Chapter 4 Gene expression Acknowledgement Centers for Disease




















































- Slides: 66

Chapter 4 Gene expression

Acknowledgement • Centers for Disease Control and Prevention- Ethiopia (CDC-E) • American Society for Clinical Pathology (ASCP) • Addiss Ababa University • University of Gondar • University of Hawassa • Jimma University • Haromaya University

Learning objectives At the end of this chapter, students will be able to: • Discuss transcription, translation and gene expression control • Describe the genetic code • List the enzymes associated with transcription, translation & describe their specific functions.

Outline 1. Introduction to gene expression 2. Transcription – Introduction • components of transcription – Steps in transcription 3. Translation – Introduction – Steps in translation – The genetic code 4. The control gene expression in Eukaryotes and prokaryotes 5. Review questions 6. References

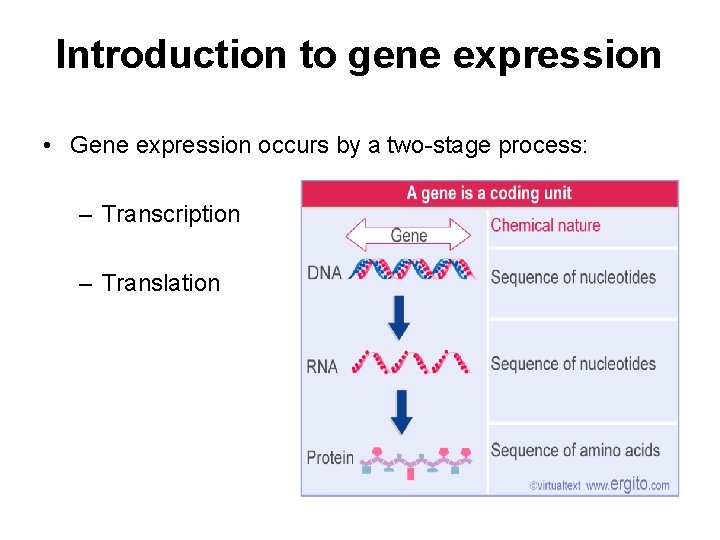
Introduction to gene expression • Gene expression occurs by a two-stage process: – Transcription – Translation

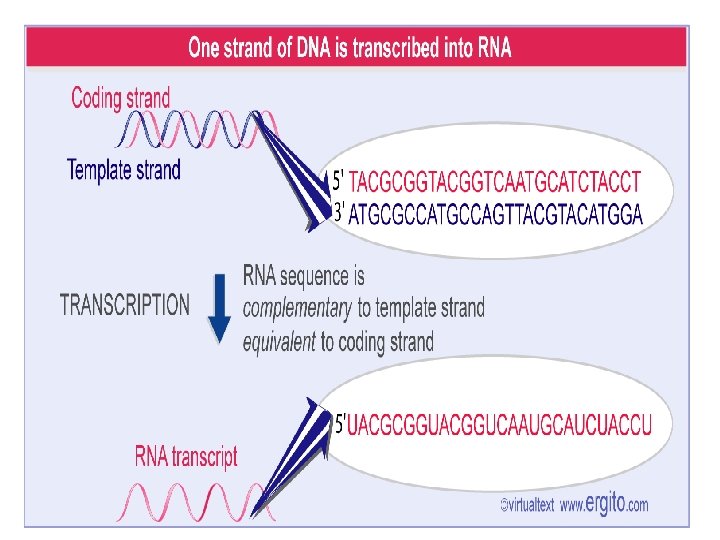
4. 1 Transcription: Introduction • Is the synthesis of an RNA chain representing one strand of a DNA duplex • Transcription is the first step in the expression of the genetic information stored in the genome: from gene to gene product (frequently a protein). • It became clear that DNA could not be directly converted into protein, there had to be an intermediate step. • This unstable intermediate product was identified as messenger RNA (m. RNA) • RNA is identical in sequence with one strand of the DNA, which is called the coding strand


