Chapter 3 Historical Methods Fundamentals of Forensic DNA


























- Slides: 54

Chapter 3 Historical Methods Fundamentals of Forensic DNA Typing Slides prepared by John M. Butler June 2009

“If you want to understand today, you have to search yesterday. ” Pearl Buck – Attributed to Pearl Buck (http: //www. quotegarden. com/history. html) The Nobel Prize in Literature 1938 http: //nobelprize. org/nobel_prizes/literature/laureates/1938/buck. jpg Value of a Historical Review

Chapter 3 – Historical Methods Chapter Summary Over the past several decades, methods for DNA testing have improved in terms of sensitivity, speed, and strength of result. The first DNA tests involved Southern blotting of restriction fragment length polymorphisms (RFLPs) separated on agarose gels and detected with radioactive probes. RFLP techniques required substantial amounts of intact DNA and could take weeks to obtain results especially with single-locus probes. Early polymerase chain reaction (PCR) methods, such as HLA-DQα and D 1 S 80, appeared in the early 1990 s and improved sensitivity but were initially less informative than single locus or multi-locus RFLP methodologies. In the mid-1990 s, short tandem repeat markers (STRs) became available in commercial kit formats, first as silver-stained monoplexes or simple multiplexes and eventually in multi-colored fluorescence detection megaplex assays that are widely used today. Simultaneously, mitochondrial DNA (mt. DNA) sequencing was implemented in the forensic DNA community to aid recovery of information from samples containing limited or highly degraded DNA such as hair and skeletal remains.

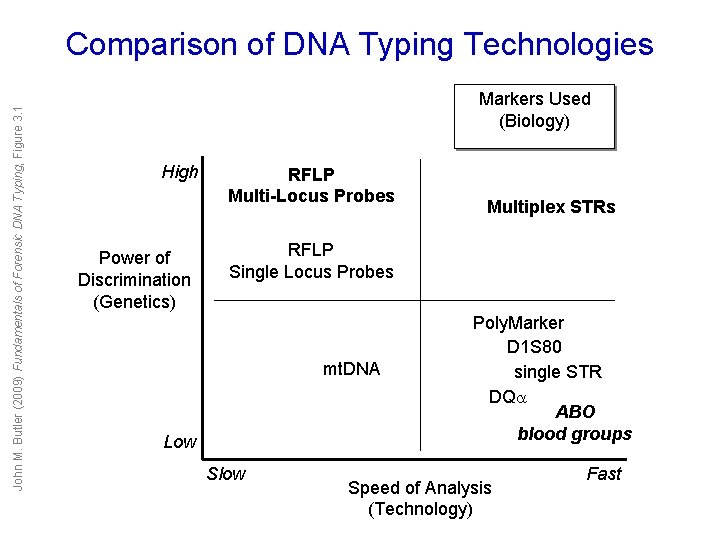
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 1 Comparison of DNA Typing Technologies Markers Used (Biology) High Power of Discrimination (Genetics) RFLP Multi-Locus Probes Multiplex STRs RFLP Single Locus Probes mt. DNA Low Slow Poly. Marker D 1 S 80 single STR DQ ABO blood groups Speed of Analysis (Technology) Fast

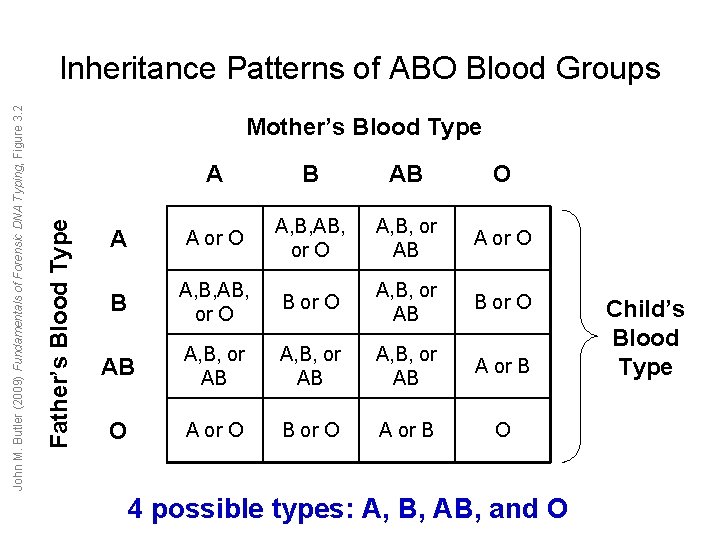
Mother’s Blood Type Father’s Blood Type John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 2 Inheritance Patterns of ABO Blood Groups A B AB O A A or O A, B, AB, or O A, B, or AB A or O B A, B, AB, or O B or O A, B, or AB B or O AB A, B, or AB A or B O A or O B or O A or B O 4 possible types: A, B, AB, and O Child’s Blood Type

Blood Group Typing Advantages 1. Rapid, simple tests. 2. Only test available for many years. Limitation 1. Poor power of discrimination (≈1 in 10) with such few alleles (i. e. , inclusions are not very meaningful). John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

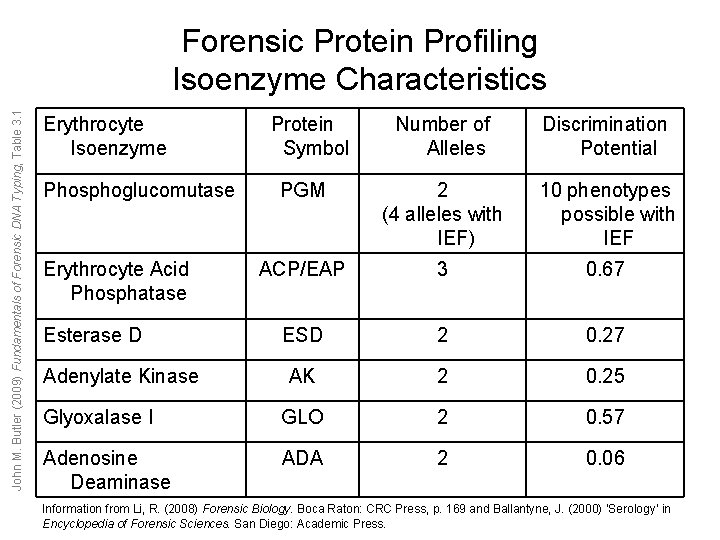
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 1 Forensic Protein Profiling Isoenzyme Characteristics Erythrocyte Isoenzyme Number of Alleles Discrimination Potential PGM 2 (4 alleles with IEF) 10 phenotypes possible with IEF ACP/EAP 3 0. 67 ESD 2 0. 27 AK 2 0. 25 Glyoxalase I GLO 2 0. 57 Adenosine Deaminase ADA 2 0. 06 Phosphoglucomutase Erythrocyte Acid Phosphatase Esterase D Adenylate Kinase Protein Symbol Information from Li, R. (2008) Forensic Biology. Boca Raton: CRC Press, p. 169 and Ballantyne, J. (2000) ‘Serology’ in Encyclopedia of Forensic Sciences. San Diego: Academic Press.

