Chapter 16 The Molecular Basis of Inheritance Power

Chapter 16 The Molecular Basis of Inheritance Power. Point® Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from Joan Sharp Copyright © 2008 Pearson Education, Inc. , publishing as Pearson Benjamin Cummings

The Search for the Genetic Material • Early in the 20 th century, the identification of the molecules of inheritance loomed as a major challenge to biologists – When T. H. Morgan’s group showed that genes are located on chromosomes, the two components of chromosomes—DNA and protein—became candidates for the genetic material – The key factor in determining the genetic material was choosing appropriate experimental organisms – The role of DNA in heredity was first discovered by studying bacteria and the viruses that infect them Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

Evidence That DNA Can Transform Bacteria • The discovery of the genetic role of DNA began with research by Frederick Griffith in 1928 • Griffith worked with two strains of a bacterium, one pathogenic and one harmless • When he mixed heat-killed remains of the pathogenic strain with living cells of the harmless strain, some living cells became pathogenic • He called this phenomenon transformation, now defined as a change in genotype and phenotype due to assimilation of foreign DNA Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
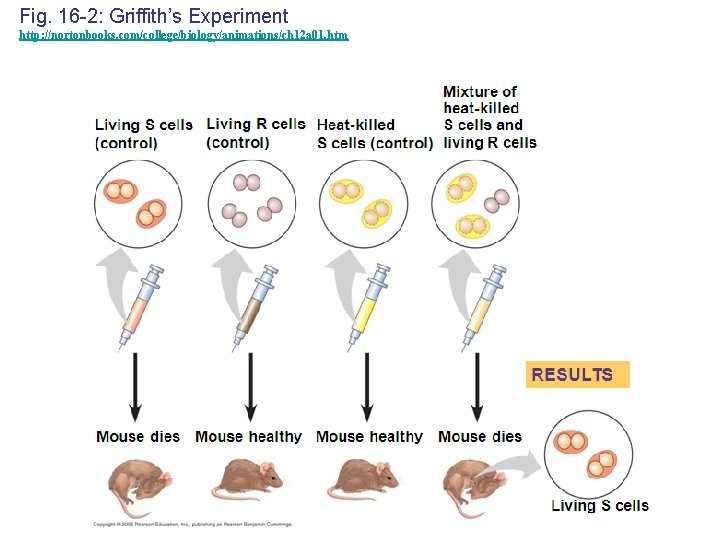
Fig. 16 -2: Griffith’s Experiment http: //nortonbooks. com/college/biology/animations/ch 12 a 01. htm

The Search for the Genetic Material • In 1944, Oswald Avery, Maclyn Mc. Carty, and Colin Mac. Leod announced that the transforming substance was DNA • Their conclusion was based on experimental evidence that only DNA worked in transforming harmless bacteria into pathogenic bacteria • Many biologists remained skeptical, mainly because little was known about DNA Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
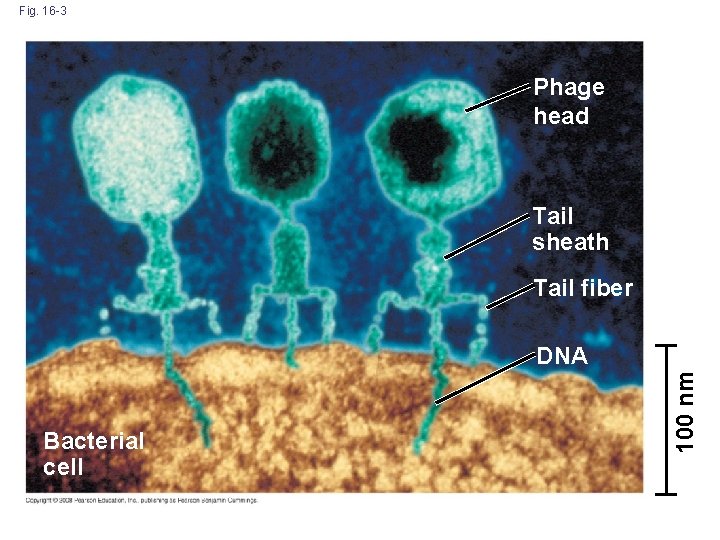
Fig. 16 -3 Phage head Tail sheath Tail fiber Bacterial cell 100 nm DNA

Evidence That Viral DNA Can Program Cells • In 1952, Alfred Hershey and Martha Chase performed experiments showing that DNA is the genetic material of a phage known as T 2 • To determine the source of genetic material in the phage, they designed an experiment showing that only one of the two components of T 2 (DNA or protein) enters an E. coli cell during infection • They concluded that the injected DNA of the phage provides the genetic information Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
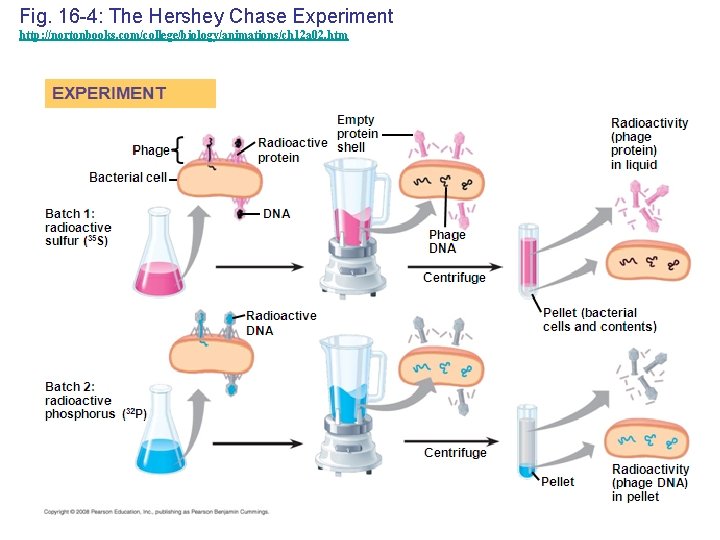
Fig. 16 -4: The Hershey Chase Experiment http: //nortonbooks. com/college/biology/animations/ch 12 a 02. htm

