CHAPTER 5 DNA HYBRIDIZATION INTRODUCTION DNA PROBE LABELING

CHAPTER 5 DNA HYBRIDIZATION -INTRODUCTION -DNA PROBE, LABELING, SIGNAL MISS NUR SHALENA SOFIAN

INTRODUCTION • DNA hybridization – ability of ss. DNA of one strand to form ds. DNA of another ss. DNA by complementary sequences • Probe is used to identify a desired clone among thousands of irrelevant ones : a. polynucleotides / oligonucleotides - Must have a probe to screen the library b. antibodies - Must have antibodies specific for your protein

ANTIBODY PROBE • E. g. λgt 11 vector for making and screening c. DNA libraries • Group of clones are screen via expression of the right protein therefore need antiserum directed against protein of interest • When c. DNA inserts are plated, proteins released by each clone are blotted onto nitrocellulose • Proteins that have transferred onto nitrocellulose can be probed with antiserum
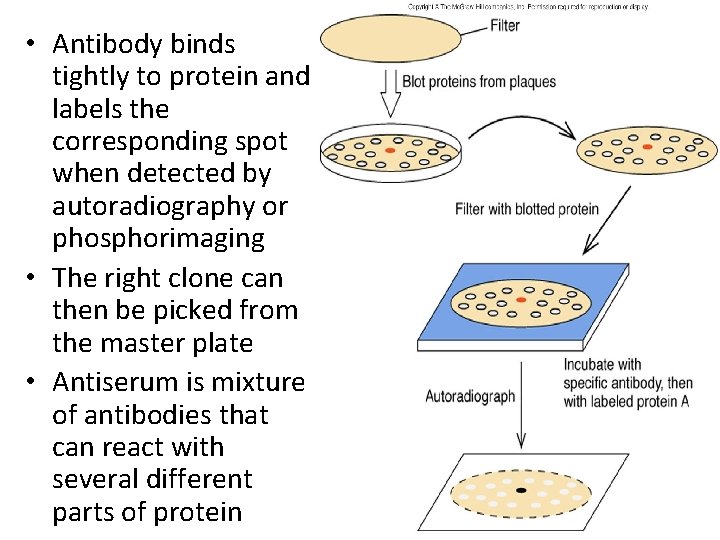
• Antibody binds tightly to protein and labels the corresponding spot when detected by autoradiography or phosphorimaging • The right clone can then be picked from the master plate • Antiserum is mixture of antibodies that can react with several different parts of protein

POLYNUCLEOTIDE PROBE • Using homologous gene from another organism to probe for gene of interest • Lowering stringency of hybridization conditions so it can tolerate mismatches in base sequence between the probe and the cloned gene e. g. high temperature, organic solvents, salt concentrations
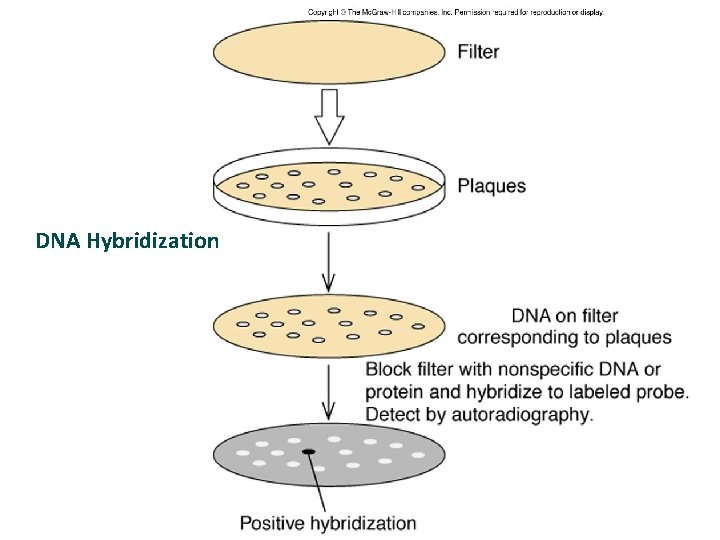
DNA Hybridization

Labeled Tracers • Radioactive tracers allow small quantities of substances to be detected • E. g. RNA quantities of less than pg – not suitable using UV light absorption or staining with dyes • Most widely tracers used: - autoradiography - phosphorimaging - liquid scintillation - scintillation counting

a. Autoradiography • Detecting radioactive compounds with photographic emulsion – x-ray film • Radioactive DNA fragments underwent electrophoresis. Later the gel is placed in contact with x-ray film, leaves in dark • Radioactive emissions from the bands of DNA producing dark bands • Estimating the intensity of the bands by scanning with densitometer • Measures the absorbance of light by the sample – the more intensify the bands, the more it will most light
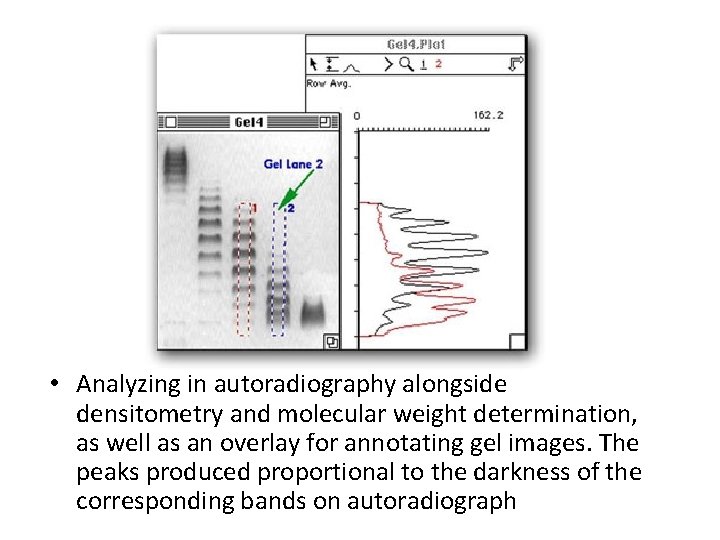
• Analyzing in autoradiography alongside densitometry and molecular weight determination, as well as an overlay for annotating gel images. The peaks produced proportional to the darkness of the corresponding bands on autoradiograph

b. Phosphorimaging • Responding to radioactivity far more linear than of x-ray film • A radioactive sample (blotting DNA bands that have been hybridized with labeled probes) • Sample is placed in contact with phosphorimager plate, absorbing β-rays • The rays excite the molecules until phorimager scans the plate with laser • β-rays energy is released, monitored by computerized detector • Computer converts energy to an image in false color

Nucleosome formation during DNA repair in Xenopus extracts. The picture is a phosphor. Image of a native polyacrylamide gel showing DNA and nucleosome complexes before and after repair. Kosmoski, J. V, Ackerman, E. J. and Smerdon, M. J. (2001). DNA repair of a single photoproduct in a design nucleosome. PNAS. 98

a. DNA Hybridization: Simple Dot Blot • To detect biomolecules • Represents a simplification of the Northern blot, Southern blot, or Western blot methods • The biomolecules to be detected are not first separated by chromatography. Instead, a mixture containing the molecule to be detected is applied directly on a membrane as a dot • Detect by either nucleotide probes (for a Northern blot and Southern blot) or antibodies (for a Western blot)

• However, it offers no information on the size of the target biomolecule • If two molecules of different sizes are detected, they will still appear as a single dot • Dot blots therefore can only confirm the presence or absence of a biomolecule(s) which can be detected by the DNA probes or the antibody
- Slides: 13