Chapter 17 AP Biology From Gene to Protein

Chapter 17 AP Biology From Gene to Protein

Central Dogma of Molecular Biology DNA RNA Protein

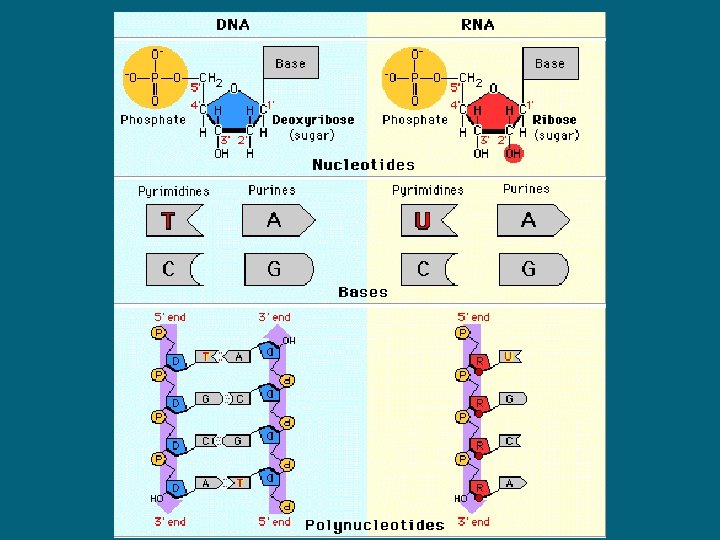
What is RNA? • Contains the bases A, C, G, and U instead of T • single-stranded (often folds onto itself) • Three types of RNA: messenger RNA (m. RNA), transfer RNA (t. RNA) and ribosomal RNA (r. RNA)
Protein Synthesis 1. Transcription - DNA message is transcribed into m. RNA and sent to ribosomes. 2. Translation - m. RNA is translated into a Protein by a ribosome. DNA RNA Protein
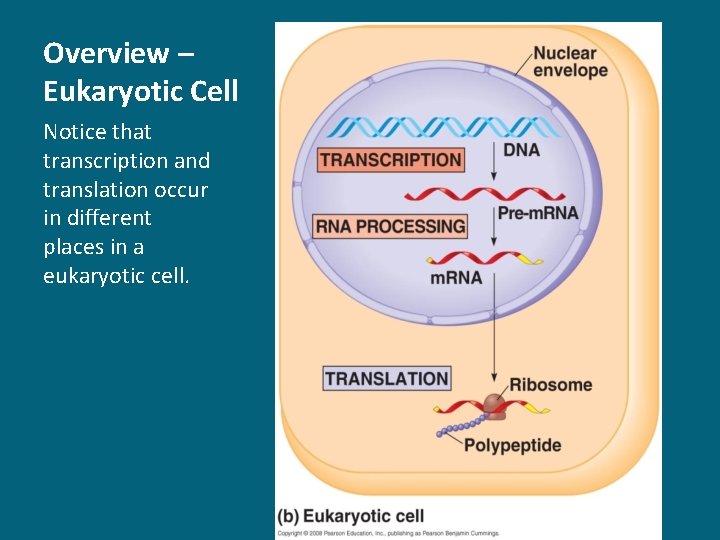
Overview – Eukaryotic Cell Notice that transcription and translation occur in different places in a eukaryotic cell.
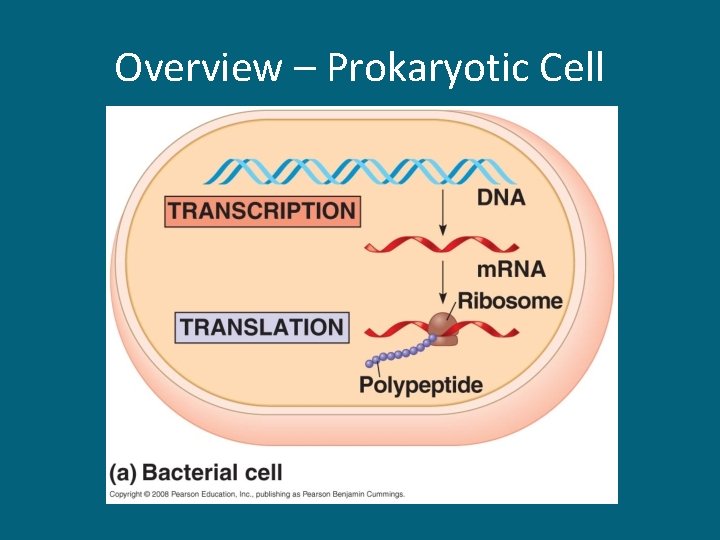
Overview – Prokaryotic Cell
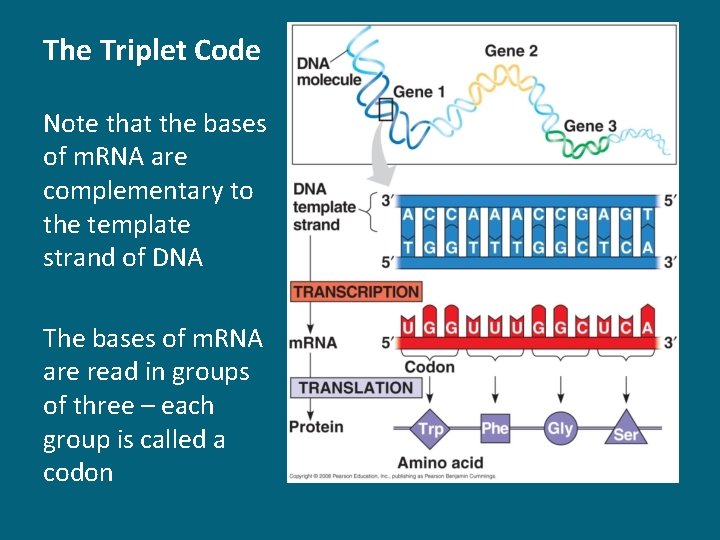
The Triplet Code Note that the bases of m. RNA are complementary to the template strand of DNA The bases of m. RNA are read in groups of three – each group is called a codon

