Chapter 25 Phylogeny Systematics AP Biology Investigating the

Chapter 25: Phylogeny & Systematics AP Biology

Investigating the Tree of Life Phylogeny w The evolutionary history of a species or group of related species Systematics u Analytical approach to understanding diversity and relationships of organisms, extinct present The fossil record helps classify organisms u u Provides information about ancient organisms Sedimentary fossils are the most common 1 Rivers carry sediment to the ocean. Sedimentary rock layers containing fossils form on the ocean floor. 2 Over time, new strata are deposited, containing fossils from each time period. 3 As sea levels change and the seafloor is pushed upward, sedimentary rocks are exposed. Erosion reveals strata and fossils. AP Biology Younger stratum ���with more recent fossils Older stratum with older fossils

Though sedimentary fossils are the most common u Paleontologists study a wide variety of fossils (c) Leaf fossil, about 40 million years old (b) Petrified tree in Arizona, about 190 million years old (a) Dinosaur bones being excavated from sandstone (d) Casts of ammonites, about 375 million years old (f) Insects preserved whole in amber Figure 25. 4 a–g AP Biology (g) Tusks of a 23, 000 -year-old mammoth, frozen whole in Siberian ice (e) Boy standing in a 150 -million-year-old dinosaur track in Colorado

Phylogenic Tree Shows the evolutionary relationship between different species based on common ancestry Scientists classify organisms based on: Ø Structure (Morphology) Ø Biochemistry Ø Cell Type Ø Embryological data Ø Behavior Ø Fossils AP Biology

Morphological and Molecular Homologies In general, organisms that share very similar morphologies or similar DNA sequences u Are likely to be more closely related than organisms with vastly different structures or sequences AP Biology
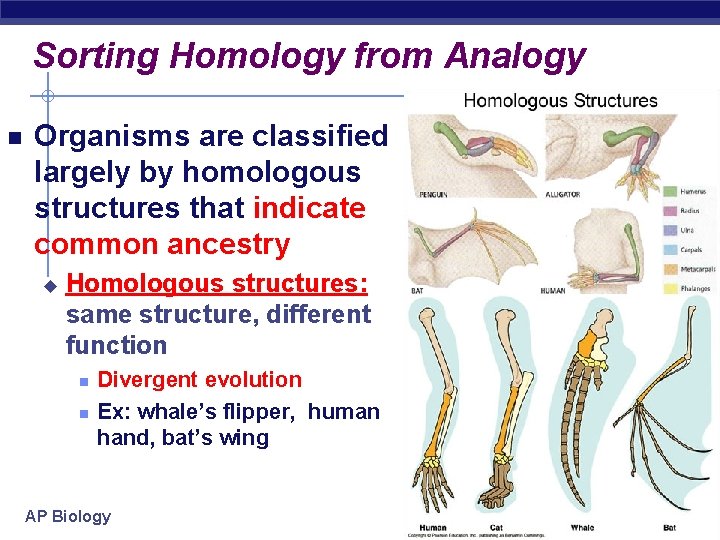
Sorting Homology from Analogy Organisms are classified largely by homologous structures that indicate common ancestry u Homologous structures: same structure, different function Divergent evolution Ex: whale’s flipper, human hand, bat’s wing AP Biology
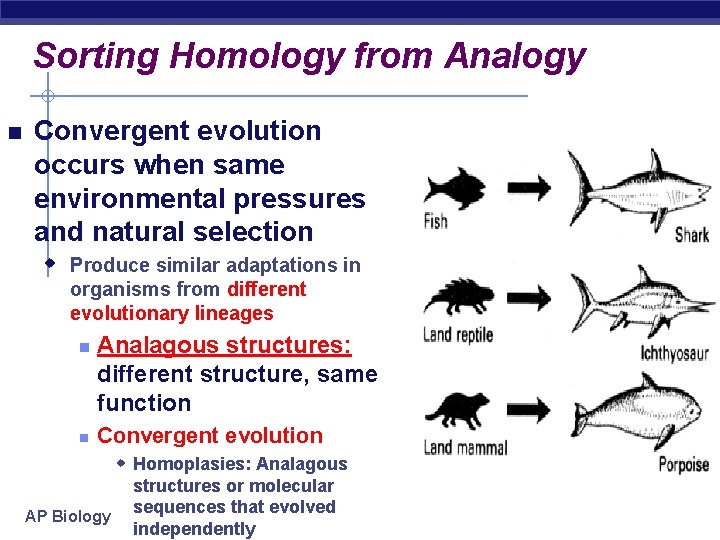
Sorting Homology from Analogy Convergent evolution occurs when same environmental pressures and natural selection w Produce similar adaptations in organisms from different evolutionary lineages Analagous structures: different structure, same function Convergent evolution w Homoplasies: Analagous AP Biology structures or molecular sequences that evolved independently

