Chapter 16 The Molecular Basis of Inheritance Scientific

Chapter 16~ The Molecular Basis of Inheritance

Scientific History • The march to understanding that DNA is the genetic material – T. H. Morgan (1908) – Frederick Griffith (1928) – Avery, Mc. Carty & Mac. Leod (1944) – Erwin Chargaff (1947) – Hershey & Chase (1952) – Watson & Crick (1953) – Meselson & Stahl (1958)

The “Transforming 1928 Principle” • Frederick Griffith – Streptococcus pneumonia bacteria • was working to find cure for pneumonia – harmless live bacteria (“rough”) mixed with heat-killed pathogenic bacteria (“smooth”) causes fatal disease in mice – a substance passed from dead bacteria to live bacteria to change their phenotype • “Transforming Principle”
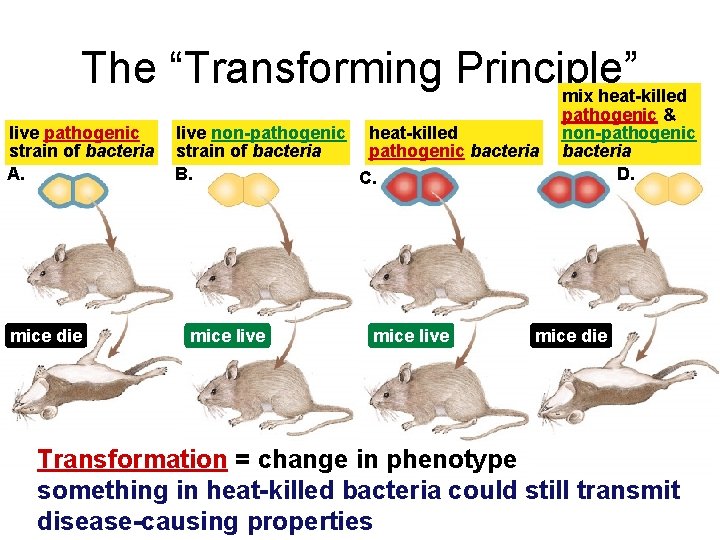
The “Transforming Principle” mix heat-killed live pathogenic strain of bacteria A. mice die live non-pathogenic heat-killed strain of bacteria pathogenic bacteria B. C. mice live pathogenic & non-pathogenic bacteria D. mice die Transformation = change in phenotype something in heat-killed bacteria could still transmit disease-causing properties
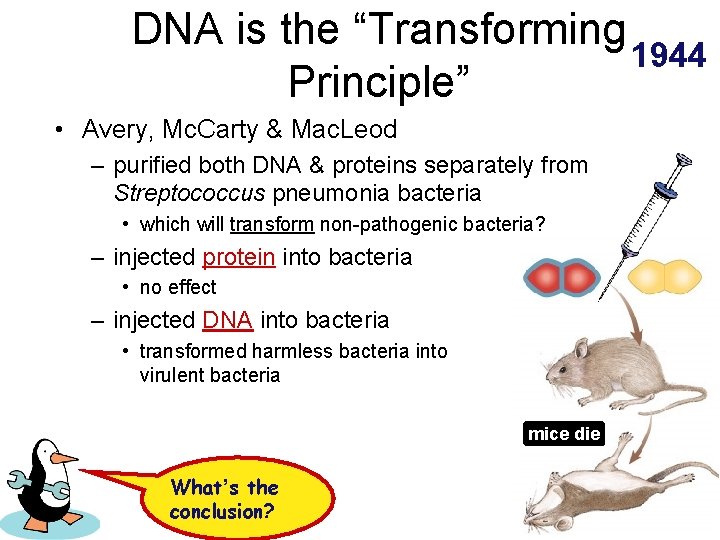
DNA is the “Transforming 1944 Principle” • Avery, Mc. Carty & Mac. Leod – purified both DNA & proteins separately from Streptococcus pneumonia bacteria • which will transform non-pathogenic bacteria? – injected protein into bacteria • no effect – injected DNA into bacteria • transformed harmless bacteria into virulent bacteria mice die What’s the conclusion?