From m. RNA to Amino Acids • The m. RNA base triplets are called codons • m. RNA is written in the 5’ to 3’ direction • The codons code for each of the 20 amino acids • The genetic code is redundant – More than one codon codes for each of the 20 amino acids
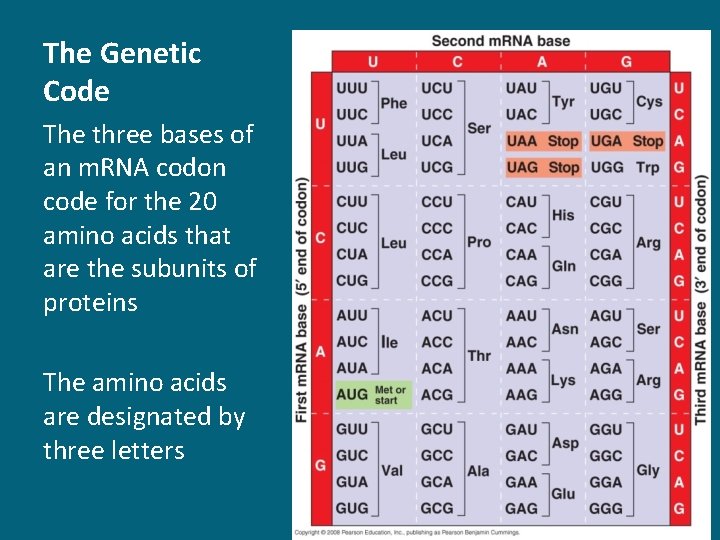
The Genetic Code The three bases of an m. RNA codon code for the 20 amino acids that are the subunits of proteins The amino acids are designated by three letters

Universal Nature of the Genetic Code
Transcription Promoter • The DNA sequence at which RNA polymerase attaches is called the promoter • RNA polymerase adds RNA nucleotides to a growing RNA strand in the 5’ 3’ direction