Additional Evidence That DNA Is the Genetic Material • It was known that DNA is a polymer of nucleotides, each consisting of a nitrogenous base, a sugar, and a phosphate group • In 1950, Erwin Chargaff reported that DNA composition varies from one species to the next • This evidence of diversity made DNA a more credible candidate for the genetic material • Chargaff’s rules state that in any species there is an equal number of A and T bases, and an equal number of G and C bases Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
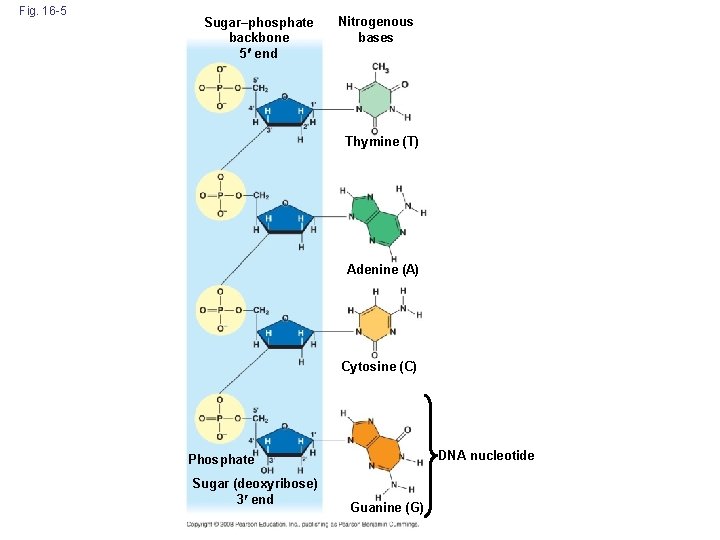
Fig. 16 -5 Sugar–phosphate backbone 5 end Nitrogenous bases Thymine (T) Adenine (A) Cytosine (C) DNA nucleotide Phosphate Sugar (deoxyribose) 3 end Guanine (G)

Practice Question • Which of the following is LEAST related to the others? – Transformation – Phage – DNA – Griffeth – Avery • Answer: phage

Practice Question • What was the contribution of the following scientists regarding the discovery of our present knowledge of the nature of genes and/or the shape of the DNA? – Griffith – Hershey & Chase – Avery, Mac. Leod, and Mc. Carty – Chargaff – Meselson & Stahl

Building a Structural Model of DNA • After most biologists became convinced that DNA was the genetic material, the challenge was to determine how its structure accounts for its role • Maurice Wilkins and Rosalind Franklin were using a technique called X-ray crystallography to study molecular structure • Franklin produced a picture of the DNA molecule using this technique Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

Fig. 16 -6 (a) Rosalind Franklin (b) Franklin’s X-ray diffraction photograph of DNA
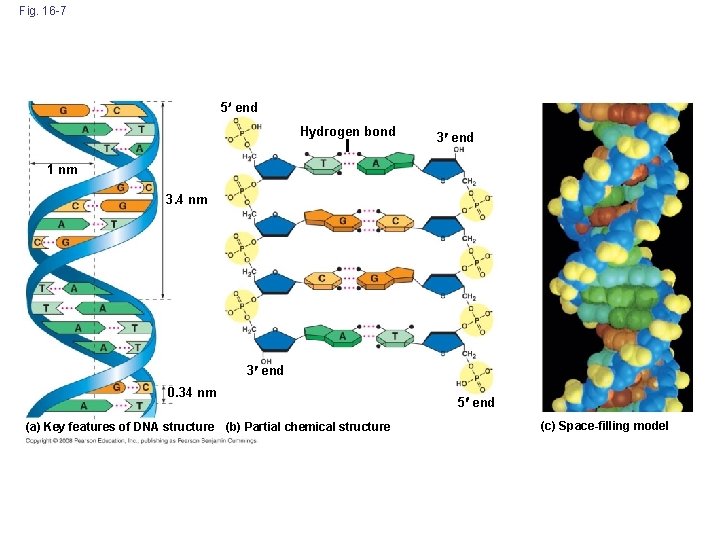
Fig. 16 -7 5 end Hydrogen bond 3 end 1 nm 3. 4 nm 3 end 0. 34 nm (a) Key features of DNA structure (b) Partial chemical structure 5 end (c) Space-filling model

Practice Question • The strands that make up DNA are antiparallel. What does this mean? • Answer: the 5’ to 3’ direction of one strand runs counter to the 5’ to 3’ direction of the other strand

Life’s Operating Instructions • Francis Watson and James Crick built models of a double helix to conform to the X-rays and chemistry of DNA • Franklin had concluded that there were two antiparallel sugar-phosphate backbones, with the nitrogenous bases paired in the molecule’s interior • In 1953, James Watson and Francis Crick introduced an elegant double -helical model for the structure of deoxyribonucleic acid, or DNA • DNA, the substance of inheritance, is the most celebrated molecule of our time • Hereditary information is encoded in DNA and reproduced in all cells of the body • This DNA program directs the development of biochemical, anatomical, physiological, and (to some extent) behavioral traits Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

