fixation in Chemoautotroph aerobic Oxidation of inorganic electron donors generates ATP by Oxidative Phosphorylation electron donors: donors H, reducing nitride, reducing sulphide and Fe 2+. Use CO 2 Fixation of Calvin cycle as carbon resources CO 2 reduced to [CH 2 O] ——consume much energy and reducing power

1. Energy metabolism of nitribacteria Oxidation Nitrobacter: NH 3 NO 2 Oxidation Nitrosomonas: NO 2 - NO 3 - When two genera such Nitrobacter and Nitrosomonas together in a niche, ammonia is grow converted to nitrate, a process called nitrification

2. Hydrogen oxidizing bacteria Main strains: Alcaligenes, Flavobacterium Aquaspirillum Mycobacterium Nocardia and so on. generates energy: 2 H 2 + O 2 — 2 H 2 O synthesize reaction: 2 H 2 + CO 2 — [ CH 2 O ] + H 2 O。
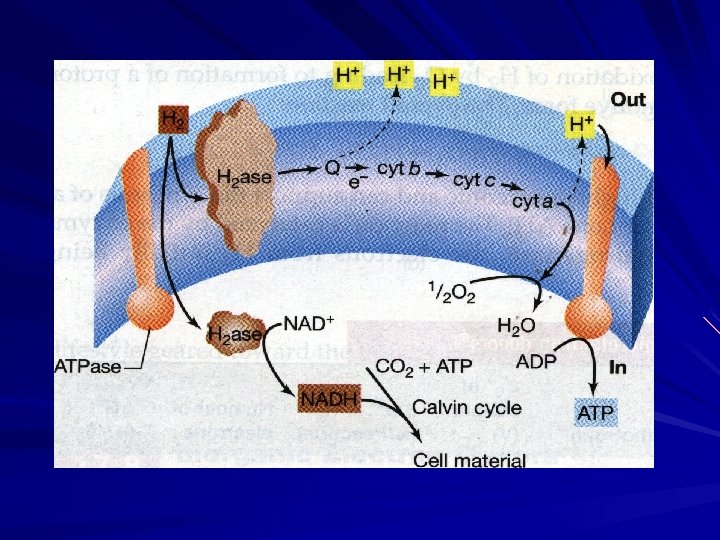

3. Sulfur Bacteria Energy Metabolism of Sulfur Bacteria Thiobacillus Energy source: Thiosulfate freely soluble in water and in the neutral condition. The respiratory chain of Thiobacilli: NADH 2 dehydrogenase, Fumaric reductase, flacoprotein( FP), ubliquinone(Co. Q), cyt b, Cytochrome oxidase aa 3
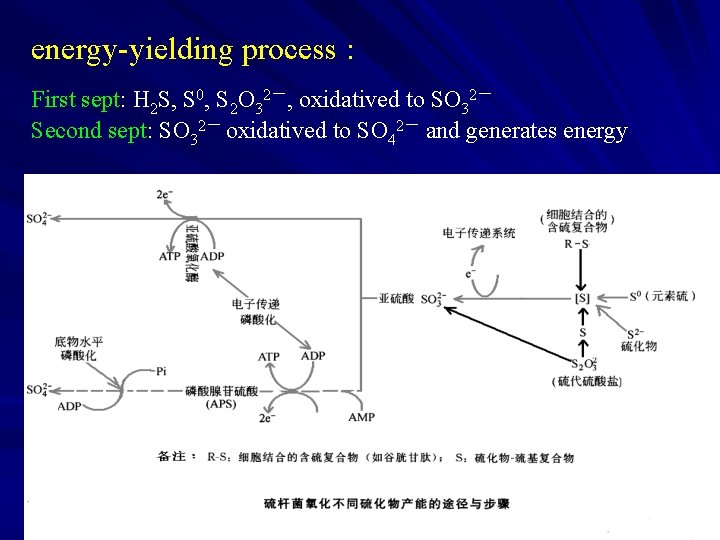
energy-yielding process : First sept: H 2 S, S 0, S 2 O 32-, oxidatived to SO 32- Second sept: SO 32- oxidatived to SO 42- and generates energy

4 iron bacteria and bacterial leaching (1) iron bacteria (iron oxidizing bacteria) oxidative Fe 2+into Fe 3 + and generates energy. For example, Ferrobacillus, Gallionella, Leptothrix, Crenothrix and Sphaerotilus; Thiobacillus frrooxidans: oxidative, S 0 and reducing sulphide, and Fe 2+ oxidatived to Fe 3+,so both Sulfur Bacteria and iron bacteria. mostly obligate Chemoautotrophic, few facultative Chemoautotrophic bacteria, acidophilic bacteria.