Forensic Protein Profiling Advantage 1. Improved power of discrimination over blood group typing. Limitations 1. Poor power of discrimination (≈1 in 100) even with multiple systems. 2. Poor sensitivity. 3. Proteins are not always stable in forensic stains or found in every sample. John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

Historical Methods for DNA Testing • RFLP – Multi-locus probes – Single-locus probes • PCR based sequence polymorphisms – HLA-DQ alpha – PM + DQA 1 • Amp. FLPs – D 1 S 80 • Silver-stained STRs • Fluorescently detected STRs

The Principles of RFLP Testing • Cut the DNA with “biological scissors” (Restriction enzymes) • Separate Fragments of differing Length by gel electrophoresis • Detect length-based differences (Polymorphisms) in DNA fragments of interest

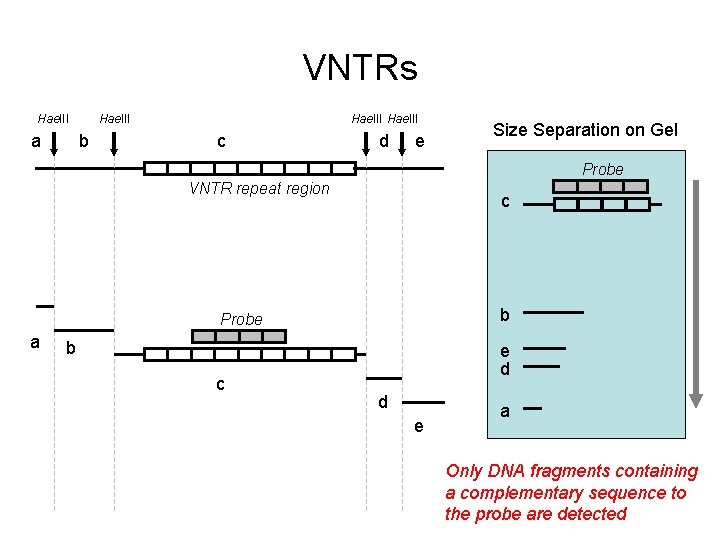
VNTRs Hae. III a Hae. III b Hae. III c d e Size Separation on Gel Probe VNTR repeat region c b Probe a b c e d d e a Only DNA fragments containing a complementary sequence to the probe are detected

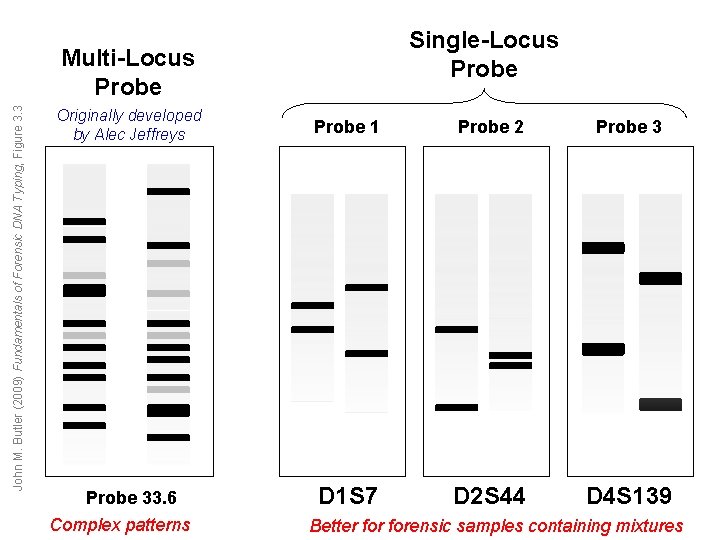
Single-Locus Probe John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 3 Multi-Locus Probe Originally developed by Alec Jeffreys Probe 1 Probe 2 Probe 33. 6 D 1 S 7 D 2 S 44 D 4 S 139 Complex patterns Better forensic samples containing mixtures

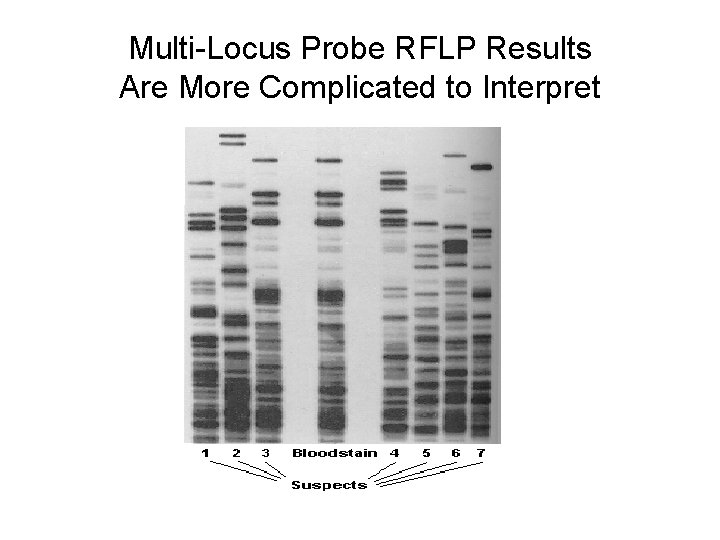
Multi-Locus Probe RFLP Results Are More Complicated to Interpret

Restriction Enzyme Cut Sites

Electrophoresis of DNA Fragments

Transfer of DNA Fragments to a Nylon Membrane - “Southern Blotting”

Probing of Membrane with Radioactive or Labeled DNA Probes

Exposure of Labeled Membrane to Autoradiographic Film

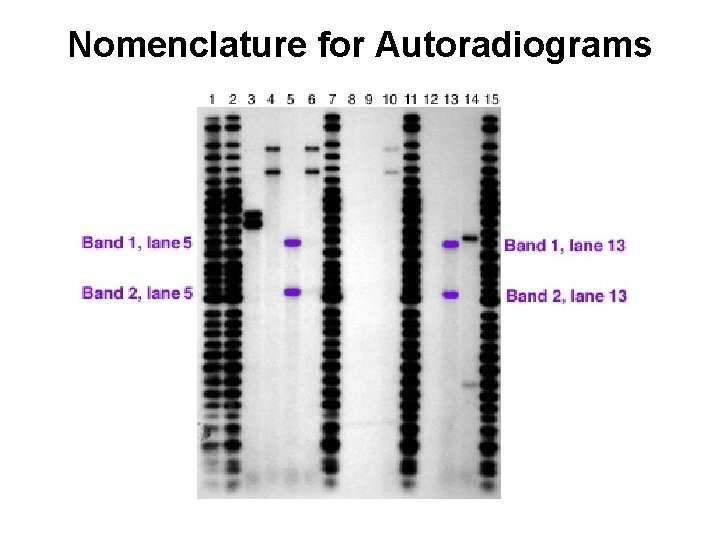
Nomenclature for Autoradiograms

Repeated Probing of Same Membrane to Yield a Series of Autoradiograms Sequential Detection of RFLP Single Locus Probes

