Chapter 4 Clusters and repeats b Gene clusters

Chapter 4 Clusters and repeats

b Gene clusters are formed by duplication and divergence b Sequence divergence is the basis for the evolutionary clock b Pseudogenes are dead ends of evolution b Unequal crossing-over rearranges gene clusters b Genes for r. RNA form tandem repeats b ( The repeated genes for r. RNA maintain constant sequence) b Crossover fixation could maintain identical repeats b Satellite DNAs often lie in heterochromatin b Arthropod satellites have very short identical repeats b Mammalian satellites consist of hierarchical repeats b Minisatellites are useful for genetic mapping

4. 1 Introduction Gene cluster is a group of adjacent genes that are identical or related. Gene family consists of a set of genes whose exons are related; the members were derived by duplication and variation from some ancestral gene. Satellite DNA consists of many tandem repeats (identical or related) of a short basic repeating unit. Unequal crossing-over describes a recombination event in which the two recombining sites lie at nonidentical locations in the two parental DNA molecules.
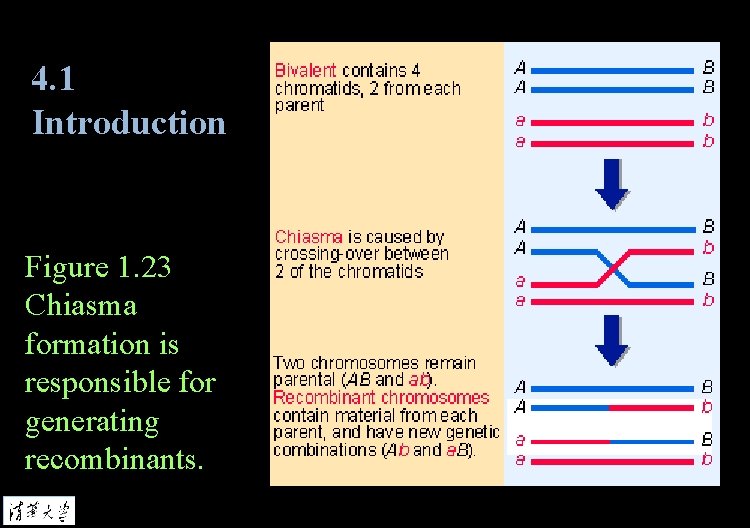
4. 1 Introduction Figure 1. 23 Chiasma formation is responsible for generating recombinants.
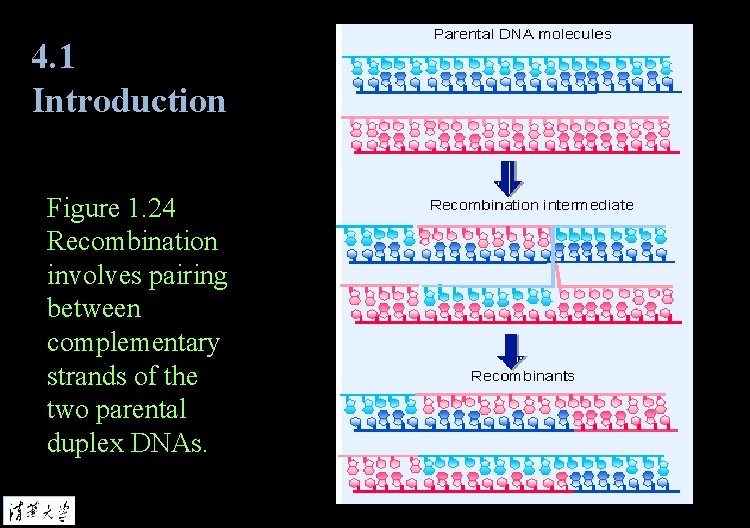
4. 1 Introduction Figure 1. 24 Recombination involves pairing between complementary strands of the two parental duplex DNAs.


4. 2 Gene clusters are formed by duplication and divergence Figure 2. 13 All functional globin genes have an interrupted structure with three exons. The lengths indicated in the figure apply to the mammalian bglobin genes.
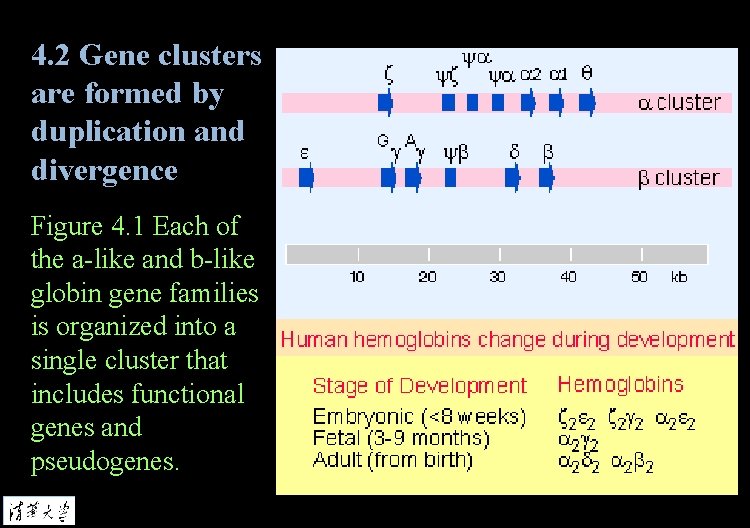
4. 2 Gene clusters are formed by duplication and divergence Figure 4. 1 Each of the a-like and b-like globin gene families is organized into a single cluster that includes functional genes and pseudogenes.