Transcription Unit • The entire stretch of DNA that is transcribed into an RNA molecule • A transcription unit may code for a polypeptide or an RNA (like t. RNA or r. RNA) – RNA Polymerase II makes m. RNA

Terminator • The DNA sequence that signals the end of transcription is called the terminator
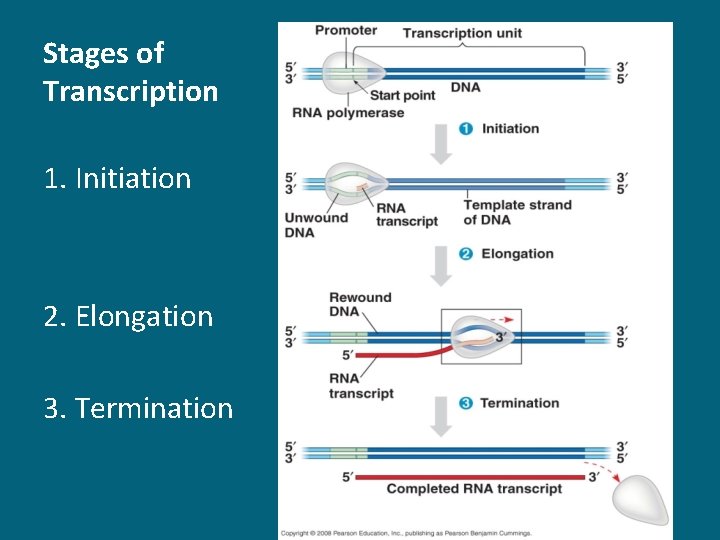
Stages of Transcription 1. Initiation 2. Elongation 3. Termination
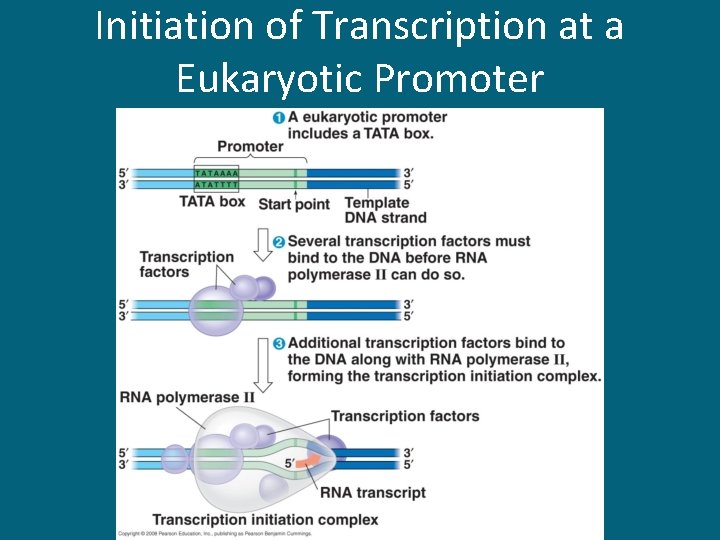
Initiation of Transcription at a Eukaryotic Promoter

RNA Processing • In eukaryotes, transcription results in prem. RNA – Pre-m. RNA undergoes processing to yield final m. RNA which leaves the nucleus and goes to the ribosomes • In prokaryotes transcription results in m. RNA – Processing of m. RNA does not occur in prokaryotes – Transcription and translation can occur simultaneously

Processing of Pre-m. RNA • Addition of a 5’ cap – A modified form of a guanine nucleotide • Addition of a poly-A tail – 50 -250 adenine (A) nucleotides are added – This is referred to as the poly-A tail • RNA splicing – Editing of the initial strand of m. RNA (a cut and paste job)

Addition of 5’ cap and poly-A tail

5’ cap and poly-A tail • Both facilitate export of m. RNA from the nucleus • Both help protect m. RNA from degradation by enzymes • Both facilitate the attachment of m. RNA to the ribosome