Iron oxidizing bacteria: bacteria energy-yielding process by Oxidative Phosphorylation

(2) bacterial leaching Principle: a. 2 S + 3 O 2 + 2 H 2 O — 2 H 2 SO 4 4 Fe. SO 4 + 2 H 2 SO 4 + O 2 — 2 Fe 2(SO 4)3 + 2 H 2 O b. Cu. S + 2 Fe 2(SO 4)3 + 2 H 2 O + O 2 — Cu. SO 4 + 4 Fe. SO 4 +2 H 2 SO 4 c. Cu. SO 4 + Fe — Fe. SO 4 + Cu

3 Biologic Fixation of Nitrogen 1. Kinds of nitrogen fixing microorganism Free-living nitrogen fixing bacteria Symbiosis nitrogen fixing bacteria Association nitrogen fixing bacteria
![2. The necessary condition of Nitrogen fixation Supplying ATP Reducing Force [H] and its 2. The necessary condition of Nitrogen fixation Supplying ATP Reducing Force [H] and its](http://slidetodoc.com/presentation_image_h2/4fb94879317d0da44888d73675633538/image-54.jpg)
2. The necessary condition of Nitrogen fixation Supplying ATP Reducing Force [H] and its Carrier Nitrogenase (molebdoferredoxin, azoferredoxin) N 2 Mg 2+ Strict Anoxy microenvironment
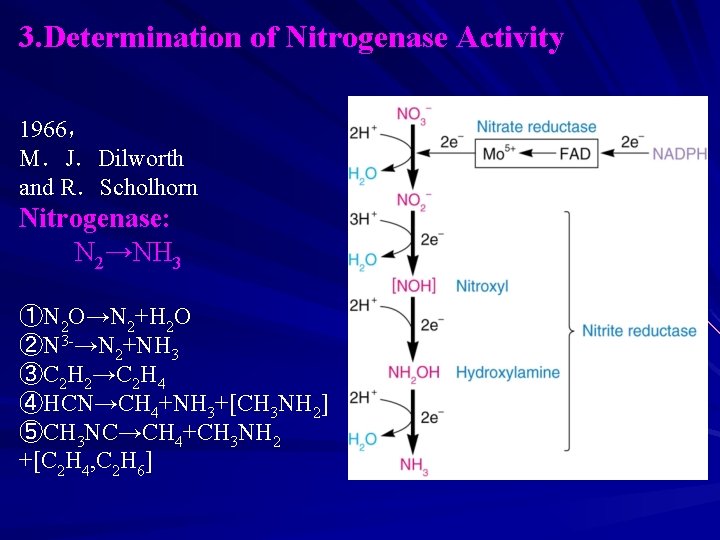
3. Determination of Nitrogenase Activity 1966, M.J.Dilworth and R.Scholhorn Nitrogenase: N 2→NH 3 ①N 2 O→N 2+H 2 O ②N 3 -→N 2+NH 3 ③C 2 H 2→C 2 H 4 ④HCN→CH 4+NH 3+[CH 3 NH 2] ⑤CH 3 NC→CH 4+CH 3 NH 2 +[C 2 H 4, C 2 H 6]
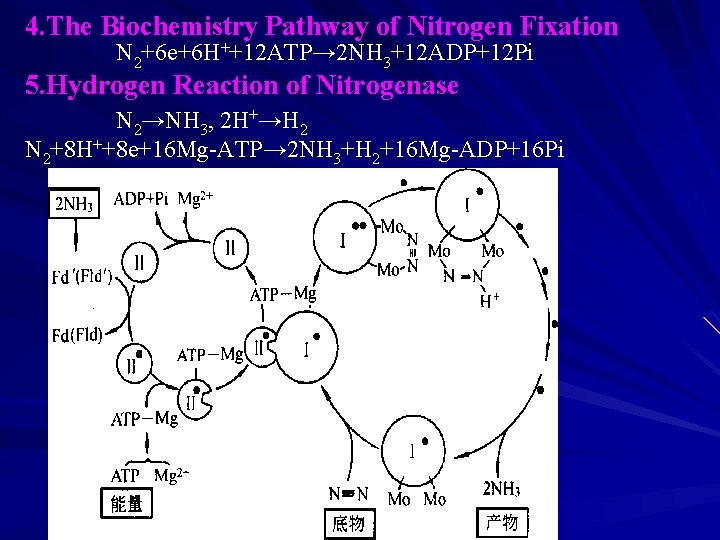
4. The Biochemistry Pathway of Nitrogen Fixation N 2+6 e+6 H++12 ATP→ 2 NH 3+12 ADP+12 Pi 5. Hydrogen Reaction of Nitrogenase N 2→NH 3, 2 H+→H 2 N 2+8 H++8 e+16 Mg-ATP→ 2 NH 3+H 2+16 Mg-ADP+16 Pi

