CHAPTER 18 THE STRUCTUREAL BASIS OF CELLULAR INFORMATION

CHAPTER 18 THE STRUCTUREAL BASIS OF CELLULAR INFORMATION: DNA, CHROMOSOMES, AND THE NUCLEUS 1 The chemical nature of the genetic material 2 DNA structure 3 The organization of DNA in genomes 4 DNA packaging 5 the nucleus

18. 3 The organization of DNA in genomes Genome: the DNA (or RNA in the case of some viruses) that contains one complete copy of all the genetic information of an organism or virus. Comparison of genomes between viruses/prokaryotes and eukaryotic cells: Viruses and prokaryotes Both virus and prokaryotes have small number of genomes. Virus: genome resides in a single linear or circular DNA/RNA molecules. Prokaryotes: genome are circular DNA molecules. Eukaryotic cells: have variety of genomes: (1) nuclear genome (linear) (2) mitochondrial genome (single, circular) (3) chloroplast genome (single circular).
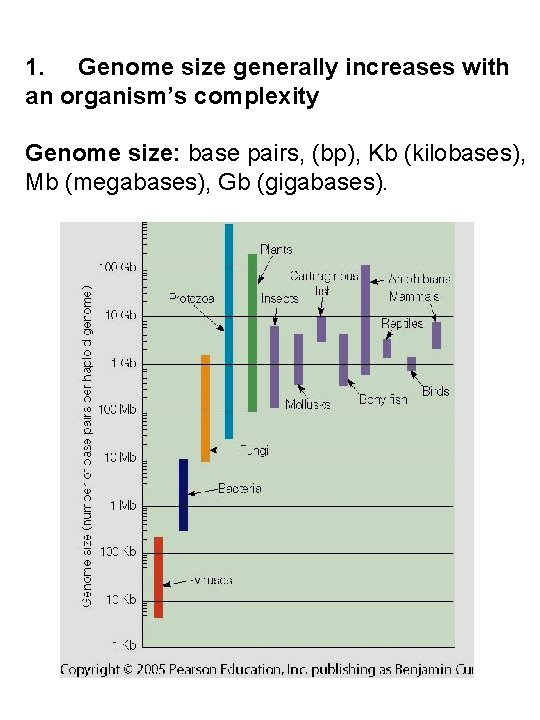
1. Genome size generally increases with an organism’s complexity Genome size: base pairs, (bp), Kb (kilobases), Mb (megabases), Gb (gigabases).

2. Restriction Enzymes: Enzymes isolated from bacteria that cut foreign DNA molecules at specific sites. (enzymes and methyl groups: -CH 3) Restriction Site: Each restriction enzyme recognizes a specific nucleotide sequence and cuts the DNA molecule at only that specific sequence. Stick ends and blunt ends: Many restriction enzymes leave a short length of unpaired bases after