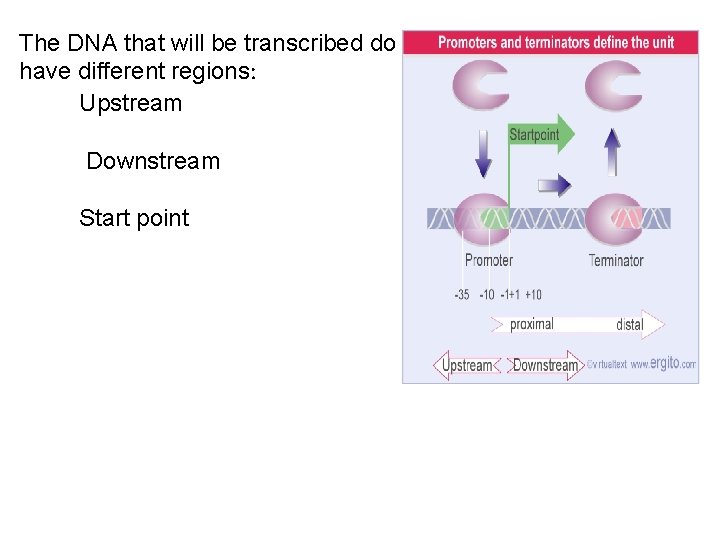
The DNA that will be transcribed do have different regions: Upstream Downstream Start point

• Transcription starts when RNA polymerase binds to a special region, the promoter, at the start of the gene. The promoter surrounds the first base pair that is transcribed into RNA, the startpoint. From this point, RNA polymerase moves along the template, synthesizing RNA, until it reaches a terminator sequence. • A transcription unit is a sequence of DNA • transcribed into a single RNA, starting at the promoter and ending at the terminator

• Base positions are numbered in both directions away from the start point, which is assigned the value +1; numbers are increased going downstream. • The base before the start point is numbered – 1, and the negative numbers increase going upstream. (There is no base assigned the number 0). • The immediate product of transcription is called the primary transcript (from promoter to terminator).

• Transcription consists in the production of an RNA copy of one of the two strands of a double-stranded DNA template only, by a process that relies on the complementary pairing of bases. • In the chromosome overall, both DNA strands are used as template but in any one gene, only one strand is used • Transcription is chemically and enzymatically very similar to DNA replication. Both processes involve enzymes that synthesize a new strand of nucleic acid complementary to a DNA template strand (synthesis always proceeds 5' to 3'). • But, there are some very important differences:

(i) in transcription the new strand is made of ribonucleotides (instead of deoxyribonucleotides) and thymine is replaced by uracil (ii) RNA polymerase is used to synthesize RNA: • Bacteria and Archaea generally have a single RNA polymerase that synthesizes the unstable m. RNA molecules and the stable functional RNAs: ribosomal RNA (r. RNA) and transfer RNA (t. RNA). • Eukaryotes have three RNA polymerases: Pol I that produces the r. RNA precursor, Pol II that produces all the m. RNAs and some sn. RNAs (small nuclear RNA), and Pol III that synthesizes t. RNAs, 5 S r. RNA and some sn. RNAs. • RNA polymerase does not require a primer (in contrast to DNA polymerases). It can initiate RNA synthesis de novo.

(iii) The RNA product does not remain base-paired to the template DNA: • The enzyme displaces the growing chain only a few nucleotides behind where each nucleotide is added. – This displacement is crucial for subsequent translation of the m. RNA into protein – Multiple RNA polymerase molecules can transcribe the same gene at the same time. (iv)Though transcription is a rather accurate process (about one mistake in 104 nucleotides added) – it is less accurate than DNA replication (about one mistake in every 107 nucleotides added).

(v) Transcription selectively copies only certain parts of the genome, and proceeds with very different efficiencies for different genes: • Specific signals and mechanisms are used to define the start and termination sites of transcripts, and the frequency of transcription is highly regulated and proceeds in function of the cell's needs for the corresponding gene products. • In contrast, DNA replication must copy the entire genome and make exactly one copy every cell division.

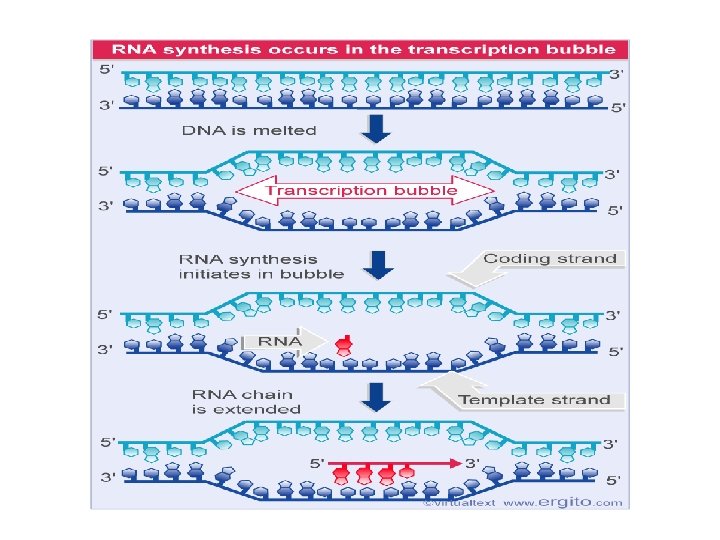
Steps in transcription Initiation • binding of RNA polymerase to double-stranded DNA (no priming required). • involves a transition to single-strandedness in the region of binding. • RNA polymerase binds at a sequence of DNA called the promoter. Elongation • the covalent addition of nucleotides to the 3' end of the growing polynucleotide chain. • involves the development of a short stretch of DNA that is transiently single-stranded Termination • the recognition of the transcription termination sequence and the release of RNA polymerase

