Chapter 6 Biotransformation of Xenobiotics II Dr Arthur

Chapter 6: Biotransformation of Xenobiotics II Dr. Arthur Roberts 1

Xenobiotic Biotransformation 1. 2. 3. 4. hydrolytic metabolism (H 2 O) reductive metabolism oxidative metabolism conjugation reactions 2

Oxidative Metabolism • • • Cytochromes P 450 Catalase Flavin-Containing Monooxygenases (FMO) Monoamine Oxidase (MAO) Diamine Oxidase (DAO) (a. k. a. Amine Oxidase and Histaminase) Xanthine Oxidase (XO, XAO) Xanthine Dehydrogenase (XDH) Aldehyde Oxidase (AO) Aldehyde Dehydrogenase (ALDH) 3
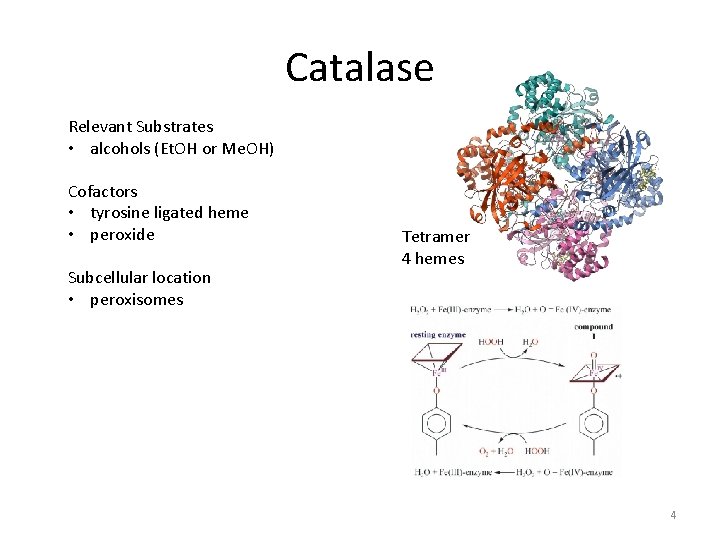
Catalase Relevant Substrates • alcohols (Et. OH or Me. OH) Cofactors • tyrosine ligated heme • peroxide Subcellular location • peroxisomes Tetramer 4 hemes 4
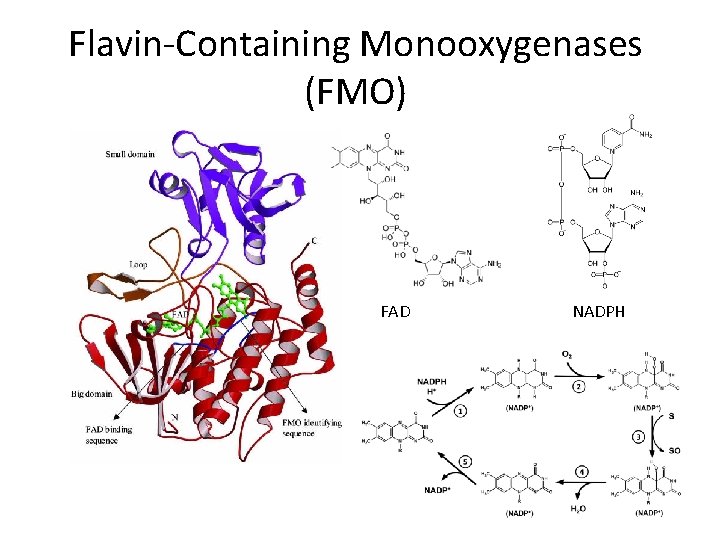
Flavin-Containing Monooxygenases (FMO) FAD NADPH
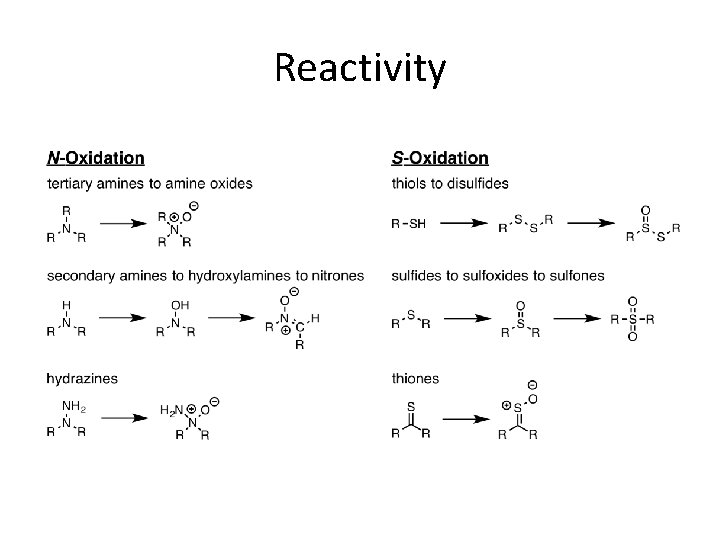
Reactivity
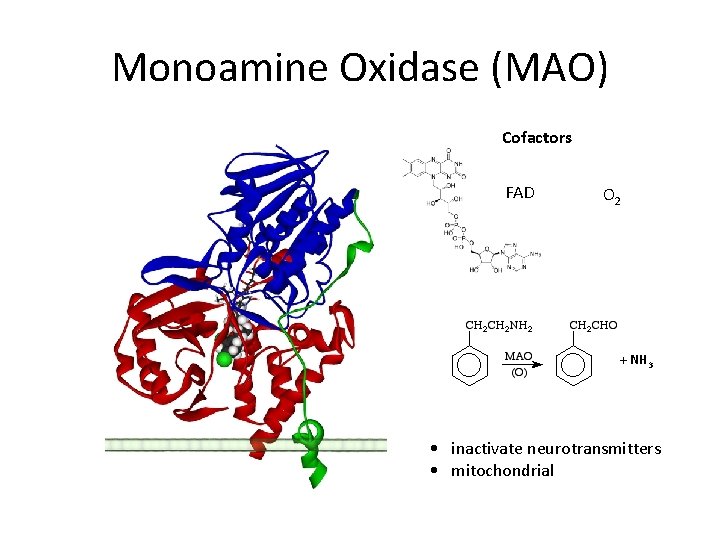
Monoamine Oxidase (MAO) Cofactors FAD O 2 + NH 3 • inactivate neurotransmitters • mitochondrial
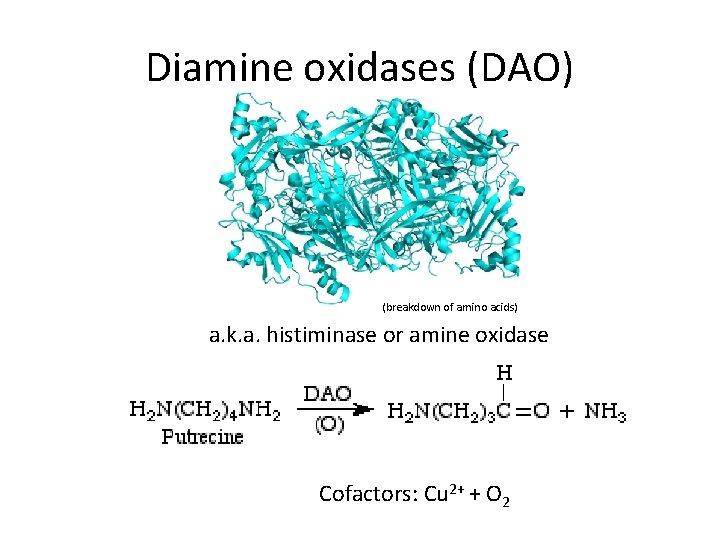
Diamine oxidases (DAO) (breakdown of amino acids) a. k. a. histiminase or amine oxidase Cofactors: Cu 2+ + O 2
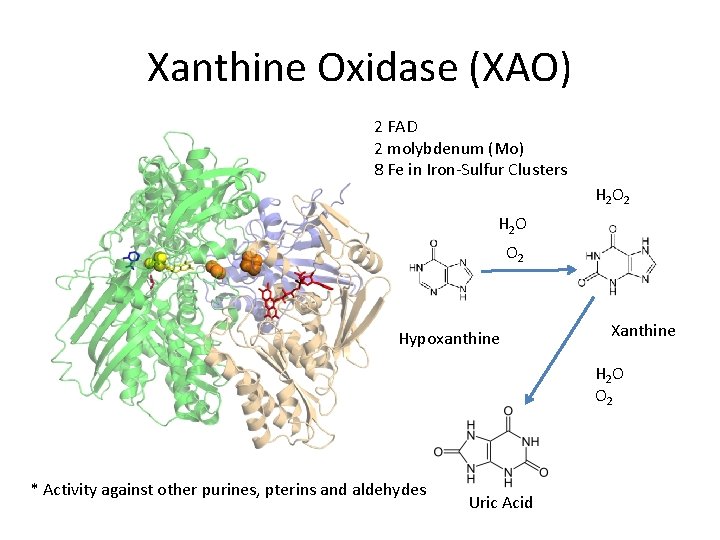
Xanthine Oxidase (XAO) 2 FAD 2 molybdenum (Mo) 8 Fe in Iron-Sulfur Clusters H 2 O 2 H 2 O O 2 Hypoxanthine Xanthine H 2 O O 2 * Activity against other purines, pterins and aldehydes Uric Acid
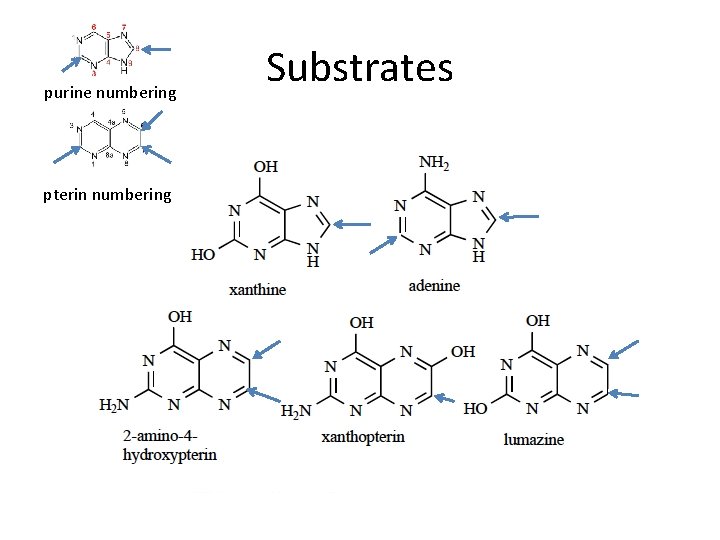
purine numbering pterin numbering Substrates

