Chapter 4 DistanceBased Methods of Phylogenetics HUANG GuanShieng

Chapter 4 Distance–Based Methods of Phylogenetics 暨南大學資訊 程學系 黃光璿 (HUANG, Guan-Shieng) 2004/03/29 1

Motivation n Evolution events on genomes: q q n substitutions insertions deletions rearrangements We focus on cluster analysis in this chapter. 2


8

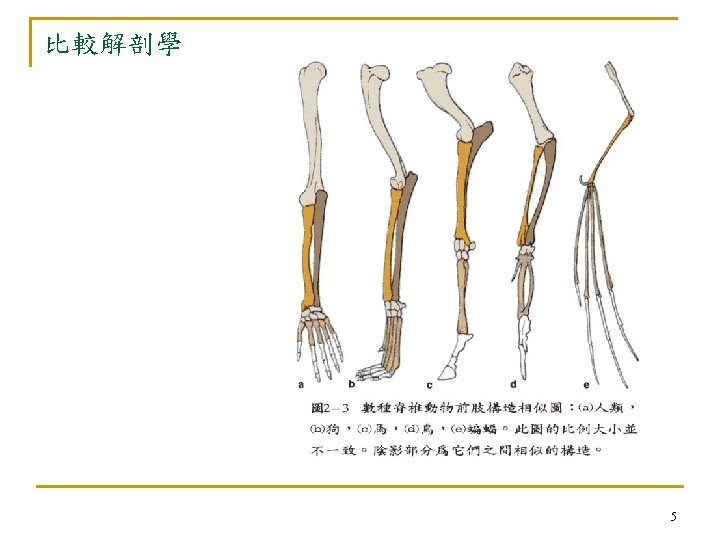
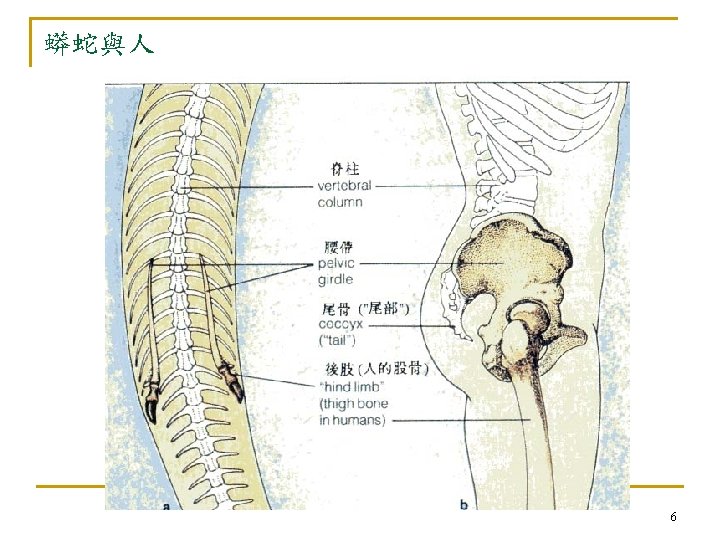
4. 2 Advantages of Molecular Phylogenies n fundamental q q n evolution is defined as genetic changes molecular clock hypothesis (Chap. 3) In early days, taxonomists inferred genotypes from phenotypes. q q phenotypes(表現型): how organisms looks genotypes: the genes that gave rise to their physical appearance 9

n And then q q q behavior (行為) ultrastructural (超顯微結構) biochemical characteristics were studied. 10