6. The antioxygen mechamism of Nitrogenase in Aerobic Nitrogen Fixing Bacteria A. The Protection Mechanism in Aerobic Freeliving Nitrogen Fixting Bacteria a. Breathing Protection b. Conformation Protection B. The Protection Mechanism in Cyanobacteria a. Special Reducing Heterocysts Differentiated b. The Protection of Nitrogenase in Cyanobacteria that don’t form heterocysts C. The Antioxygen Protection of Nitrogenase in Root Nodule Bacteria a. Symbiosis Rhizobia in Leguminous Plant b. Symbiosis Rhizobia in nonleguminous Plant

4 The Synthesis of Peptidoglycan A The Synthesis in Bacterial Cytoplasm a G — G-UDP,M--UDP G — — 6 -P-fructose— 6 -P- glucosamine— — N-acetoglucosamine -1 - P — N-acetylglucosamine-UDP — N-acetylmuramic acid -UDP b M — “Park” nucleotide (UDP-M-pentapeptide) UDP as carrier sugar D-Val-D-Val are repressed by cycloserine;
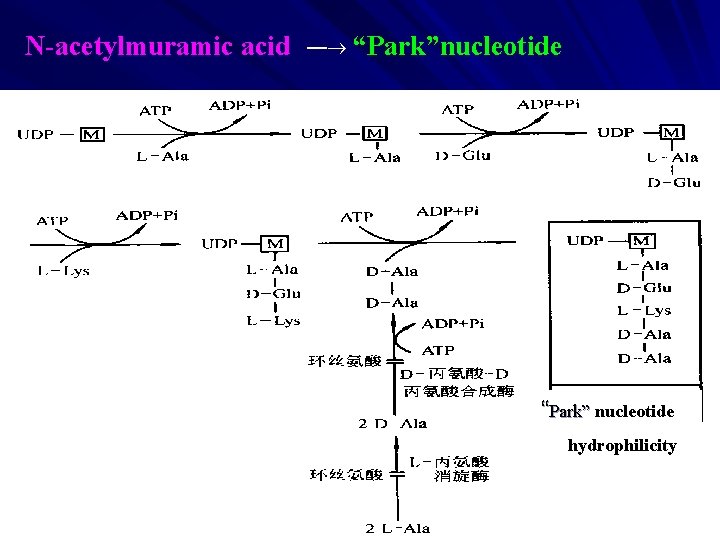
N-acetylmuramic acid — “Park”nucleotide “Park” nucleotide hydrophilicity