Steps in transcription---Initiation • Describes the synthesis of the first nucleotide bonds in RNA. • The enzyme remains at the promoter while it synthesizes the first ~9 nucleotide bonds. • The initiation phase is protracted by the occurrence of abortive events, in which the enzyme makes short transcripts, releases them, and then starts synthesis of RNA again. • The initiation phase ends when the enzyme succeeds in extending the chain and clears the promoter. • The sequence of DNA needed for RNA polymerase to bind to the template and accomplish the initiation reaction defines the promoter. • Initiation is accomplished if and when the enzyme manages to move along the template to move the next region of the DNA into the active site.

Steps in transcription---Initiation • Although transcription is performed by RNA Polymerase, the enzyme needs other proteins (transcription factors) to produce the transcript. • Transcription factor - any protein other than RNA Polymerase that is required for transcription. Functions of Transcription Factors – bind to RNA Polymerase – bind another transcription factor – bind to cis-acting DNA sequences

Steps in transcription---Initiation • RNA Polymerase and the group of protein that directly interact with it are called the basal transcription apparatus. • Basal transcription apparatus - RNA polymerase + general factors; both needed to initiate transcription. • Upstream factors - ubiquitous factors that increase the efficiency of transcription initiation; set of factors unique to each promoter

Steps in transcription---Initiation Functions of Upstream Factors – Influence the initiation of transcription by contacting members of the basal apparatus – Promotes assembly of the apparatus – May bind co activators that interact with the basal apparatus – Some are inducible factors. • Promoter - all the DNA sequences containing binding sites for RNA polymerase and the transcription factors necessary for normal transcription

Steps in transcription---Initiation • Promoter recognition depends on consensus Sequences: • There are four (perhaps five) conserved features in a bacterial promoter: – the start point – the – 10 sequence – the – 35 sequence – the separation between the – 10 and – 35 sequences; and (sometimes) the UP element

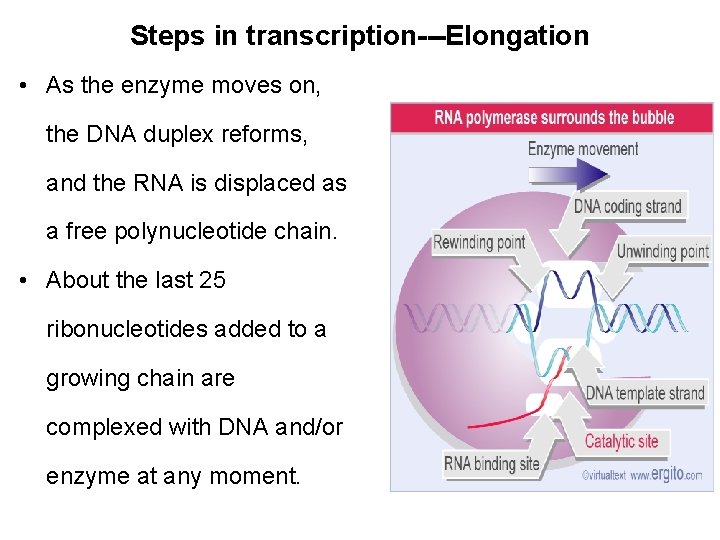
Steps in transcription---Elongation • As the enzyme moves on, the DNA duplex reforms, and the RNA is displaced as a free polynucleotide chain. • About the last 25 ribonucleotides added to a growing chain are complexed with DNA and/or enzyme at any moment.