cutting DNA, which is called a “sticky” end, whereas other restriction enzymes make a cut across both strands creating double stranded DNA fragments with “blunt” ends. In general, restriction sites are palindromic, meaning they read the same sequence of bases forwards and backwards on the opposite DNA strand. Name of restriction enzymes: They are named after the bacteria from which restriction enzymes are obtained. Eco. RI: from E. coli strain R Hae. III: from Hemophilus aegyptius
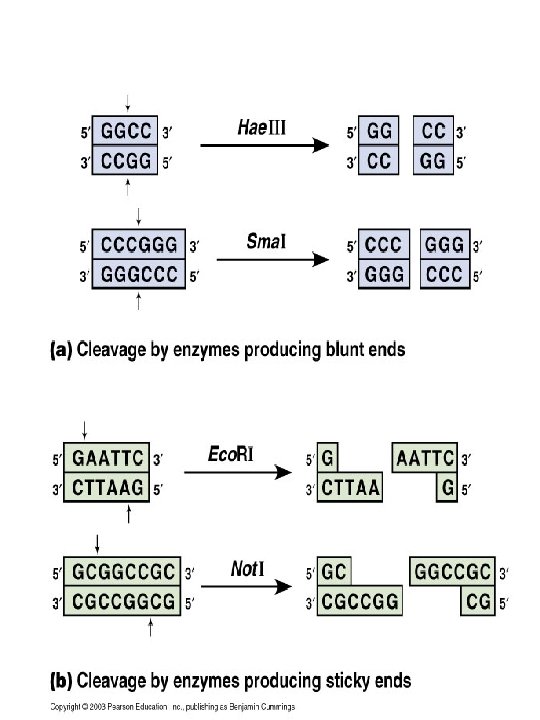
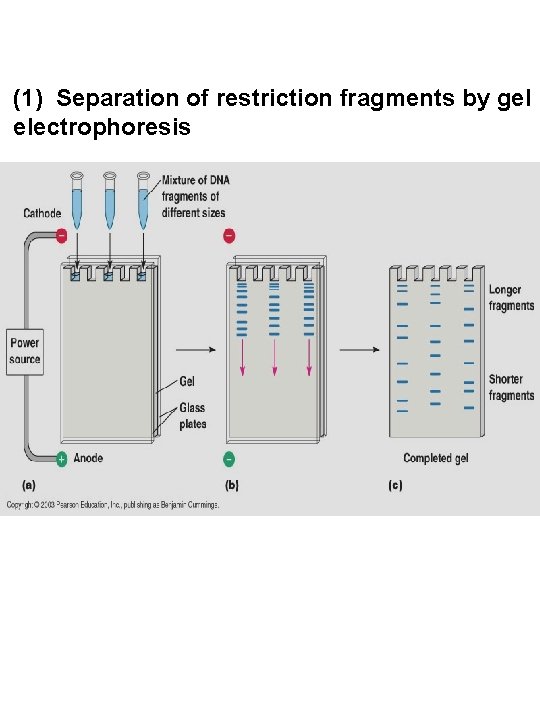
(1) Separation of restriction fragments by gel electrophoresis
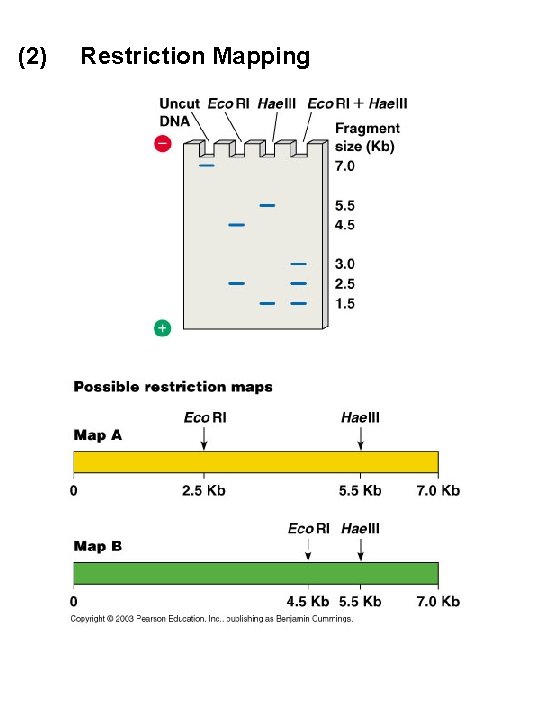
(2) Restriction Mapping
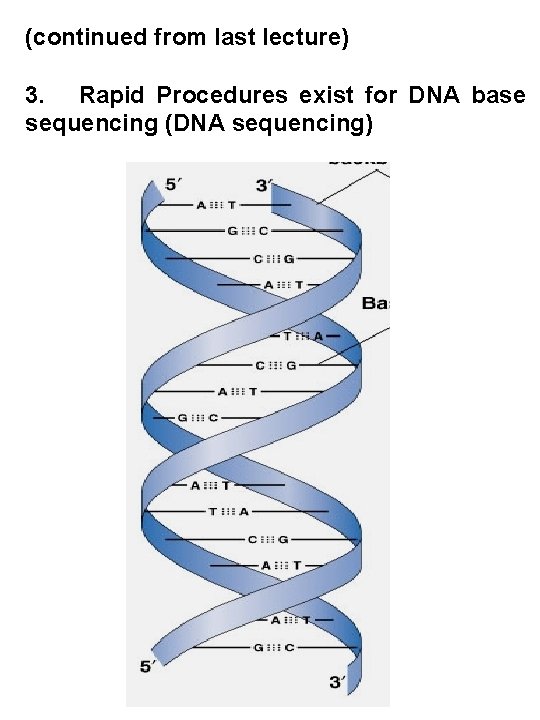
(continued from last lecture) 3. Rapid Procedures exist for DNA base sequencing (DNA sequencing)

Sample: Nucleic Acid Sequence of GM-CSF receptor (alpha subunit. 1) agcaggtggaaggagaggaagcggatgcc 29 gtggggtttacagcaggaaaatccgtggagacagcagatccgagaagcggcgatgtttgc 89 gtagaaccctgtacgtgcttcggcctgtcgctcttcccttctctctgaccagcacc 149 atgcttctcctggtgacaagccttctgctctgtgagttaccacacccagcattcctcctg 209 atcccagagaaatcggatctgcgaacagtggcaccagcctctagtctcaatgtgaggttt gactccaggacgatgaatttaagctgggactgccaagaaaacacaaccttcagcaagtgt 329 ttcttaactgacaagaagaacagagtcgtggaacccaggctcagtaacaacgaatgttcg 389 tgcacatttcgtgaaatttgtctgcatgaaggagtcacatttgaggttcacgtgaatact 449 agtcaaagaggagttcaacagaaactgctttatccaaattcaggaagggtaccgct 509 