n 傳統研究方法有以下問題無法解決 q convergent evolution n q many organisms do not have easily studied phenotypic features n q 眼睛:humans, flies, mollusks (軟體動物) bacteria (細菌) comparing distantly related organisms n n bacteria, worms, mammals few characteristics in common! 11
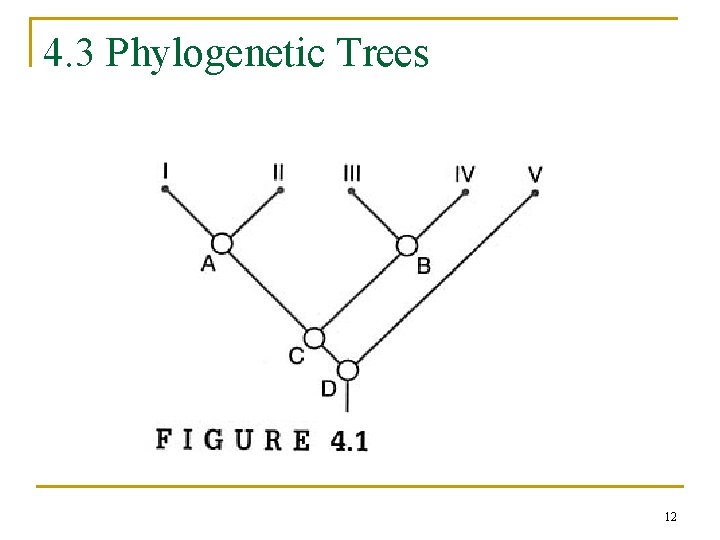
4. 3 Phylogenetic Trees 12
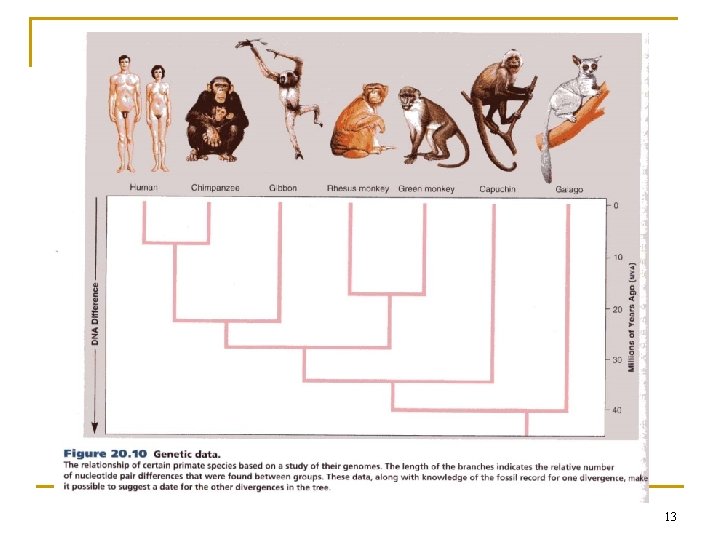
13

4. 3. 1 Terminology of Tree Reconstruction n phylogenetic tree, or dendrogram q q q nodes: taxonomical units branches terminal nodes n q internal nodes n n collected data (I, III, IV, V) inferred ancestors (A, B, C, D) Newick format q (((I, II), (III, IV)), V) 14
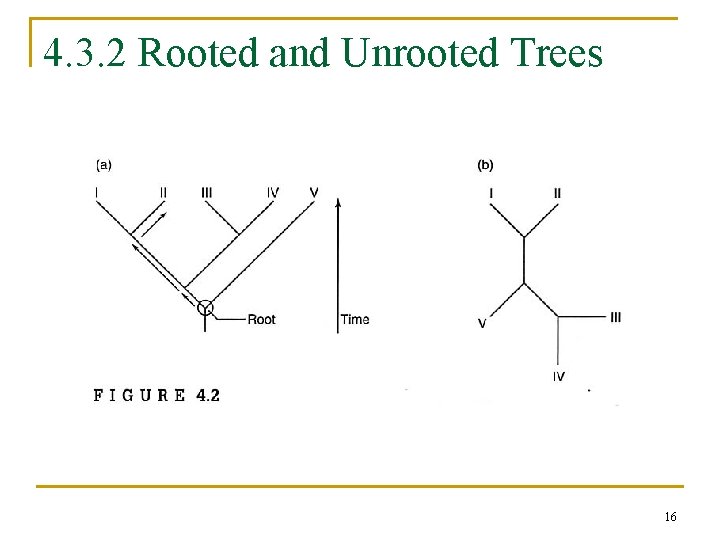
4. 3. 2 Rooted and Unrooted Trees 16
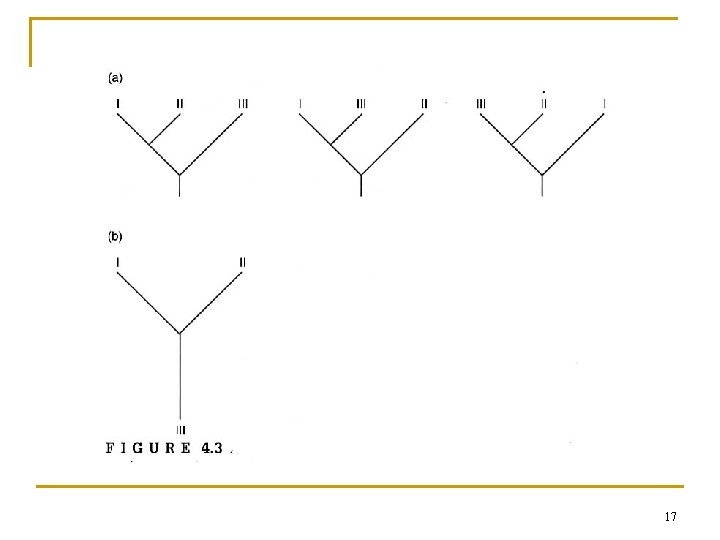
17