Steps in transcription---Elongation

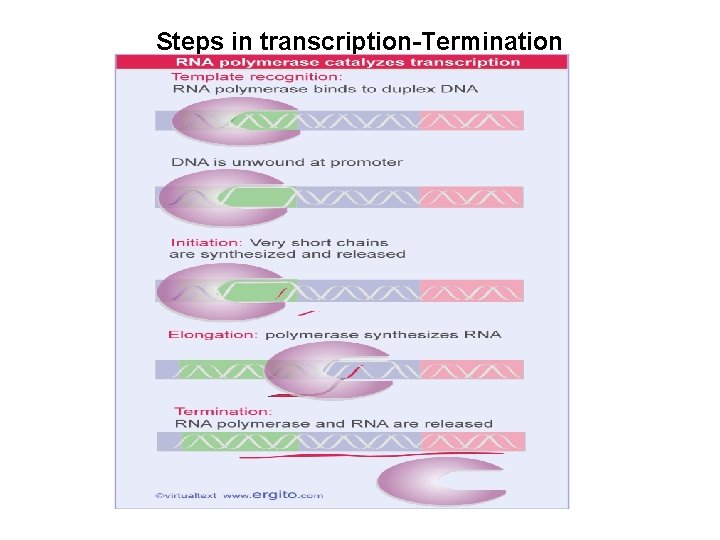
Steps in transcription-Termination • When polymerase meets a termination structure (hairpin followed by a stretch of T residues, or protein dependent terminator) : – The ternary complex becomes unstable and dissociates – And hence, both the DNA and the RNA dissociate from the complex. – The DNA reforms in duplex state – The enzyme and RNA are both released

Steps in transcription-Termination

4. 2 Translation Introduction • It is the process that translates the m. RNA sequence to protein • Translation can be seen to occur in two phases: 1. information transfer, in which RNA base sequence of the m. RNA determines the sequence of amino acids and 2. chemical processes, in which the peptide bonds between the adjacent amino acids are formed.

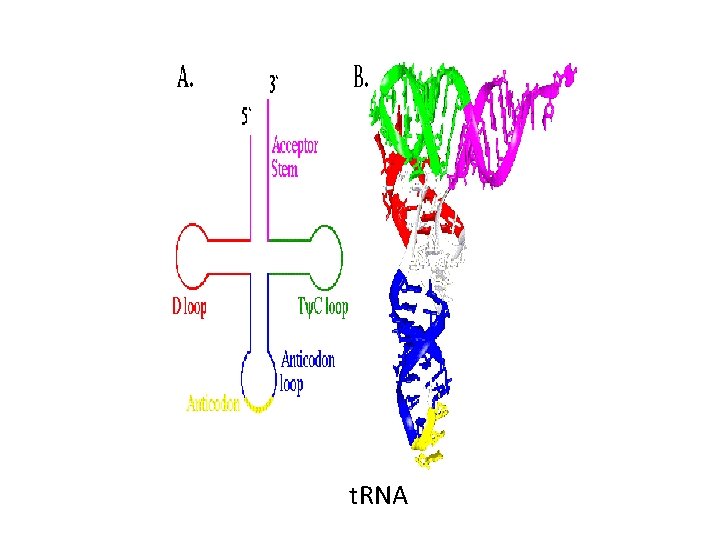
Translation cont’d-- • The components required for translation include: – m. RNA, – ribosomes, – t. RNA, – aminoacyl t. RNA synthetases, – Initiation, elongation and termination factors

t. RNA

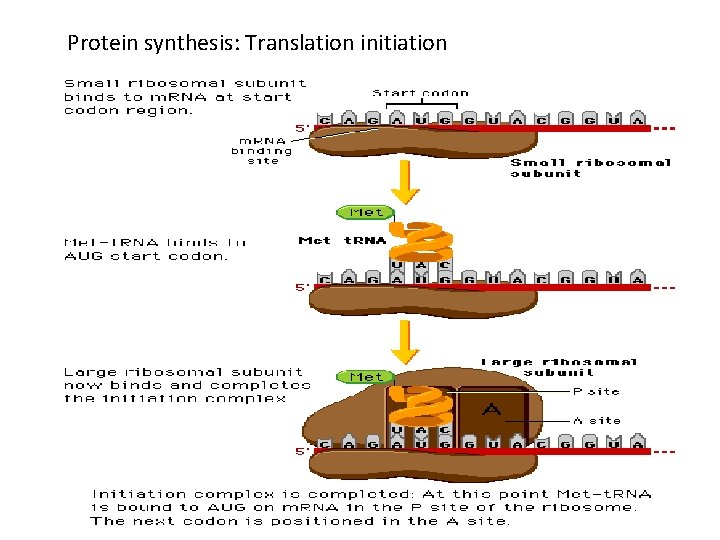
Translation: initiation • Ribosome small subunit binds to m. RNA • Charged t. RNA anticodon forms base pairs with the m. RNA codon • Small subunit interacts with initiation factors and special initiator t. RNA that is charged with methionine • m. RNA-small subunit-t. RNA complex recruits the large subunit • Eukaryotic and prokaryotic initiation differ slightly