Toxicology/Clinical Significance • superoxide producer – oxidative eye injury RH + H 2 O + 2 O 2 ROH + 2 O 2 - + 2 H+ • Xanthinuria – Type II – accumulation of xanthine – renal failure • Gout – allopurinol Uric Acid Crystals
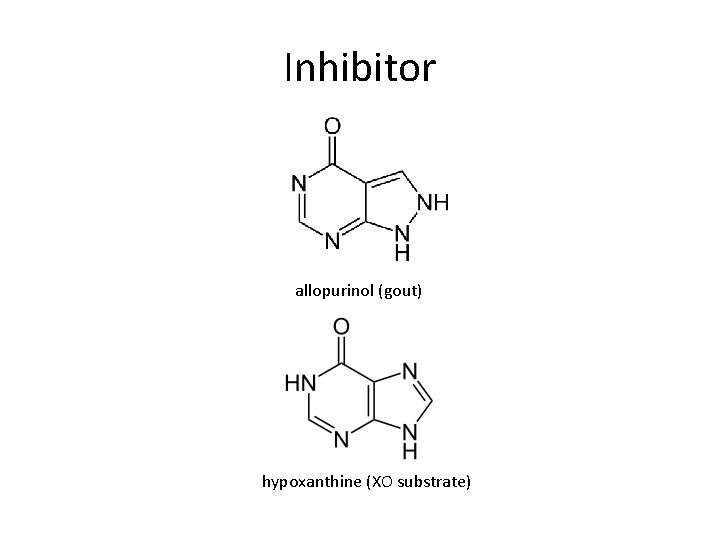
Inhibitor allopurinol (gout) hypoxanthine (XO substrate)
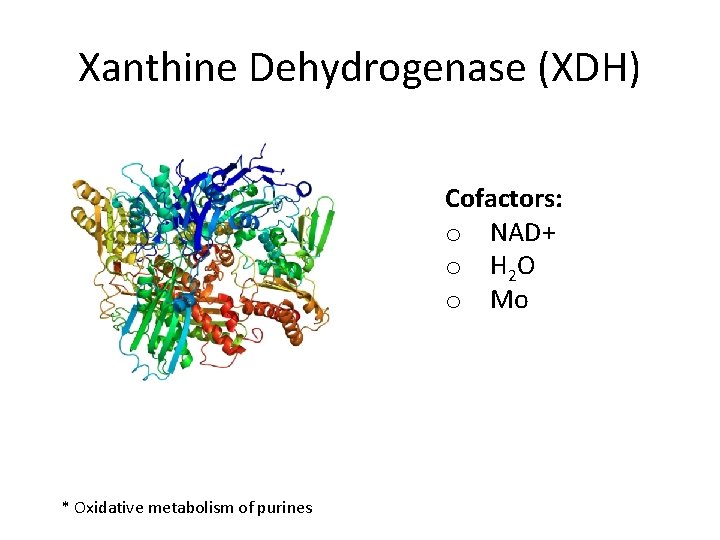
Xanthine Dehydrogenase (XDH) Cofactors: o NAD+ o H 2 O o Mo * Oxidative metabolism of purines
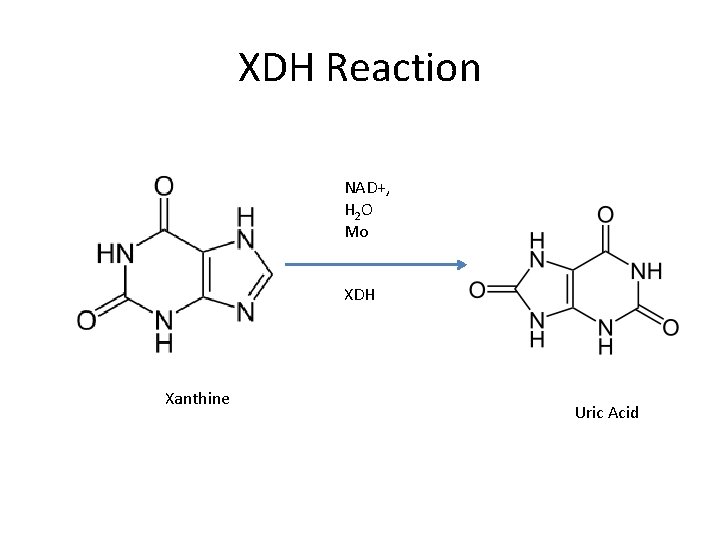
XDH Reaction NAD+, H 2 O Mo XDH Xanthine Uric Acid
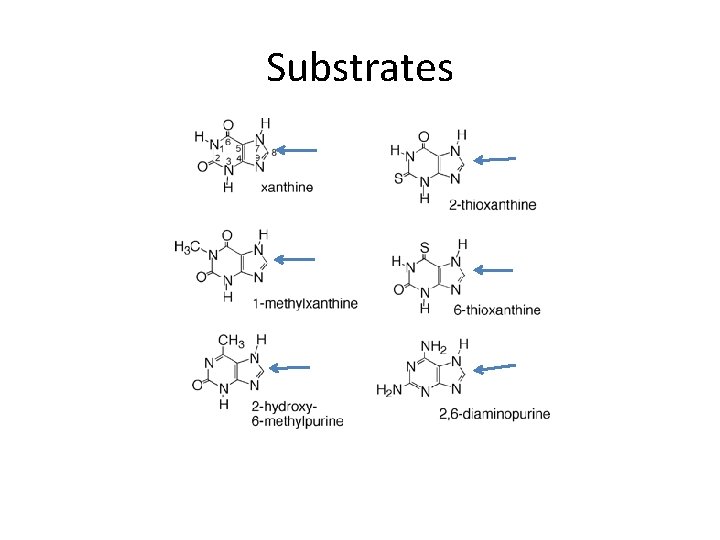
Substrates