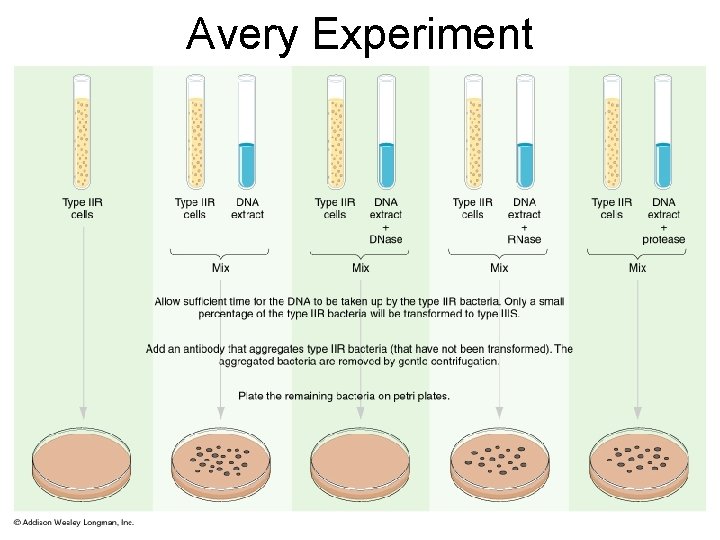
Avery Experiment

1944 | ? ? !! Avery, Mc. Carty & Mac. Leod • Conclusion – first experimental evidence that DNA was the genetic material Oswald Avery Maclyn Mc. Carty Colin Mac. Leod
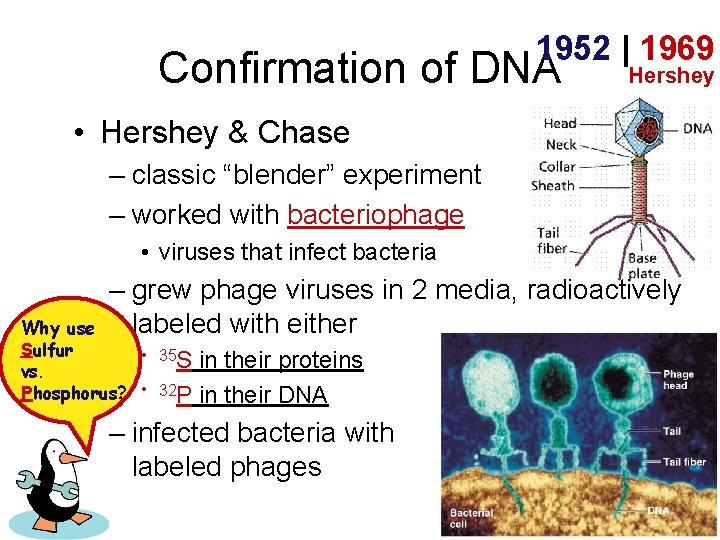
1952 | 1969 Confirmation of DNA Hershey • Hershey & Chase – classic “blender” experiment – worked with bacteriophage • viruses that infect bacteria – grew phage viruses in 2 media, radioactively labeled with either Why use Sulfur • 35 S vs. Phosphorus? • 32 P in their proteins in their DNA – infected bacteria with labeled phages
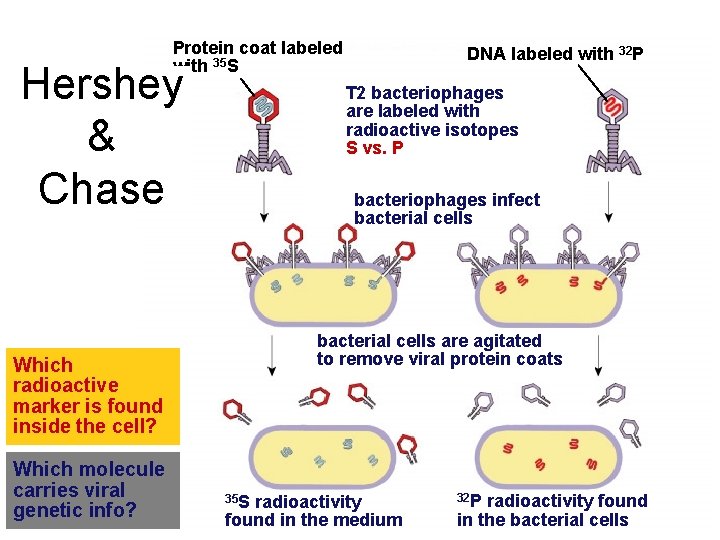
Protein coat labeled with 35 S Hershey & Chase T 2 bacteriophages are labeled with radioactive isotopes S vs. P bacteriophages infect bacterial cells are agitated to remove viral protein coats Which radioactive marker is found inside the cell? Which molecule carries viral genetic info? DNA labeled with 32 P 35 S radioactivity found in the medium 32 P radioactivity found in the bacterial cells
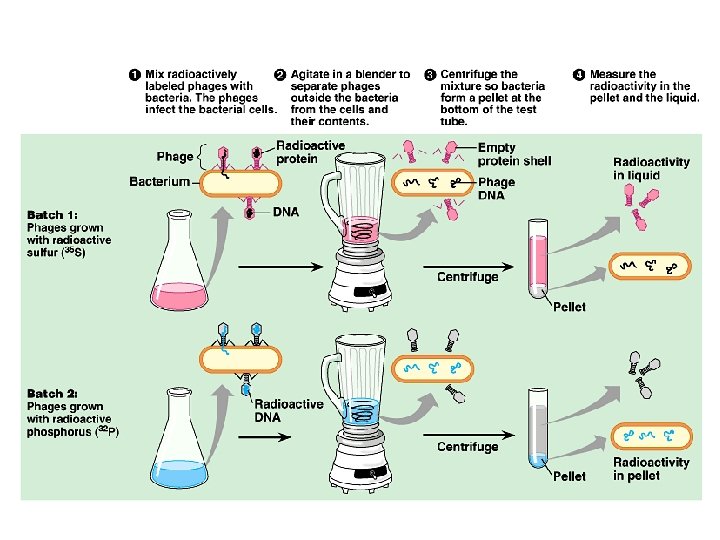