4. 2 Gene clusters are formed by duplication and divergence Figure 4. 2 Clusters of b-globin genes and pseudogenes are found in vertebrates. Seven mouse genes include 2 early embryonic, 1 late embryonic, 2 adult genes, and 2 pseudogenes. Rabbit and chick each have four genes.

4. 2 Gene clusters are formed by duplication and divergence Figure 4. 3 All globin genes have evolved by a series of duplications, transpositions, and mutations from a single ancestral gene.

4. 2 Gene clusters are formed by duplication and divergence Figure 4. 3 All globin genes have evolved by a series of duplications, transpositions, and mutations from a single ancestral gene.

4. 3 Sequence divergence is the basis for the evolutionary clock Divergence is the percent difference in nucleotide sequence between two related DNA sequences or in amino acid sequences between two proteins. Evolutionary clock is defined by the rate at which mutations accumulate in a given gene. Replacement sites in a gene are those at which mutations alter the amino acid that is coded.
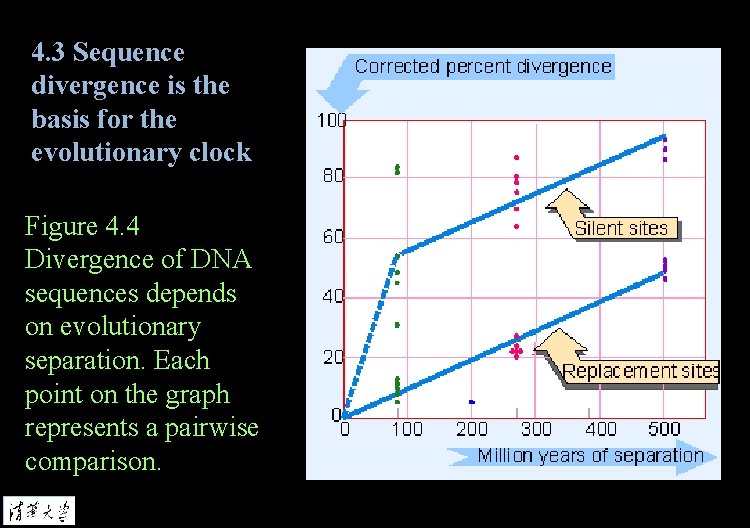
4. 3 Sequence divergence is the basis for the evolutionary clock Figure 4. 4 Divergence of DNA sequences depends on evolutionary separation. Each point on the graph represents a pairwise comparison.
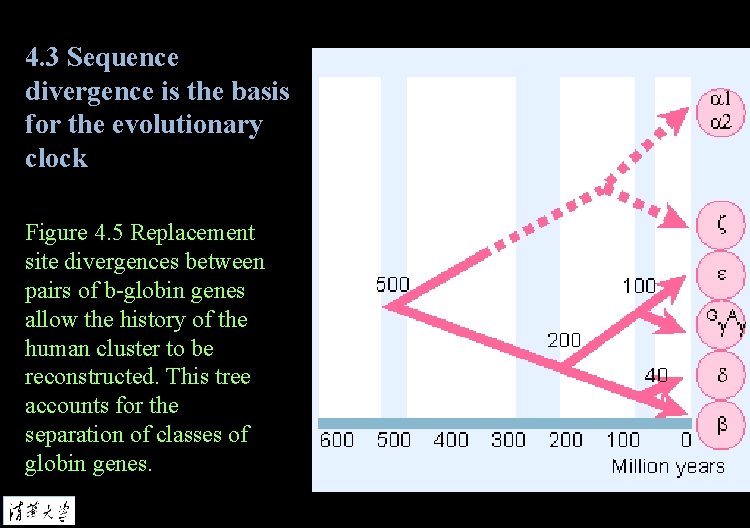
4. 3 Sequence divergence is the basis for the evolutionary clock Figure 4. 5 Replacement site divergences between pairs of b-globin genes allow the history of the human cluster to be reconstructed. This tree accounts for the separation of classes of globin genes.