XO vs. XDH XO • Cofactors: 2 FAD, 2 Mo, 2 Fe, H 2 O, O 2 • Specificity: purines, pterins and aldehydes • Toxic Products: superoxide • Disease: Type II xanthinuria, gout XDH • Cofactors: NAD+, Mo, H 2 O • Specificity: xanthinesque • Toxic Products: None • Disease: Type I xanthinuria
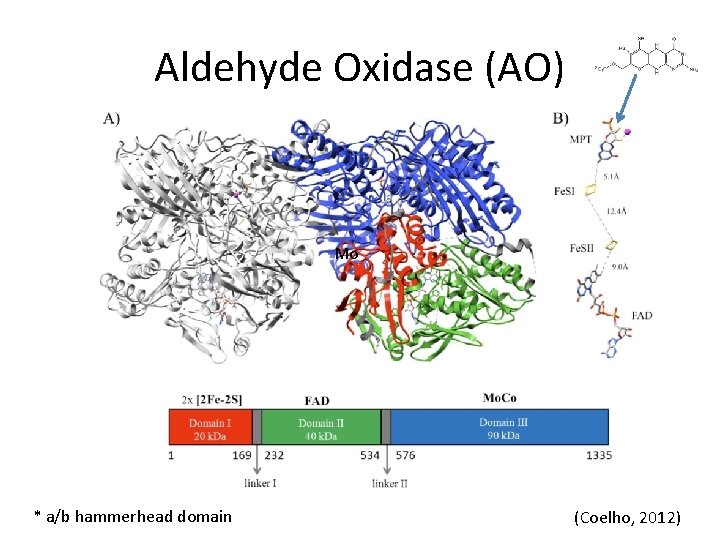
Aldehyde Oxidase (AO) Mo * a/b hammerhead domain (Coelho, 2012)
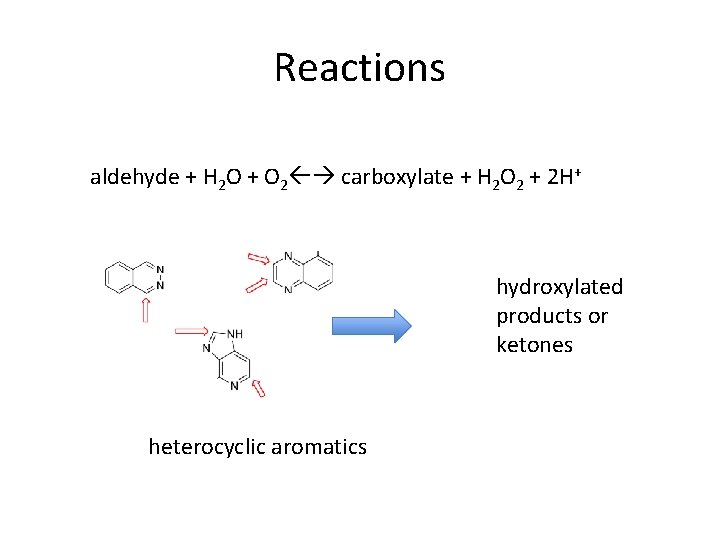
Reactions aldehyde + H 2 O + O 2 carboxylate + H 2 O 2 + 2 H+ hydroxylated products or ketones heterocyclic aromatics
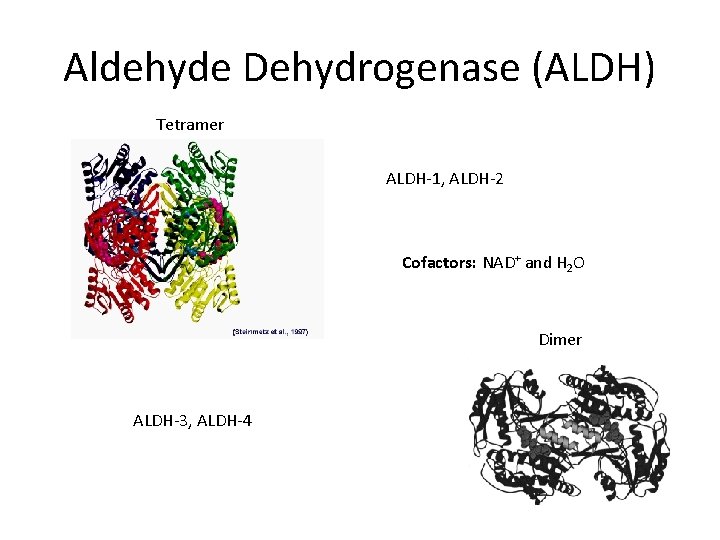
Aldehyde Dehydrogenase (ALDH) Tetramer ALDH-1, ALDH-2 Cofactors: NAD+ and H 2 O Dimer ALDH-3, ALDH-4
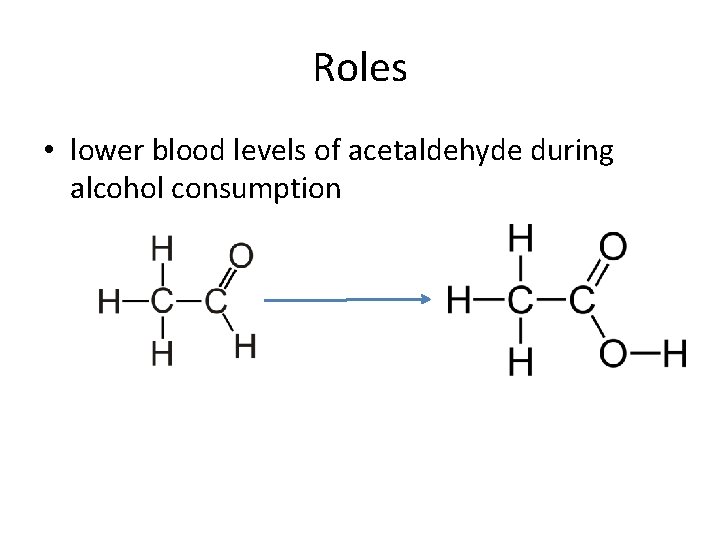
Roles • lower blood levels of acetaldehyde during alcohol consumption

Genetics • ALDH 2*2 (K 487 E) – 8% activity – lower alcohol tolerance • Ethnic Groups deficient in ALDH – North American Indians 5% – South American Indians 40% – Asians 40% – Europeans 0% – Africans 0%

AO vs. ALDH AO • Cofactors: MPT, Mo, 2 x 2 Fe 2 S clusters, FAD, O 2 • Substrate Specificity: nitrogen-containing heterocyclic aromatics and aldehydes • Diseases: No major ALDH • Cofactors: NAD+, H 2 O • Substrate Specificity: aldehydes • Diseases: Lower alcohol tolerance

Oxidative Metabolism • • • Cytochromes P 450 Catalase Flavin-Containing Monooxygenases (FMO) Monoamine Oxidase (MAO) Diamine Oxidase (DAO) (a. k. a. Amine Oxidase and Histaminase) Xanthine Oxidase (XO, XAO) Xanthine Dehydrogenase (XDH) Aldehyde Oxidase (AO) Aldehyde Dehydrogenase (ALDH) 23

