Chapter 4 Recombinant DNA Technology Recombinant DNA Technology

Chapter 4 Recombinant DNA Technology

Recombinant DNA Technology n Generation of recombinant DNA molecule ¨ Cloning vector-insert DNA construct (DNA construct) n Cut DNA from a donor organism ¨ n n n Ligation to a cloning vector DNA Transformation ¨ n Cloned DNA, insert DNA, target DNA, foreign DNA Introduction and maintain the DNA construct within a host cell Selection of transformed cells Production of the foreign protein in the host (optional)
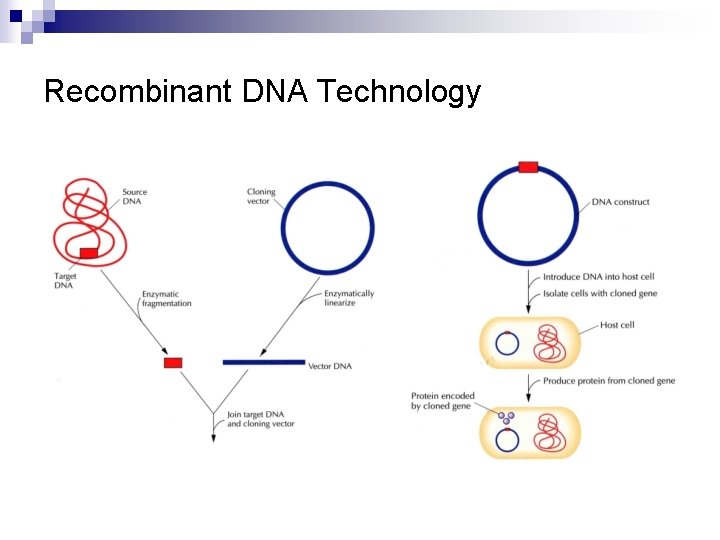
Recombinant DNA Technology

Restriction Endonucleases n Type I: ¨ recognizes n a specific sequence but makes cut elsewhere Type II: ¨ makes n n cut only within the recognition site Eco. RI ¨ E: the genus of the source organism ¨ co: the first two of the species of this organism ¨ R: the strain of origin (Capital, RY 13) ¨ I: the order of discovery (Roman numerals) Bgl. II ¨ B: Bacillus ¨ gl: globigii