4. 4 Pseudogenes are dead ends of evolution Processed pseudogene is an inactive gene copy that lacks introns, contrasted with the interrupted structure of the active gene. Such genes presumably originate by reverse transcription of m. RNA and insertion of a duplex copy into the genome.
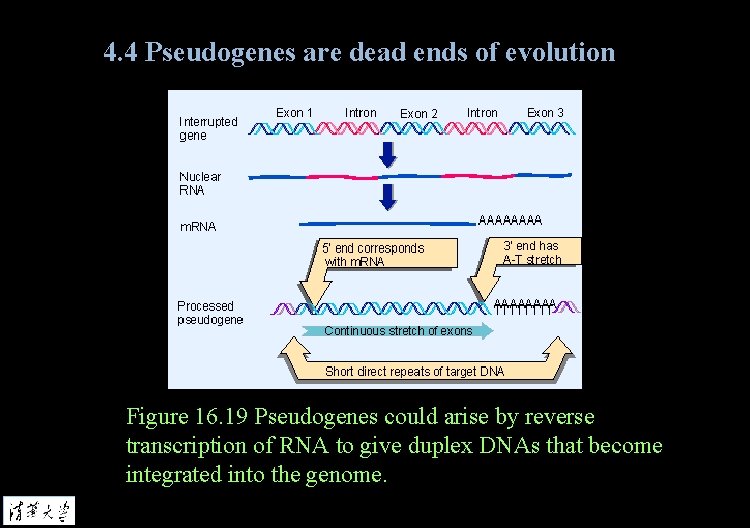
4. 4 Pseudogenes are dead ends of evolution Figure 16. 19 Pseudogenes could arise by reverse transcription of RNA to give duplex DNAs that become integrated into the genome.

4. 5 Unequal crossing-over rearranges gene clusters Thalassemia is disease of red blood cells resulting from lack of either a or b globin. Unequal crossing-over describes a recombination event in which the two recombining sites lie at nonidentical locations in the two parental DNA molecules.

4. 5 Unequal crossing-over rearranges gene clusters Figure 4. 2 Clusters of b-globin genes and pseudogenes are found in vertebrates. Seven mouse genes include 2 early embryonic, 1 late embryonic, 2 adult genes, and 2 pseudogenes. Rabbit and chick each have four genes.
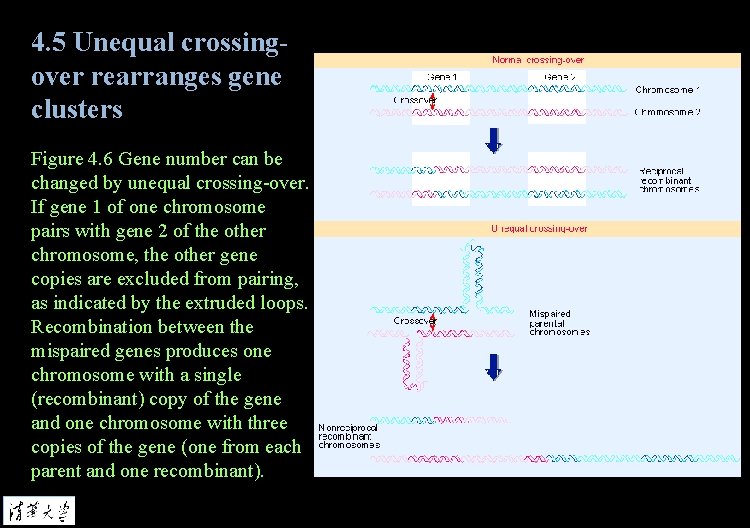
4. 5 Unequal crossingover rearranges gene clusters Figure 4. 6 Gene number can be changed by unequal crossing-over. If gene 1 of one chromosome pairs with gene 2 of the other chromosome, the other gene copies are excluded from pairing, as indicated by the extruded loops. Recombination between the mispaired genes produces one chromosome with a single (recombinant) copy of the gene and one chromosome with three copies of the gene (one from each parent and one recombinant).

4. 5 Unequal crossing-over rearranges gene clusters Figure 4. 7 Thalassemias result from various deletions in the a-globin gene cluster.

4. 5 Unequal crossing-over rearranges gene clusters Figure 4. 8 Deletions in the b -globin gene cluster cause several types of thalassemia.

4. 6 Genes for r. RNA form tandem repeats Nontranscribed spacer is the region between transcription units in a tandem gene cluster. Nucleolar organizer is the region of a chromosome carrying genes coding for r. RNA. Nucleolus is a discrete region of the nucleus created by the transcription of r. RNA genes.
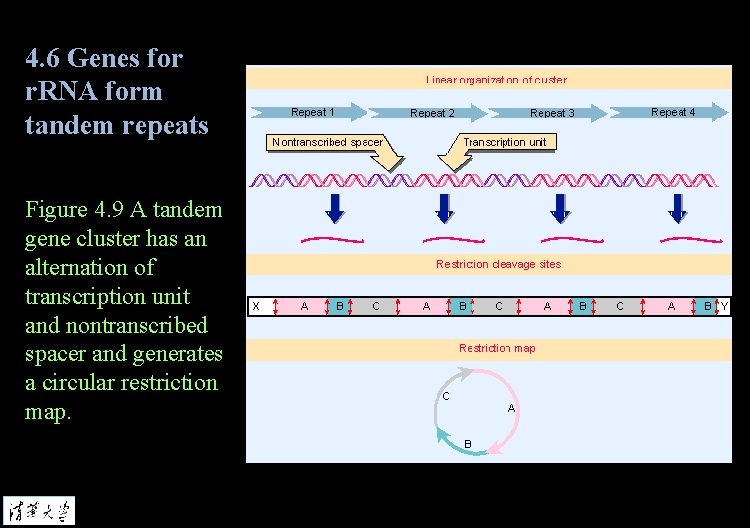
4. 6 Genes for r. RNA form tandem repeats Figure 4. 9 A tandem gene cluster has an alternation of transcription unit and nontranscribed spacer and generates a circular restriction map.