Xenobiotic Biotransformation 1. Both Oxidative and Reductive Metabolism 2. Oxidation Reduction 24
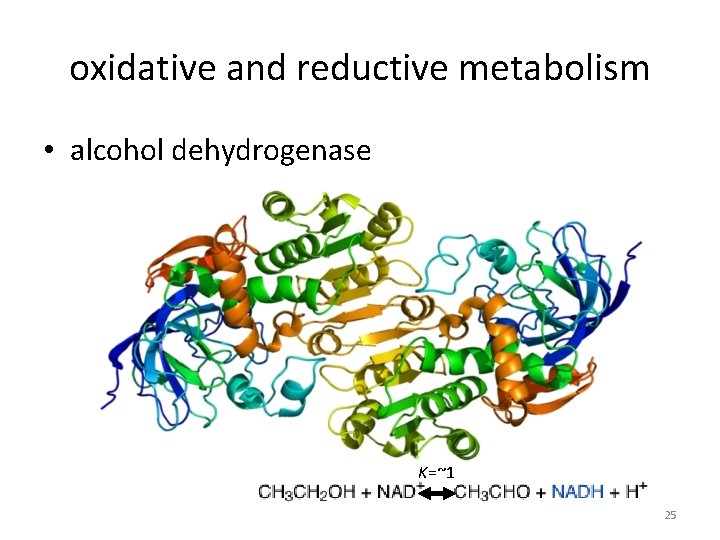
oxidative and reductive metabolism • alcohol dehydrogenase K=~1 25
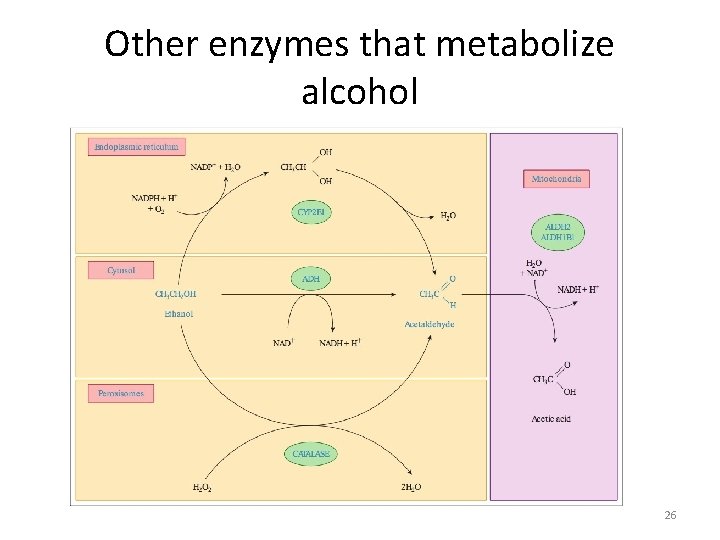
Other enzymes that metabolize alcohol 26

oxidative reductive metabolism • anaerobic – aldehyde oxidase – cytochromes P 450 27
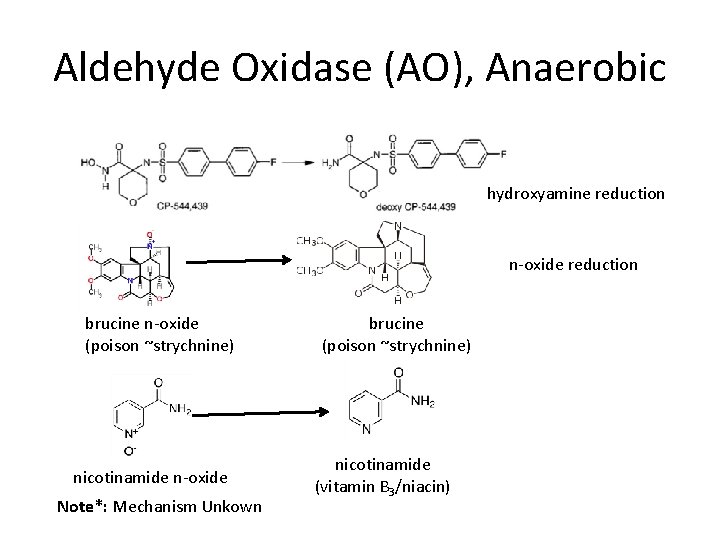
Aldehyde Oxidase (AO), Anaerobic hydroxyamine reduction n-oxide reduction brucine n-oxide (poison ~strychnine) nicotinamide n-oxide Note*: Mechanism Unkown brucine (poison ~strychnine) nicotinamide (vitamin B 3/niacin)
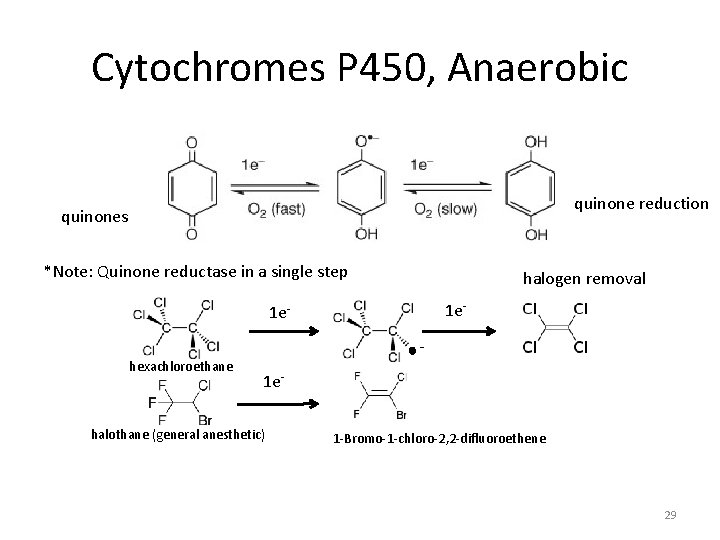
Cytochromes P 450, Anaerobic quinone reduction quinones *Note: Quinone reductase in a single step halogen removal 1 e- 1 ehexachloroethane 1 e- halothane (general anesthetic) 1 -Bromo-1 -chloro-2, 2 -difluoroethene 29