RNA Splicing • Only takes place in eukaryotic cells • Large sections are spliced out; these are called introns • The sections that remain are called exons – “exons EXIT the nucleus” • These exons are spliced together by a spliceosome to form the m. RNA that leaves the nucleus and travels to the ribosomes
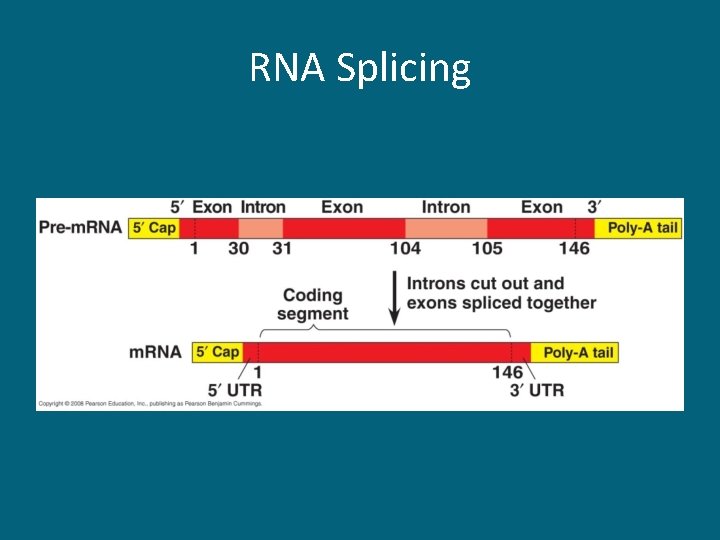
RNA Splicing
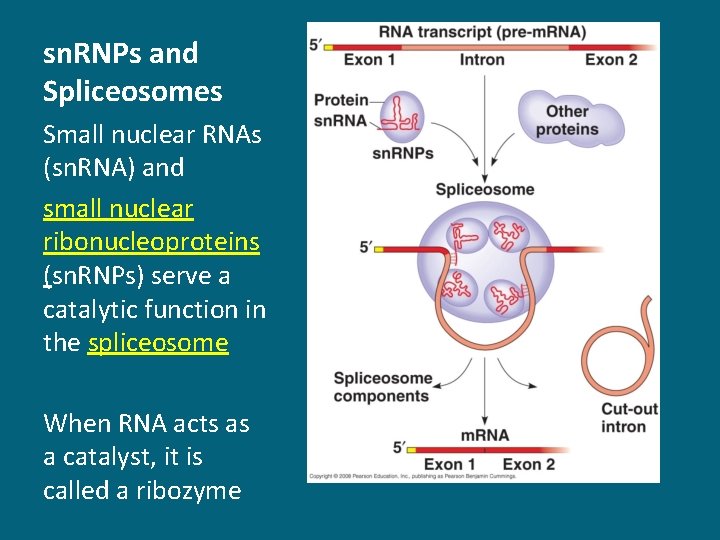
sn. RNPs and Spliceosomes Small nuclear RNAs (sn. RNA) and small nuclear ribonucleoproteins (sn. RNPs) serve a catalytic function in the spliceosome When RNA acts as a catalyst, it is called a ribozyme

Alternative RNA Splicing • Different regulatory proteins in different cells splice the pre-m. RNA in different ways • This is called alternative gene splicing • This allows for different combinations of exons • This results in more than one polypeptide per gene • This explains why we have fewer genes in our genome than what was expected

Transcription Animation • https: //www. youtube. com/watch? v=SMt. Wv. D bf. HLo

17. 4 Translation • t. RNA functions in transferring amino acids from the cytoplasm to a ribosome • r. RNA complexes with proteins to form the two subunits that make up ribosomes