4. 6 Genes for r. RNA form tandem repeats Figure 4. 10 The nucleolar core identifies r. DNA under transcription, and the surrounding granular cortex consists of assembling ribosomal subunits. This thin section shows the nucleolus of the newt Notopthalmus viridescens. Photograph kindly provided by Oscar Miller.
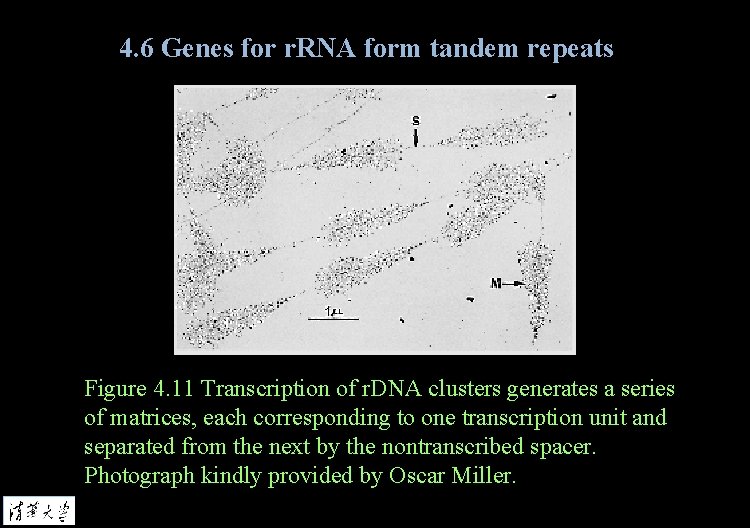
4. 6 Genes for r. RNA form tandem repeats Figure 4. 11 Transcription of r. DNA clusters generates a series of matrices, each corresponding to one transcription unit and separated from the next by the nontranscribed spacer. Photograph kindly provided by Oscar Miller.
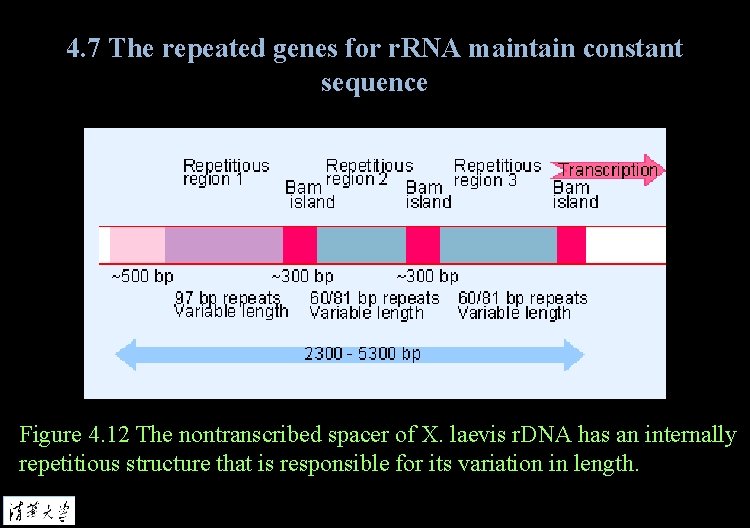
4. 7 The repeated genes for r. RNA maintain constant sequence Figure 4. 12 The nontranscribed spacer of X. laevis r. DNA has an internally repetitious structure that is responsible for its variation in length.