conjugation reactions • • • UDP-glucuronosyltransferases (UGTs) Sulfotransferases (SULTs) Glutathione-S-Transferases (GSTs) N-acetyltransferases (NATs) Methyltransferases – Catechol O-Methyltransferases (COMTs) – Histamine Methyltransferases (HMTs) – Thiopurine Methyltransferases (TPMTs)
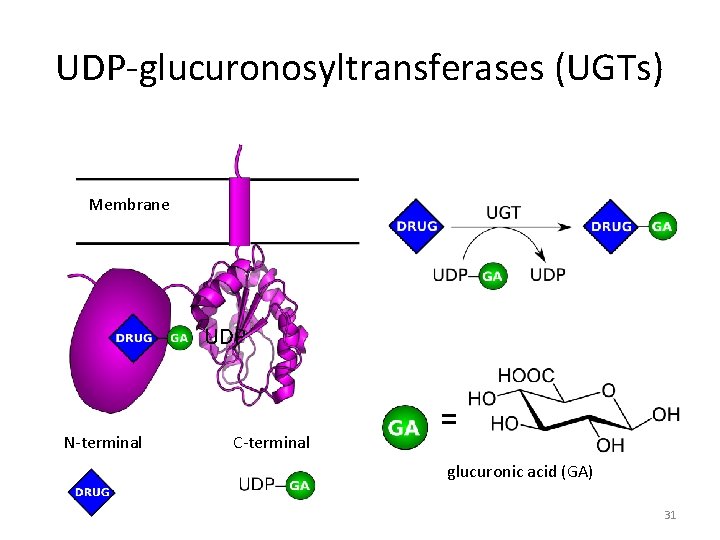
UDP-glucuronosyltransferases (UGTs) Membrane UDP N-terminal C-terminal = glucuronic acid (GA) 31
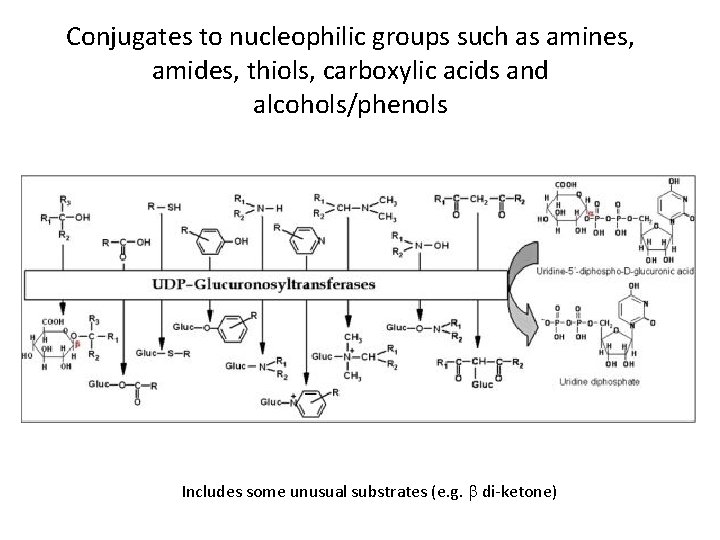
Conjugates to nucleophilic groups such as amines, amides, thiols, carboxylic acids and alcohols/phenols Includes some unusual substrates (e. g. b di-ketone)
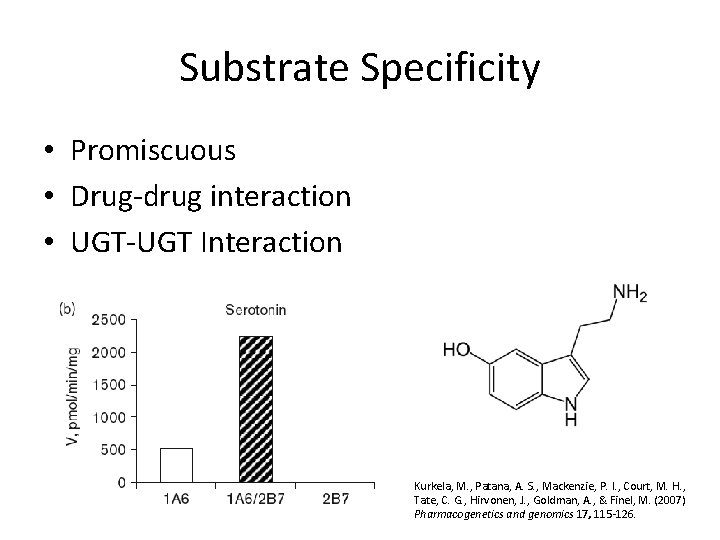
Substrate Specificity • Promiscuous • Drug-drug interaction • UGT-UGT Interaction Kurkela, M. , Patana, A. S. , Mackenzie, P. I. , Court, M. H. , Tate, C. G. , Hirvonen, J. , Goldman, A. , & Finel, M. (2007) Pharmacogenetics and genomics 17, 115 -126.
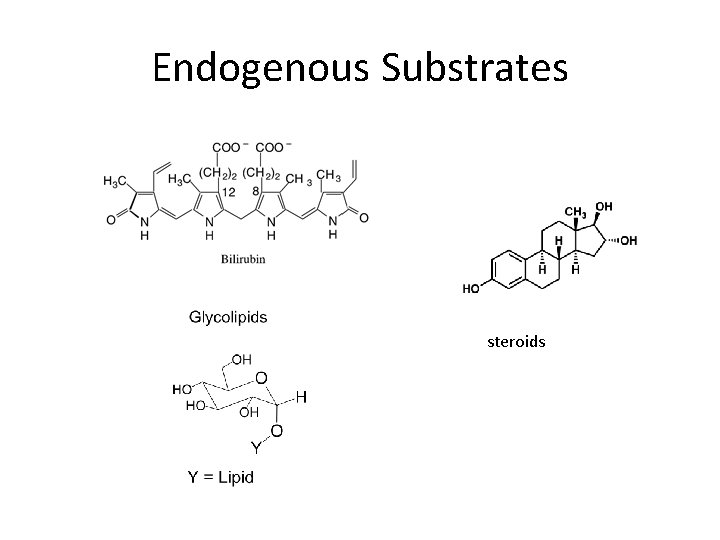
Endogenous Substrates steroids

Families • UGT 1 A – Planar and bulky phenols (e. g. estrogens), amines and unsaturated carboxylic acids – UGT 1 A 1: bilirubin and morphine • UGT 2 B – drugs, steroids, bile acids and unsaturated carboxylic acids – UGT 2 B 7: morphine Note: bilirubin and morphine are not planar.

Diseases • Hyperbilirubinemia – Gilbert’s syndrome (20 -30% UGT 1 A 1 Activity) – Crigler-Najjar syndrome • Type I (0% UGT 1 A 1 Activity) • Type II (10% UGT 1 A 1 Activity) • Gray Baby Syndrome (Chloramphenicol) • Neonatal Jaundice – UGT 1 A 1*28/*28 – light therapy • Autoimmune Hepatitis
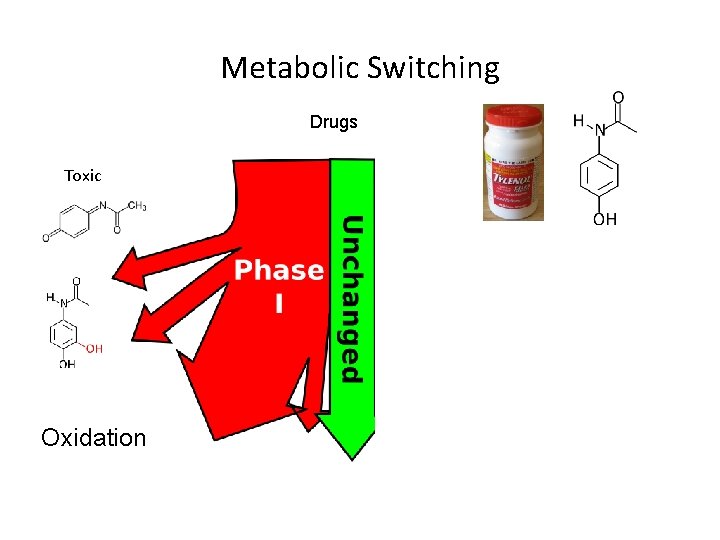
Metabolic Switching Drugs Toxic Oxidation Conjugation
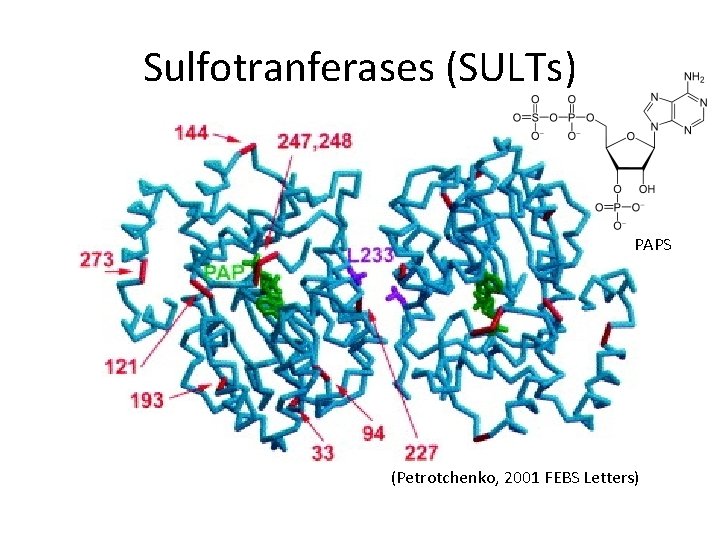
Sulfotranferases (SULTs) PAPS (Petrotchenko, 2001 FEBS Letters)
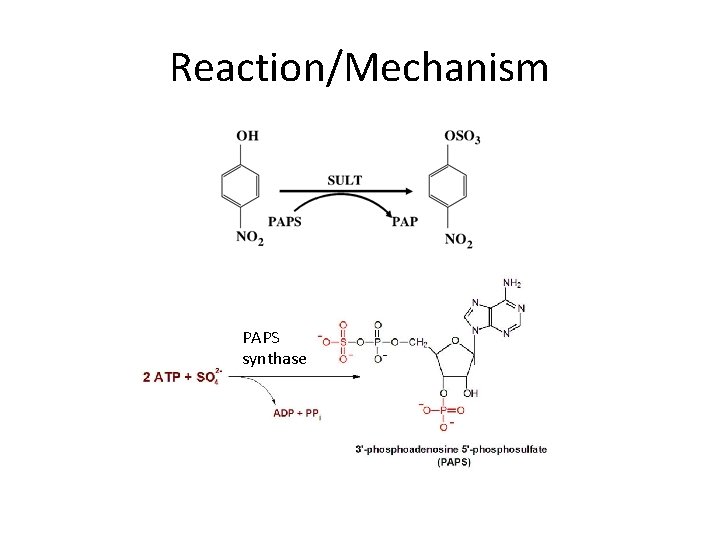
Reaction/Mechanism PAPS synthase
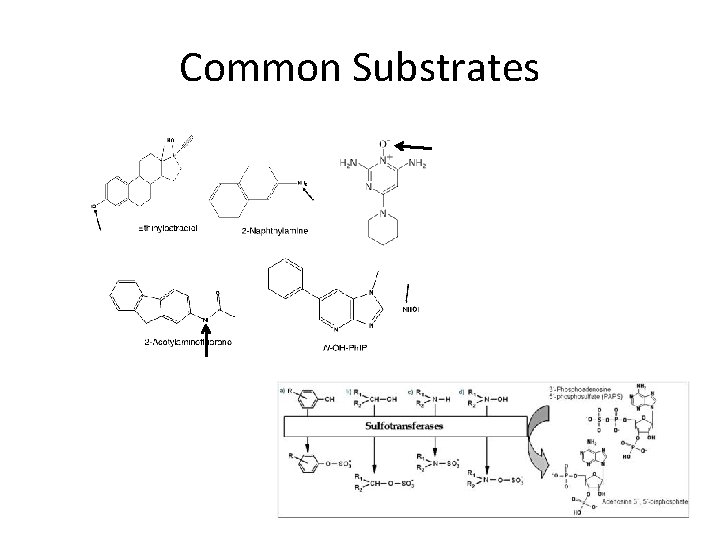
Common Substrates
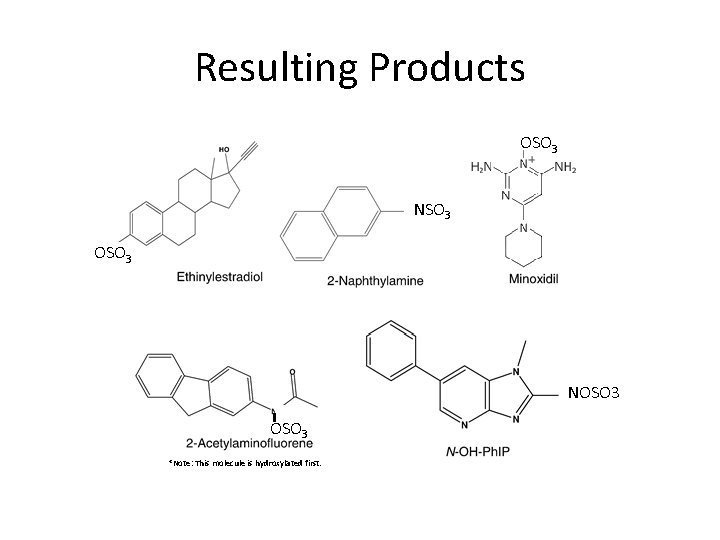
Resulting Products OSO 3 NSO 3 OSO 3 NOSO 3 *Note: This molecule is hydroxylated first.