Translation: initiation • The large subunit of the ribosome contains three binding sites – Amino acyl (A site) – Peptidyl (P site) – Exit (E site) • At initiation, – The t. RNAf. Met occupies the P site – A second, charged t. RNA complementary to the next codon binds the A site.

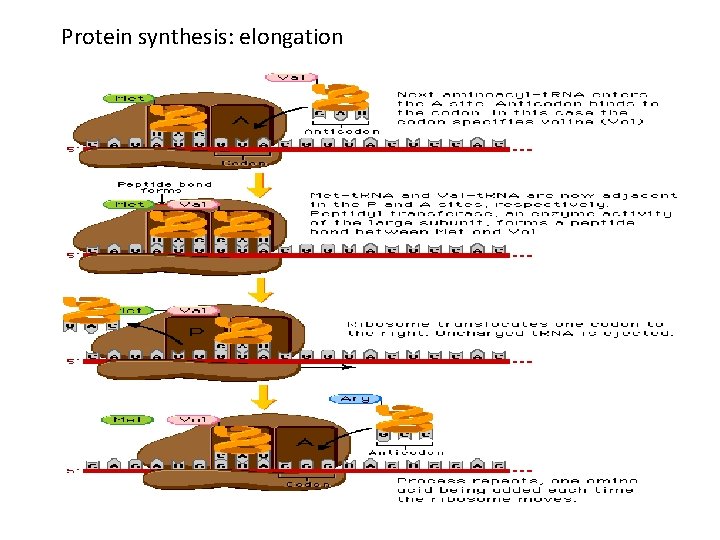
Translation: elongation • Ribosome translocates by three bases after peptide bond formed • New charged t. RNA aligns in the A site • Peptide bond between amino acids in A and P sites is formed • Ribosome translocates by three more bases • The uncharged t. RNA in the A site is moved to the E site.

Translation: elongation • EF-Tu recruits charged t. RNA to A site. Requires hydrolysis of GTP • Peptidyl transferase catalyzes peptide bond formation (bond between aa and t. RNA in the P site converted to peptide bond between the two amino acids) • Peptide bond formation requires RNA and may be a ribozyme-catalyzed reaction

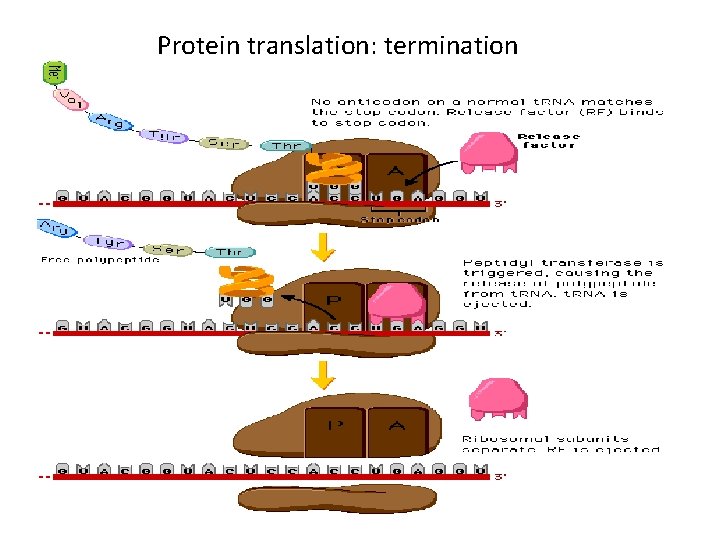
Translation: termination • Elongation proceeds until STOP codon reached – UAA, UAG, UGA • No t. RNA normally exists that can form base pairing with a STOP codon; recognized by a release factor • t. RNA charged with last amino acid will remain at P site • Release factors cleave the amino acid from the t. RNA • Ribosome subunits dissociate from each other