Type II Recognition Sequences • Cohesive ends (sticky ends, protruding ends) 5’ overhang: Major, Eco. RI, Bam. HI, etc. 5’ G/AATTC 3’ 3’ CTTAA/G 5’ 5’ G 3’ CTTAA AATTC 3’ G 5’ 3’ overhang: Pst. I, Kpn. I 5’ CTGCA/G 3’ 3’ G/ACGTC 5’ CTGCA 3’ G G 3’ ACGTC 5’ • Blunt ends (flush ends): Sma. I, Eco. RV 5’ CCC/GGG 3’ 3’ GGG/CCC 5’ 5’ CCC 3’ GGG 3’ CCC 5’
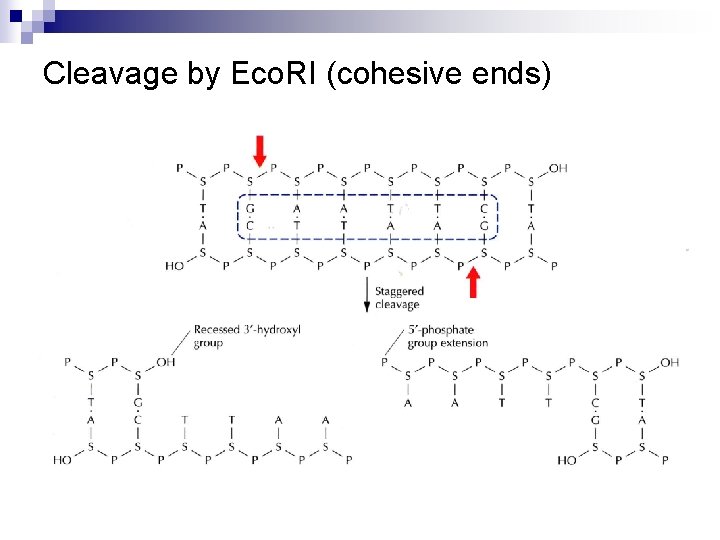
Cleavage by Eco. RI (cohesive ends)
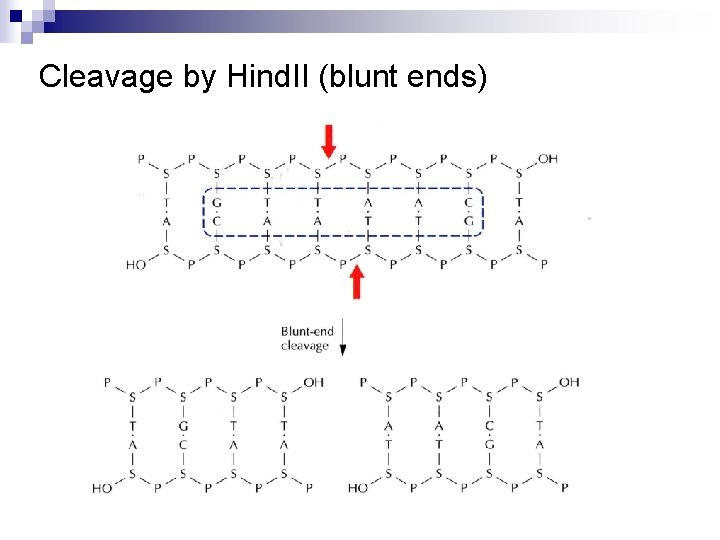
Cleavage by Hind. II (blunt ends)

Palindromic sequence --- The two strands are identical when either is read in the same polarity (5’ 3’). e ye 스위스 소주만병만주소 다시합창합시다 과학은좋은학과 R ace Car Madam , I’m Adam I l ov e Tevol i

Palindrome n AD 79, Pompei ¨ “Sator Arepo Tenet Opera Rotas” SA TOR AREPO TENE T OPERA ROTAS i i i

Number of Restriction enzymes >3000 enzymes from 10, 000 species http: //rebase. neb. com/rebase. html n 4 -Base Cutters Sau 3 AI, Hae. III n 6 -Base Cutters Eco. RI, Bgl. II, Pvu. II n 8 -Base Cutters Not. I, Sbf 1