Convergent evolution occurs when similar environmental pressures and natural selection u Produce similar (analogous) adaptations in organisms from different evolutionary lineages Figure 25. 5 AP Biology
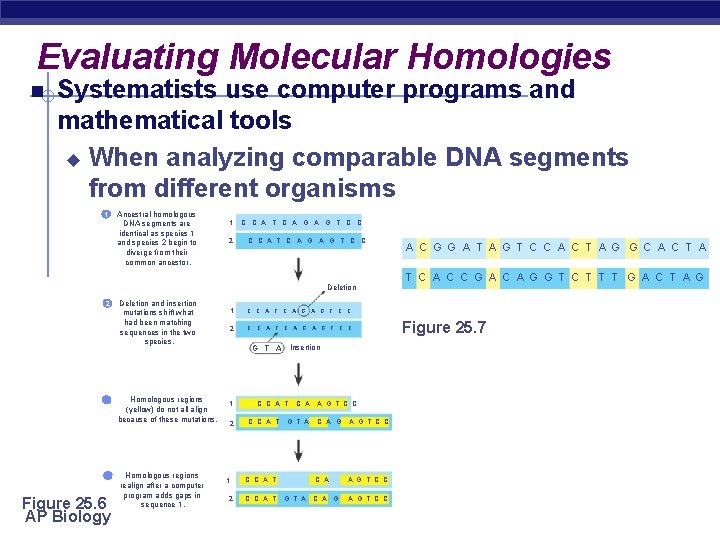
Evaluating Molecular Homologies Systematists use computer programs and mathematical tools u When analyzing comparable DNA segments from different organisms Ancestral homologous DNA segments are identical as species 1 and species 2 begin to diverge from their common ancestor. 1 1 2 C C A T C A G A G T C C Deletion and insertion mutations shift what had been matching sequences in the two species. 2 3 Homologous regions (yellow) do not all align because of these mutations. 4 Figure 25. 6 AP Biology Homologous regions realign after a computer program adds gaps in sequence 1. 1 C C A T C A G T C C 2 C C A T C A G T C C Insertion G T A 1 2 C C A T C A G T A C C A T G T A A G T C C C A A G T C C C A G T C C A C G G A T A G T C C A C T A G G C A C T A T C A C C G A C A G G T C T T T G A C T A G Figure 25. 7

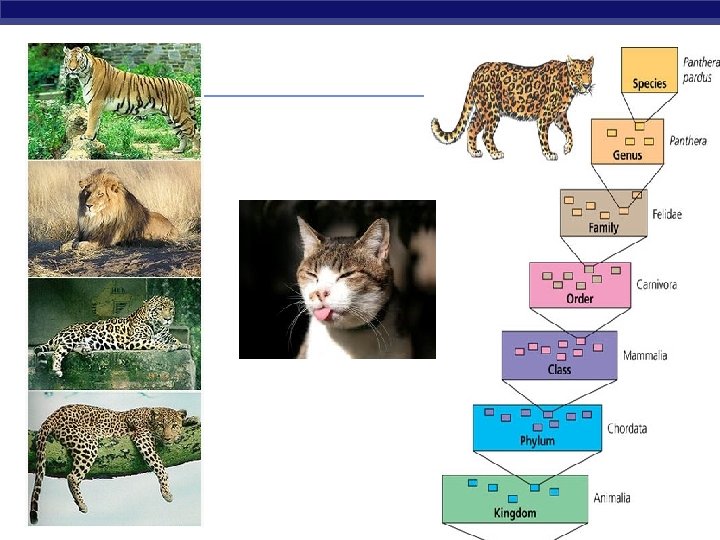
Hierarchical Classification Taxonomy- classifying organisms and assigning each organism a scientific name Carolus Linnaeus AP Biology § Kingdom § Phylum § Class § Order § Family § Genus § Species
AP Biology