gctcagaatttctcctgtttcatctacaatgcggatttaatgaactgtacctgggcgagg 569 ggtccgacggccccccgtgacgtccagtattttttgtacatacgaaactcaaagagaagg 629 agggagatccggtgtccttattacaagactcaggaacccatgtgggatgtcacctg 689 gataacctgtcaggattaacgtctcgcaattactttctggttaacggaaccagccgagaa 749 attggcatccaattctttgattcacttttggacacaaagaaaatagaacgattcaaccct 809 cccagcaatgtcaccgtacgttgcaacacgacgcactgcctcgtacggtggaaacagccc 869 aggacctatcagaagctgtcgtacctggactttcagtaccagctggacgtccacagaaag 929 aatacccagcctggcacggaaaacctactcattaatgtttctggtgatttggaaaataga 989 tacaactttccaagctctgagcccagagcaaaacacagtgtgaagatcagagctgcagac 1049 gtccgcatcttgaattggagctcctggagtgaagccattgaatttggttctgacgacggg 1109 aacctcggctctgtgtacatttatgtgctcctaatcgtgggaacccttgtctgtggcatc 1169 gtcctcggcttcctctttaaaaggttccttaggatacagcggctgttcccgccagttcca 1229 cagatcaaagacaaactgaatgataaccatgaggtggaagacgagatcatctgggaggaa 1289 ttcaccccagaggaagggaaaggctaccgcgaagaggtcttgaccgtgaaggaaattacc 1349 tgagacccagagggtgtaggaatggcatggacatctccgcgacacgggggaact 1409 ttttcttgatgatgctgtgaacctttatatcattttctatgtttttaaaaacatg 1469
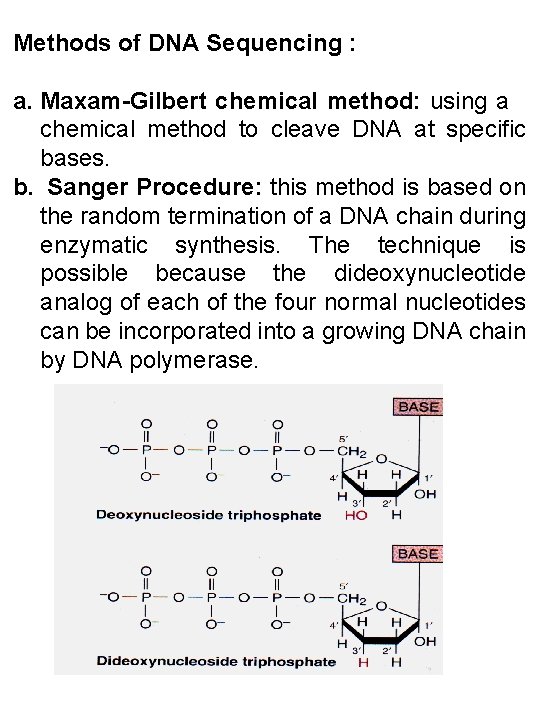
Methods of DNA Sequencing : a. Maxam-Gilbert chemical method: using a chemical method to cleave DNA at specific bases. b. Sanger Procedure: this method is based on the random termination of a DNA chain during enzymatic synthesis. The technique is possible because the dideoxynucleotide analog of each of the four normal nucleotides can be incorporated into a growing DNA chain by DNA polymerase.