4. 8 Crossover fixation could maintain identical repeats Concerted evolution describes the ability of two related genes to evolve together as though constituting a single locus. Crossover fixation refers to a possible consequence of unequal crossing-over that allows a mutation in one member of a tandem cluster to spread through the whole cluster (or to be eliminated). Gene conversion is the alteration of one strand of a heteroduplex DNA to make it complementary with the other strand at any position(s) where there were mispaired bases.
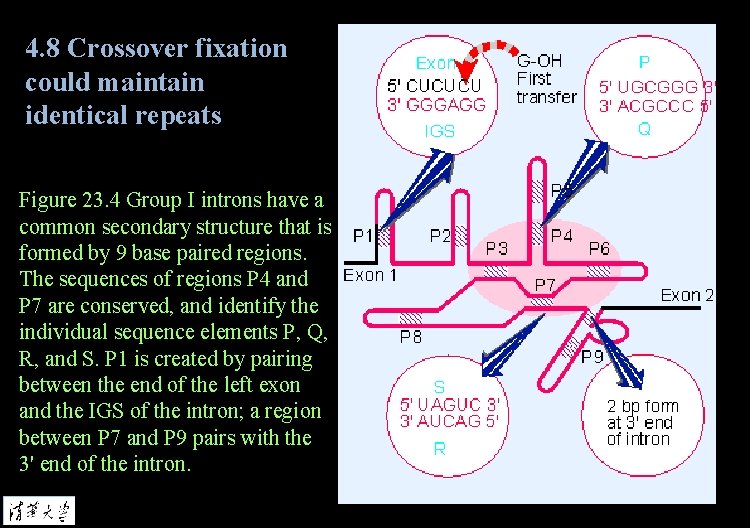
4. 8 Crossover fixation could maintain identical repeats Figure 23. 4 Group I introns have a common secondary structure that is formed by 9 base paired regions. The sequences of regions P 4 and P 7 are conserved, and identify the individual sequence elements P, Q, R, and S. P 1 is created by pairing between the end of the left exon and the IGS of the intron; a region between P 7 and P 9 pairs with the 3' end of the intron.
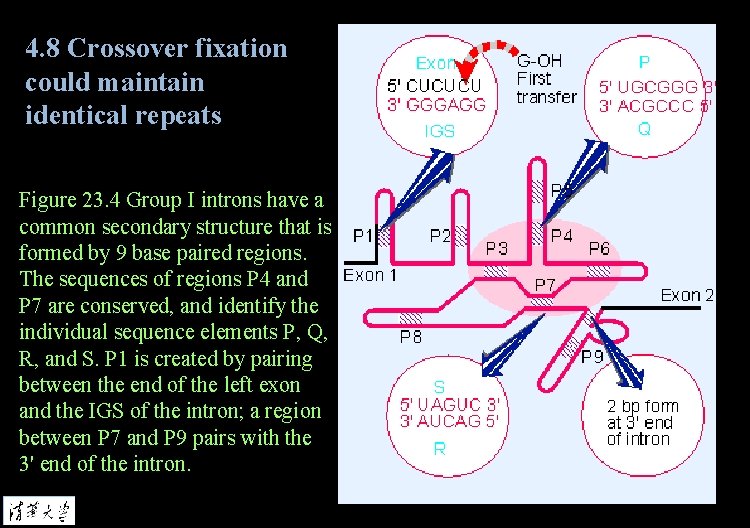
4. 8 Crossover fixation could maintain identical repeats Figure 23. 4 Group I introns have a common secondary structure that is formed by 9 base paired regions. The sequences of regions P 4 and P 7 are conserved, and identify the individual sequence elements P, Q, R, and S. P 1 is created by pairing between the end of the left exon and the IGS of the intron; a region between P 7 and P 9 pairs with the 3' end of the intron.
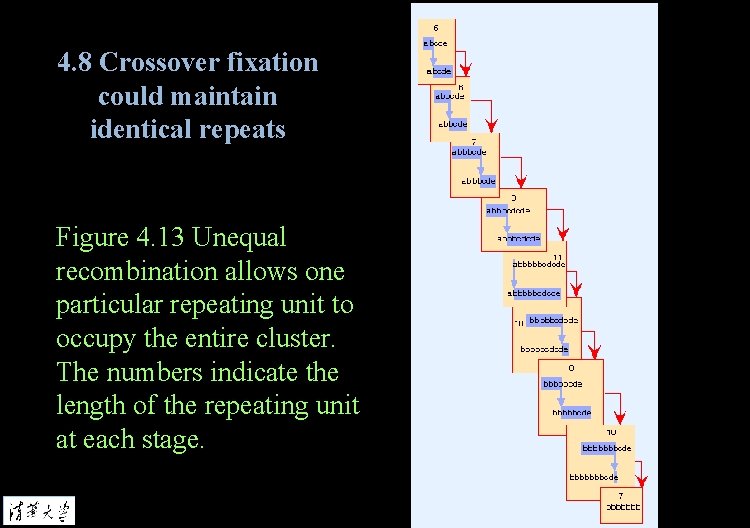
4. 8 Crossover fixation could maintain identical repeats Figure 4. 13 Unequal recombination allows one particular repeating unit to occupy the entire cluster. The numbers indicate the length of the repeating unit at each stage.

4. 9 Satellite DNAs often lie in heterochromatin Cryptic satellite is a satellite DNA sequence not identified as such by a separate peak on a density gradient; that is, it remains present in main-band DNA. Euchromatin comprises all of the genome in the interphase nucleus except for the heterochromatin. Heterochromatin describes regions of the genome that are permanently in a highly condensed condition, are not transcribed, and are late-replicating. May be constitutive or facultative.