Blender experiment • Radioactive phage & bacteria in blender – 35 S phage • radioactive proteins stayed in supernatant • therefore viral protein did NOT enter bacteria – 32 P phage • radioactive DNA stayed in pellet • therefore viral DNA did enter bacteria – Confirmed DNA is “transforming factor” Taaa-Daaa!

Hershey & Chase Martha Chase 1952 | 1969 Alfred Hershey

Chargaff • DNA composition: “Chargaff’s rules” – varies from species to species – all 4 bases not in equal quantity – bases present in characteristic ratio • humans: A = 30. 9% T = 29. 4% G = 19. 9% C = 19. 8% That’s interesting! What do you notice? Rules A = T C = G 1947

Structure of 1953 | 1962 DNA • Watson & Crick – developed double helix model of DNA • other leading scientists working on question: – Rosalind Franklin – Maurice Wilkins – Linus Pauling Franklin Wilkins Pauling

1953 article in Nature Watson and Crick Watson Crick

Rosalind Franklin (1920 -1958)
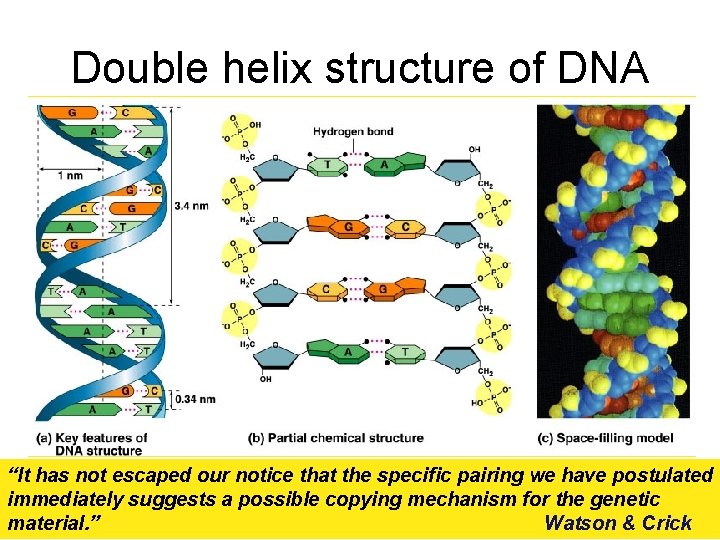
Double helix structure of DNA “It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material. ” Watson & Crick
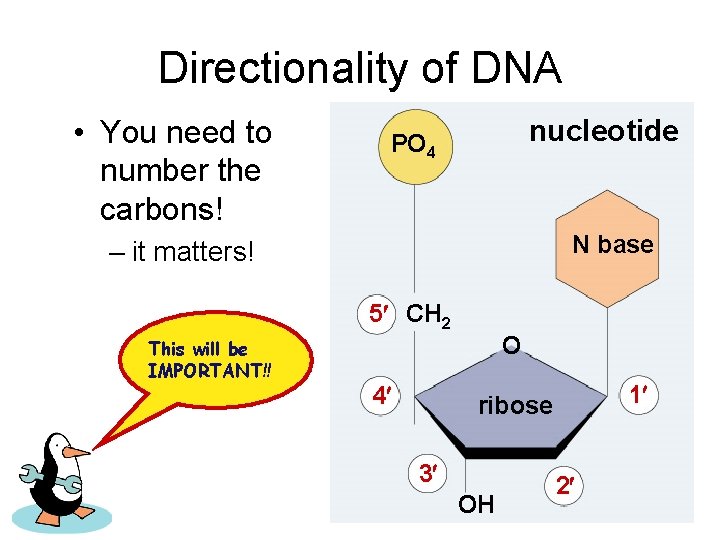
Directionality of DNA • You need to number the carbons! nucleotide PO 4 N base – it matters! 5 CH 2 This will be IMPORTANT!! 4 O 1 ribose 3 OH 2
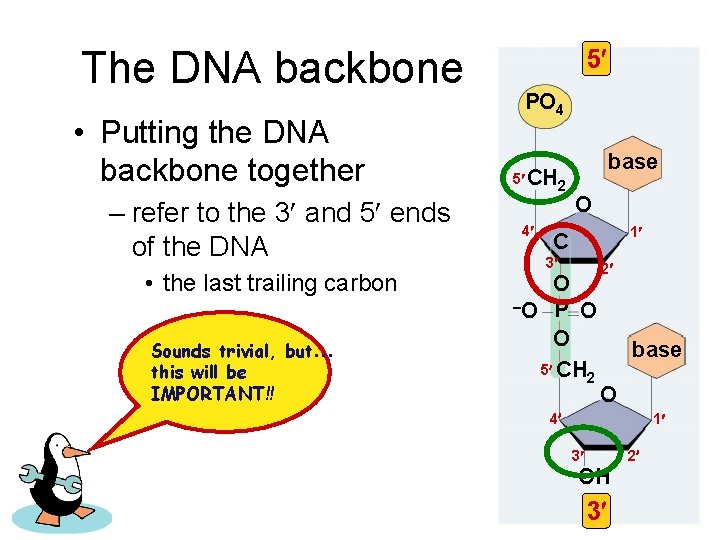
The DNA backbone • Putting the DNA backbone together – refer to the 3 and 5 ends of the DNA • the last trailing carbon Sounds trivial, but… this will be IMPORTANT!! 5 PO 4 5 CH 2 4 base O 1 C 3 O –O P O O 5 CH 2 2 base O 4 1 2 3 OH 3