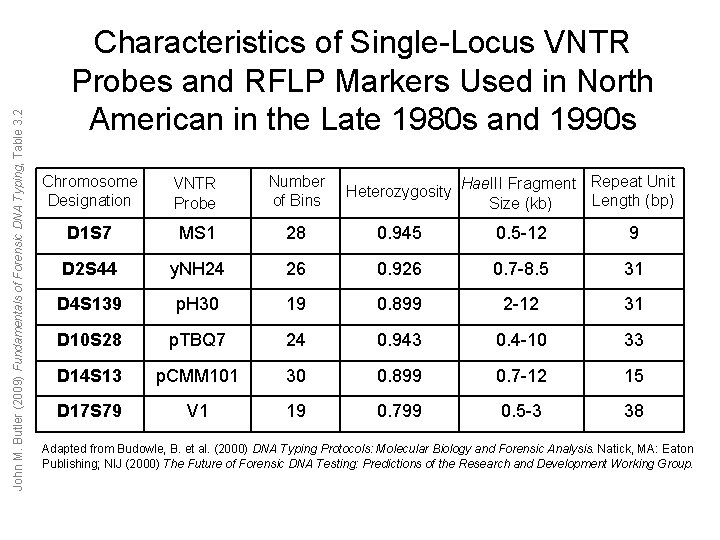
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 2 Characteristics of Single-Locus VNTR Probes and RFLP Markers Used in North American in the Late 1980 s and 1990 s Chromosome Designation VNTR Probe Number of Bins Repeat Unit Heterozygosity Hae. III Fragment Length (bp) Size (kb) D 1 S 7 MS 1 28 0. 945 0. 5 -12 9 D 2 S 44 y. NH 24 26 0. 926 0. 7 -8. 5 31 D 4 S 139 p. H 30 19 0. 899 2 -12 31 D 10 S 28 p. TBQ 7 24 0. 943 0. 4 -10 33 D 14 S 13 p. CMM 101 30 0. 899 0. 7 -12 15 D 17 S 79 V 1 19 0. 799 0. 5 -3 38 Adapted from Budowle, B. et al. (2000) DNA Typing Protocols: Molecular Biology and Forensic Analysis. Natick, MA: Eaton Publishing; NIJ (2000) The Future of Forensic DNA Testing: Predictions of the Research and Development Working Group.

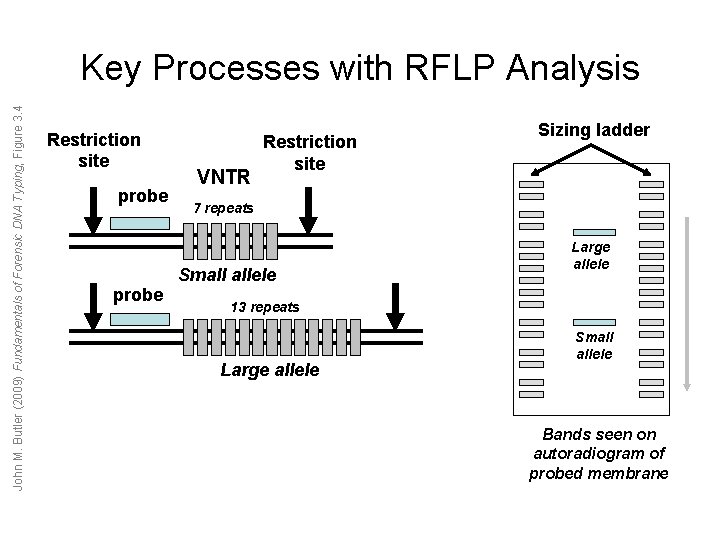
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 4 Key Processes with RFLP Analysis Restriction site probe VNTR Restriction site 7 repeats Small allele probe Sizing ladder Large allele 13 repeats Large allele Small allele Bands seen on autoradiogram of probed membrane

John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 5 Restriction Enzyme Cut Sites Hae. III 4 base cutter Hinf. I 5 base cutter Pst. I 6 base cutter TGCAGG ACGTCC CCTAACG GGATTGC FBI, RCMP, U. S. public labs Cuts on average every ~256 bases TGCAG ANTCTAACG ACGTCTNA GATTGC Cellmark and European labs Cuts on average every ~1024 bases TGCACTGCA GTAACG ACGTCATTGC Lifecodes Cuts on average every ~4096 bases Use of different cut sites by various labs meant that results could not be compared

John M. Butler (2009) Fundamentals of Forensic DNA Typing, D. N. A. Box 3. 1 DNA Evidence and Monica Lewinsky’s Blue Dress August 17, 1998 FBI Report on Analysis of Stain on Monica Lewinsky’s Blue Dress http: //www. law. umkc. edu/faculty/projects/ftrials/clinton/lewinskydress. html

Restriction Fragment Length Polymorphism (RFLP) with single locus VNTR probes Advantages Limitations 1. Limited sensitivity (>50 ng to 500 ng 1. Excellent powers of required). discrimination (≈1 in millions or 2. Time-consuming process (days to greater with four loci). weeks) that cannot be automated. 2. Large number of alleles (20 to 30 3. Not suitable with degraded DNA bins) at each locus which samples due to high molecular facilitates mixed-sample analysis. weight needed. 4. Essentially continuous allele sizes which requires grouping alleles into bins. Binning introduces statistical complications and sometimes difficulties of interpretation. 5. Limited number of validated loci (4 to 6 loci commonly used) which meant that these VNTRs were of limited value in distinguishing between siblings. John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

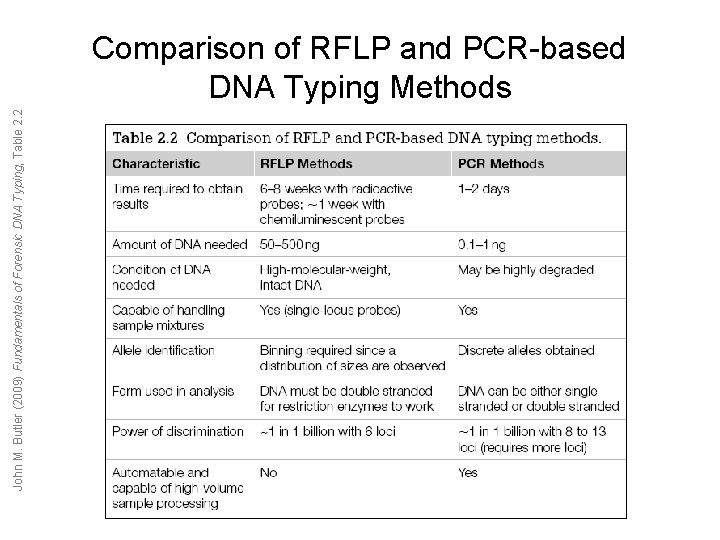
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 2. 2 Comparison of RFLP and PCR-based DNA Typing Methods