t. RNA • Each type of t. RNA is specific for a particular amino acid • At one end t. RNA loosely binds the amino acid and at the other end it has a nucleotide triplet called the anticodon • The anticodon allows it to pair specifically with a complementary codon on m. RNA
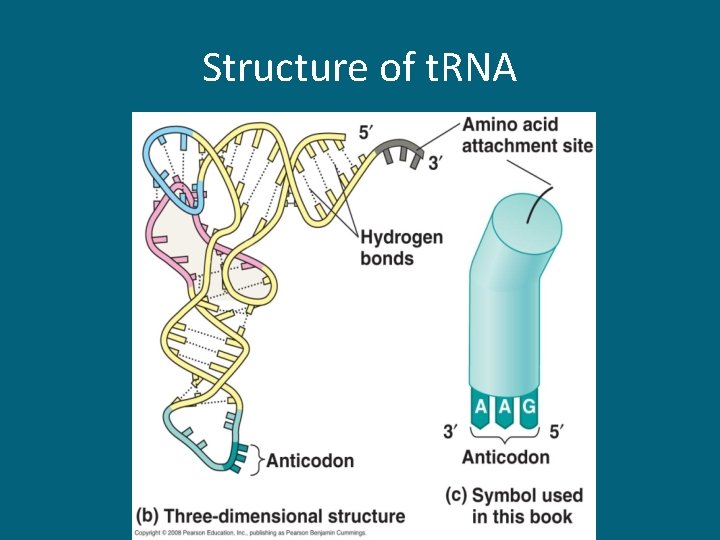
Structure of t. RNA
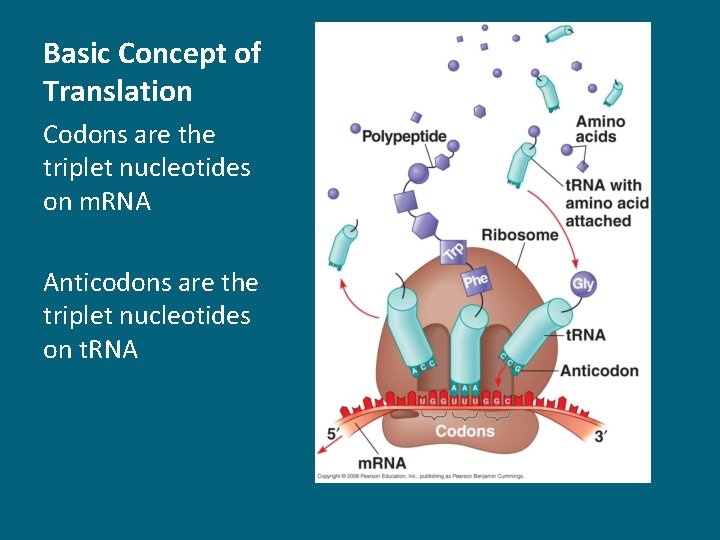
Basic Concept of Translation Codons are the triplet nucleotides on m. RNA Anticodons are the triplet nucleotides on t. RNA

Wobble • If one t. RNA variety existed for each codon, there would need to be 61 t. RNAs, there are only about 45, some can bind to more than codon • The rule for base pairing is not as strict between the third base of a codon and an anticodon, this relaxation of base-pairing rules is called wobble • Ex: a t. RNA with the anticodon 3’-UCU-5’ can base pair with either the m. RNA codon 5’-AGA-3’ or 5’AGG-3’, both code for Arg