Fig. 16 -1
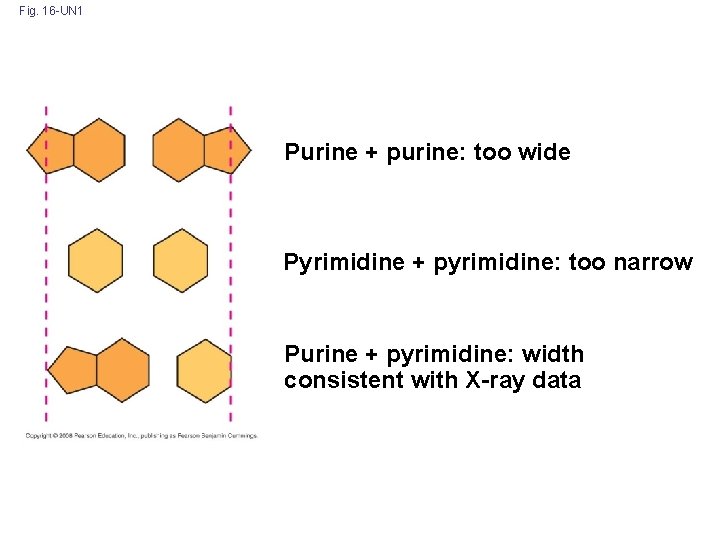
Fig. 16 -UN 1 Purine + purine: too wide Pyrimidine + pyrimidine: too narrow Purine + pyrimidine: width consistent with X-ray data
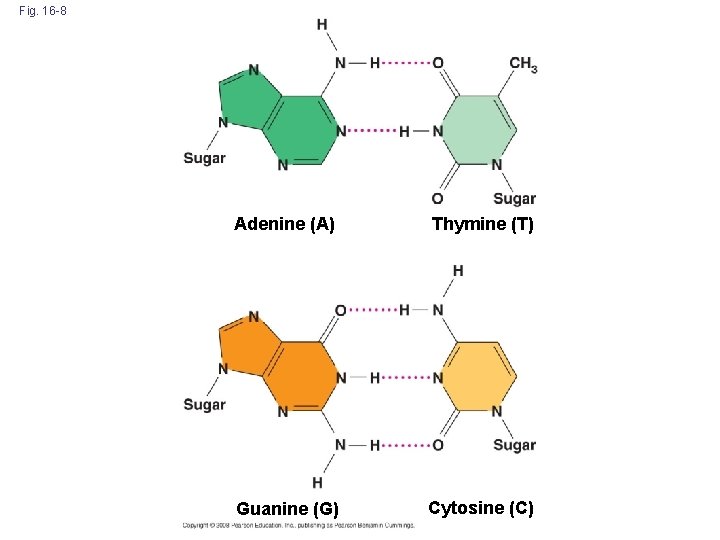
Fig. 16 -8 Adenine (A) Thymine (T) Guanine (G) Cytosine (C)
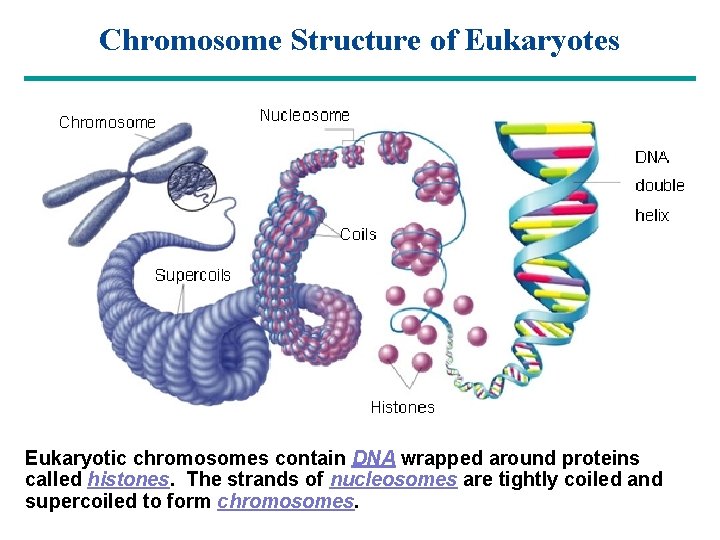
Chromosome Structure of Eukaryotes Eukaryotic chromosomes contain DNA wrapped around proteins called histones. The strands of nucleosomes are tightly coiled and to form chromosomes. Gosupercoiled to Section:

Practice Question • All of the following elements are present in DNA except: – Oxygen – Nitrogen – Carbon – Sulfur – Phosphorus • Answer: sulfur

Practice Question • Cytosine makes up 38% of the nucleotides in a sample of DNA from an organism. What percent of the nucleotides in this sample will be thymine? – 12 % – 24 % – 31 % – 38 % – Cannot be determined with information provided • Answer: 12 %

Practice Question • In an analysis of the nucleotide composition of DNA, which of the following is TRUE? – A=C – A = G and C = T – A+C=G+T – A+T=G+C – Both B and C are true • Answer: A + C = G + T

Practice Question • All of the following were determined directly from X-ray diffraction photographs of crystallized DNA except: – The diameter of the double helix – The helical shape of DNA – The sequence of nucleotides – The linear distance required for one full turn of the double helix – The width of the double helix • Answer: the sequence of nucleotides

The Basic Principle: Base Pairing to a Template Strand • The relationship between structure and function is manifest in the double helix • Watson and Crick noted that the specific base pairing suggested a possible copying mechanism for genetic material • Since the two strands of DNA are complementary, each strand acts as a template for building a new strand in replication • In DNA replication, the parent molecule unwinds, and two new daughter strands are built following the rules of complimentary base pairing Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
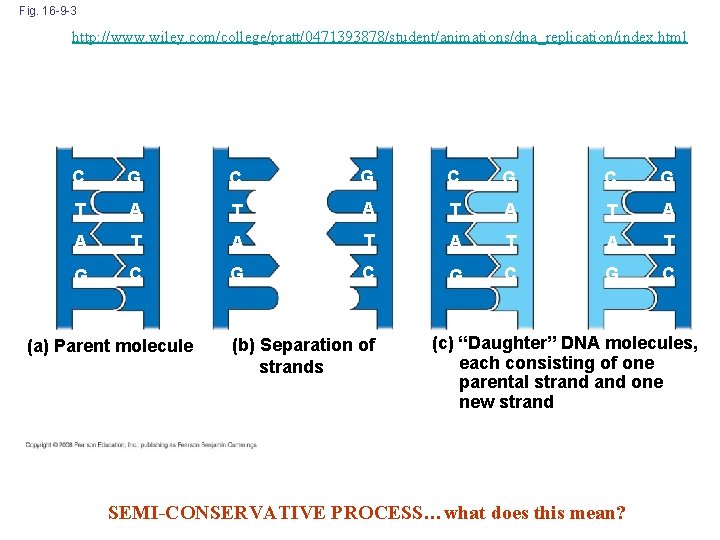
Fig. 16 -9 -3 http: //www. wiley. com/college/pratt/0471393878/student/animations/dna_replication/index. html A T A T C G C G T A T A T G C G C (a) Parent molecule (b) Separation of strands (c) “Daughter” DNA molecules, each consisting of one parental strand one new strand SEMI-CONSERVATIVE PROCESS…what does this mean?
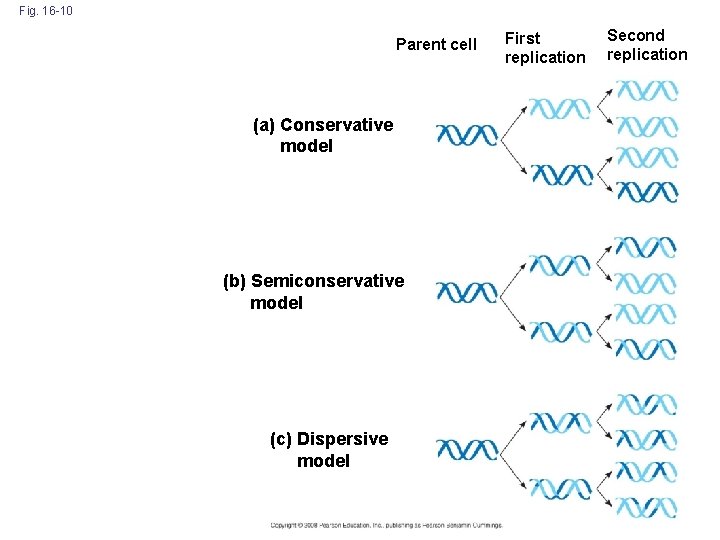
Fig. 16 -10 Parent cell (a) Conservative model (b) Semiconservative model (c) Dispersive model First replication Second replication