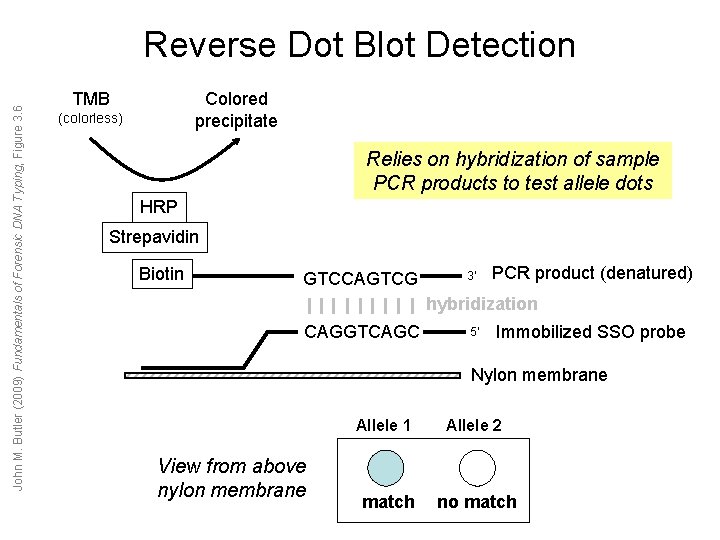
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 6 Reverse Dot Blot Detection TMB Colored precipitate (colorless) Relies on hybridization of sample PCR products to test allele dots HRP Strepavidin Biotin GTCCAGTCG 3’ PCR product (denatured) hybridization CAGGTCAGC 5’ Immobilized SSO probe Nylon membrane View from above nylon membrane Allele 1 Allele 2 match no match

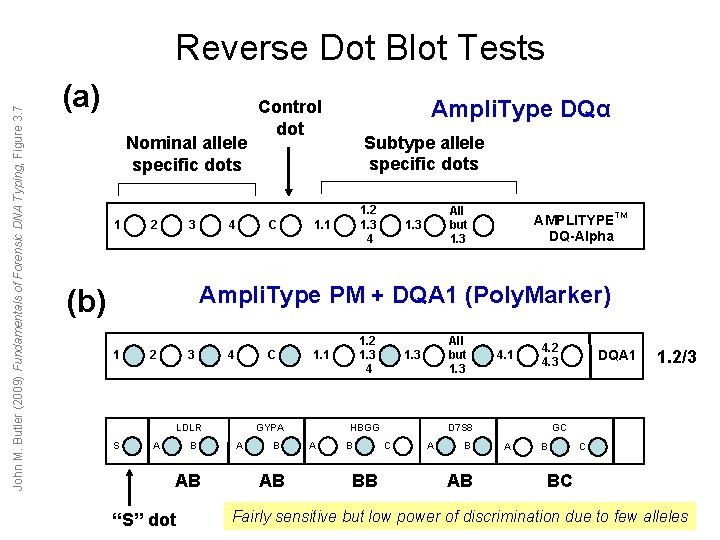
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 7 Reverse Dot Blot Tests (a) Nominal allele specific dots 1 2 3 4 Ampli. Type DQα Control dot C Subtype allele specific dots 1. 2 1. 3 4 1. 1 All but 1. 3 AMPLITYPETM DQ-Alpha Ampli. Type PM + DQA 1 (Poly. Marker) (b) 1 2 3 4 C LDLR S A B AB “S” dot GYPA A B AB 1. 2 1. 3 4 1. 1 All but 1. 3 HBGG A B BB 4. 1 4. 2 4. 3 D 7 S 8 C A B AB DQA 1 1. 2/3 GC A B C BC Fairly sensitive but low power of discrimination due to few alleles

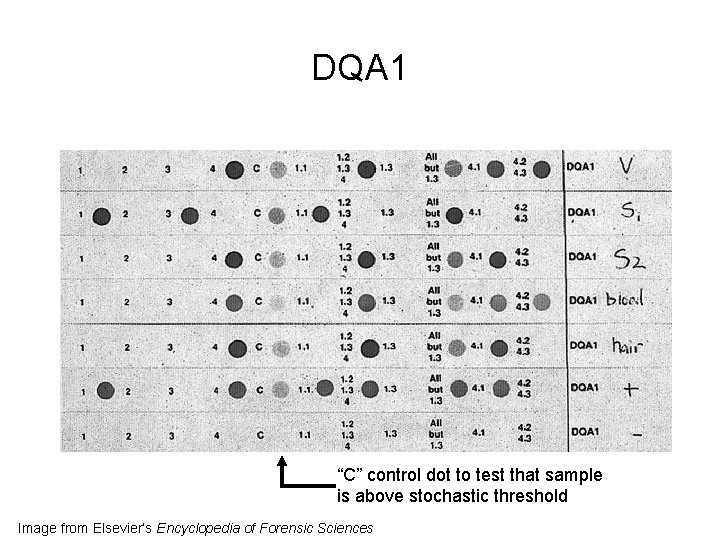
DQA 1 “C” control dot to test that sample is above stochastic threshold Image from Elsevier’s Encyclopedia of Forensic Sciences

Ampli. Type PM (Poly. Marker) “S” dot to test that sample is above stochastic threshold Image from Elsevier’s Encyclopedia of Forensic Sciences

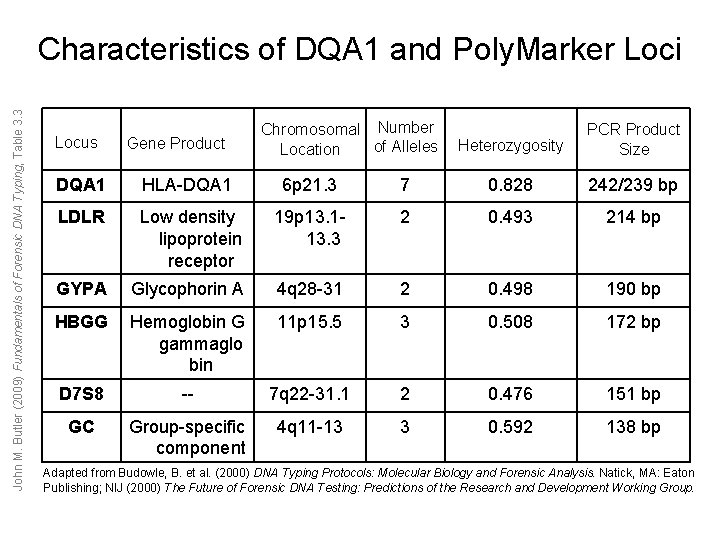
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 3 Characteristics of DQA 1 and Poly. Marker Loci Locus Gene Product Chromosomal Number of Alleles Location Heterozygosity PCR Product Size DQA 1 HLA-DQA 1 6 p 21. 3 7 0. 828 242/239 bp LDLR Low density lipoprotein receptor 19 p 13. 113. 3 2 0. 493 214 bp GYPA Glycophorin A 4 q 28 -31 2 0. 498 190 bp HBGG Hemoglobin G gammaglo bin 11 p 15. 5 3 0. 508 172 bp D 7 S 8 -- 7 q 22 -31. 1 2 0. 476 151 bp GC Group-specific component 4 q 11 -13 3 0. 592 138 bp Adapted from Budowle, B. et al. (2000) DNA Typing Protocols: Molecular Biology and Forensic Analysis. Natick, MA: Eaton Publishing; NIJ (2000) The Future of Forensic DNA Testing: Predictions of the Research and Development Working Group.