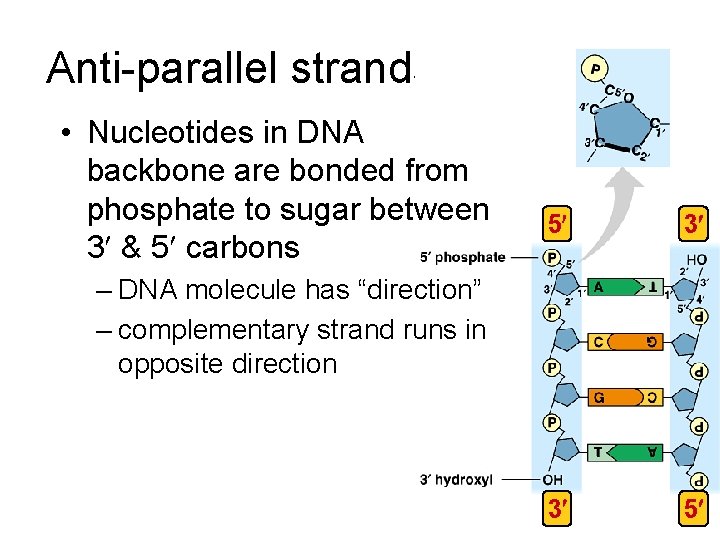
Anti-parallel strands • Nucleotides in DNA backbone are bonded from phosphate to sugar between 3 & 5 carbons 5 3 3 5 – DNA molecule has “direction” – complementary strand runs in opposite direction
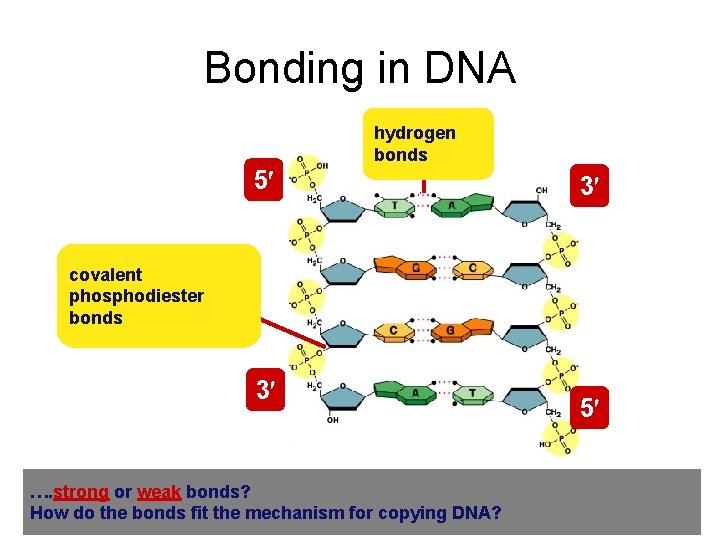
Bonding in DNA hydrogen bonds 5 3 covalent phosphodiester bonds 3 …. strong or weak bonds? How do the bonds fit the mechanism for copying DNA? 5
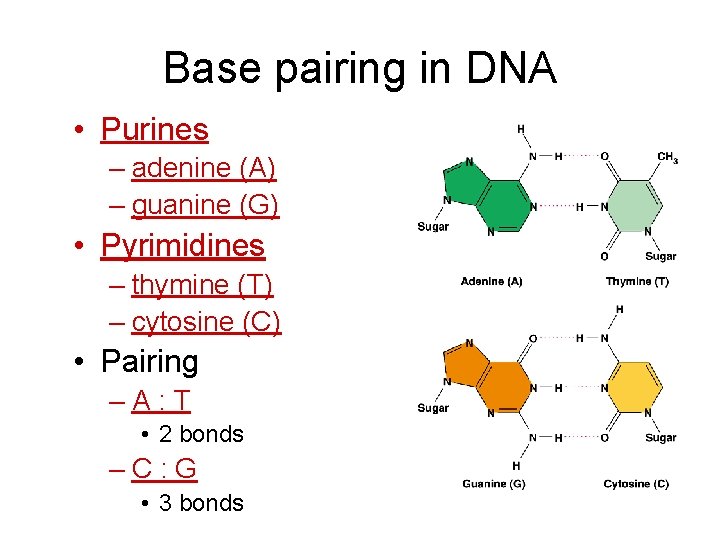
Base pairing in DNA • Purines – adenine (A) – guanine (G) • Pyrimidines – thymine (T) – cytosine (C) • Pairing –A: T • 2 bonds –C: G • 3 bonds
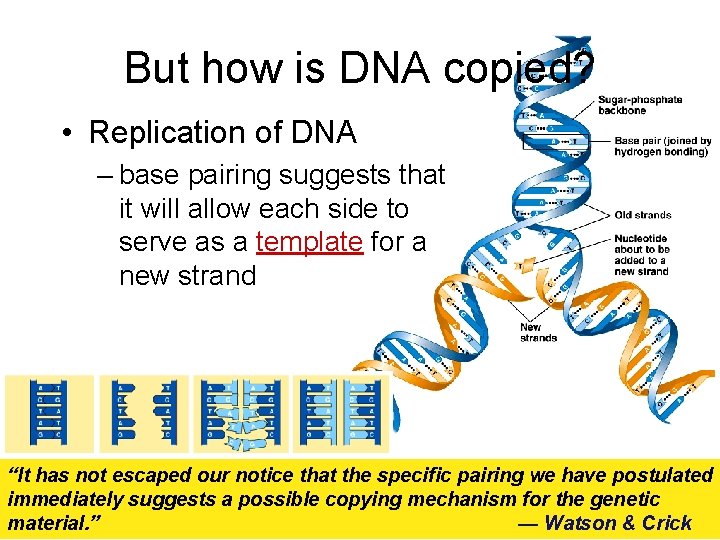
But how is DNA copied? • Replication of DNA – base pairing suggests that it will allow each side to serve as a template for a new strand “It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material. ” — Watson & Crick
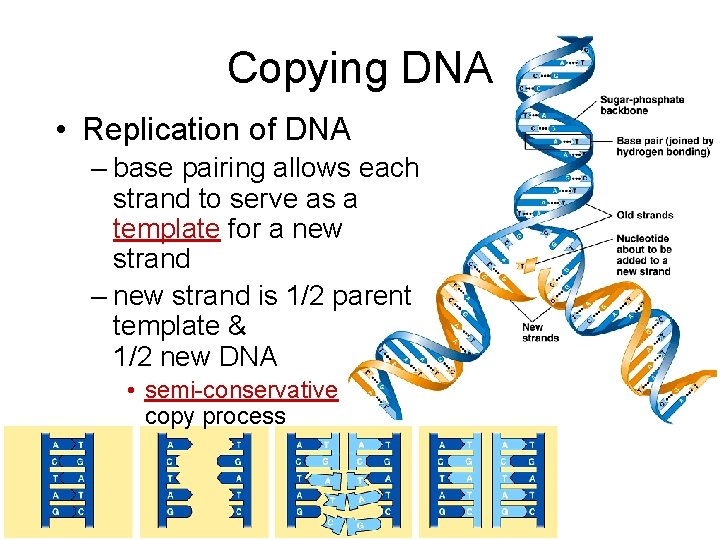
Copying DNA • Replication of DNA – base pairing allows each strand to serve as a template for a new strand – new strand is 1/2 parent template & 1/2 new DNA • semi-conservative copy process
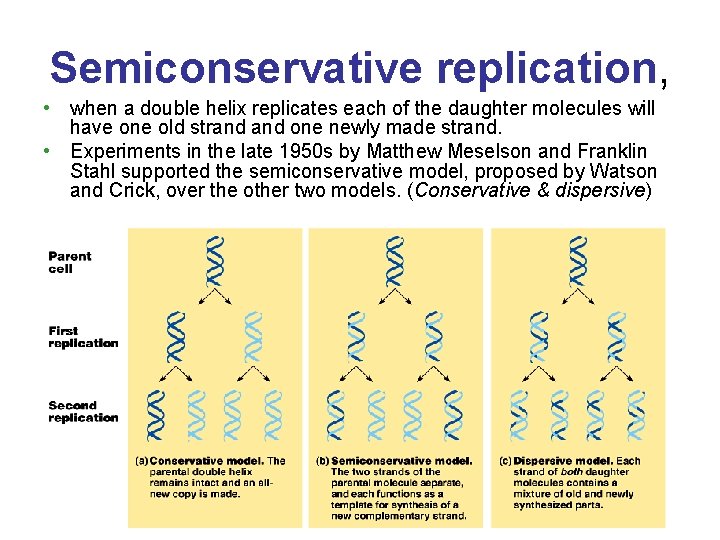