Modern Taxonomy Based on Theory of Evolution u New species arise, evolve, over long periods of time from preexisting species SPECIES: a group of similar organisms that interbreed in nature AP Biology

Binomial Nominclature Ursus maritimus Latinized, two- word naming system RULES: Genus species Ursus americanus or Genus species or G. species AP Biology Ailuropoda melanoleuca
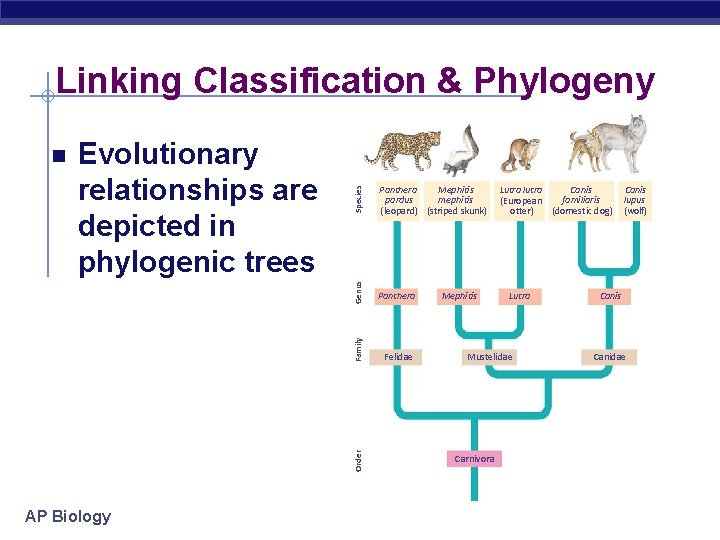
Species Panthera Mephitis pardus mephitis (leopard) (striped skunk) Genus Evolutionary relationships are depicted in phylogenic trees Panthera Felidae Order Family Linking Classification & Phylogeny AP Biology Mephitis Canis Lutra lutra familiaris (European (domestic dog) otter) Lutra Mustelidae Carnivora Canis lupus (wolf) Canis Canidae
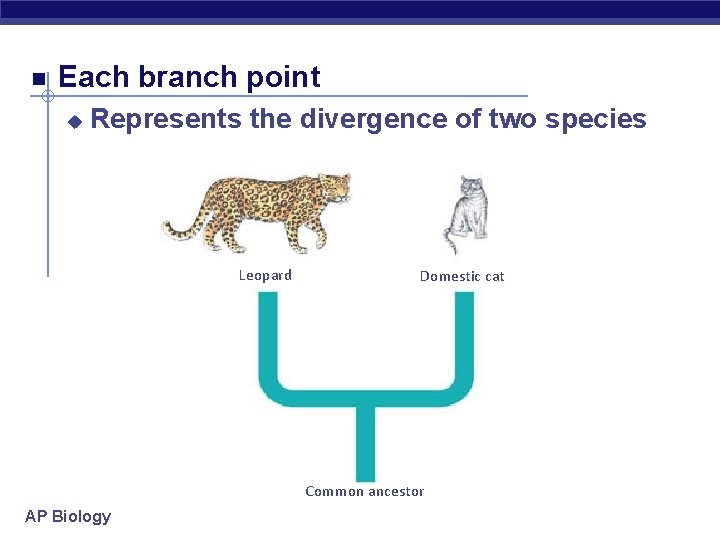