DQA 1 + PM Advantages Limitations 1. Fast, simple method (compared 1. Poor power of discrimination (≈1 to RFLP). in 1000) with only six loci 2. Capable of analyzing small or developed each containing only degraded samples because it a few alleles. uses PCR. 2. Mixture interpretation difficult 3. No instrumentation needed after with limited number of alleles PCR. per locus. John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

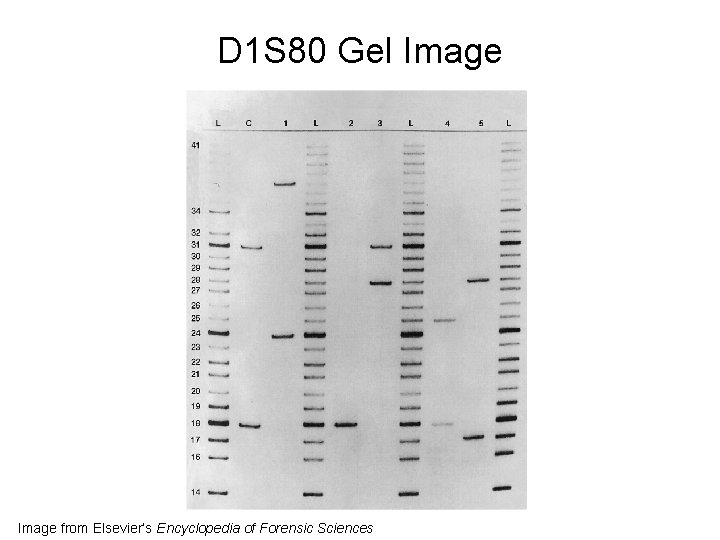
D 1 S 80 • Amp. FLP = amplified fragment length polymorphism • 16 bp repeat sequence • PCR products in 400 -800 bp range • Separated on a polyacrylamide gel • Detected with silver staining • Sometimes combined with amelogenin to facilitate sex-typing

D 1 S 80 Gel Image from Elsevier’s Encyclopedia of Forensic Sciences

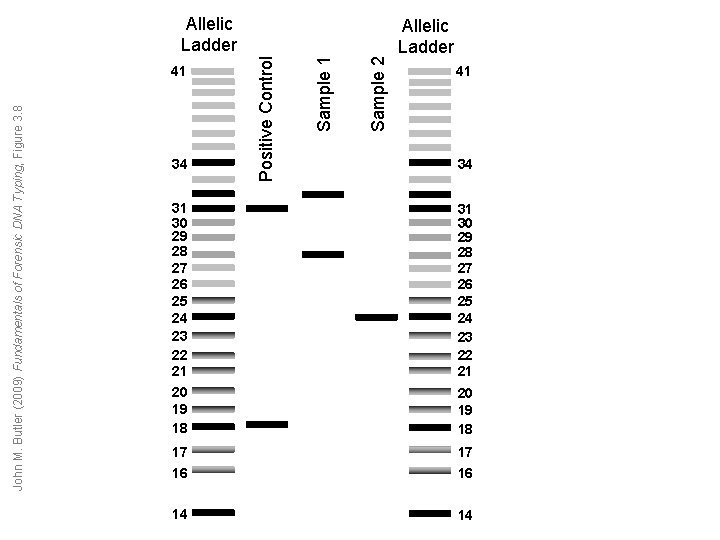
34 Sample 2 Sample 1 John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 8 41 Positive Control Allelic Ladder 41 34 31 30 29 28 27 26 25 24 23 22 21 20 19 18 17 17 16 16 14 14

D 1 S 80 Advantages Limitations 1. Improved sensitivity compared to RFLP because it uses PCR. 2. Many alleles which facilitates mixed-sample analysis. 3. Discrete allele calling possible using allelic ladder, which also simplifies statistical interpretation. 1. Large allele range making it difficult to multiplex with other loci and giving rise to preferential amplification of smaller alleles. 2. Poor power of discrimination as a single locus (≈1 in 50). 3. Allele dropout seen with highly degraded DNA. 4. Gel separation and silver-stain detection not amenable to automation or high-throughput sample processing. John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

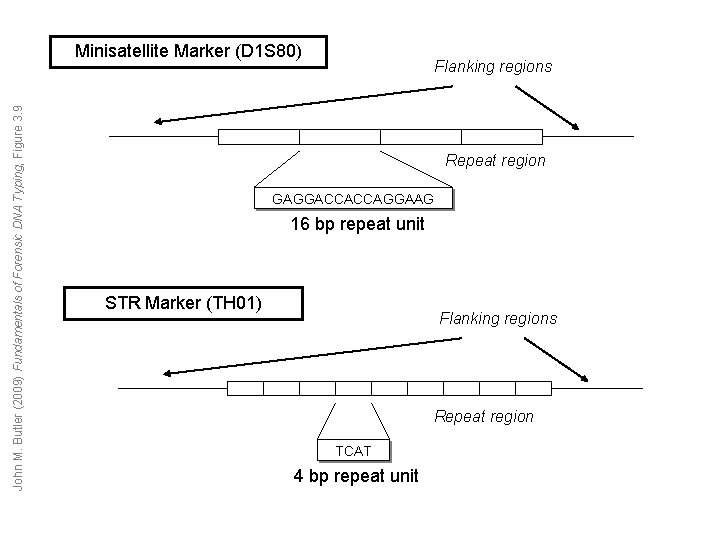
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 9 Minisatellite Marker (D 1 S 80) Flanking regions Repeat region GAGGACCACCAGGAAG 16 bp repeat unit STR Marker (TH 01) Flanking regions Repeat region TCAT 4 bp repeat unit