Three Stages of Translation • Initiation – Begins with the start codon AUG (always!) • Elongation – Codon recognition – Peptide bond formation (between 2 a. a. ) • Termination – A stop codon is reached and translation stops
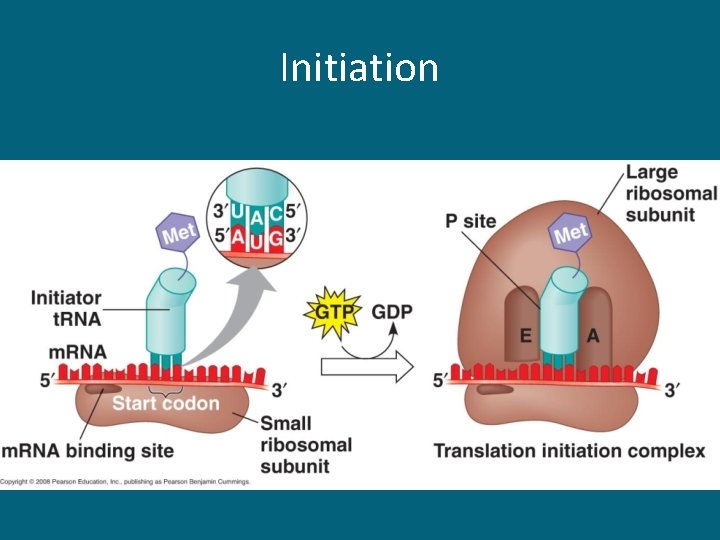
Initiation
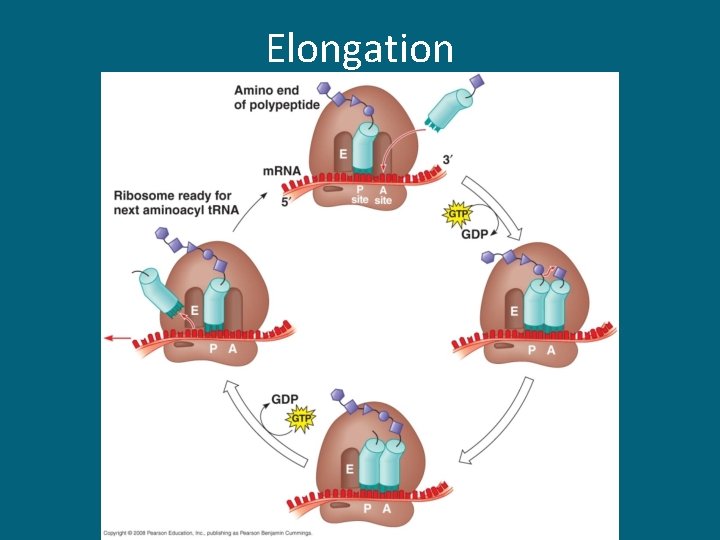
Elongation
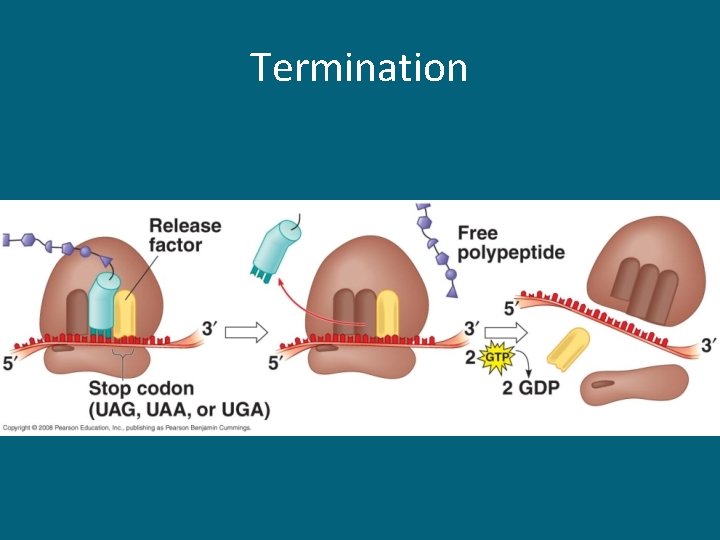
Termination