B. The Synthesis in Bacterial Cytomembrane peptidoglycan unit synthesis Carrier:bactoprenol(carrier lipoid) ——C 55 -isoprenol(contain 11 isoprenoid units) combinate to G during transmembrane ,link into interpeptide bridge; carrier lipoid: concerned with the synthesize of polysaccharide and lipoidglycan in outer cell ( cellulose, polymannan, and so on) vancomycin,bacitracin inhibite some reactins
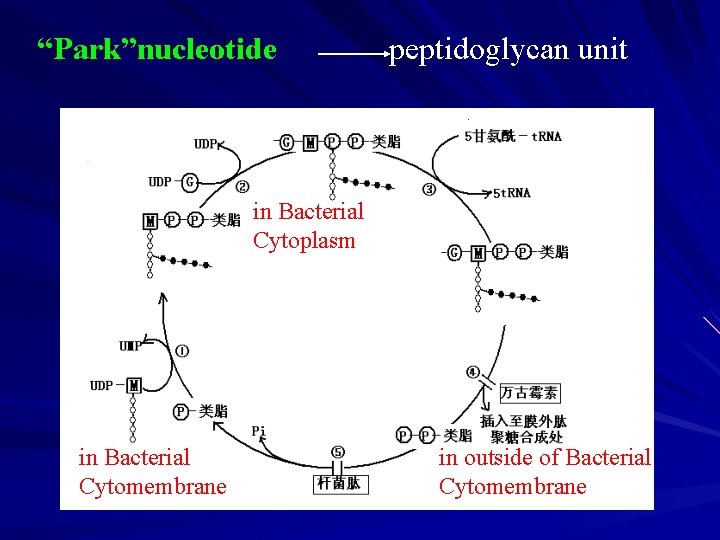
“Park”nucleotide peptidoglycan unit in Bacterial Cytoplasm in Bacterial Cytomembrane in outside of Bacterial Cytomembrane

C. The Synthesis in outside of Bacterial Cytomembrane a. glycan chain extension(transverse link): peptidoglycan monomer+primer extense one disaccharide unit — glycan chain transverse Transglycosyl(form -1,4 glucosidic bond) b. Adjacent glycan chain connect(vertical link): 2 D-Val — M-tetrapeptide + D-Val — cross-links occur between the distalamino group of the diamino acid in the positon 3 of one stem peptide and D-alanine in the positon 4 of another stem peptide. Transpeptidation : repressed by penicillins, penicillins : D-Val-D-Val analog, competitive transpeptidase.
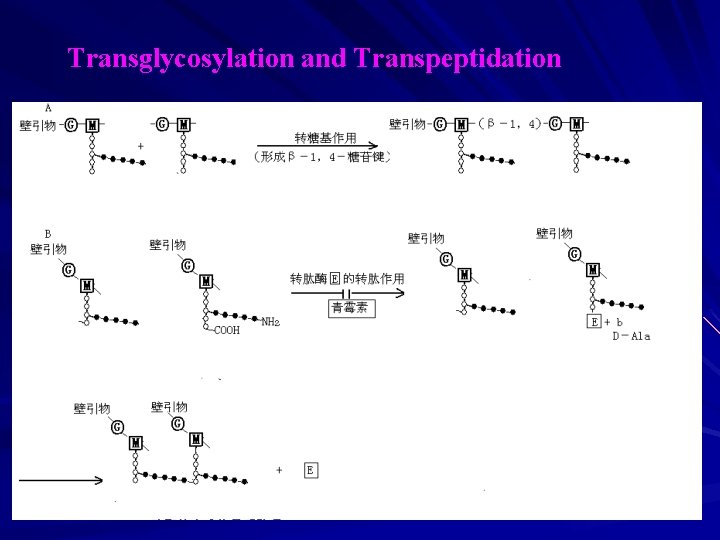
Transglycosylation and Transpeptidation

Section 3 microbial secondary metabolism and its product There are many kinds of secondary metabolite which primary metabolic products serve as substrate, some of these products have a considerable importance in fermentation industry. Metabolic adjustment of secondary metabolite is similar to primary metabolism, influenced by many factors.

Microbial secondary metabolism and its product Regulation of secondary metabolism 1 primary metabolism 2 nitric compound 3 induction and feedback inhibition

Section 4 Regulation of metabolism and ferment product practise. Metabolic regulation kinds are various, microorganism produce metabolic product to offer service for fermentation industry and to benefit for mankind with their relevant knowledge of adjusting control.