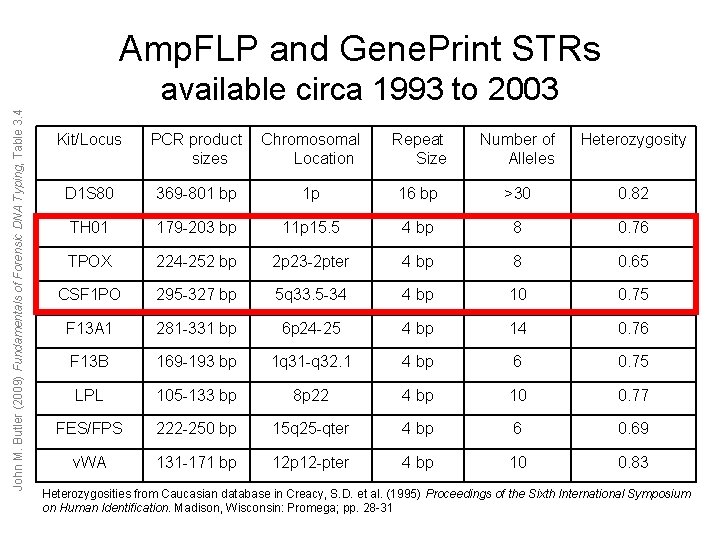
Amp. FLP and Gene. Print STRs John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 4 available circa 1993 to 2003 Kit/Locus PCR product sizes Chromosomal Location Repeat Size Number of Alleles Heterozygosity D 1 S 80 369 -801 bp 1 p 16 bp >30 0. 82 TH 01 179 -203 bp 11 p 15. 5 4 bp 8 0. 76 TPOX 224 -252 bp 2 p 23 -2 pter 4 bp 8 0. 65 CSF 1 PO 295 -327 bp 5 q 33. 5 -34 4 bp 10 0. 75 F 13 A 1 281 -331 bp 6 p 24 -25 4 bp 14 0. 76 F 13 B 169 -193 bp 1 q 31 -q 32. 1 4 bp 6 0. 75 LPL 105 -133 bp 8 p 22 4 bp 10 0. 77 FES/FPS 222 -250 bp 15 q 25 -qter 4 bp 6 0. 69 v. WA 131 -171 bp 12 -pter 4 bp 10 0. 83 Heterozygosities from Caucasian database in Creacy, S. D. et al. (1995) Proceedings of the Sixth International Symposium on Human Identification. Madison, Wisconsin: Promega; pp. 28 -31

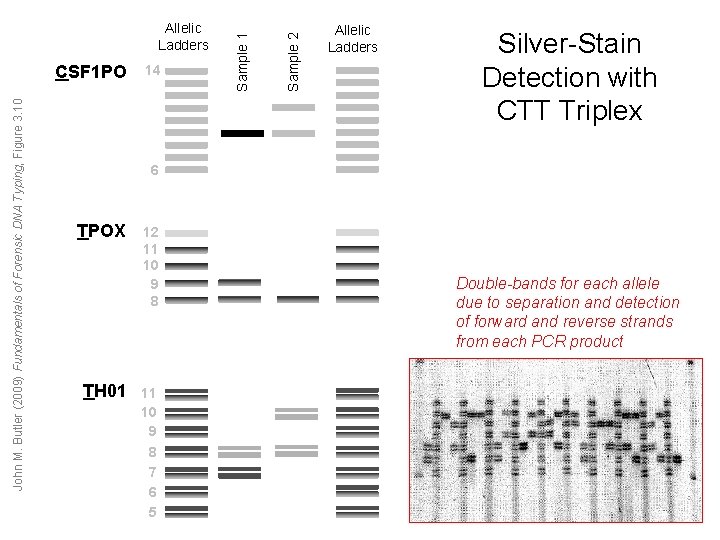
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 10 14 Sample 2 CSF 1 PO Sample 1 Allelic Ladders Silver-Stain Detection with CTT Triplex 6 TPOX TH 01 12 11 10 9 8 7 6 5 Double-bands for each allele due to separation and detection of forward and reverse strands from each PCR product

Silver-Stained STRs Advantages Limitations 1. Sensitive due to PCR. 1. Because only a single ‘color’ 2. Relatively rapid process (a day channel is available, multiplex or two). amplification and detection is 3. Works well with degraded DNA limited to 3 to 4 loci samples since shorter 2. Both strands of DNA are fragments of DNA can be detected leading to double analyzed (compared to D 1 S 80). bands with some loci that can 4. A lower start-up cost compared complicate interpretation. to fluorescent STRs John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 5 Early fluorescent STR multiplex systems available during the 1990 s Assay/Kit Lab or Supplier STR Locus British Home Office Quadruplex Forensic Science Service TH 01, VWA, F 13 A 1, FES/FPS Second Generation Multiplex (SGM) Forensic Science Service TH 01, VWA, FGA, D 8 S 1179, D 18 S 51, D 21 S 11, amelogenin CTTv Promega Corporation CSF 1 PO, TPOX, TH 01, VWA FFFL Promega Corporation F 13 A 1, F 13 B, FES/FPS, LPL Gamma. STR Promega Corporation D 16 S 539, D 13 S 317, D 7 S 820, D 5 S 818 Ampl. STR Blue Applied Biosystems D 3 S 1358 VWA, FGA Ampl. STR Green Applied Biosystems CSF 1 PO, TPOX, TH 01, amelogenin

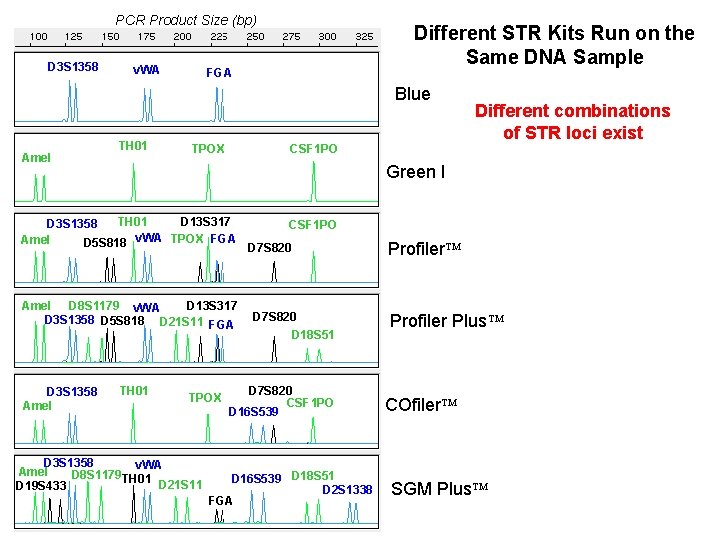
PCR Product Size (bp) D 3 S 1358 v. WA Different STR Kits Run on the Same DNA Sample FGA Blue Amel TH 01 CSF 1 PO TPOX Green I TH 01 D 13 S 317 D 3 S 1358 Amel D 5 S 818 v. WA TPOX FGA D 13 S 317 Amel D 8 S 1179 v. WA D 3 S 1358 D 5 S 818 D 21 S 11 FGA D 3 S 1358 Amel Different combinations of STR loci exist TH 01 TPOX D 3 S 1358 v. WA Amel D 8 S 1179 TH 01 D 21 S 11 D 19 S 433 CSF 1 PO D 7 S 820 D 18 S 51 D 7 S 820 CSF 1 PO D 16 S 539 D 18 S 51 D 2 S 1338 FGA Profiler Plus COfiler SGM Plus