Translation Animation • https: //www. youtube. com/watch? v=Tf. Yf_r. P WUd. Y

Folding of the Polypeptide • Following release from the ribosome, the polypeptide then folds to its specific conformation • Chaperonins are the proteins that help with this folding process • The first 20 amino acids of the polypeptide serve as a signal peptide and act as a cellular zip code, directing the polypeptide to its final destination
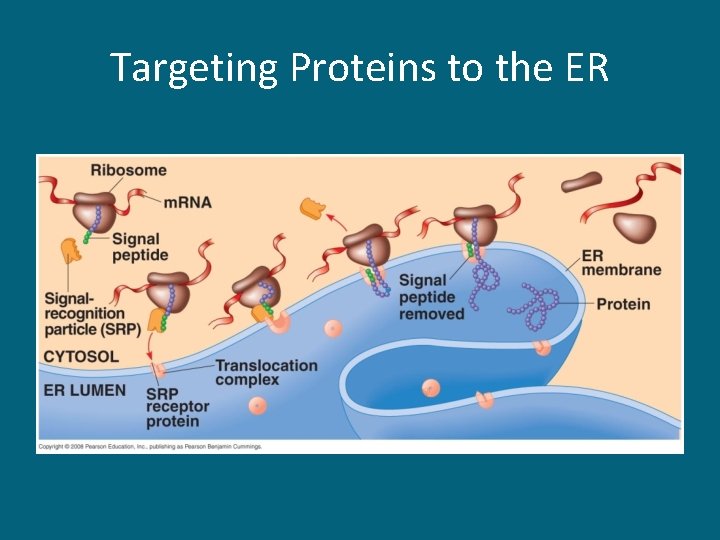
Targeting Proteins to the ER

17. 5 Point Mutations • Mutations are alterations in the genetic material of the cell caused by mutagens • Point mutations are alterations of just 1 base pair in a gene – Base-pair substitution • Silent mutations – have no effect on the encoded protein • Missense mutations – change one amino acid to another; might still code for the correct amino acid • Nonsense mutations – change a regular amino acid codon into a stop codon – Insertions & deletions • Frameshift mutation – m. RNA is read incorrectly
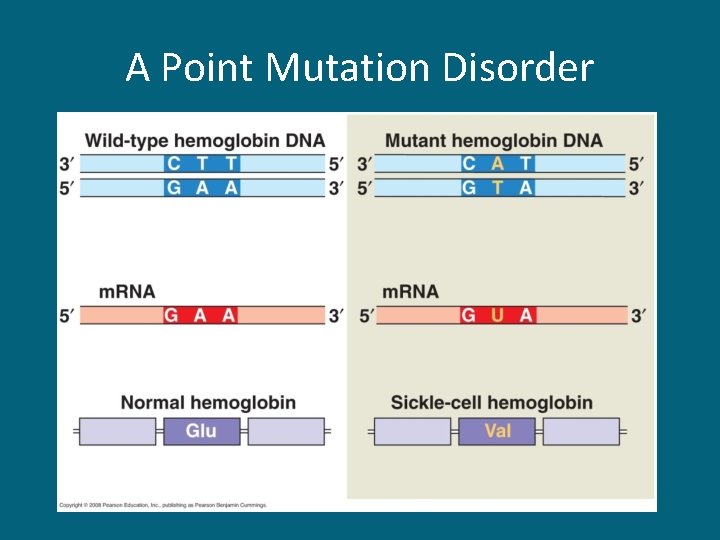
A Point Mutation Disorder
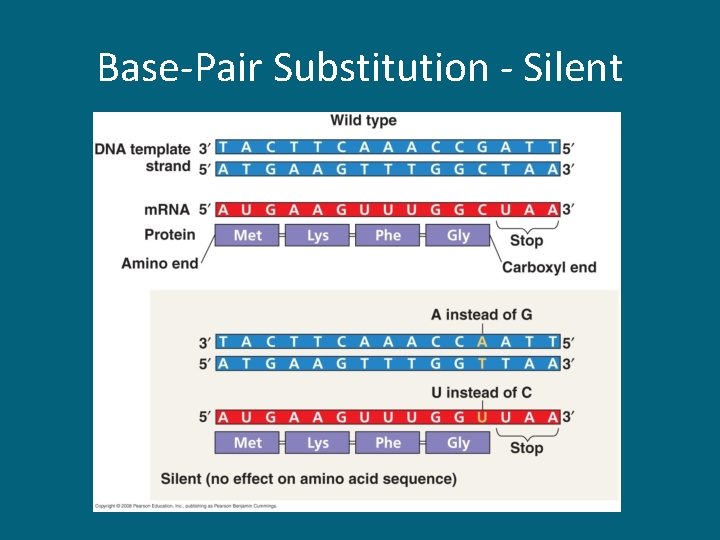
Base-Pair Substitution - Silent
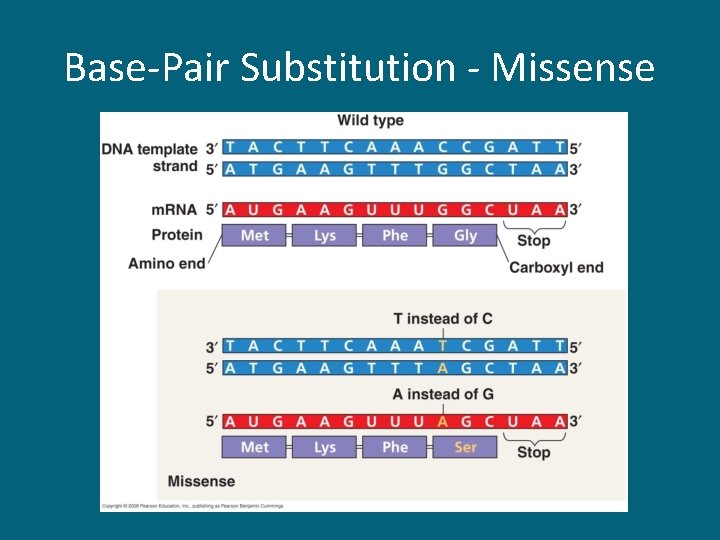
Base-Pair Substitution - Missense