DNA synthesis in normal: Template primer d. ATP, d. CTP, d. GTP, and d. TTP DNA polymerase DNA synthesis in Sanger procedure: Template primer d. ATP, d. CTP, d. GTP, and d. TTP dd. ATP, dd. CTP, dd. GTP, and dd. TTP DNA polymerase

Radioactive Sanger Method 1. Denature DNA into single strands. The single strand is mixed with primers designed to anneal the 3’ end of each single strand 2. The samples of the primer bound singlestranded DNA are distributed into four tubes. Add DNA polymerase and four d. NTPs (d. ATP, d. CTP, d. GTP, and d. TTP). 3. Then, small amounts of dd. ATP, dd. CTP, dd. GTP, or d. TTP will be added to corresponding tubes, respectively. 4. Run gel, develop film, and DNA sequence is visualized
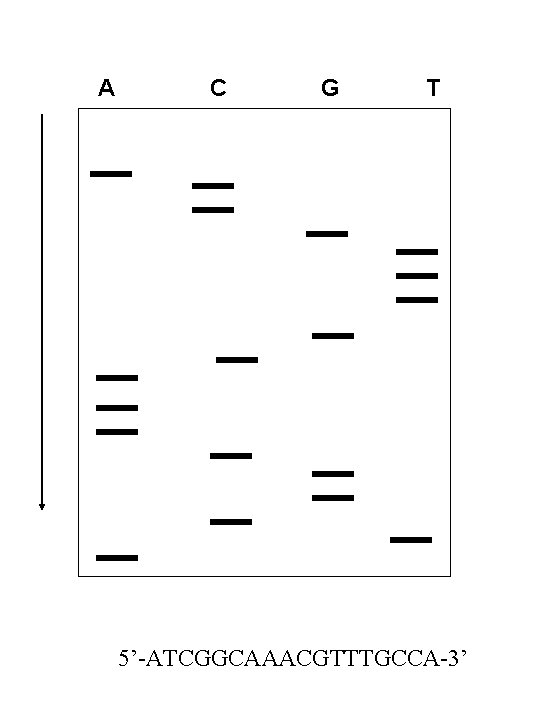
A C G T 5’-ATCGGCAAACGTTTGCCA-3’
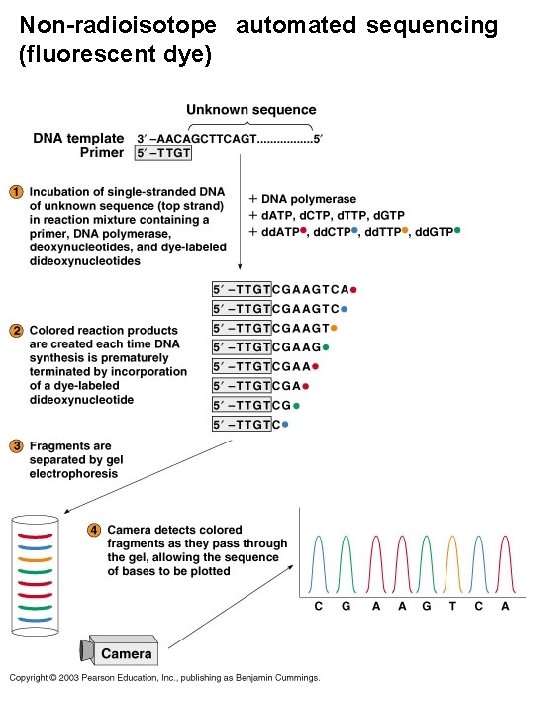
Non-radioisotope automated sequencing (fluorescent dye)

4. Single nucleotide polymorphisms and Tandemly repeated DNA. (SNPs), (1) Human nuclear genome contains about 3. 2 billion bases (about 23000 protein-cording genes) (2) On average, about 99. 7% of DNA bases in our body will match perfectly with this published sequence. The remaining 0. 3% of the bases vary from person to person, creating features that make us unique individuals. Differences involving single base changes are called single nucleotide polymorphisms (SNPs). (3) Most SNPs are not located in the protein coding regions of genes. They are usually located near each other on the same chromosome. (4) Genome contains repeated DNA sequences (Variable Number Tandemly Repeats-VNTR), which is located in intron areas. A given person's VNTRs come from the genetic information donated by his or her parents; he or she could have VNTRs inherited from his or her mother or father, or a combination, but never a VNTR either of his or her parents do not have.