Each branch point u Represents the divergence of two species Leopard Domestic cat Common ancestor AP Biology
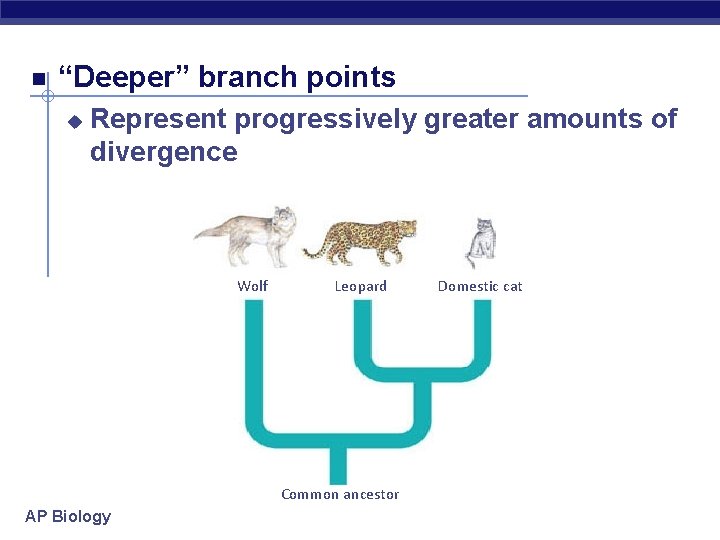
“Deeper” branch points u Represent progressively greater amounts of divergence Wolf Leopard Common ancestor AP Biology Domestic cat
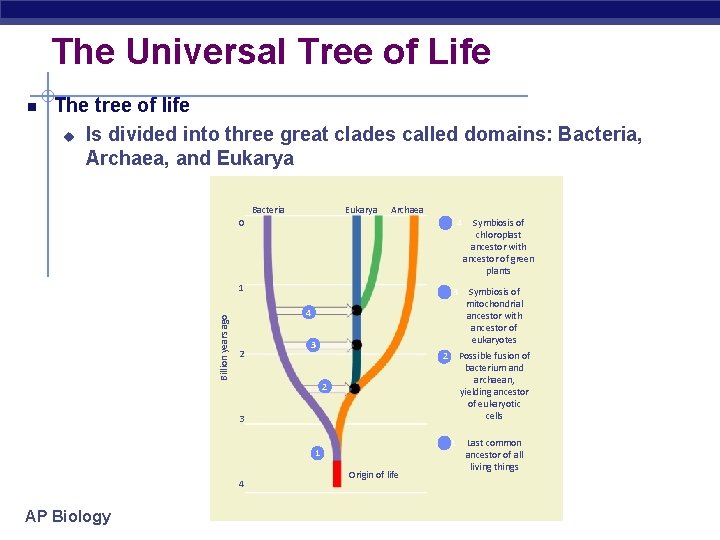
The Universal Tree of Life The tree of life u Is divided into three great clades called domains: Bacteria, Archaea, and Eukarya Bacteria Eukarya Archaea 4 0 Billion years ago 1 3 4 2 3 4 1 AP Biology Origin of life Symbiosis of mitochondrial ancestor with ancestor of eukaryotes Possible fusion of bacterium and archaean, yielding ancestor of eukaryotic cells 2 1 Symbiosis of chloroplast ancestor with ancestor of green plants Last common ancestor of all living things