Restriction Enzymes n Isoschizomer Enzymes that recognize the same target DNA sequence and cleave it in the same way ¨ e. g. Sph. I and Bbu. I (CGTAC/G) ¨ n Neoschizomer Enzymes that recognizes the same target DNA sequence but cleave at different points ¨ e. g. Sma. I (CCC/GGG) and Xma. I (C/CCGGG) ¨ n Isocaudomers Enzymes that produce the same nucleotide extensions but have different recognition sites ¨ e. g. Bam. HI (G/GATCC) and Sau 3 AI (/GATC) ¨
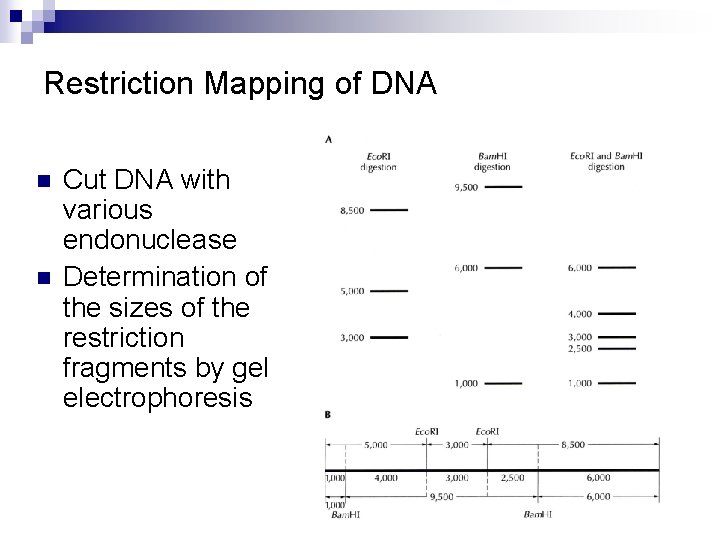
Restriction Mapping of DNA n n Cut DNA with various endonuclease Determination of the sizes of the restriction fragments by gel electrophoresis
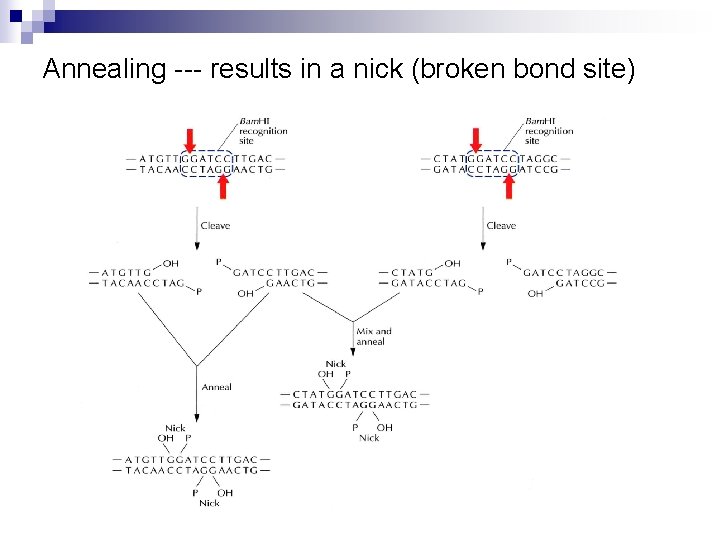
Annealing --- results in a nick (broken bond site)
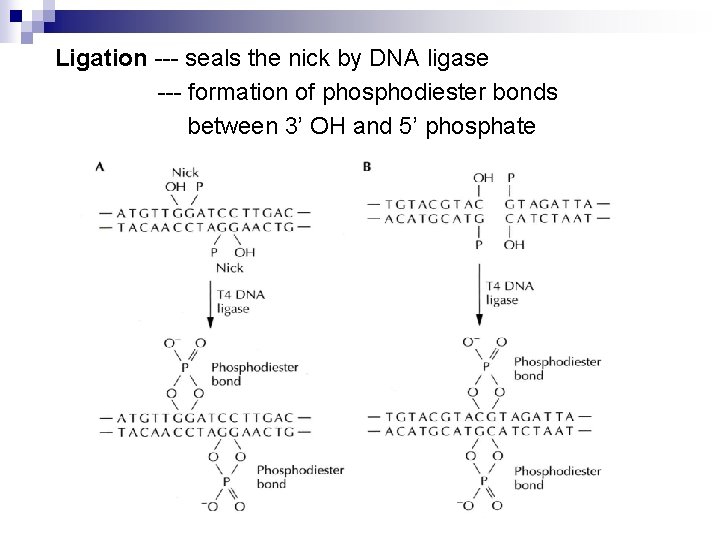
Ligation --- seals the nick by DNA ligase --- formation of phosphodiester bonds between 3’ OH and 5’ phosphate

Ligation n Sticky-end ligation ¨ at low temp for long period ¨ for base pairing n Blunt-end ligation ¨ at room temp ¨ stable base pairing is not required ¨ Blunt-end ligations require 10 to 100 times more DNA ligase than sticky-end ligation.

Ligation Conditions n Temperature Consider enzyme activity and base pairing of cohesive termini ¨ Cohesive ends: 4 -15 o. C: ensure base pairing ¨ Blunt ends: 18 o. C, use 10 to 100 times higher concentration of T 4 DNA ligase ¨ n DNA concentration Dilute concentration favors circulization of linear fragment ¨ Insert : Vector = 2 : 1 molar ratio ¨ n Phosphatase treatment ¨ Prevention of self ligation of vector
- Slides: 16