4. 9 Satellite DNAs often lie in heterochromatin In situ hybridization is performed by denaturing the DNA of cells squashed on a microscope slide so that reaction is possible with an added single-stranded RNA or DNA; the added preparation is radioactively labeled and its hybridization is followed by autoradiography. Satellite DNA consists of many tandem repeats (identical or related) of a short basic repeating unit.
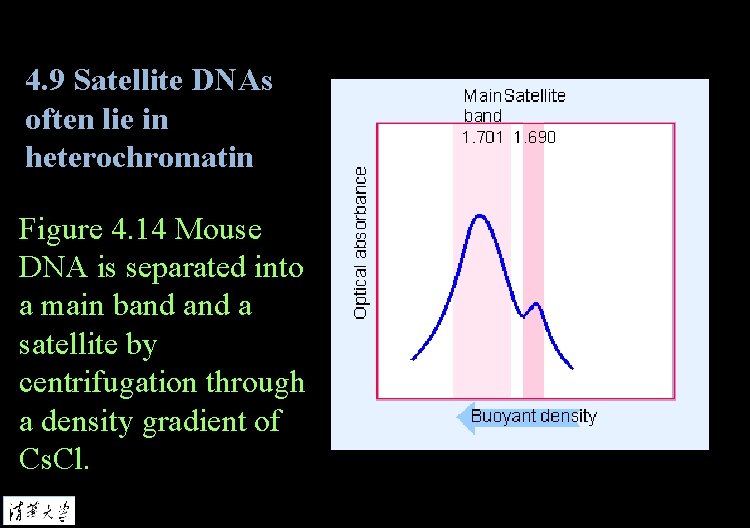
4. 9 Satellite DNAs often lie in heterochromatin Figure 4. 14 Mouse DNA is separated into a main band a satellite by centrifugation through a density gradient of Cs. Cl.

4. 9 Satellite DNAs often lie in heterochromatin Figure 18. 16 Individual bands containing particular genes can be identified by in situ hybridization.
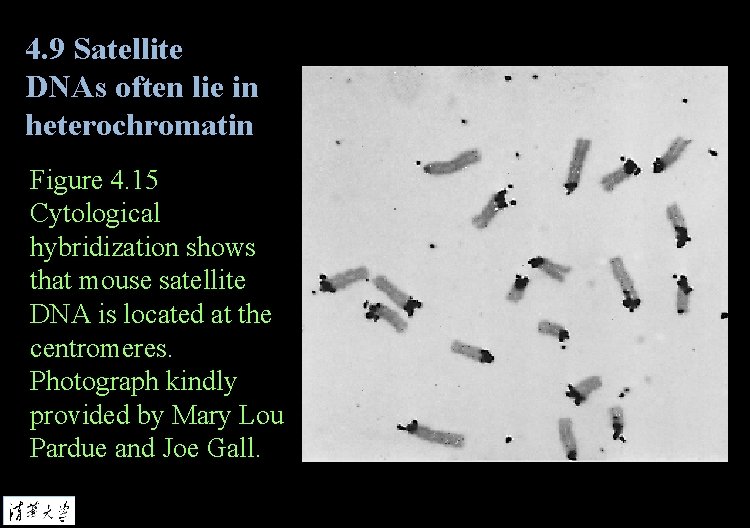
4. 9 Satellite DNAs often lie in heterochromatin Figure 4. 15 Cytological hybridization shows that mouse satellite DNA is located at the centromeres. Photograph kindly provided by Mary Lou Pardue and Joe Gall.