Depicting Shared Characteristics A cladogram u A clade within a cladogram u u Is a depiction of patterns of shared characteristics among taxa Is defined as a group of species that includes an ancestral species and all its descendants Can be nested within later clades Cladistics u u Is the study of resemblances among clades Clades are defined by evolutionary relationships AP Biology

A valid clade is monophyletic u Signifying that it consists of the ancestor species and all its descendants Grouping 1 E D J H G F C K I B A Figure 25. 10 a AP Biology (a) Monophyletic. In this tree, grouping 1, consisting of the seven species B–H, is a monophyletic group, or clade. A monophyletic group is made up of an ancestral species (species B in this case) and all of its descendant species. Only monophyletic groups qualify as legitimate taxa derived from cladistics.

A paraphyletic clade u Is a grouping that consists of an ancestral species and some, but not all, of the descendants Grouping 2 G E D C J H K I F B A Figure 25. 10 b AP Biology (b) Paraphyletic. Grouping 2 does not meet the cladistic criterion: It is paraphyletic, which means that it consists of an ancestor (A in this case) and some, but not all, of that ancestor’s descendants. (Grouping 2 includes the descendants I, J, and K, but excludes B–H, which also descended from A. )

A polyphyletic grouping u Includes numerous types of organisms that lack a common ancestor Grouping 3 D E G J H I F C K B A Figure 25. 10 c AP Biology (c) Polyphyletic. Grouping 3 also fails the cladistic test. It is polyphyletic, which means that it lacks the common ancestor of (A) the species in the group. Furthermore, a valid taxon that includes the extant species G, H, J, and K would necessarily also contain D and E, which are also descended from A.

A shared primitive character Is a homologous structure that predates the branching of a particular clade from other members of that clade u Is shared beyond the taxon we are trying to define u A shared derived character u Is an evolutionary novelty unique to a particular clade AP Biology

Outgroups Systematists use a method called outgroup comparison u To differentiate between shared derived and shared primitive characteristics AP Biology

As a basis of comparison we need to designate an outgroup u which is a species or group of species that is closely related to the ingroup, the various species we are studying Outgroup comparison Is based on the assumption that homologies present in both the outgroup and ingroup must be primitive characters that predate the divergence of both groups from a common ancestor AP Biology u

The outgroup comparison Enables us to focus on just those characters that were derived at the various branch points in the evolution of a clade Lamprey Tuna Salamander Turtle Leopard TAXA Hair 0 0 0 1 Amniotic (shelled) egg 0 0 1 1 Four walking legs 0 0 0 1 1 1 Hinged jaws 0 0 1 1 Vertebral column (backbone) 0 1 1 1 CHARACTERS u Lancelet (outgroup) Turtle Salamander Tuna Lamprey Lancelet (outgroup) Figure 25. 11 a, b AP Biology (a) Character table. A 0 indicates that a character is absent; a 1 indicates that a character is present. Leopard Hair Amniotic egg Four walking legs Hinged jaws Vertebral column (b) Cladogram. Analyzing the distribution of these derived characters can provide insight into vertebrate phylogeny.

Phylogenetic Trees and Timing Any chronology represented by the branching pattern of a phylogenetic tree u Is relative rather than absolute in terms of representing the timing of divergences AP Biology

Phylograms ia n ph ib Figure 25. 12 AP Biology ou se M at R um H rd Bi an Am let ce n La sh In a phylogram u The length of a branch in a cladogram reflects the number of genetic changes that have taken place in a particular DNA or RNA sequence in that lineage Fi Dr ila ph o os

ou se M rd Hu ma n R at Bi Am Fi sh el e t ph ib la 251 Millions of years ago 542 Paleozoic Proterozoic Figure 25. 13 AP Biology La nc ph i ro so D 65. 5 Cenozoic In an ultrametric tree u The branching pattern is the same as in a phylogram, but all the branches that can be traced from the common ancestor to the present are of equal length Mesozoic ian Ultrametric Trees

Maximum Parsimony and Maximum Likelihood Systematists Can never be sure of finding the single best tree in a large data set u Narrow the possibilities by applying the principles of maximum parsimony and maximum likelihood u AP Biology

Among phylogenetic hypotheses u The most parsimonious tree is the one that requires the fewest evolutionary events to have occurred in the form of shared derived characters AP Biology

Applying parsimony to a problem in molecular systematics Human Mushroom 0 Mushroom Tulip 30% 0 Tulip Figure 25. 14 AP Biology (a) Percentage differences between sequences 40% 0
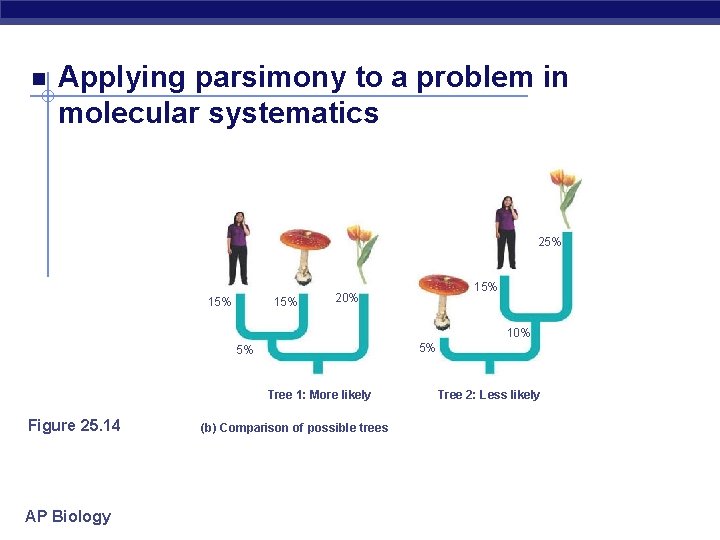
Applying parsimony to a problem in molecular systematics 25% 15% 15% 20% 10% 5% 5% Tree 1: More likely Figure 25. 14 AP Biology (b) Comparison of possible trees Tree 2: Less likely