Families and General Substrate Specificities • SULT 1 – Phenols, Catechols, Benzylic Alcohols, Amines, Estrogens • SULT 2 – Steroids
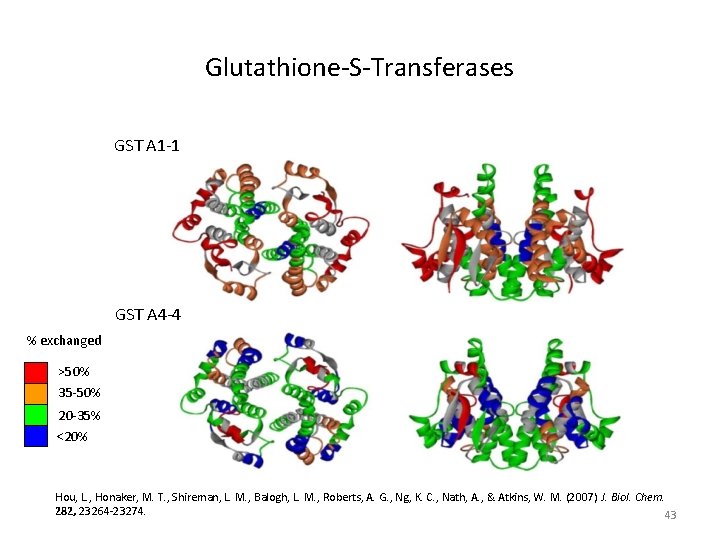
Glutathione-S-Transferases GST A 1 -1 GST A 4 -4 % exchanged >50% 35 -50% 20 -35% <20% Hou, L. , Honaker, M. T. , Shireman, L. M. , Balogh, L. M. , Roberts, A. G. , Ng, K. C. , Nath, A. , & Atkins, W. M. (2007) J. Biol. Chem. 282, 23264 -23274. 43
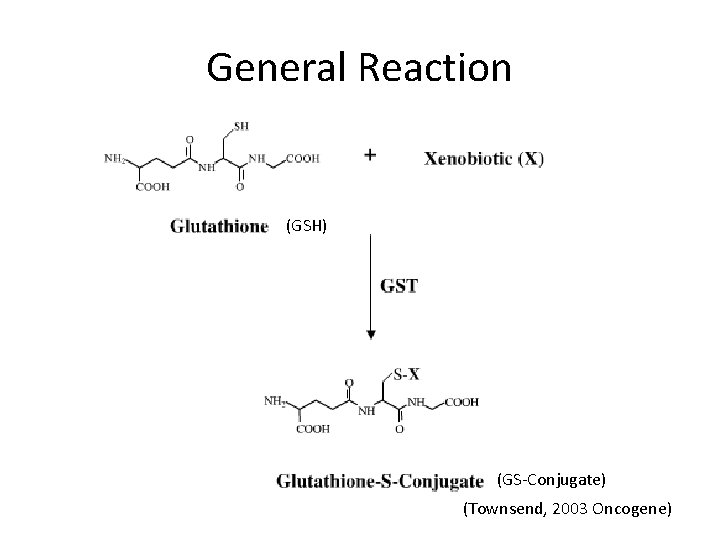
General Reaction (GSH) (GS-Conjugate) (Townsend, 2003 Oncogene)
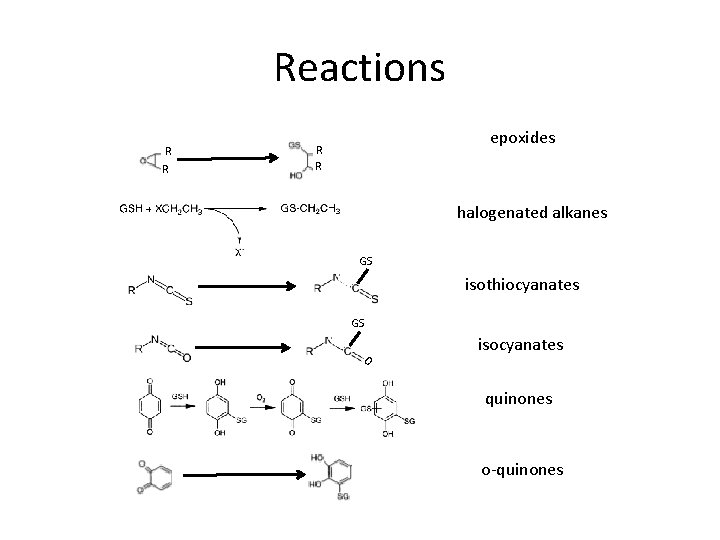
Reactions R R epoxides R R halogenated alkanes GS isothiocyanates GS O isocyanates quinones o-quinones
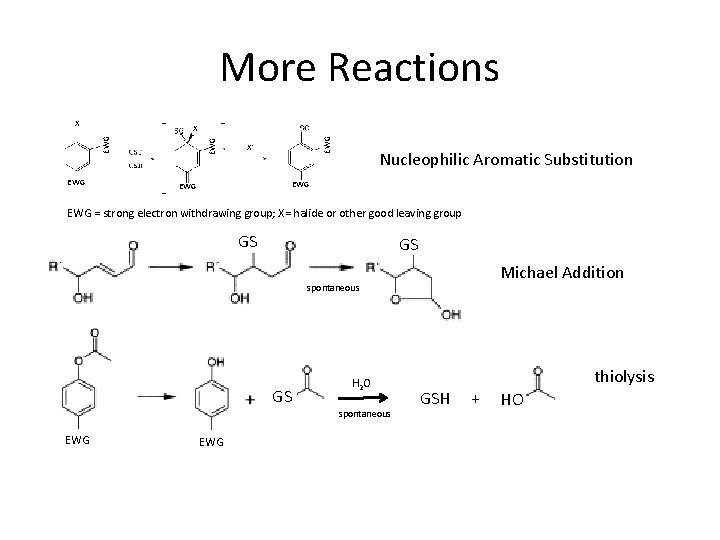
More Reactions X EWG EWG X X- Nucleophilic Aromatic Substitution EWG EWG = strong electron withdrawing group; X= halide or other good leaving group GS GS Michael Addition spontaneous GS H 2 O spontaneous EWG thiolysis GSH + HO