Meselson & Stahl: Semiconservative Model of Replication • Experiments by Matthew Meselson and Franklin Stahl supported the semiconservative model • They labeled the nucleotides of the old strands with a heavy isotope of nitrogen, while any new nucleotides were labeled with a lighter isotope • The first replication produced a band of hybrid DNA, eliminating the conservative model • A second replication produced both light and hybrid DNA, eliminating the dispersive model and supporting the semiconservative model Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
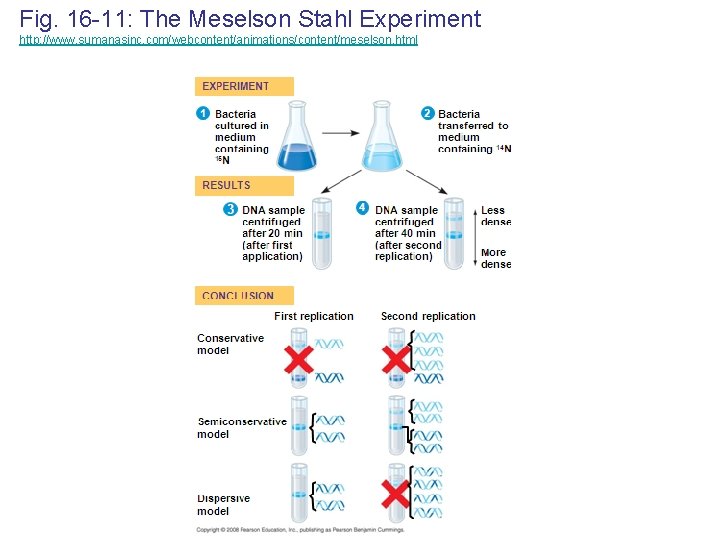
Fig. 16 -11: The Meselson Stahl Experiment http: //www. sumanasinc. com/webcontent/animations/content/meselson. html

DNA Replication: A Closer Look http: //bcs. whfreeman. com/thelifewire/content/chp 11/1102002. html • Replication begins at special sites called origins of replication, where the two DNA strands are separated, opening up a replication “bubble” • A eukaryotic chromosome may have hundreds or even thousands of origins of replication • Replication proceeds in both directions from each origin, until the entire molecule is copied Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
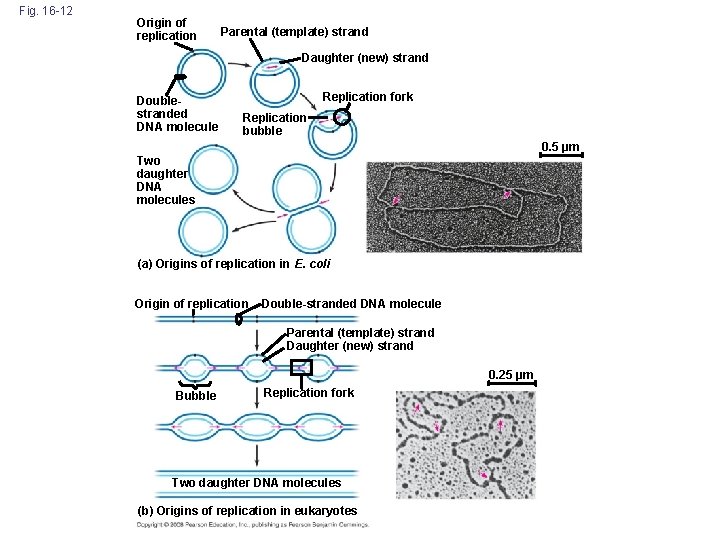
Fig. 16 -12 Origin of replication Parental (template) strand Daughter (new) strand Doublestranded DNA molecule Replication fork Replication bubble 0. 5 µm Two daughter DNA molecules (a) Origins of replication in E. coli Origin of replication Double-stranded DNA molecule Parental (template) strand Daughter (new) strand 0. 25 µm Bubble Replication fork Two daughter DNA molecules (b) Origins of replication in eukaryotes
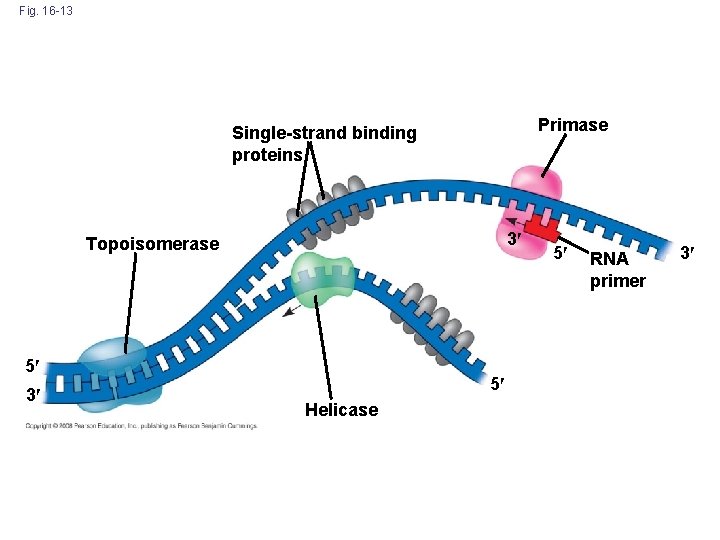
Fig. 16 -13 Primase Single-strand binding proteins 3 Topoisomerase 5 3 5 Helicase 5 RNA primer 3
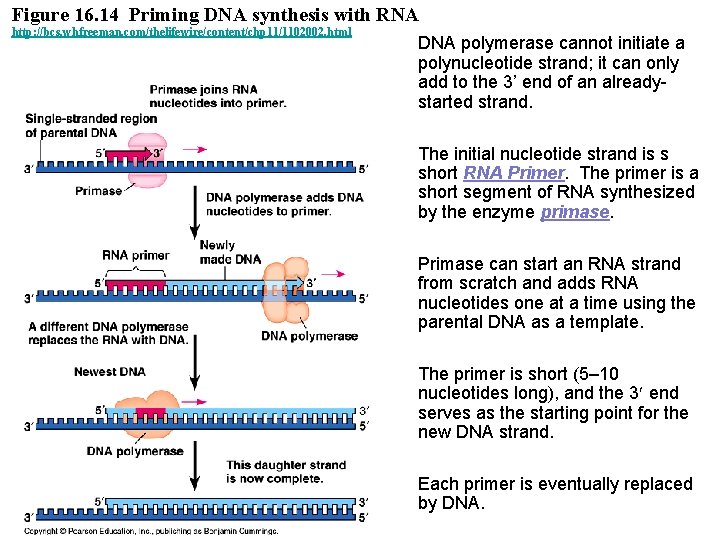
Figure 16. 14 Priming DNA synthesis with RNA http: //bcs. whfreeman. com/thelifewire/content/chp 11/1102002. html DNA polymerase cannot initiate a polynucleotide strand; it can only add to the 3’ end of an alreadystarted strand. The initial nucleotide strand is s short RNA Primer. The primer is a short segment of RNA synthesized by the enzyme primase. Primase can start an RNA strand from scratch and adds RNA nucleotides one at a time using the parental DNA as a template. The primer is short (5– 10 nucleotides long), and the 3 end serves as the starting point for the new DNA strand. Each primer is eventually replaced by DNA.