Protein synthesis: Translation initiation

Protein synthesis: elongation

Protein translation: termination

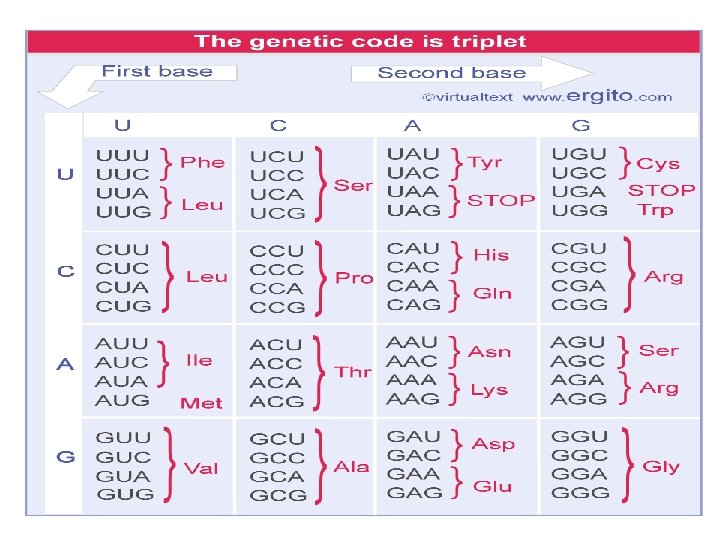
The Genetic Code • The sequence of a coding strand of DNA, read in the direction from 5ʹ to 3ʹ , consists of nucleotide triplets (codons) corresponding to the amino acid sequence of a protein read from N-terminus to C-terminus. • There are 64 codons (each of 4 possible nucleotides can occupy each of the three positions of the codon, making 43 = 64 possible trinucleotide sequences). • Each of these codons has a specific meaning in protein synthesis: – 61 codons represent amino acids; – 3 codons cause the termination of protein synthesis.


THE GENETIC CODE … • Most amino acids are represented by more than one codon • The multiple codons for an amino acid are usually related • Related amino acids often have related codons, minimizing the effects of mutation.

Prokaryotic transcription and translation • Takes place in the same compartment. • Transcription and translation happens almost at the same time • The m. RNA do have short half life as it will get degraded immediately. • Prokaryotic m. RNA is polycistronic(includes coding regions representing more than one gene). • Initiation of translation begins with N-formyl-methionine • Initiation of translation at special sequence in on m. RNA called the ribosome binding site. • No introns • Co-linearity between DNA/RNA/Protein

Eukaryotic transcription and translation • Transcription and translation takes place in different compartments: – Transcription in nucleus – Translation in cytolaplasm • The first transcription results in primary transcript (m. RNA precursor that still contains introns copied from the gene). • m. RNA mature before being translated through posttranslational modification – Splicing- removal of introns – 5’ capping – 3’ polyadenylation

Eukaryotic transcription and translation: 5’ Capping • A 5 cap is formed by adding a G to the terminal base of the transcript via a 5 – 5 link Functions of capping: • At least four functions – Protect m. RNA from degradation – Enhance the translatability of m. RNAs – Enhance the transport of m. RNAs from the nucleus into cytoplasm – Enhance the efficiency of splicing of m. RNAs

Eukaryotic transcription and translation : 3’ Polyadenylation • Most eukaryotic m. RNAs and their precursors have a chain of AMP residues about 250 nt long at their 3’ – end. • It is added by poly (A) polymerase. Function: • Protection – When poly. A tail too short, m. RNA degraded • Translatability – Synergistic stimulation with Cap

Eukaryotic transcription and translation • The mature m. RNA will be transported from nucleus to cytoplasm. • Eukaryotic initiator t. RNA is a Met-t. RNA that is different from the Met-t. RNA used in elongation, but the methionine is not formylated • Eukaryotic m. RNA is monocistronic. • Exons separated by introns ( a sequence present in the gene but absent in m. RNA). • No co-linearity between DNA and m. RNA/Protein

Eukaryotic transcription and translation: Post translational modifications • Most of the proteins that are translated from m. RNA undergo chemical modifications before becoming functional in different body cells. • Post Translational modifications occurring at the peptide terminus of the amino acid chain play an important role in translocating them across biological membranes.

• Can also take place in secretory proteins of prokaryotes and eukaryotes and also proteins that are intended to be incorporated in various cellular and organelle membranes such as lysosomes, chloroplast, mitochondria and plasma membranes. • The amino terminal sequences are removed by proteolytic cleavage when the proteins cross the membranes. These amino terminal sequences target the proteins /transporting them /to their actual point of action in the cell

Some examples of Post translational modifications in Eukaryotic cells • Glycosylation: the addition of carbohydrate moiety. • Acetylation: the addition of an acetyl group, usually at the N -terminus of the protein. • Alkylation: The addition of an alkyl group (e. g. methyl, ethyl). • Methylation: The addition of a methyl group, usually at lysine or arginine residues. (This is a type of alkylation. ) • etc…. .