5. Restriction digestion and Southern blot are used to analyze DNA fingerprinting pattern. (1) (2) (3) (4) (5) (6) Isolate DNA from individuals Restriction enzyme digestion. Run gel Transfer to a membrane Hybridization with radioactive probe (VNPR probe). Verify VNTR pattern. Note: (1) since restriction enzyme digestion will produced different fragments in size due to single nucleotide polymorphisms, it also called restriction fragment length polymorphisms (RFLP) (2) the procedure described above is called Southern blot. (3) DNA fingerprinting pattern can be used for diagnosing genetic diseases and solving violent crimes, or determining family linking.
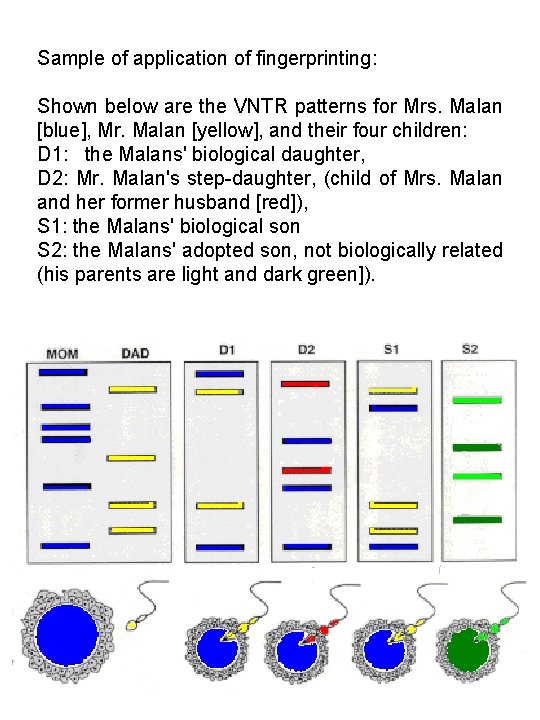
Sample of application of fingerprinting: Shown below are the VNTR patterns for Mrs. Malan [blue], Mr. Malan [yellow], and their four children: D 1: the Malans' biological daughter, D 2: Mr. Malan's step-daughter, (child of Mrs. Malan and her former husband [red]), S 1: the Malans' biological son S 2: the Malans' adopted son, not biologically related (his parents are light and dark green]).