The principle of maximum likelihood u States that, given certain rules about how DNA changes over time, a tree can be found that reflects the most likely sequence of evolutionary events APPLICATION In considering possible phylogenies for a group of species, systematists compare molecular data for the species. The most efficient way to study the various phylogenetic hypotheses is to begin by first considering the most parsimonious—that is, which hypothesis requires the fewest total evolutionary events (molecular changes) to have occurred. TECHNIQUE 1 Follow the numbered steps as we apply the principle of parsimony to a hypothetical phylogenetic problem involving four closely related bird species. First, draw the possible phylogenies for the species (only 3 of the 15 possible trees relating these four species are shown here). Species I I II I IV Species III Species II II I IV II IV Three possible phylogenetic hypothese 1 2 Tabulate the molecular data for the species (in this simplified example, the data represent a DNA sequence consisting of just seven nucleotide bases). Species Figure 25. 15 a AP Biology 3 Now focus on site 1 in the DNA sequence. A single basechange event, marked by the crossbar in the branch leading to species I, is sufficient to account for the site 1 data. Sites in DNA sequence 2 5 6 3 4 7 I A G G G T II G G G A G G G III G A G G A A T IV G G A A G I A II G III G G Base-change event IV G G G Bases at site 1 for each species III
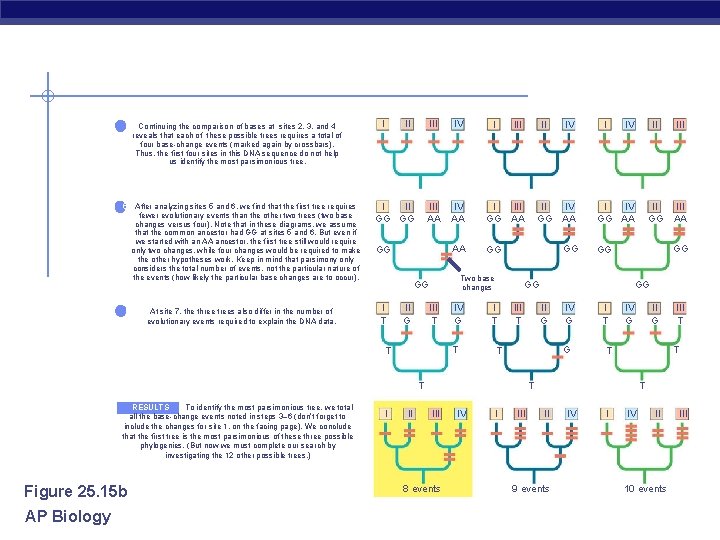
4 Continuing the comparison of bases at sites 2, 3, and 4 reveals that each of these possible trees requires a total of four base-change events (marked again by crossbars). Thus, the first four sites in this DNA sequence do not help us identify the most parsimonious tree. 5 After analyzing sites 5 and 6, we find that the first tree requires fewer evolutionary events than the other two trees (two base changes versus four). Note that in these diagrams, we assume that the common ancestor had GG at sites 5 and 6. But even if we started with an AA ancestor, the first tree still would require only two changes, while four changes would be required to make the other hypotheses work. Keep in mind that parsimony only considers the total number of events, not the particular nature of the events (how likely the particular base changes are to occur). 6 At site 7, the three trees also differ in the number of evolutionary events required to explain the DNA data. I II GG GG III IV III AA IV AA I III GG AA AA GG GG III Two base changes GG I T I III T IV G T T Figure 25. 15 b AP Biology I II IV II GG IV AA I IV GG AA GG GG I IV I T III T 8 events IV II G IV G G T I III II GG III AA GG I T IV G III T T III II GG GG T RESULTS To identify the most parsimonious tree, we total all the base-change events noted in steps 3– 6 (don’t forget to include the changes for site 1, on the facing page). We conclude that the first tree is the most parsimonious of these three possible phylogenies. (But now we must complete our search by investigating the 12 other possible trees. ) II T II 9 events IV II 10 events III