Synthesizing a New DNA Strand • Enzymes called DNA polymerases catalyze the elongation of new DNA at a replication fork • Most DNA polymerases require a primer and a DNA template strand • The rate of elongation is about 500 nucleotides per second in bacteria and 50 per second in human cells • The elongation of a DNA strand is a process that requires energy Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
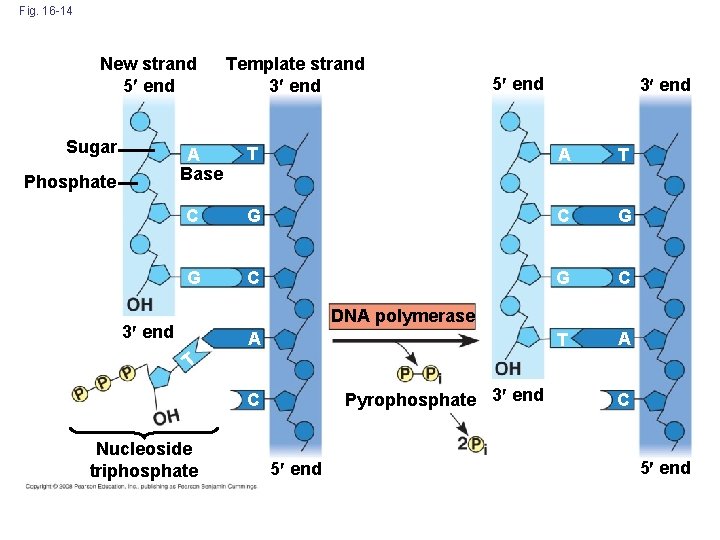
Fig. 16 -14 New strand 5 end Sugar 5 end 3 end T A T C G G C T A A Base Phosphate Template strand 3 end DNA polymerase 3 end A T Pyrophosphate 3 end C Nucleoside triphosphate 5 end C 5 end
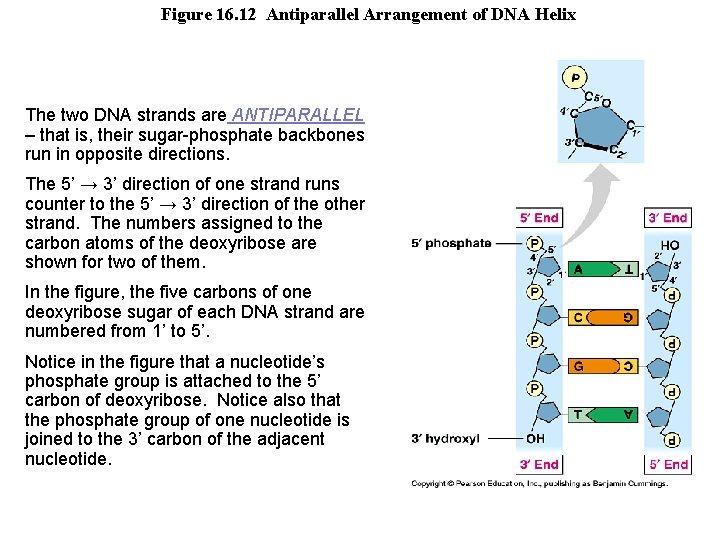
Figure 16. 12 Antiparallel Arrangement of DNA Helix The two DNA strands are ANTIPARALLEL – that is, their sugar-phosphate backbones run in opposite directions. The 5’ → 3’ direction of one strand runs counter to the 5’ → 3’ direction of the other strand. The numbers assigned to the carbon atoms of the deoxyribose are shown for two of them. In the figure, the five carbons of one deoxyribose sugar of each DNA strand are numbered from 1’ to 5’. Notice in the figure that a nucleotide’s phosphate group is attached to the 5’ carbon of deoxyribose. Notice also that the phosphate group of one nucleotide is joined to the 3’ carbon of the adjacent nucleotide.