4. 10 Arthropod satellites have very short identical repeats Figure 4. 16 Satellite DNAs of D. virilis are related. More than 95% of each satellite consists of a tandem repetition of the predominant sequence.
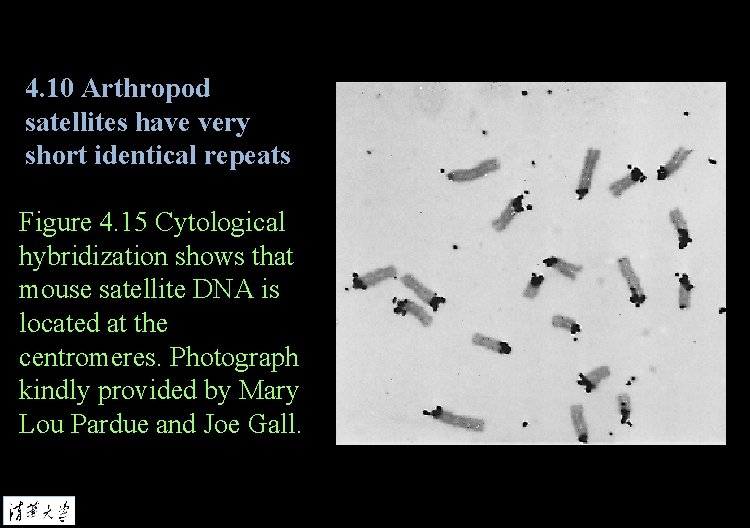
4. 10 Arthropod satellites have very short identical repeats Figure 4. 15 Cytological hybridization shows that mouse satellite DNA is located at the centromeres. Photograph kindly provided by Mary Lou Pardue and Joe Gall.
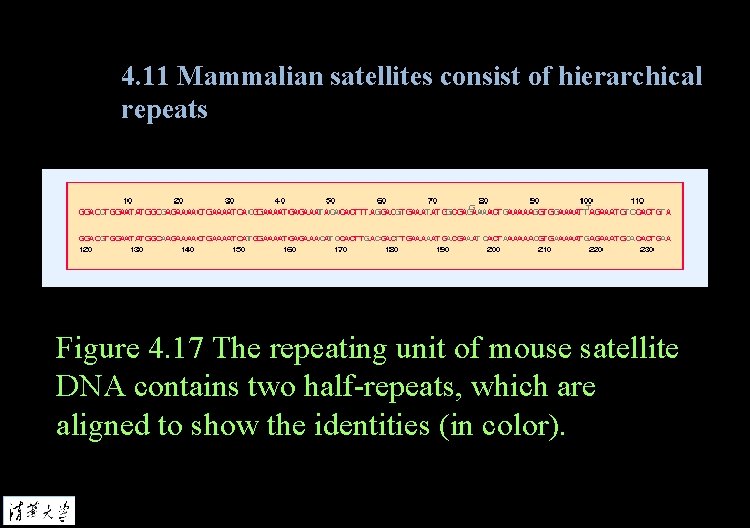
4. 11 Mammalian satellites consist of hierarchical repeats Figure 4. 17 The repeating unit of mouse satellite DNA contains two half-repeats, which are aligned to show the identities (in color).
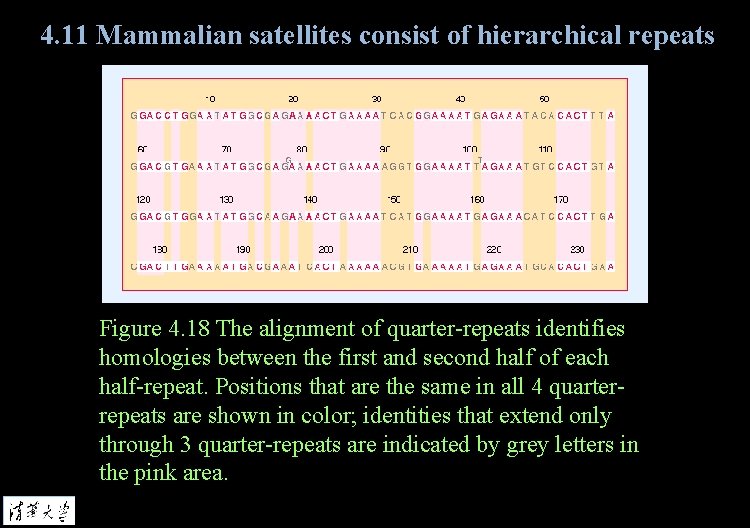
4. 11 Mammalian satellites consist of hierarchical repeats Figure 4. 18 The alignment of quarter-repeats identifies homologies between the first and second half of each half-repeat. Positions that are the same in all 4 quarterrepeats are shown in color; identities that extend only through 3 quarter-repeats are indicated by grey letters in the pink area.

4. 11 Mammalian satellites consist of hierarchical repeats Figure 4. 19 The alignment of eighth-repeats shows that each quarter-repeat consists of an a and a b half. The consensus sequence gives the most common base at each position. The "ancestral" sequence shows a sequence very closely related to the consensus sequence, which could have been the predecessor to the a and b units. (The satellite sequence is continuous, so that for the purposes of deducing the consensus sequence, we can treat it as a circular permutation, as indicated by joining the last GAA triplet to the first 6 bp. )