Families and Tissue Locations • alpha (A) - Liver • kappa (K) • mu (M) – brain, lung , testes, lymphoblasts, skeletal muscle • omega (O) • pi (P) – all tissues except Liver • sigma (S) – macrophages, placenta and fat – prostaglandin metabolism • theta (T) • zeta (Z) • microsomal – ER and mitochondria

Genetics (Often Missing) • GST M 1 – <50% Caucasians and East Asians – >90% Polynesians and Micronesians – >50% Africans – absence increases risk for lung and bladder cancer • GST T 1 – 20% missing in Caucasians – 50% missing in Asians – 40% missing in Africans
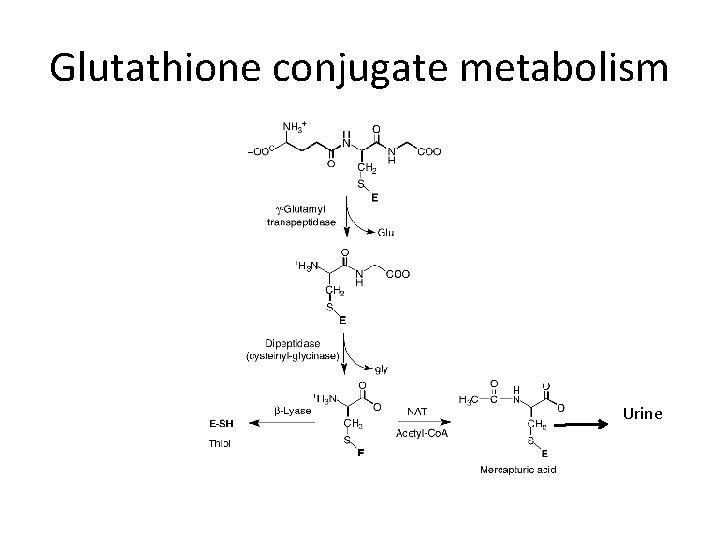
Glutathione conjugate metabolism Urine
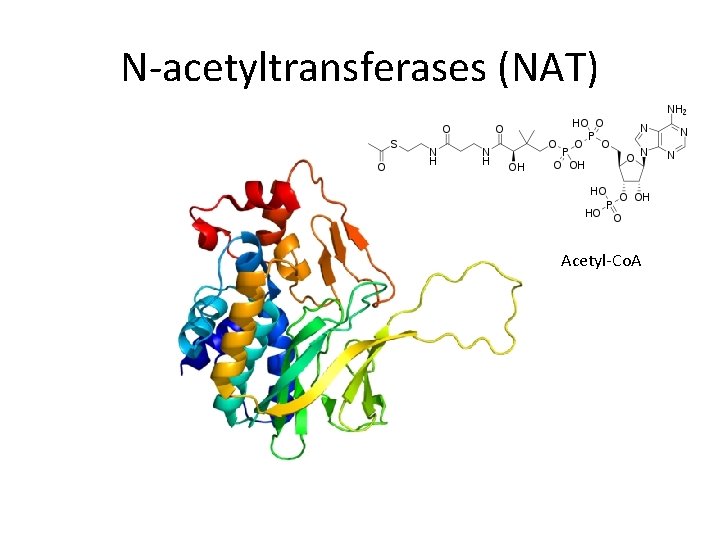
N-acetyltransferases (NAT) Acetyl-Co. A
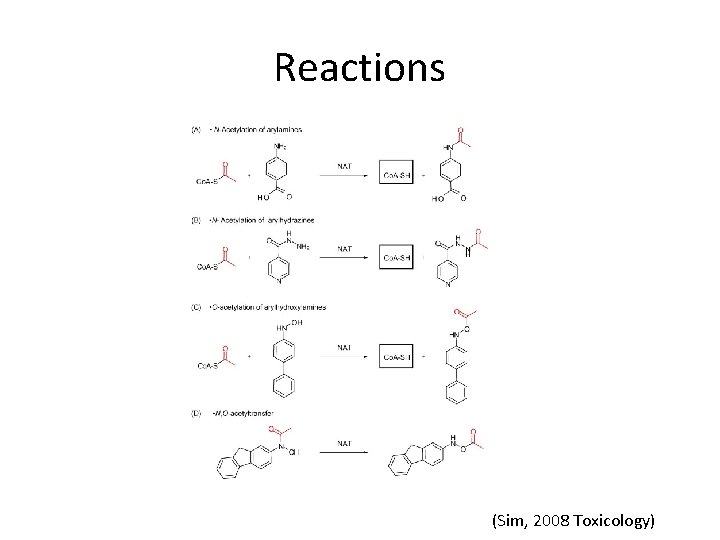
Reactions (Sim, 2008 Toxicology)
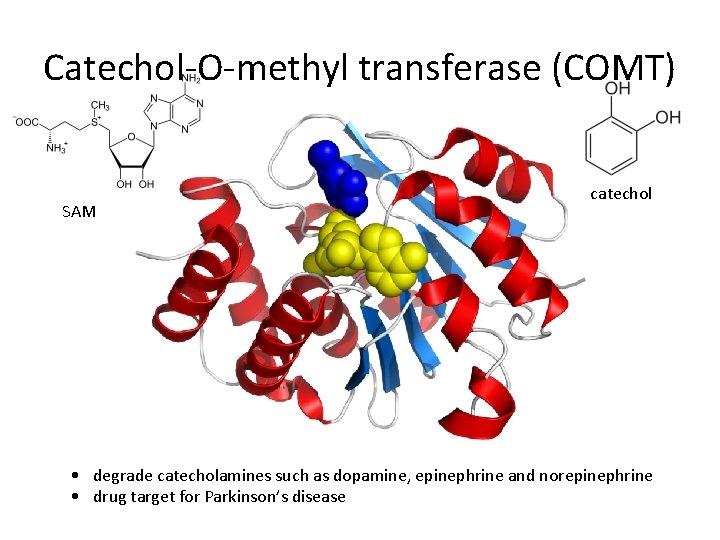
Catechol-O-methyl transferase (COMT) SAM catechol • degrade catecholamines such as dopamine, epinephrine and norepinephrine • drug target for Parkinson’s disease
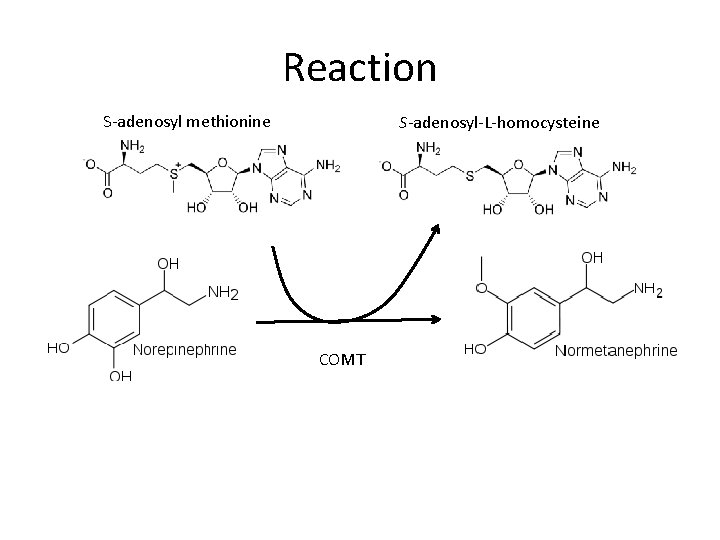
Reaction S-adenosyl methionine S-adenosyl-L-homocysteine COMT