Antiparallel Elongation – The Leading & Lagging Strand • The antiparallel structure of the double helix (two strands oriented in opposite directions) affects replication • DNA polymerases add nucleotides only to the free 3 end of a growing strand; therefore, a new DNA strand can elongate only in the 5 to 3 direction • Along one template strand of DNA, the DNA polymerase synthesizes a leading strand continuously, moving toward the replication fork • Along the other template strand of DNA, the DNA polymerase synthesize a lagging strand discontinuously in segments called Okazaki fragments, moving away from the replication fork Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
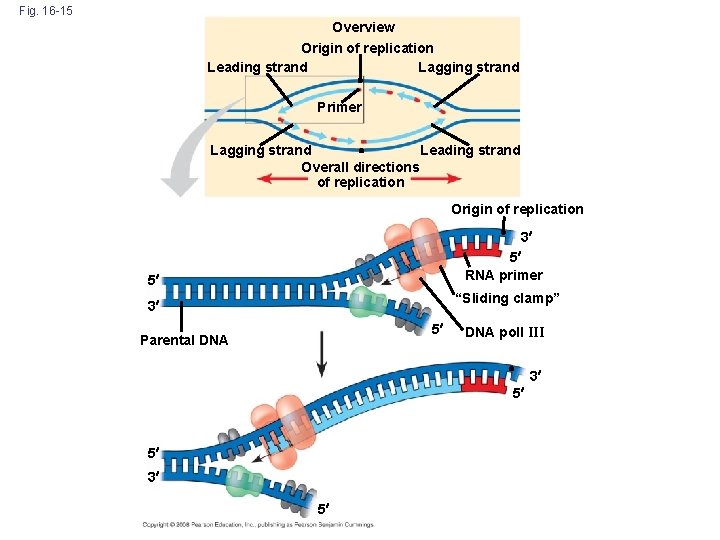
Fig. 16 -15 Overview Origin of replication Leading strand Lagging strand Primer Lagging strand Leading strand Overall directions of replication Origin of replication 3 5 RNA primer 5 “Sliding clamp” 3 5 Parental DNA poll III 3 5 5 3 5
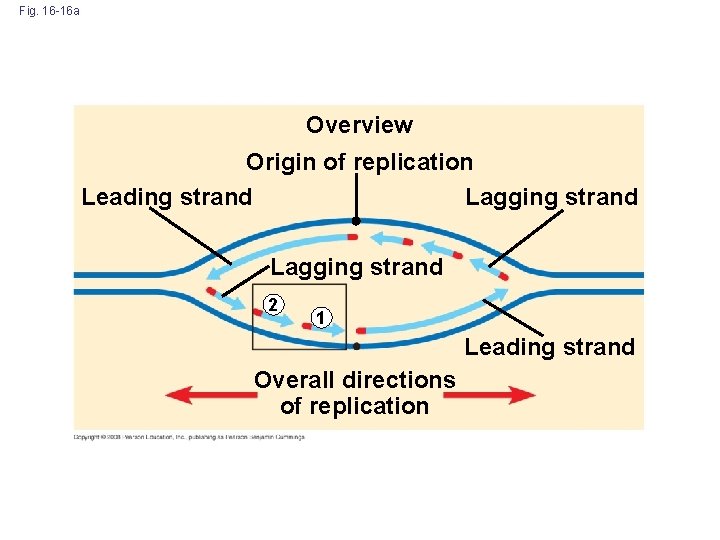
Fig. 16 -16 a Overview Origin of replication Leading strand Lagging strand 2 1 Leading strand Overall directions of replication
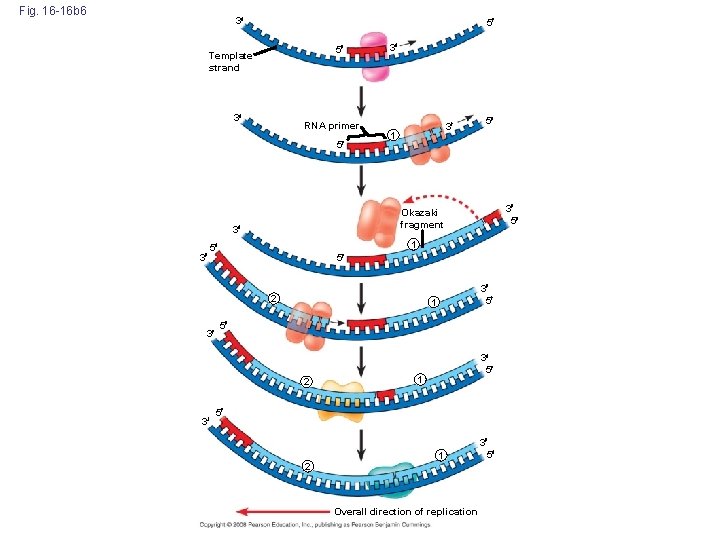
Fig. 16 -16 b 6 3 5 5 Template strand 3 RNA primer 5 3 1 5 2 3 5 1 5 2 3 3 5 1 5 3 5 Okazaki fragment 3 3 3 3 5 1 5 2 1 Overall direction of replication 3 5

Table 16 -1
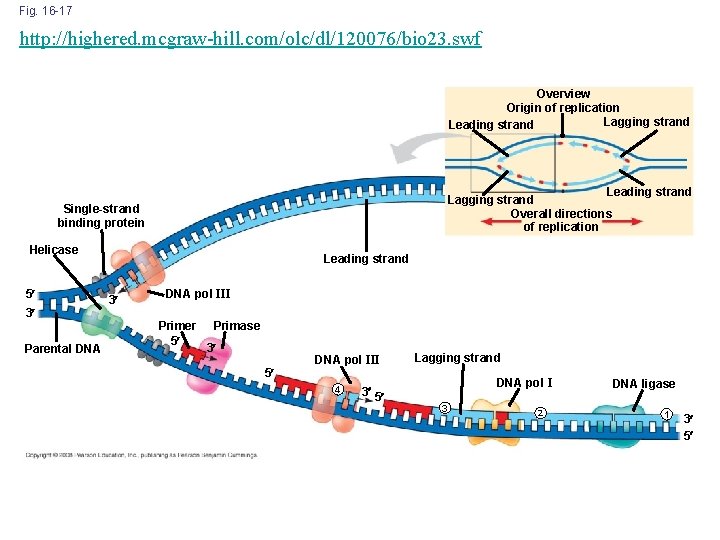
Fig. 16 -17 http: //highered. mcgraw-hill. com/olc/dl/120076/bio 23. swf Overview Origin of replication Lagging strand Leading strand Lagging strand Overall directions of replication Single-strand binding protein Helicase 5 3 Parental DNA Leading strand 3 DNA pol III Primer 5 Primase 3 5 DNA pol III 4 3 5 Lagging strand DNA pol I 3 2 DNA ligase 1 3 5
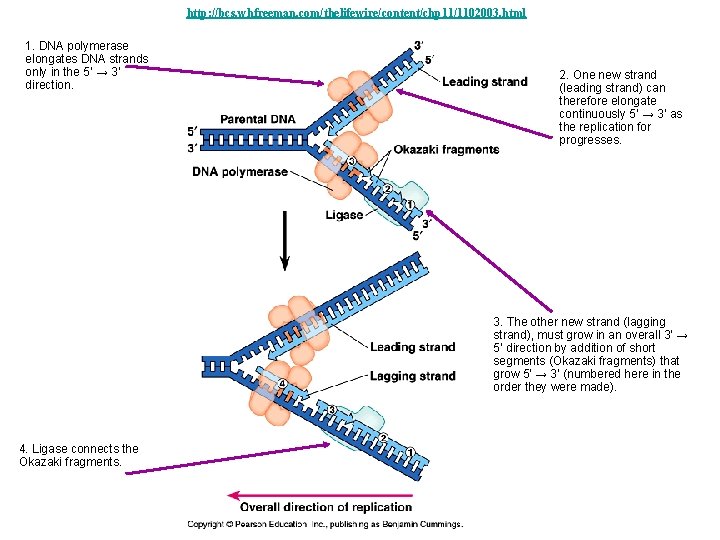
http: //bcs. whfreeman. com/thelifewire/content/chp 11/1102003. html 1. DNA polymerase elongates DNA strands only in the 5’ → 3’ direction. 2. One new strand (leading strand) can therefore elongate continuously 5’ → 3’ as the replication for progresses. 3. The other new strand (lagging strand), must grow in an overall 3’ → 5’ direction by addition of short segments (Okazaki fragments) that grow 5’ → 3’ (numbered here in the order they were made). 4. Ligase connects the Okazaki fragments.