4. 3. 4 Character and Distance Data n characters (特質) q n distance q n n n DNA sequences, protein sequences, color, behavior, response time, …… overall, pairwise difference character data distance data pheneticist: prefers distance based methods cladist: prefer character based methods 21

4. 4 Distance Matrix Methods n n UPGMA (Unweighted-Pair-Group Method with Arithmetic mean) Transformed Distance Method Neighbor’s Relation Method Neighbor-Joining Method 22
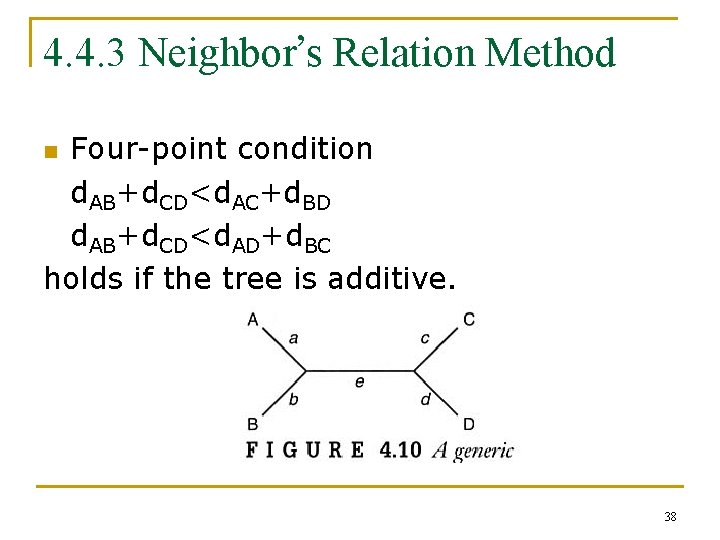
4. 4. 3 Neighbor’s Relation Method Four-point condition d. AB+d. CD<d. AC+d. BD d. AB+d. CD<d. AD+d. BC holds if the tree is additive. n 38

Given any four points, say A, B, C, D, we have d. AB+d. CD d. AC+d. BD d. AD+d. BC. The smallest indicates how to pair up. n 39

n n S. Sattath & A. Tversky, 1977 For any four points, say A, B, C, D, compute d. AB+d. CD d. AC+d. BD d. AD+d. BC. The smallest should be paired, and wins a score 1 for each pair. After trying all possible quadruples, the pair wins the highest scores is grouped. 40