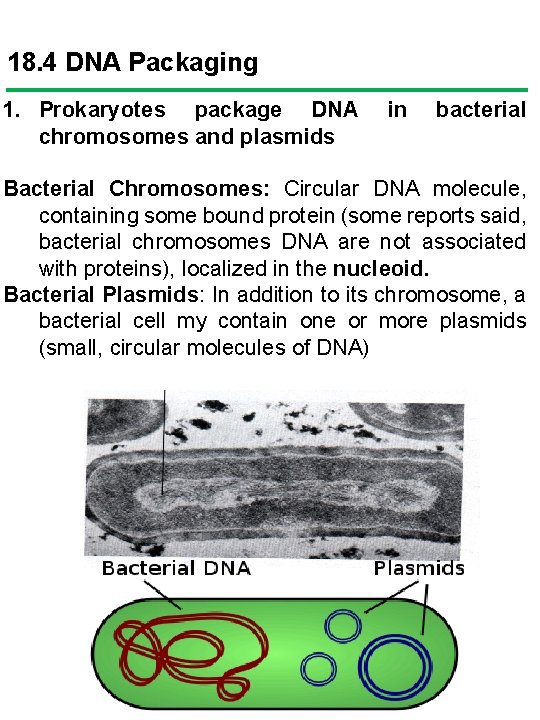
18. 4 DNA Packaging 1. Prokaryotes package DNA chromosomes and plasmids in bacterial Bacterial Chromosomes: Circular DNA molecule, containing some bound protein (some reports said, bacterial chromosomes DNA are not associated with proteins), localized in the nucleoid. Bacterial Plasmids: In addition to its chromosome, a bacterial cell my contain one or more plasmids (small, circular molecules of DNA)
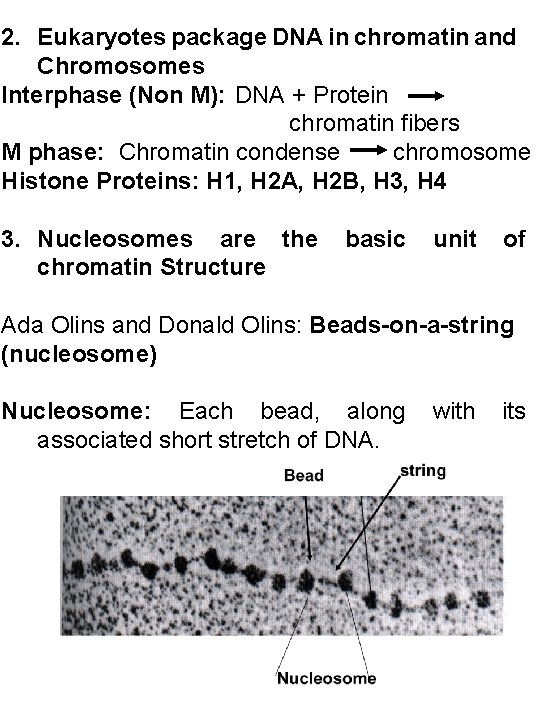
2. Eukaryotes package DNA in chromatin and Chromosomes Interphase (Non M): DNA + Protein chromatin fibers M phase: Chromatin condense chromosome Histone Proteins: H 1, H 2 A, H 2 B, H 3, H 4 3. Nucleosomes are the chromatin Structure basic unit of Ada Olins and Donald Olins: Beads-on-a-string (nucleosome) Nucleosome: Each bead, along associated short stretch of DNA. with its
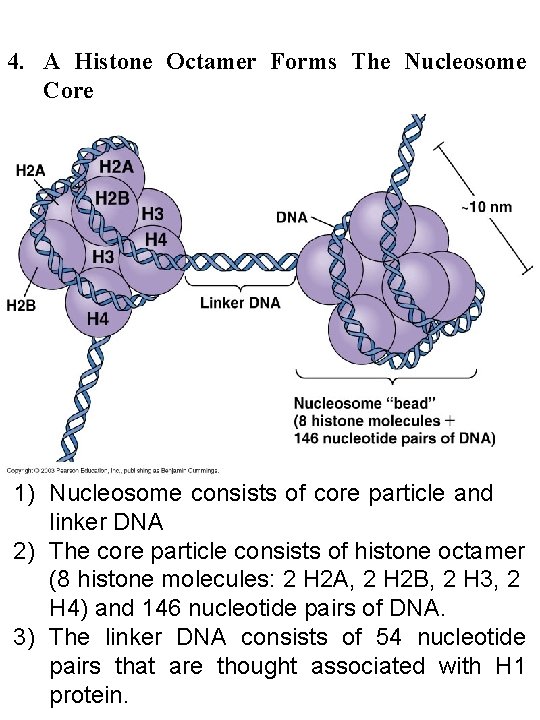
4. A Histone Octamer Forms The Nucleosome Core 1) Nucleosome consists of core particle and linker DNA 2) The core particle consists of histone octamer (8 histone molecules: 2 H 2 A, 2 H 2 B, 2 H 3, 2 H 4) and 146 nucleotide pairs of DNA. 3) The linker DNA consists of 54 nucleotide pairs that are thought associated with H 1 protein.