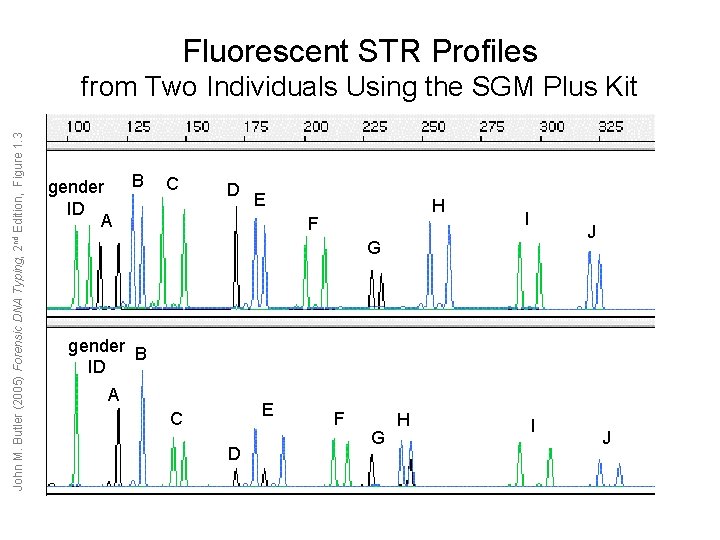
Fluorescent STR Profiles John M. Butler (2005) Forensic DNA Typing, 2 nd Edition, Figure 1. 3 from Two Individuals Using the SGM Plus Kit gender ID A B C D E H F I J G gender B ID A E C D F G H I J

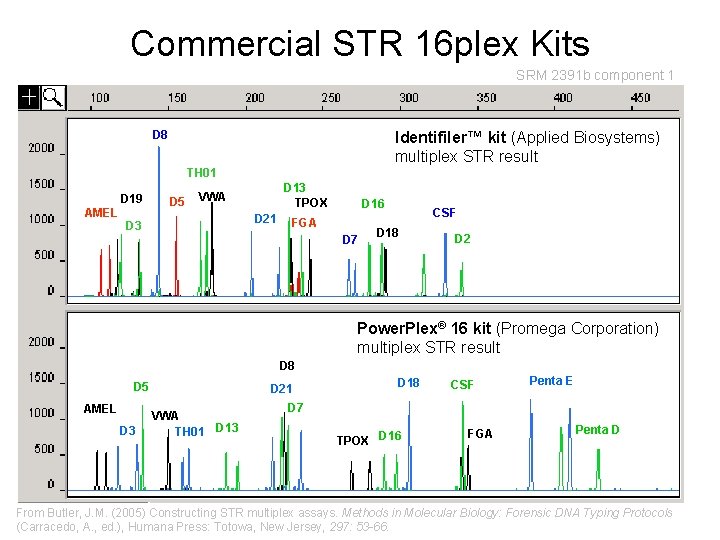
Commercial STR 16 plex Kits SRM 2391 b component 1 D 8 Identifiler™ kit (Applied Biosystems) multiplex STR result TH 01 D 19 AMEL D 5 D 13 TPOX VWA D 21 D 3 D 16 FGA D 7 CSF D 18 D 2 Power. Plex® 16 kit (Promega Corporation) multiplex STR result D 8 D 5 AMEL D 3 D 21 VWA TH 01 D 13 D 18 CSF Penta E D 7 TPOX D 16 FGA Penta D From Butler, J. M. (2005) Constructing STR multiplex assays. Methods in Molecular Biology: Forensic DNA Typing Protocols (Carracedo, A. , ed. ), Humana Press: Totowa, New Jersey, 297: 53 -66.

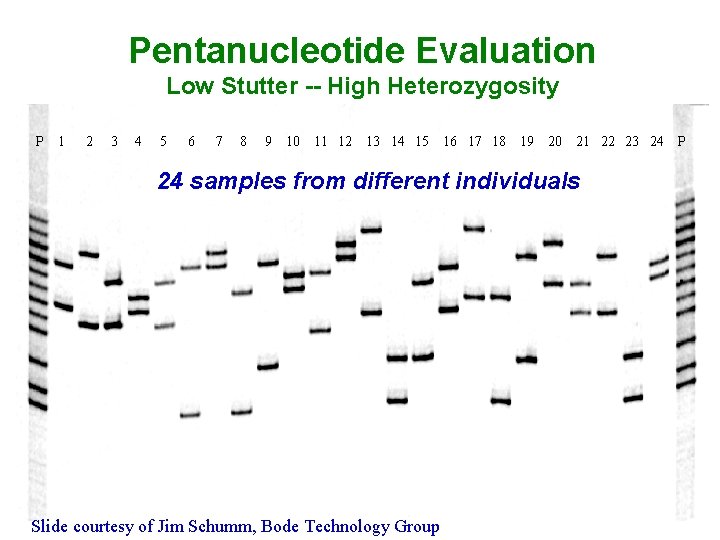
Pentanucleotide Evaluation Low Stutter -- High Heterozygosity P 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 24 samples from different individuals Slide courtesy of Jim Schumm, Bode Technology Group P

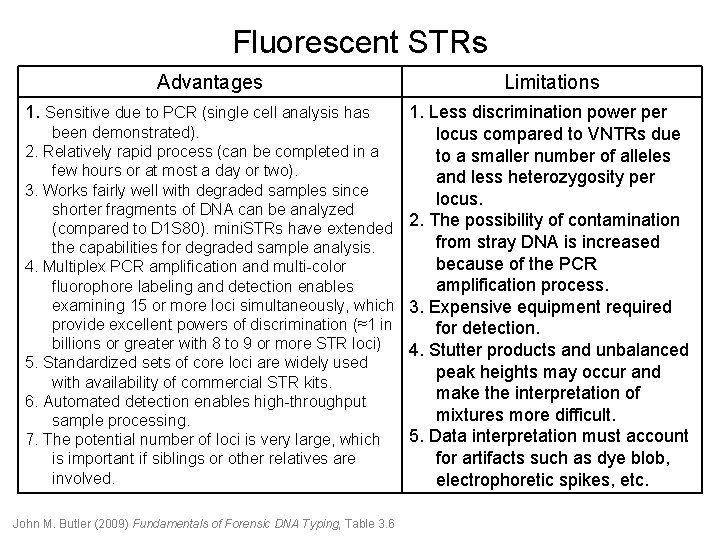
Fluorescent STRs Advantages 1. Sensitive due to PCR (single cell analysis has Limitations 1. Less discrimination power per been demonstrated). locus compared to VNTRs due 2. Relatively rapid process (can be completed in a to a smaller number of alleles few hours or at most a day or two). and less heterozygosity per 3. Works fairly well with degraded samples since locus. shorter fragments of DNA can be analyzed (compared to D 1 S 80). mini. STRs have extended 2. The possibility of contamination from stray DNA is increased the capabilities for degraded sample analysis. because of the PCR 4. Multiplex PCR amplification and multi-color amplification process. fluorophore labeling and detection enables examining 15 or more loci simultaneously, which 3. Expensive equipment required provide excellent powers of discrimination (≈1 in for detection. billions or greater with 8 to 9 or more STR loci) 4. Stutter products and unbalanced 5. Standardized sets of core loci are widely used peak heights may occur and with availability of commercial STR kits. make the interpretation of 6. Automated detection enables high-throughput mixtures more difficult. sample processing. 5. Data interpretation must account 7. The potential number of loci is very large, which for artifacts such as dye blob, is important if siblings or other relatives are involved. electrophoretic spikes, etc. John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