Example 41
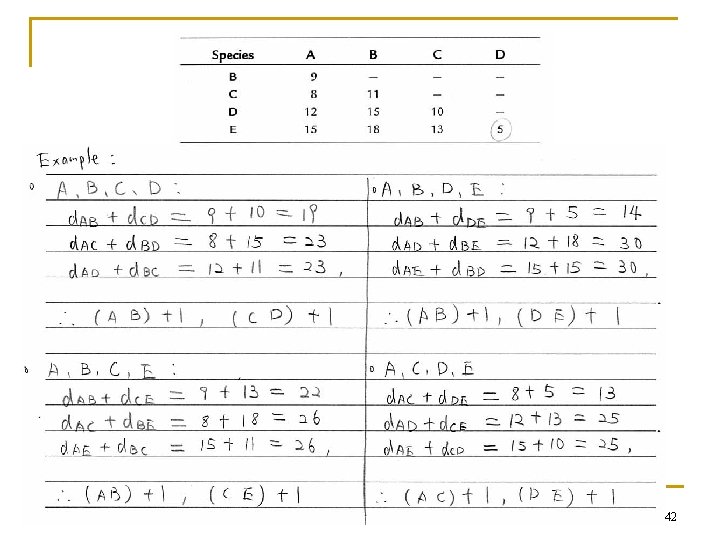
42
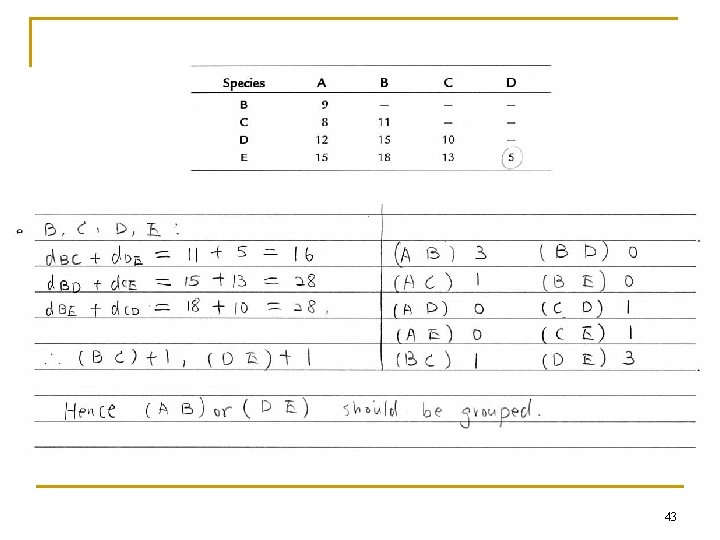
43

n The length of the branches can be determined by the outgroup method. 44

Theorem n If a matrix is additive, then its phylogenetic tree (unrooted, binary) can be reconstructed correctly and uniquely by the Neighbor’s Relation Method. 45

4. 4. 4 Neighbor-Joining Methods 46

47
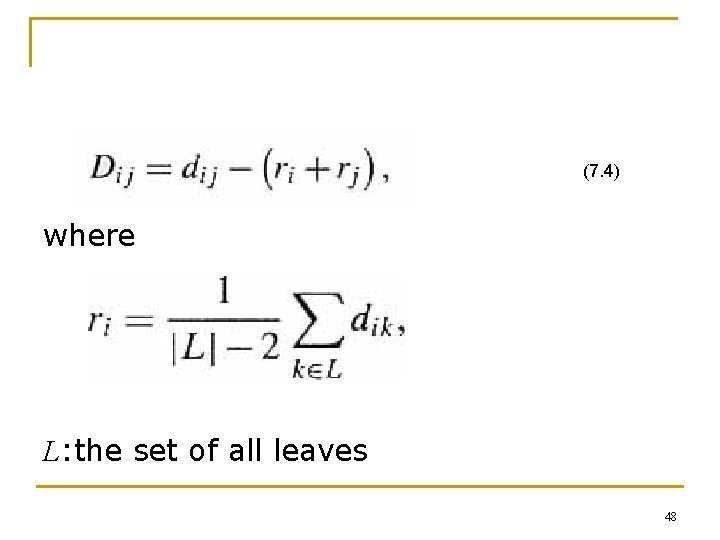
(7. 4) where L: the set of all leaves 48

49

Theorem n If a matrix is additive, then its phylogenetic tree (unrooted, binary) can be reconstructed correctly and uniquely by the Neighbor-Joining Method. 50

參考資料及圖片出處 1. 2. 3. Fundamental Concepts of Bioinformatics Dan E. Krane and Michael L. Raymer, Benjamin/Cummings, 2003. Biological Sequence Analysis – Probabilistic models of proteins and nucleic acids R. Durbin, S. Eddy, A. Krogh, G. Mitchison, Cambridge University Press, 1998. Biology, by Sylvia S. Mader, 8 th edition, Mc. Graw-Hill, 2003. 53
- Slides: 30