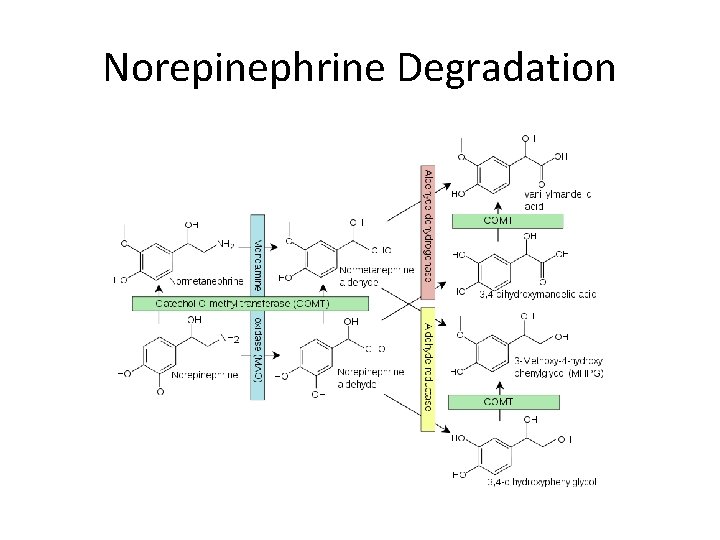
Norepinephrine Degradation
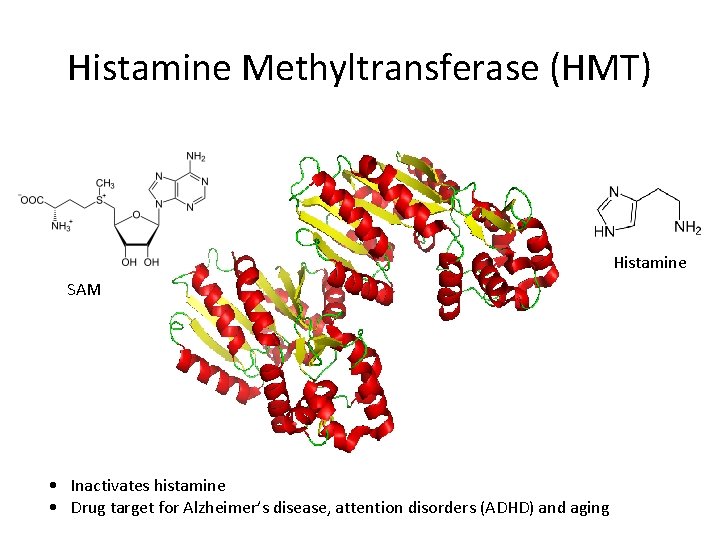
Histamine Methyltransferase (HMT) Histamine SAM • Inactivates histamine • Drug target for Alzheimer’s disease, attention disorders (ADHD) and aging
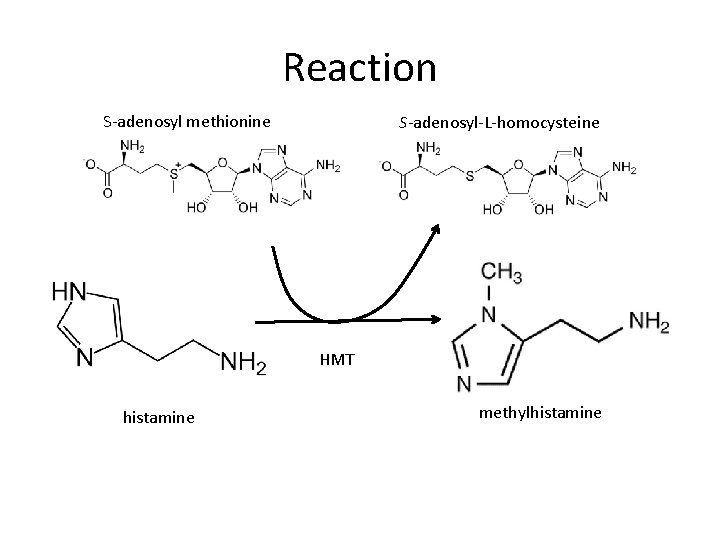
Reaction S-adenosyl methionine S-adenosyl-L-homocysteine HMT histamine methylhistamine
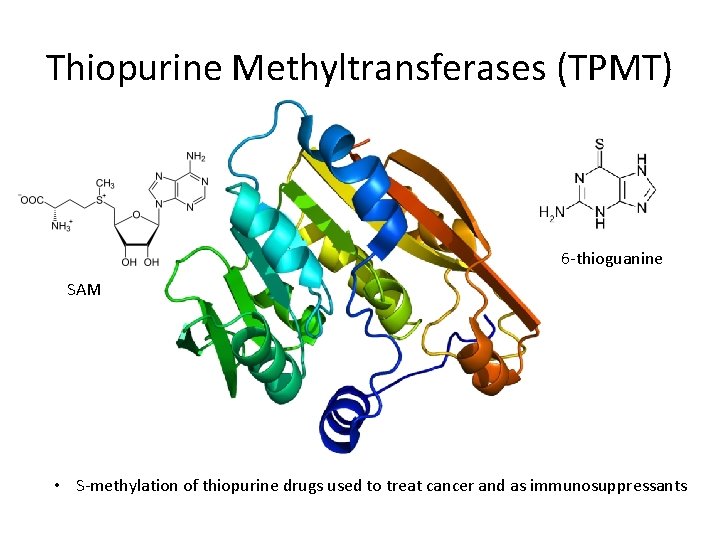
Thiopurine Methyltransferases (TPMT) 6 -thioguanine SAM • S-methylation of thiopurine drugs used to treat cancer and as immunosuppressants
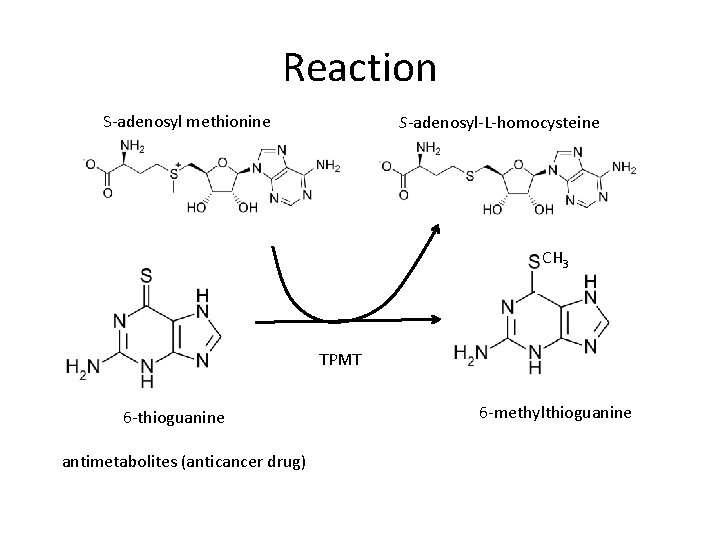
Reaction S-adenosyl methionine S-adenosyl-L-homocysteine CH 3 TPMT 6 -thioguanine antimetabolites (anticancer drug) 6 -methylthioguanine

conjugation reactions • • • UDP-glucuronosyltransferases (UGTs) Sulfotransferases (SULTs) Glutathione-S-Transferases (GSTs) N-acetyltransferases (NATs) Methyltransferases – Catechol O-Methyltransferases (COMTs) – Histamine Methyltransferases (HMTs) – Thiopurine Methyltransferases (TPMTs)
- Slides: 59