Semiconservative replication, • when a double helix replicates each of the daughter molecules will have one old strand one newly made strand. • Experiments in the late 1950 s by Matthew Meselson and Franklin Stahl supported the semiconservative model, proposed by Watson and Crick, over the other two models. (Conservative & dispersive)
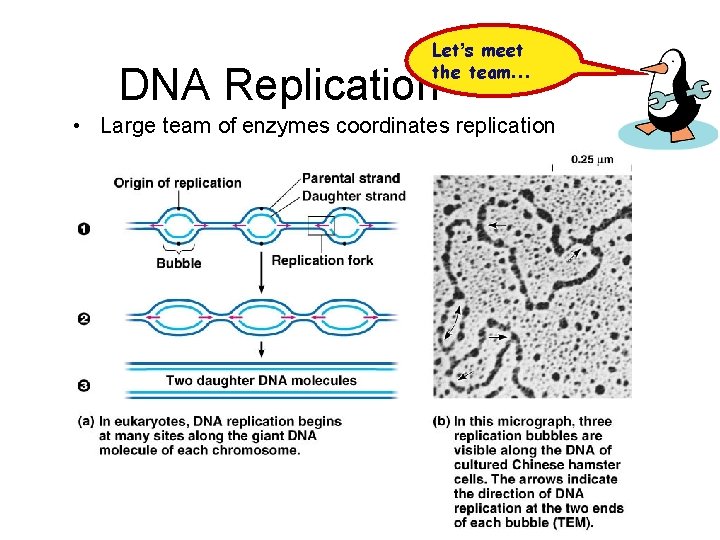
Let’s meet the team… DNA Replication • Large team of enzymes coordinates replication
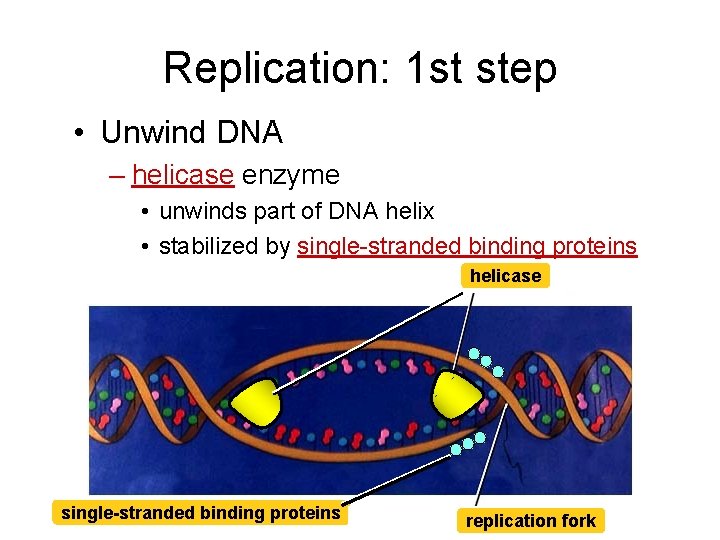
Replication: 1 st step • Unwind DNA – helicase enzyme • unwinds part of DNA helix • stabilized by single-stranded binding proteins helicase single-stranded binding proteins replication fork
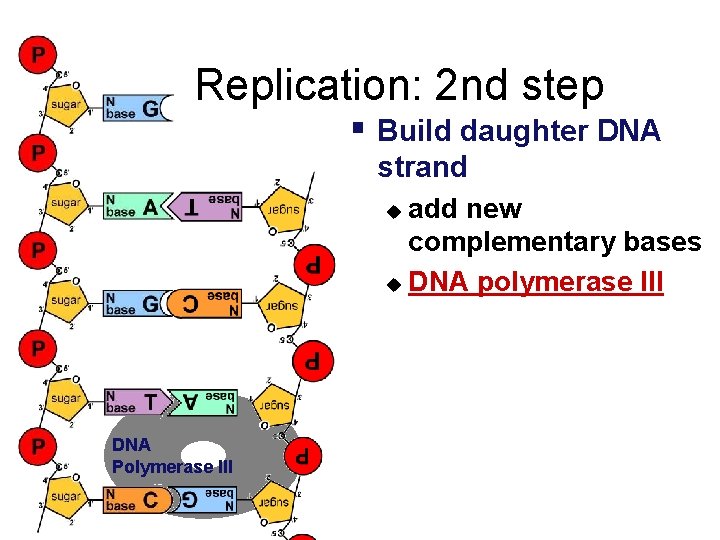
Replication: 2 nd step § Build daughter DNA strand add new complementary bases u DNA polymerase III u DNA Polymerase III
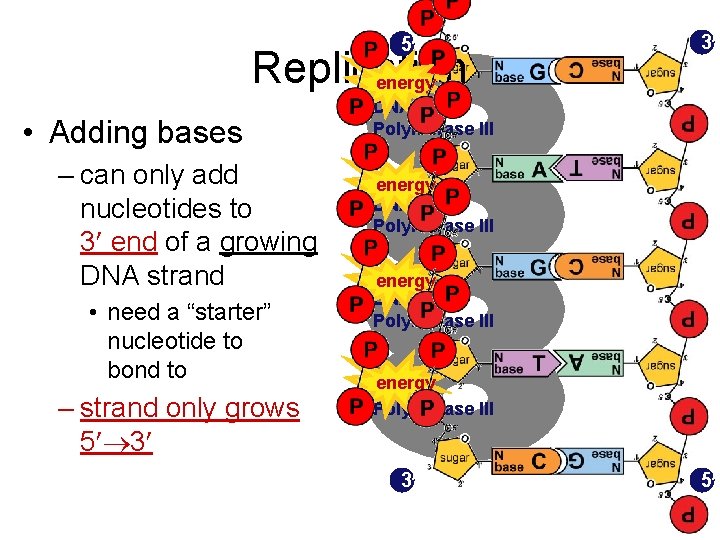
5 Replication energy • Adding bases – can only add nucleotides to 3 end of a growing DNA strand • need a “starter” nucleotide to bond to – strand only grows 5 3 3 DNA Polymerase III energy DNA Polymerase III 3 5
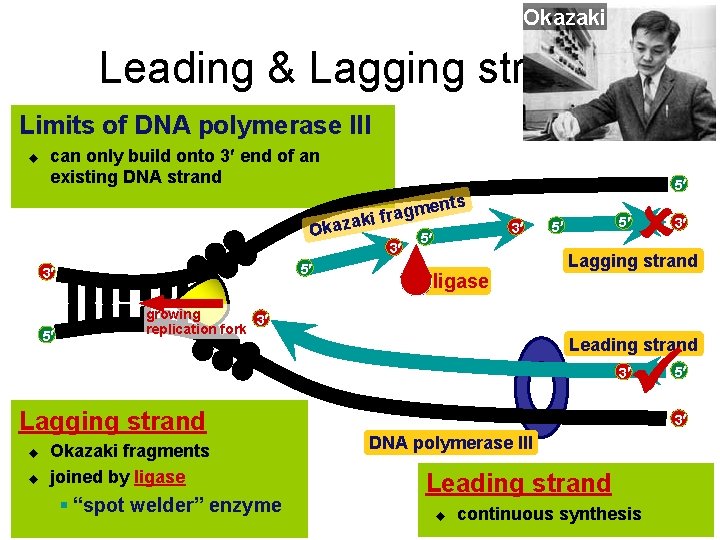
Okazaki Leading & Lagging strands Limits of DNA polymerase III u can only build onto 3 end of an existing DNA strand en m g a r f ki Okaza 3 5 3 5 5 ts 3 5 ligase growing 3 replication fork 5 5 Lagging strand Leading strand 3 Lagging strand u u Okazaki fragments joined by ligase § “spot welder” enzyme 3 5 3 DNA polymerase III Leading strand u continuous synthesis