4. 3. The control of gene expression in prokaryotes and eukaryotes Introduction • The controls of gene expression are much more complex in eukaryotes than in prokaryotes. • A major difference is the presence in eukaryotes of a nuclear membrane • In prokaryotes, control of transcriptional initiation is the major point of regulation, • In eukaryotes the regulation of gene expression is controlled nearly equivalently from many different points.

Gene Control in Prokaryotes • In bacteria, genes are clustered into operons: – gene clusters that encode the proteins necessary to perform coordinated function • RNA that is transcribed from prokaryotic operons is polycistronic • In bacteria, control of the rate of transcriptional initiation is the predominant site for control of gene expression.

Gene Control in Prokaryotes … • initiation is controlled by two DNA sequence elements, – identified as the -35 and -10 positions. – These 2 sequence elements are termed promoter sequences, – because they promote recognition oftranscriptional start sites by RNA polymerase. – The consensus sequence for the -35 position is TTGACA, and for the -10 position, TATAAT. – These promoter sequences are recognized and contacted by RNA polymerase.

Gene Control in Prokaryotes … • The activity of RNA polymerase at a given promoter is in turn regulated by interaction with accessory proteins, – which affect its ability to recognize start sites. – These regulatory proteins can act both positively (activators) and negatively (repressors). • The accessibility of promoter regions of prokaryotic DNA is in many cases regulated by the interaction of proteins with sequences termed operators. • The operator region is adjacent to the promoter elements in most operons and in most cases the sequences of the operator bind a repressor protein.

Gene Control in Prokaryotes …

Gene Control in Prokaryotes … • As indicated above, prokaryotic genes that encode the proteins necessary to perform coordinated function are clustered into operons. • Two major modes of transcriptional regulation function in bacteria (E. coli) to control the expression of operons. Both mechanisms involve repressor proteins: • One mode of regulation is exerted upon operons that produce gene products necessary for the utilization of energy; these are catabolite-regulated operons. • The other mode regulates operons that produce gene products necessary for the synthesis of small biomolecules such as amino acids. Expression from this class of operons is attenuated by sequences within the transcribed RNA.

Gene Control in Prokaryotes … • A classic example of a catabolite-regulated operon is the lac operon, responsible for obtaining energy from βgalactosides such as lactose. • A classic example of an attenuated operon is the trp operon, responsible for the biosynthesis of tryptophan.

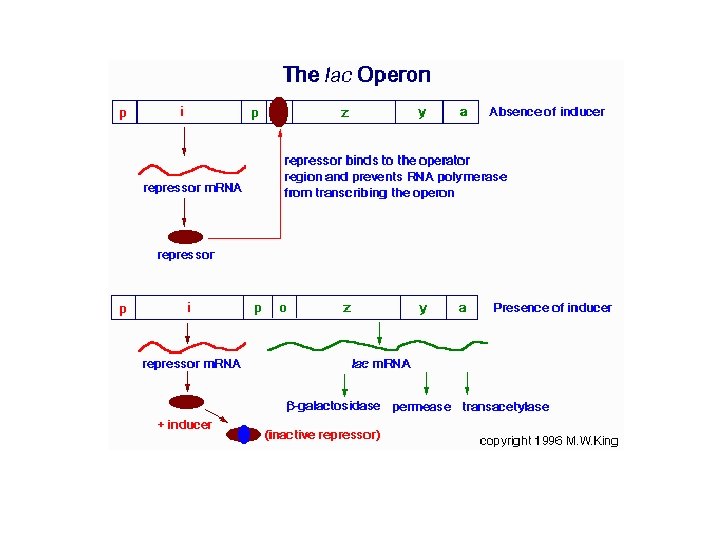
Gene Control in Prokaryotes … The lac Operon • The lac operon (see diagram below) consists of one regulatory gene (the i gene) and three structural genes (z, y, and a). – The i gene codes for the repressor of the lac operon. – The z gene codes for β-galactosidase (β-gal), for the hydrolysis of the disaccharide, lactose into its monomeric units, galactose and glucose. – The y gene codes for permease, increases permeability of the cell to β-galactosides. – The a gene encodes a transacetylase.