Using microbial metabolic product Metabolic product type: 1 primary metabolic products: amino acid, enzyme or coenzyme; 2 submetabolic products: antibiotic, hormone, alkaloid, toxin, vitamin, and so on. Fermentation type: type 1 Natural fermentation: fermentation ethanol, lacate and so on, 2 Metabolic control fermentation: fermentation terminal products: lysine, guanylic acid, adenylic acid and so on ; metabolic intermediates: citrate, α-ketoglutaric acid, succinate, inosinic acid, xanthine nucleotide and so on;

Control of ferment condiction 1)cultrue condition:temperature, p. H and so on; 2)nutrition component:glucose concentration, C/N, growth factor, and so on; 3)dissolved oxygen: aeration numbr, agitation and so on. 。
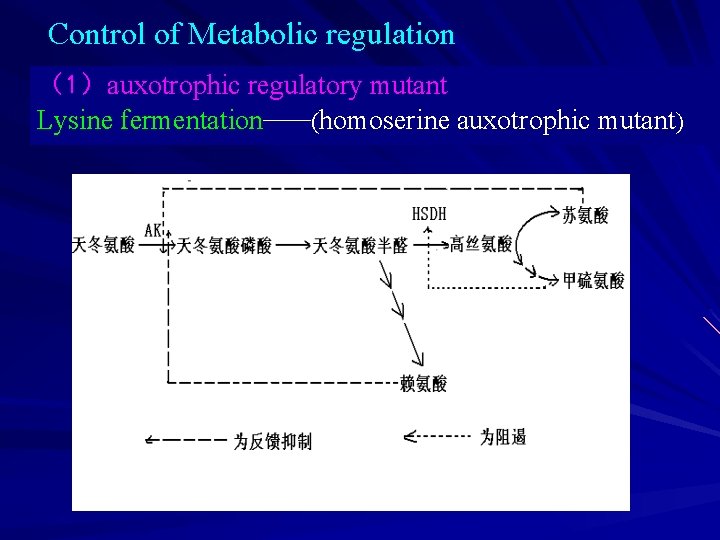
Control of Metabolic regulation (1)auxotrophic regulatory mutant Lysine fermentation——(homoserine auxotrophic mutant)
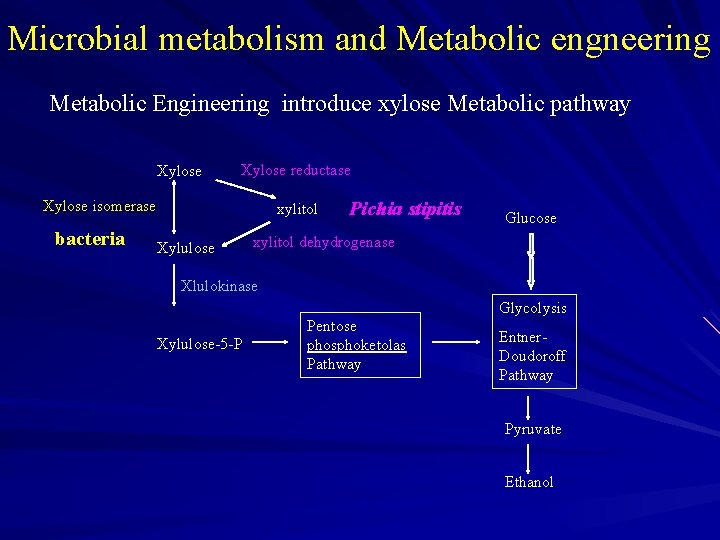
Microbial metabolism and Metabolic engneering Metabolic Engineering introduce xylose Metabolic pathway Xylose reductase Xylose isomerase bacteria xylitol Xylulose Pichia stipitis Glucose xylitol dehydrogenase Xlulokinase Glycolysis Xylulose-5 -P Pentose phosphoketolas Pathway Entner. Doudoroff Pathway Pyruvate Ethanol

Using active metabolism in microbes 1 biotransformation, 2 microbial straw xylan-degrading, 3 microbiohydrometallurgy and oil extraction, 4 biodegradation

References: Prescott LM, Harley JP, and Klein DA. Microbiology (5 th ed. ), Higher education press and Mc. Graw-Hill Companies, Inc. 2002 Michael TM, John MM, Jack P. Brock biology of micoorganisms,International edition, Pearson Education, Inc. 2003 Talaro K. P. Foundations in microbiology (Fifth Edition), Higher education press and Mc. Graw-Hill Companies, Inc. 2005
- Slides: 72