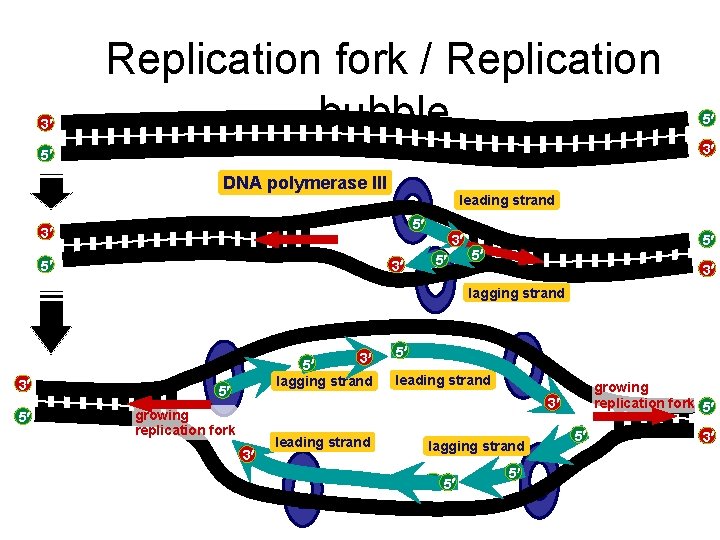
3 Replication fork / Replication bubble 5 3 5 DNA polymerase III leading strand 5 3 3 5 5 5 3 lagging strand 3 5 lagging strand 5 5 leading strand growing replication fork 5 3 growing replication fork 3 leading strand lagging strand 5 5 5 5 3
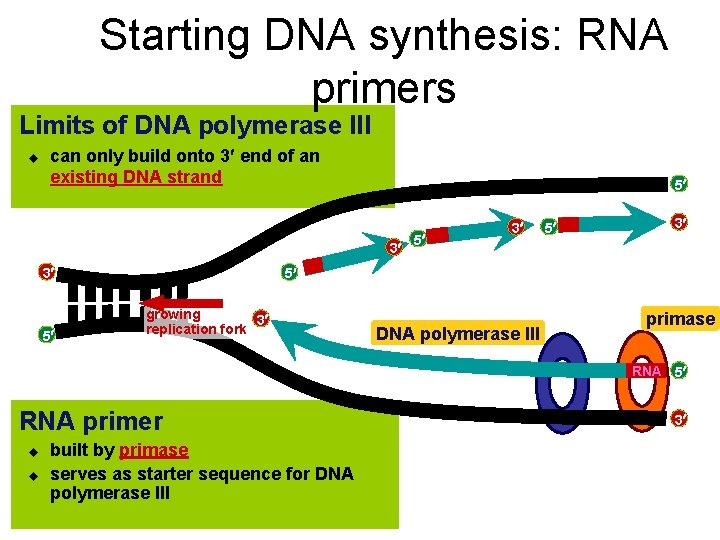
Starting DNA synthesis: RNA primers Limits of DNA polymerase III u can only build onto 3 end of an existing DNA strand 5 3 3 5 5 3 5 growing 3 replication fork DNA polymerase III primase RNA 5 RNA primer u u built by primase serves as starter sequence for DNA polymerase III 3
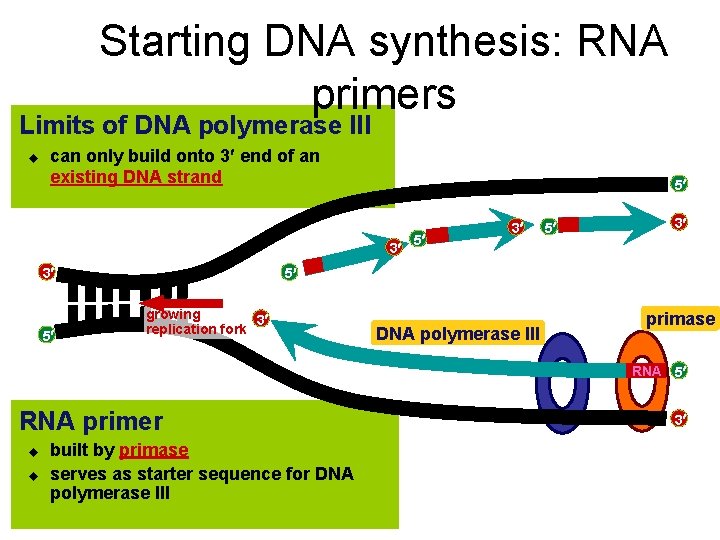
Starting DNA synthesis: RNA primers Limits of DNA polymerase III u can only build onto 3 end of an existing DNA strand 5 3 3 5 5 3 5 growing 3 replication fork DNA polymerase III primase RNA 5 RNA primer u u built by primase serves as starter sequence for DNA polymerase III 3
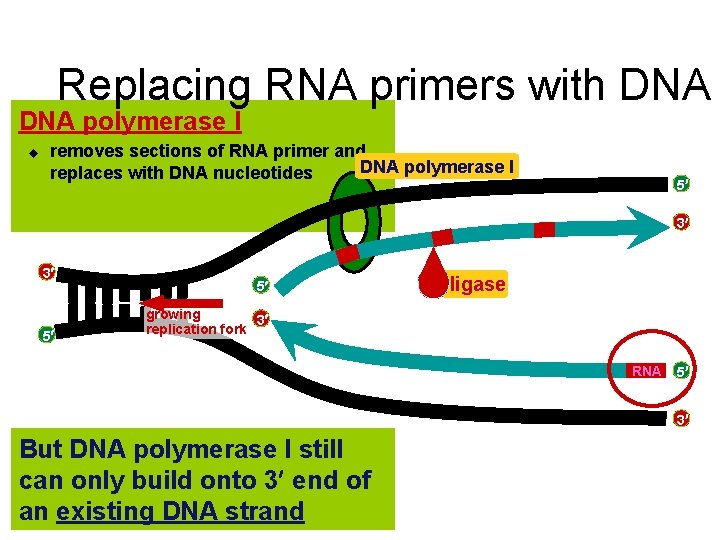
Replacing RNA primers with DNA polymerase I u removes sections of RNA primer and DNA polymerase I replaces with DNA nucleotides 5 3 3 5 5 ligase growing 3 replication fork RNA 5 3 But DNA polymerase I still can only build onto 3 end of an existing DNA strand
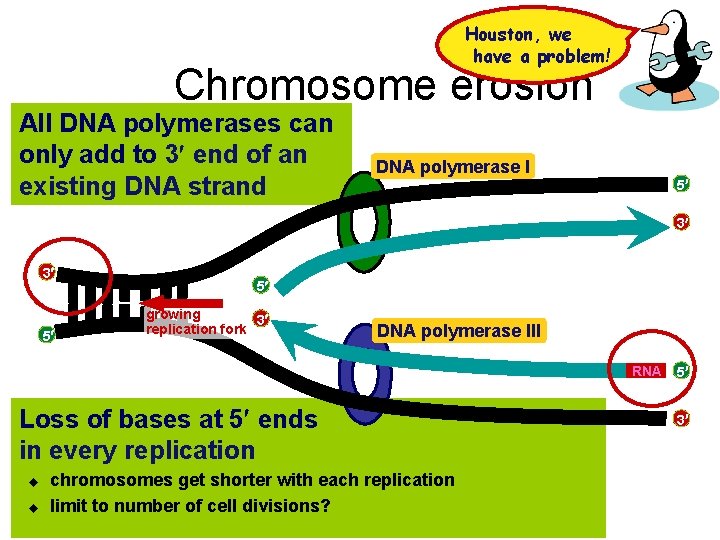