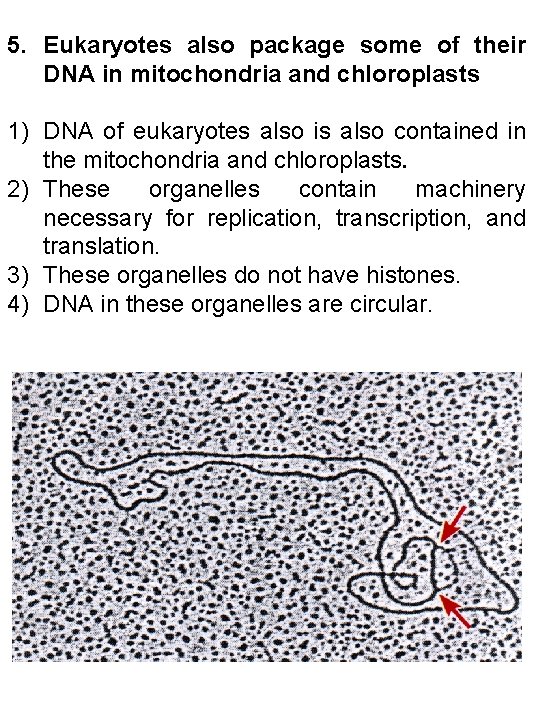
5. Eukaryotes also package some of their DNA in mitochondria and chloroplasts 1) DNA of eukaryotes also is also contained in the mitochondria and chloroplasts. 2) These organelles contain machinery necessary for replication, transcription, and translation. 3) These organelles do not have histones. 4) DNA in these organelles are circular.
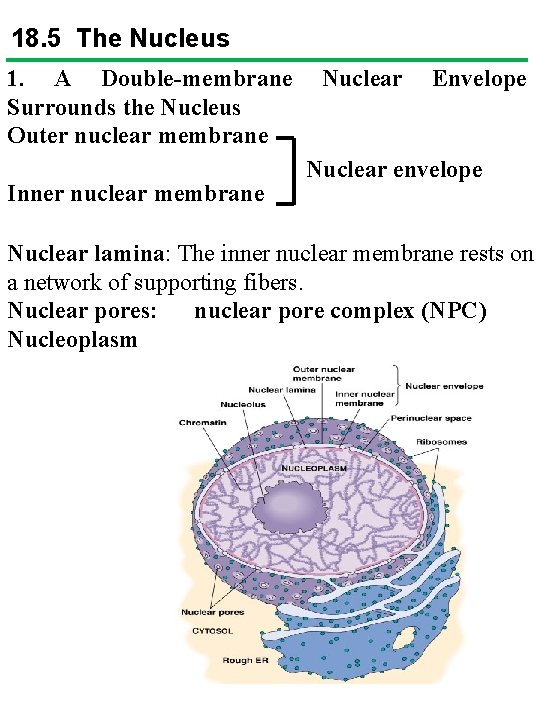
18. 5 The Nucleus 1. A Double-membrane Surrounds the Nucleus Outer nuclear membrane Inner nuclear membrane Nuclear Envelope Nuclear envelope Nuclear lamina: The inner nuclear membrane rests on a network of supporting fibers. Nuclear pores: nuclear pore complex (NPC) Nucleoplasm
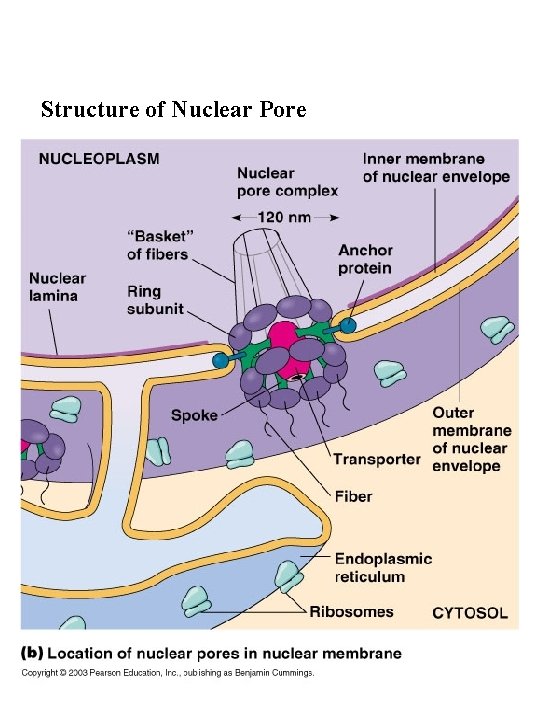
Structure of Nuclear Pore
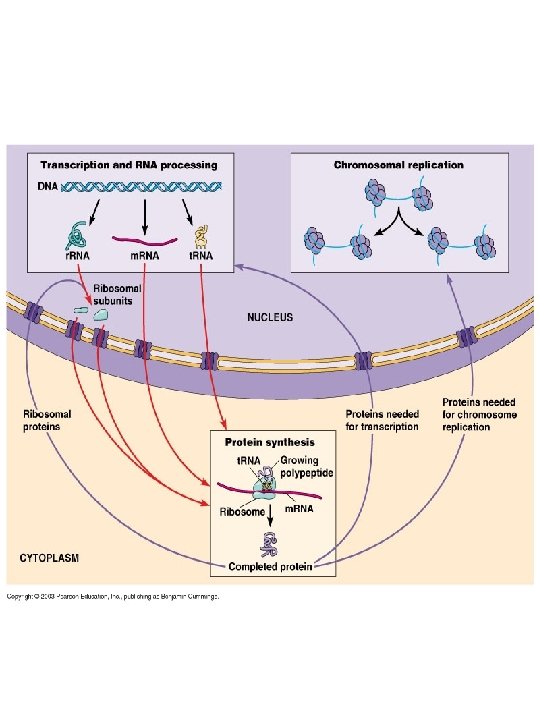
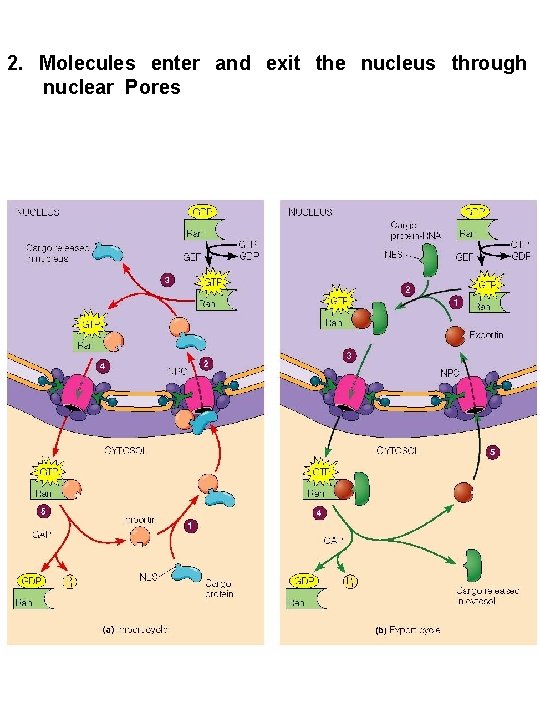
2. Molecules enter and exit the nucleus through nuclear Pores

Summary 1. The DNA (or RNA for some viruses) that makes up one complete set of an organism’s genetic information is called its genome. Eukaryotes also have a mitochondrial genome and, in the case of plants, a chloroplast genome. 2. Restriction enzyme are used to cut DNA into small fragments, and DNA sequencing techniques are used to determine the entire base sequence of a gene or the genomes of numerous organisms. 3. Genomic DNA contain large fraction that does not code for RNA or protein synthesis. 4. In prokaryotes, DNA packaged as chromosomes and plasmid complexes with relative small amounts of proteins. In eukaryotes, DNA packaged as chromatin/chromosomes complexes with large amount of proteins (histone). 5. The basic structural unit of the eukaryotic chromatin is the nucleosome. 6. Nucleus contain many nuclear pore on its membrane, which allows molecules to move between the cytosol and nucleoplasm.
- Slides: 26