4. 11 Mammalian satellites consist of hierarchical repeats Figure 4. 20 The existence of an overall consensus sequence is shown by writing the satellite sequence in terms of a 9 bp repeat.
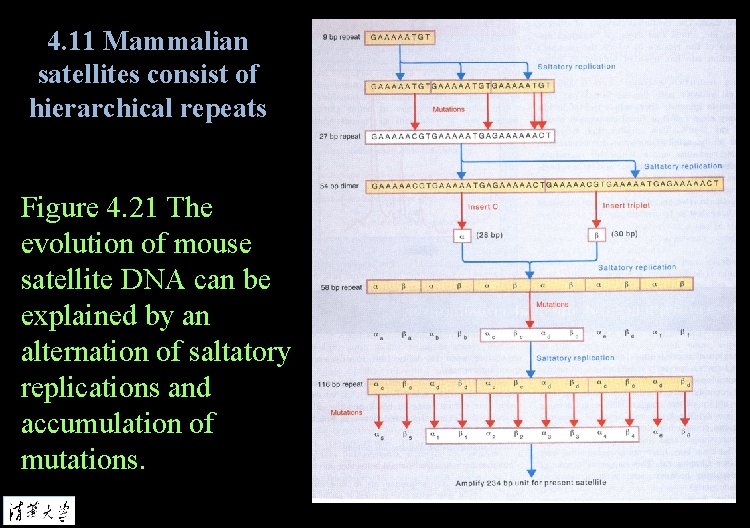
4. 11 Mammalian satellites consist of hierarchical repeats Figure 4. 21 The evolution of mouse satellite DNA can be explained by an alternation of saltatory replications and accumulation of mutations.
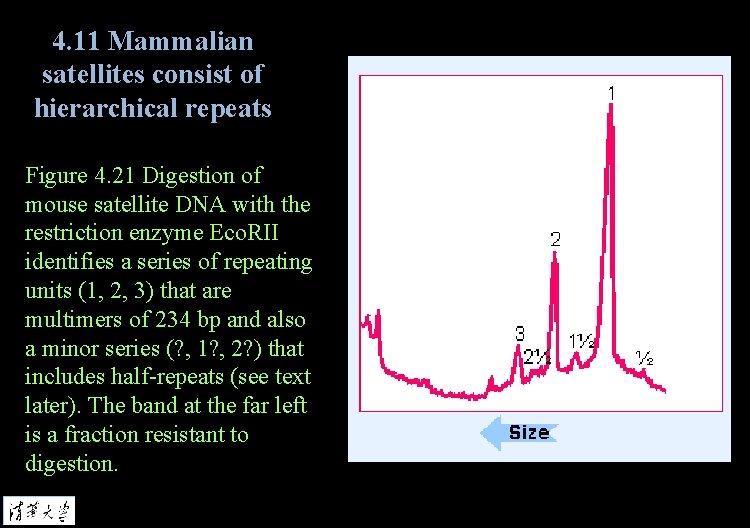
4. 11 Mammalian satellites consist of hierarchical repeats Figure 4. 21 Digestion of mouse satellite DNA with the restriction enzyme Eco. RII identifies a series of repeating units (1, 2, 3) that are multimers of 234 bp and also a minor series (? , 1? , 2? ) that includes half-repeats (see text later). The band at the far left is a fraction resistant to digestion.
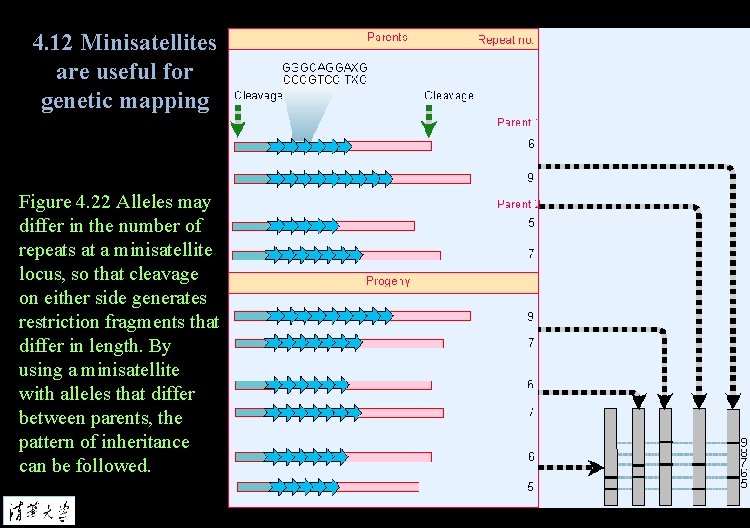
4. 12 Minisatellites are useful for genetic mapping Figure 4. 22 Alleles may differ in the number of repeats at a minisatellite locus, so that cleavage on either side generates restriction fragments that differ in length. By using a minisatellite with alleles that differ between parents, the pattern of inheritance can be followed.
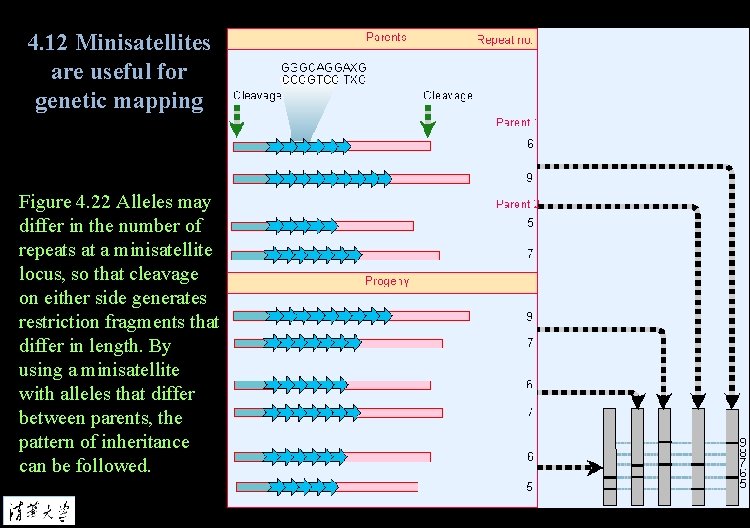
4. 12 Minisatellites are useful for genetic mapping Figure 4. 22 Alleles may differ in the number of repeats at a minisatellite locus, so that cleavage on either side generates restriction fragments that differ in length. By using a minisatellite with alleles that differ between parents, the pattern of inheritance can be followed.

4. 13 Summary 1. Almost all genes belong to families, defined by the possession of related sequences in the exons of individual members. 2. An evolving set of genes may remain together in a cluster or may be dispersed to new locations by chromosomal rearrangement. 3. Mutations accumulate more rapidly in silent sites than in replacement sites (which affect the amino acid sequence). 4. A tandem cluster consists of many copies of a repeating unit that includes the transcribed sequence(s) and a nontranscribed spacer(s). 5. Satellite DNA consists of very short sequences repeated many times in tandem. 6. Unequal crossing-over appears to have been a major determinant of satellite DNA organization.
- Slides: 46