The DNA Replication Complex • The proteins that participate in DNA replication form a large complex, a “DNA replication machine” • The DNA replication machine is probably stationary during the replication process • Recent studies support a model in which DNA polymerase molecules “reel in” parental DNA and “extrude” newly made daughter DNA molecules Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

Practice Question • Which enzymes catalyze the elongation of a DNA strand in the 5’ to 3’ direction? – Primase – DNA ligase – DNA polymerases – Topoisomerase – Helicase • Answer: DNA polymerases

Practice Question • What is the function of the following enzymes in DNA replication? – Helicase – SSB’s – Nuclease – Ligase – DNA polymerase – primase

Practice Question • In DNA, the designations 3’ and 5’ refer to what? • Answer: carbon atoms of deoxyribose to which phosphate groups may bond

Practice Question • Which of the following is LEAST related to the others on the list? – Okazaki fragment – Primer – Telomere – Leading strand – Lagging strand • Answer: telomere
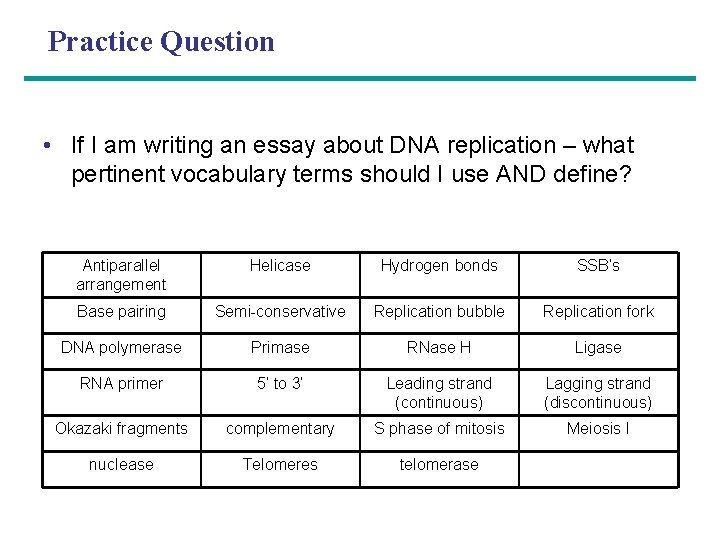
Practice Question • If I am writing an essay about DNA replication – what pertinent vocabulary terms should I use AND define? Antiparallel arrangement Helicase Hydrogen bonds SSB’s Base pairing Semi-conservative Replication bubble Replication fork DNA polymerase Primase RNase H Ligase RNA primer 5’ to 3’ Leading strand (continuous) Lagging strand (discontinuous) Okazaki fragments complementary S phase of mitosis Meiosis I nuclease Telomeres telomerase

Proofreading and Repairing DNA • DNA polymerases proofread newly made DNA, replacing any incorrect nucleotides • In mismatch repair of DNA, repair enzymes correct errors in base pairing • DNA can be damaged by chemicals, radioactive emissions, X-rays, UV light, and certain molecules (in cigarette smoke for example) • In nucleotide excision repair, a nuclease cuts out and replaces damaged stretches of DNA Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
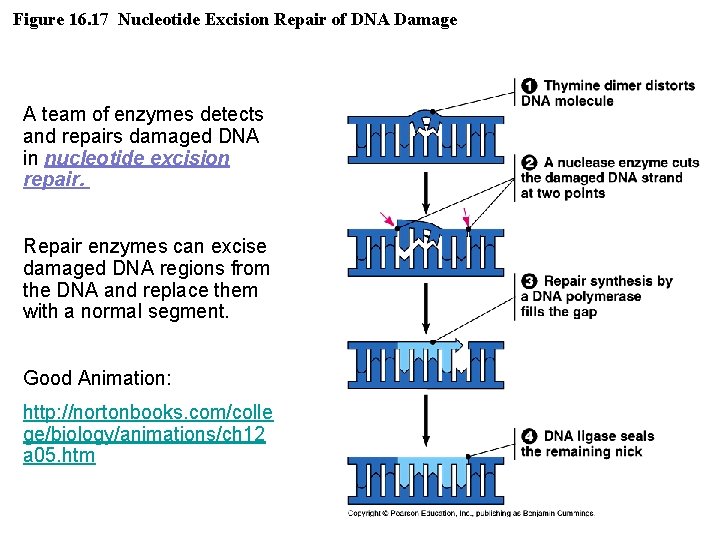
Figure 16. 17 Nucleotide Excision Repair of DNA Damage A team of enzymes detects and repairs damaged DNA in nucleotide excision repair. Repair enzymes can excise damaged DNA regions from the DNA and replace them with a normal segment. Good Animation: http: //nortonbooks. com/colle ge/biology/animations/ch 12 a 05. htm