Base-Pair Substitution - Nonsense
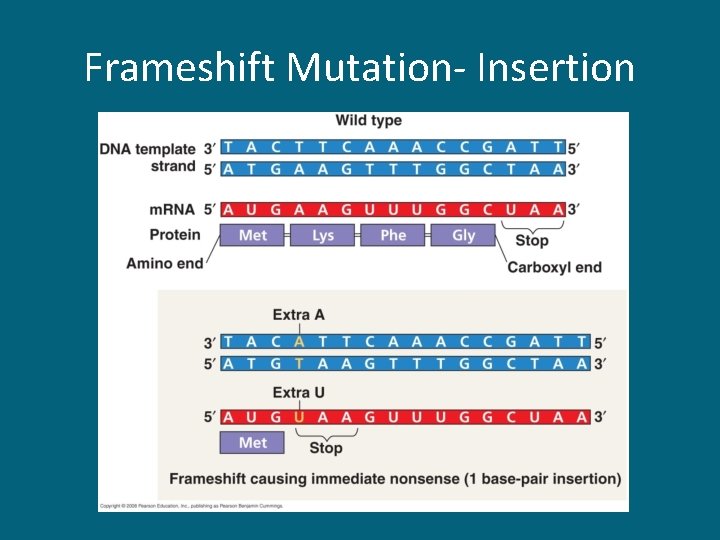
Frameshift Mutation- Insertion
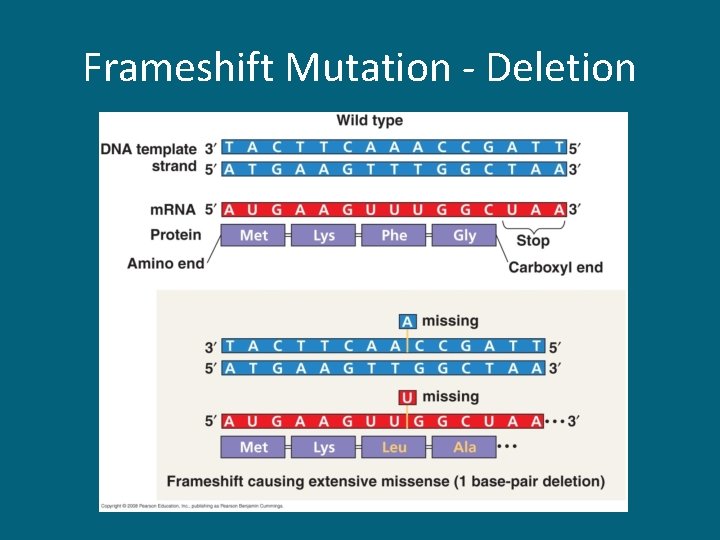
Frameshift Mutation - Deletion
- Slides: 44