Houston, we have a problem! Chromosome erosion All DNA polymerases can only add to 3 end of an existing DNA strand DNA polymerase I 5 3 3 5 5 growing 3 replication fork DNA polymerase III RNA Loss of bases at 5 ends in every replication u u chromosomes get shorter with each replication limit to number of cell divisions? 5 3
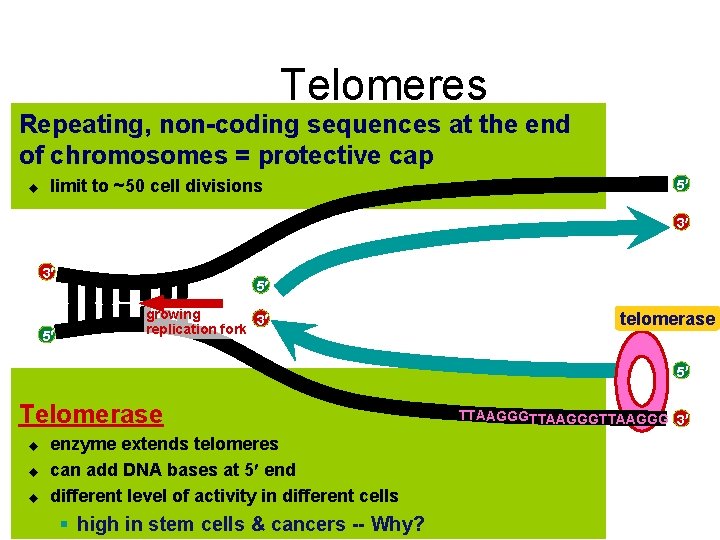
Telomeres Repeating, non-coding sequences at the end of chromosomes = protective cap u limit to ~50 cell divisions 5 3 3 5 5 growing 3 replication fork telomerase 5 Telomerase u u u enzyme extends telomeres can add DNA bases at 5 end different level of activity in different cells § high in stem cells & cancers -- Why? TTAAGGGTTAAGGG 3
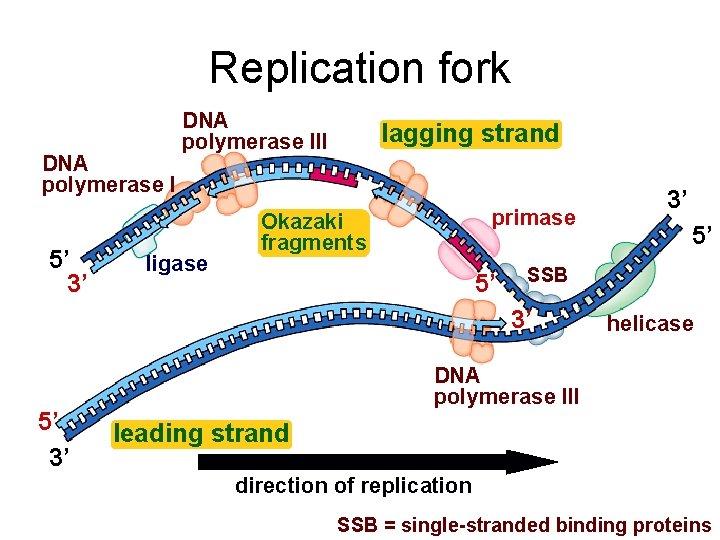
Replication fork DNA polymerase I 5’ 3’ DNA polymerase III ligase lagging strand primase Okazaki fragments 5’ 5’ SSB 3’ 5’ 3’ 3’ helicase DNA polymerase III leading strand direction of replication SSB = single-stranded binding proteins
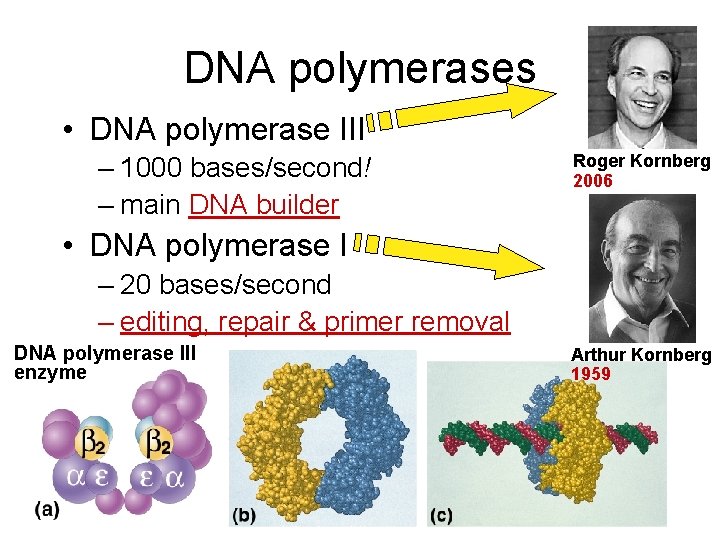
DNA polymerases • DNA polymerase III – 1000 bases/second! – main DNA builder Roger Kornberg 2006 • DNA polymerase I – 20 bases/second – editing, repair & primer removal DNA polymerase III enzyme Arthur Kornberg 1959
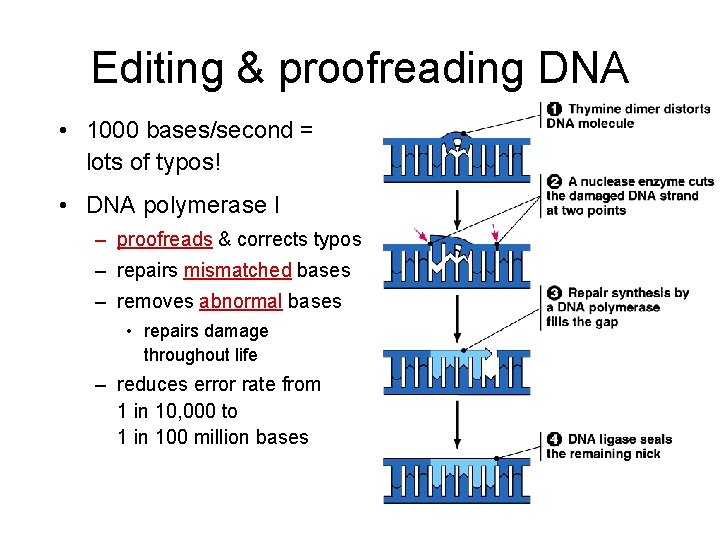