The lac Operon … • During normal growth on a glucose-based medium, the lac repressor is bound to the operator region of the lac operon, preventing transcription. • However, in the presence of an inducer of the lac operon, the repressor protein binds the inducer and is rendered incapable of interacting with the operator region of the operon. – RNA polymerase is thus able to bind at the promoter region, and transcription of the operon ensues. – The lac operon is repressed, even in the presence of lactose, if glucose is also present. This repression is maintained until the glucose supply is exhausted.

The lac Operon … • The repression of the lac operon under these conditions is termed catabolite repression and is a result of the low levels of c. AMP that result from an adequate glucose supply. – The repression of the lac operon is relieved in the presence of glucose if excess c. AMP is added. • As the level of glucose in the medium falls, the level of c. AMP increases. – Simultaneously there is an increase in inducer binding to the lac repressor. – The net result is an increase in transcription from the operon.

• The ability of c. AMP to activate expression from the lac operon results from an interaction of c. AMP with a protein termed CRP (for c. AMP receptor protein). – This protein is also called CAP (for catabolite activator protein). • The c. AMP-CRP complex binds to a region of the lac operon just upstream of the region bound by RNA polymerase and that somewhat overlaps that of the repressor binding site of the operator region. • The binding of the c. AMP-CRP complex to the lac operon stimulates RNA polymerase activity 20 -to-50 -fold.


Gene Control in Eukaryotes: • In eukaryotic cells, the ability to express biologically active proteins comes under regulation at several points: 1. Chromatin Structure: The physical structure of the DNA, as it exists compacted into chromatin • Can affect the ability of transcriptional regulatory proteins (termed transcription factors) • affect the ability of RNA polymerases to find access to specific genes and to activate transcription from them. • The presence of the histones and Cp. G methylation most affect accessibility of the chromatin to RNA polymerases and transcription factors

Gene Control in Eukaryotes … 2. Transcriptional Initiation: • This is the most important mode for control of eukaryotic gene expression. Specific factors that exert control include: • the strength of promoter elements within the DNA sequences of a given gene, • the presence or absence of enhancer sequences (which enhance the activity of RNA polymerase at a given promoter by binding specific transcription factors), • and the interaction between multiple activator proteins and inhibitor proteins.

Gene Control in Eukaryotes … 3. Transcript Processing and Modification: • Eukaryotic m. RNAs must be capped and polyadenylated, • and the introns must be accurately removed. • Several genes have been identified that undergo tissuespecific patterns of alternative splicing, which generate biologically different proteins from the same gene.

Gene Control in Eukaryotes … 4. RNA Transport: • A fully processed m. RNA must leave the nucleus in order to be translated into protein. 5. Transcript Stability: • Unlike prokaryotic m. RNAs, whose half-lives are all in the range of 1 -5 minutes, eukaryotic m. RNAs can vary greatly in their stability. – Certain unstable transcripts have sequences (predominately, but not exclusively, in the 3'-UTR) that are signals for rapid degradation. 6. Translational Initiation: • Since many m. RNAs have multiple methionine codons, the ability of ribosomes to recognize and initiate synthesis from the correct AUG codon can affect the expression of a gene product.

Gene Control in Eukaryotes … 7. Post-Translational Modification: • Common modifications include glycosylation, acetylation, fatty acylation, disulfide bond formations, etc. – Without these proper modifications, the protein may not have adequate or specific function. 8. Protein Transport: • In order for proteins to be biologically active following translation and processing, they must be transported to their site of action. 9. Control of Protein Stability: • Many proteins are rapidly degraded, whereas others are highly stable. Specific amino acid sequences in some proteins have been shown to bring about rapid degradation.

Summary • DNA functions to serve as stable storage and transmission of genetic information • RNA polymerases synthesize specific types of RNA: t. RNA, r. RNA or m. RNA by using DNA as a template • Nucleotide sequence of m. RNA is translated to protein by a process called translation

Review questions 1. List the two steps in gene expression 2. Discuss transcription and translation 3. What are the post transcription modification of m. RNA and why they are needed? 4. List important enzymes in transcription and translation

References • Robert F. weaver, Philip W. Hedrick. Genetics. • Darnel, Lodish, Baltimore. Molecular Cell Biology • James D. Watson: Recombinant DNA • Robert F. Weaver. Molecular biology • Richard J. Epistein: Human Molecular Biology • P. K. Gupta: Cell and Molecular Biology • Tarek H. EL-Metwally. Basic Medical Molecular Biology: A Comprehensive update.