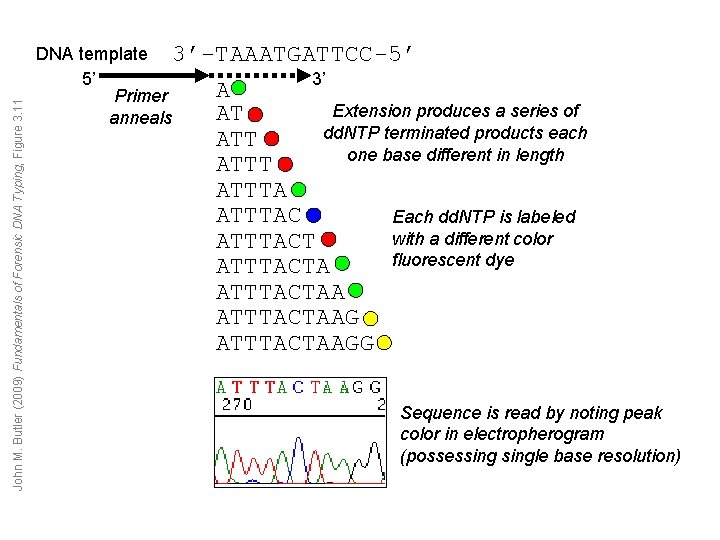
John M. Butler (2009) Fundamentals of Forensic DNA Typing, Figure 3. 11 DNA template 3’-TAAATGATTCC-5’ 5’ 3’ A Primer Extension produces a series of AT anneals dd. NTP terminated products each ATT one base different in length ATTTACT ATTTACTAAGG Each dd. NTP is labeled with a different color fluorescent dye Sequence is read by noting peak color in electropherogram (possessingle base resolution)

mt. DNA Sequencing • May be used when limited or highly degraded DNA is available such as with a hair shaft or bone specimen • Requires a great deal of labor and expertise and thus only a few forensic laboratories conduct mt. DNA analysis • Due to maternal inheritance without recombination, siblings and maternal relatives will have identical mt. DNA making it less useful in many human identity applications

Mitochondrial DNA Sequencing Advantages Limitations 1. Excellent sensitivity and success with limited or badly damaged samples due to large number of mt. DNA molecules per cell and the fact that the small mt. DNA molecule is more robust than nuclear DNA. 2. Maternal transmission from a mother to all of her children extends possible reference sample providers and enables tracing family lineages. 3. A majority of mt. DNA haplotypes occur only once in a population database making its power of discrimination better than a single nuclear locus. 1. Since all descendants through the female line are identical, mt. DNA analysis cannot be used to distinguish among members of a sibship or maternal relatives. 2. Without recombination, the discrimination power of mt. DNA is limited by the size of the population database used. 3. Heteroplasmy (the occurrence of more than one mt. DNA type in a single cell or person) can complicate the analysis. 4. Because the entire mt. Genome is inherited as a unit, it is equivalent to a single nuclear locus. Hence, the discrimination power is limited by the size of the database. John M. Butler (2009) Fundamentals of Forensic DNA Typing, Table 3. 6

Advantages of PCR Methods Overall, PCR-based methods have greatly enabled forensic DNA analysis for the following reasons: • Very small amounts of DNA template may be used – even as little as from a single cell. • DNA degraded to fragments only a few hundred base pairs in length can serve as effective templates for amplification. • Large numbers of copies of specific DNA sequences can be amplified simultaneously with multiplex PCR reactions. • Contaminant DNA, such as from fungal and bacterial sources, will not amplify because human-specific primers are used. • Commercial kits are now available for easy PCR reaction setup and amplification

Potential Pitfalls with PCR Methods 1. The target DNA template may not amplify due to the presence of PCR inhibitors in the extracted DNA. 2. Amplification may fail due to sequence mutations in the primer-binding region of the genomic DNA template – something often referred to as a ‘null allele. ’ 3. Contamination from other human DNA sources besides the forensic evidence at hand or previously amplified DNA samples is possible without the use of careful laboratory technique and validated protocols.

• Report published in Nov 2000 • Asked to estimate where DNA testing would be 2, 5, and 10 years into the future Conclusions STR typing is here to stay for a few years because of DNA databases that have grown to contain millions of profiles http: //www. ojp. usdoj. gov/nij/pubs-sum/183697. htm

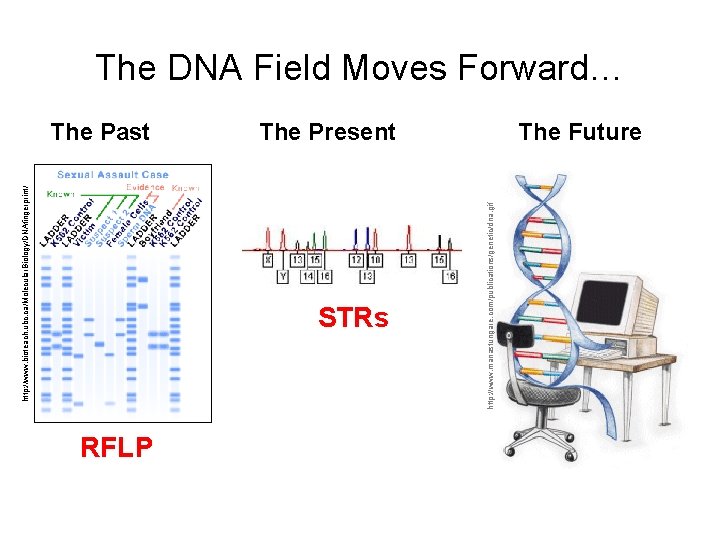
http: //www. bioteach. ubc. ca/Molecular. Biology/DNAfingerprint / The Past STRs RFLP http: //www. manastungare. com/publications/genetic/dna. gif The DNA Field Moves Forward… The Present The Future

Chapter 3 – Points for Discussion • Why were single locus probes preferred over multi-locus probes for RFLP testing? • Why did STRs replace RFLP as the method of choice in modern forensic DNA analysis? • Discuss the advantages and disadvantages of short tandem repeat (STR) markers relative to other potential genetic loci that could be used for DNA testing. • What is the difference between a mini-satellite and a micro-satellite?