Editing & proofreading DNA • 1000 bases/second = lots of typos! • DNA polymerase I – proofreads & corrects typos – repairs mismatched bases – removes abnormal bases • repairs damage throughout life – reduces error rate from 1 in 10, 000 to 1 in 100 million bases

Fast & accurate! • It takes E. coli <1 hour to copy 5 million base pairs in its single chromosome – divide to form 2 identical daughter cells • Human cell copies its 6 billion bases & divide into daughter cells in only few hours – remarkably accurate – only ~1 error per 100 million bases – ~30 errors per cell cycle
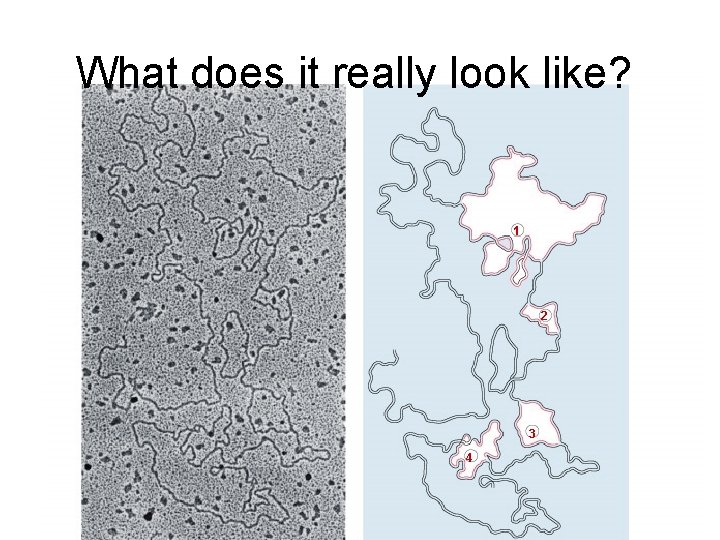
What does it really look like? 1 2 3 4
Any Questions? ? 2007 -2008
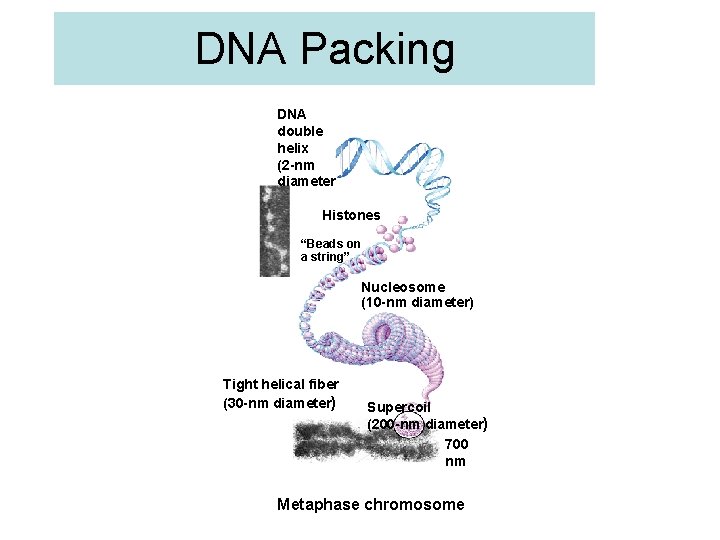
DNA Packing DNA double helix (2 -nm diameter Histones “Beads on a string” Nucleosome (10 -nm diameter) Tight helical fiber (30 -nm diameter) Supercoil (200 -nm diameter) 700 nm Metaphase chromosome
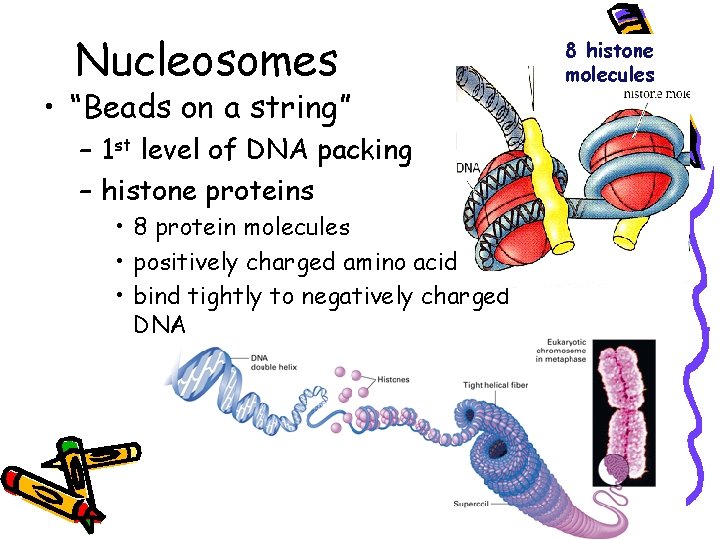
Nucleosomes • “Beads on a string” – 1 st level of DNA packing – histone proteins • 8 protein molecules • positively charged amino acids • bind tightly to negatively charged DNA 8 histone molecules
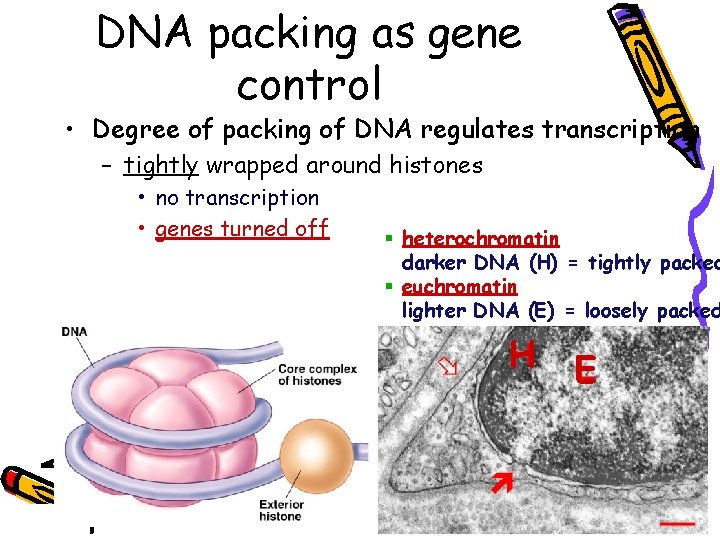
DNA packing as gene control • Degree of packing of DNA regulates transcription – tightly wrapped around histones • no transcription • genes turned off § heterochromatin darker DNA (H) = tightly packed § euchromatin lighter DNA (E) = loosely packed H E
- Slides: 45