Phylogenetic Trees as Hypotheses The best hypotheses for phylogenetic trees u Are those that fit the most data: morphological, molecular, and fossil AP Biology

Sometimes there is compelling evidence u That the best hypothesis is not the most parsimonious Bird Lizard Mammal Four-chambered heart (a) Mammal-bird clade Bird Lizard Mammal Four-chambered heart Figure 25. 16 a, b AP Biology (b) Lizard-bird clade

Evolutionary history in one’s geneone Comparing nucleic acids or other molecules to infer relatedness u Is a valuable tool for tracing organisms’ evolutionary history Gene duplication u Is one of the most important types of mutation in evolution because it increases the number of genes in the genome, providing further opportunities for evolutionary changes AP Biology
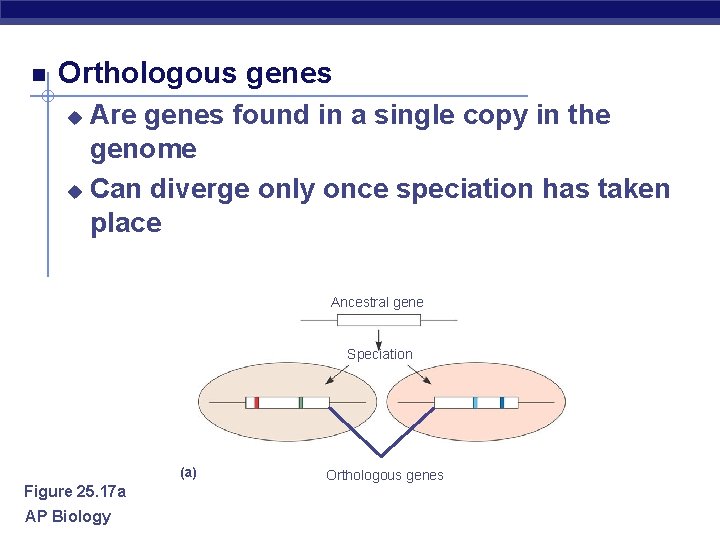
Orthologous genes Are genes found in a single copy in the genome u Can diverge only once speciation has taken place u Ancestral gene Speciation (a) Figure 25. 17 a AP Biology Orthologous genes

Paralogous genes Result from gene duplication, so they are found in more than one copy in the genome u Can diverge within the clade that carries them, often adding new functions u Ancestral gene Gene duplication Figure 25. 17 b AP Biology (b) Paralogous genes

Genome Evolution Orthologous genes are widespread u And extend across many widely varied species The widespread consistency in total gene number in organisms of varying complexity u Indicates that genes in complex organisms are extremely versatile and that each gene can perform many functions AP Biology

Molecular Clocks The molecular clock u Is a yardstick for measuring the absolute time of evolutionary change based on the observation that some genes and other regions of genomes appear to evolve at constant rates AP Biology

Neutral Theory Neutral theory states that Much evolutionary change in genes and proteins has no effect on fitness and therefore is not influenced by Darwinian selection u And that the rate of molecular change in these genes and proteins should be regular like a clock u AP Biology

Applying a Molecular Clock: The Origin of HIV Phylogenetic analysis shows that HIV u Is descended from viruses that infect chimpanzees and other primates A comparison of HIV samples from throughout the epidemic u Has shown that the virus has evolved in a remarkably clocklike fashion AP Biology
- Slides: 43