Replicating the Ends of DNA Molecules • Limitations of DNA polymerase create problems for the linear DNA of eukaryotic chromosomes • The usual replication machinery provides no way to complete the 5 ends, so repeated rounds of replication produce shorter DNA molecules • Eukaryotic chromosomal DNA molecules have at their ends nucleotide sequences called telomeres • Telomeres do not prevent the shortening of DNA molecules, but they do postpone the erosion of genes near the ends of DNA molecules • It has been proposed that the shortening of telomeres is connected to aging Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
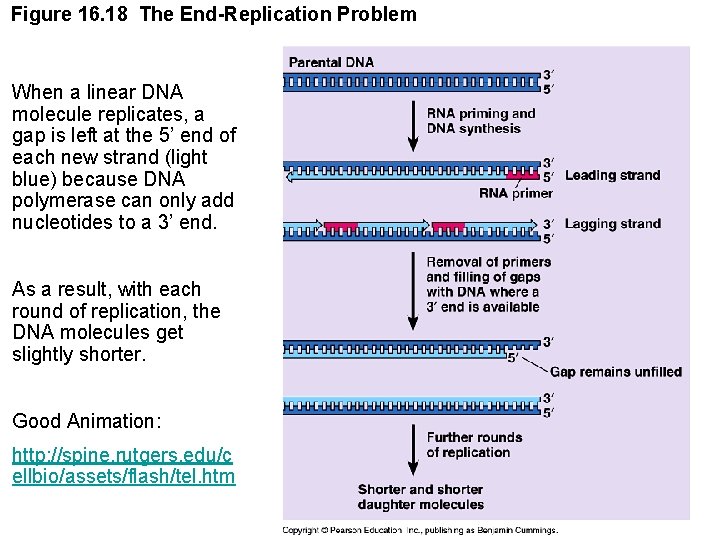
Figure 16. 18 The End-Replication Problem When a linear DNA molecule replicates, a gap is left at the 5’ end of each new strand (light blue) because DNA polymerase can only add nucleotides to a 3’ end. As a result, with each round of replication, the DNA molecules get slightly shorter. Good Animation: http: //spine. rutgers. edu/c ellbio/assets/flash/tel. htm

Telomeres & Telomerase • If chromosomes of germ cells became shorter in every cell cycle, essential genes would eventually be missing from the gametes they produce • An enzyme called telomerase catalyzes the lengthening of telomeres in germ cells • The shortening of telomeres might protect cells from cancerous growth by limiting the number of cell divisions • There is evidence of telomerase activity in cancer cells, which may allow cancer cells to persist Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings
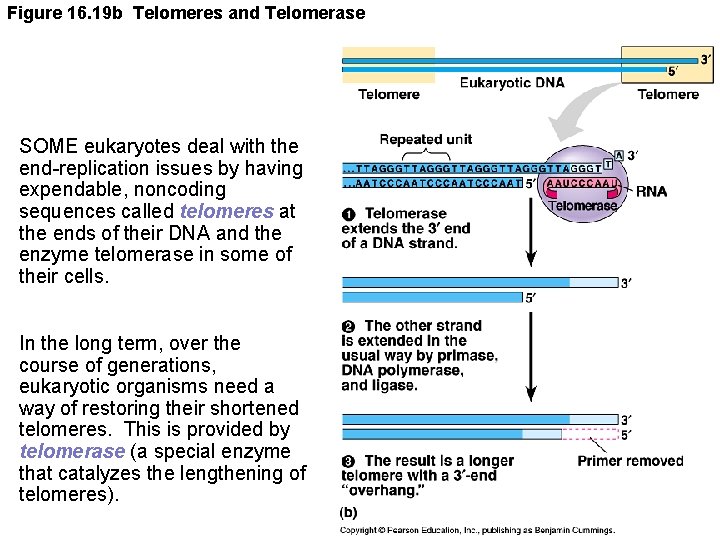
Figure 16. 19 b Telomeres and Telomerase SOME eukaryotes deal with the end-replication issues by having expendable, noncoding sequences called telomeres at the ends of their DNA and the enzyme telomerase in some of their cells. In the long term, over the course of generations, eukaryotic organisms need a way of restoring their shortened telomeres. This is provided by telomerase (a special enzyme that catalyzes the lengthening of telomeres).

Do Telomeres Limit Lifespans? • Telomerase is NOT present in most cells of multicellular organisms like ourselves, and the DNA of dividing somatic cells does tend to be shorter in older individuals and in cultured cells that have divided many times. – Thus, it is possible that telomeres are a limiting factor in the life span of certain tissues and even organisms as a whole. • Telomerase is however present in germ-line cells that give rise to gametes – and here the enzyme produces long telomeres in these cells and hence in the newborn. – Intriguingly, telomerase is also found in somatic cells that are cancerous – these usually have unusually short telomeres, which one would expect for cells that have undergone many rounds of division. – Progressive shortening would eventually lead to self-destruction of cancer unless telomerase became available to stabilize telomere length. – This is exactly what seems to happen in cancer cells. IF this is an important factor – it may well provide a useful target for cancer diagnosis and chemotherapy!

You should now be able to: 1. Describe the contributions of the following people: Griffith; Avery, Mc. Cary, and Mac. Leod; Hershey and Chase; Chargaff; Watson and Crick; Franklin; Meselson and Stahl 2. Describe the structure of DNA 3. Describe the process of DNA replication; include the following terms: antiparallel structure, DNA polymerase, leading strand, lagging strand, Okazaki fragments, DNA ligase, primer, primase, helicase, topoisomerase, single-strand binding proteins 4. Describe the function of telomeres 5. Compare a bacterial chromosome and a eukaryotic chromosome Copyright © 2008 Pearson Education Inc. , publishing as Pearson Benjamin Cummings

The Essay • Scientists seeking to determine which molecule is responsible for the transmission of characteristics from one generation to the next knew that the molecule must – (1) copy itself precisely, – (2) be stable but able to be changed, and – (3) be complex enough to determine the organism's phenotype. • Explain how DNA meets each of the three criteria stated above. • Select one of the criteria stated above and describe experimental evidence used to determine that DNA is the hereditary material.
- Slides: 59