Chapter 16 The Molecular Basis of Inheritance DNA

Chapter 16: The Molecular Basis of Inheritance (DNA) Zooming in on DNA
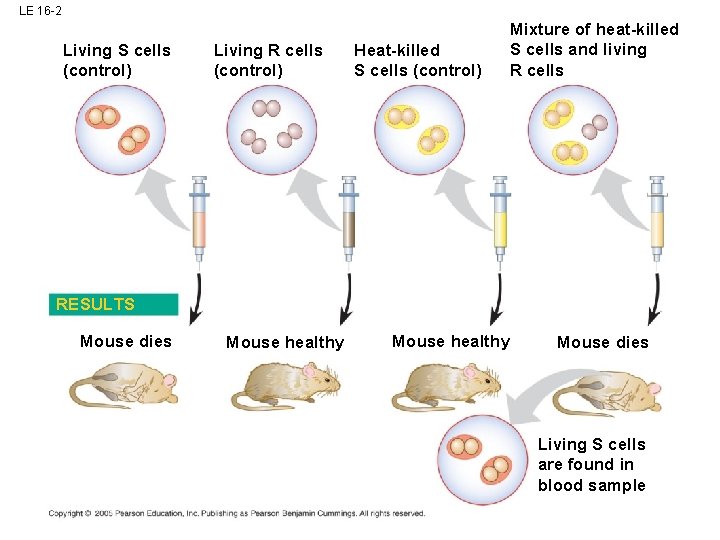
LE 16 -2 Living S cells (control) Living R cells (control) Heat-killed S cells (control) Mixture of heat-killed S cells and living R cells RESULTS Mouse dies Mouse healthy Mouse dies Living S cells are found in blood sample
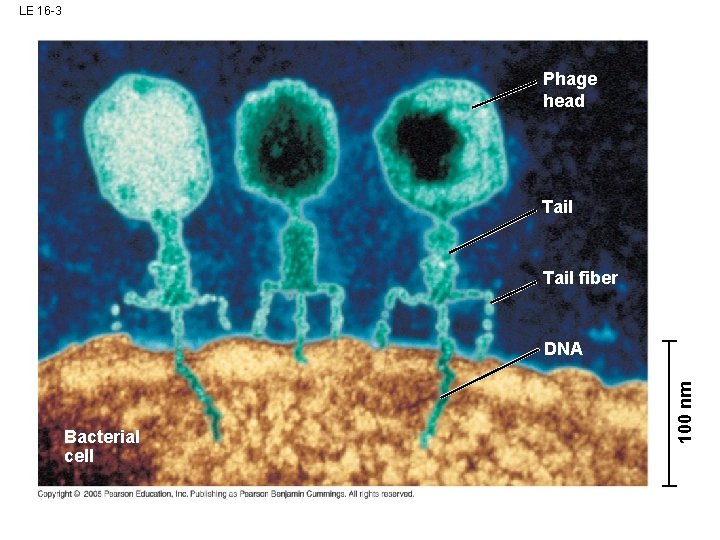
LE 16 -3 Phage head Tail fiber Bacterial cell 100 nm DNA
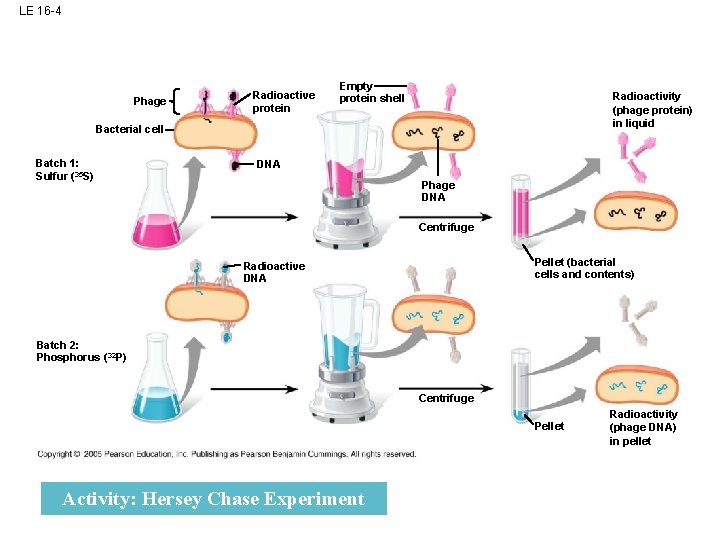
LE 16 -4 Phage Radioactive protein Empty protein shell Radioactivity (phage protein) in liquid Bacterial cell Batch 1: Sulfur (35 S) DNA Phage DNA Centrifuge Pellet (bacterial cells and contents) Radioactive DNA Batch 2: Phosphorus (32 P) Centrifuge Pellet Activity: Hersey Chase Experiment Radioactivity (phage DNA) in pellet
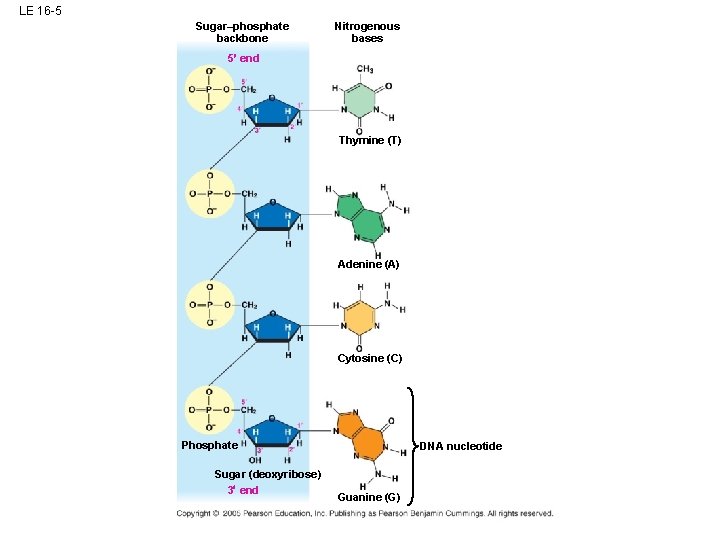
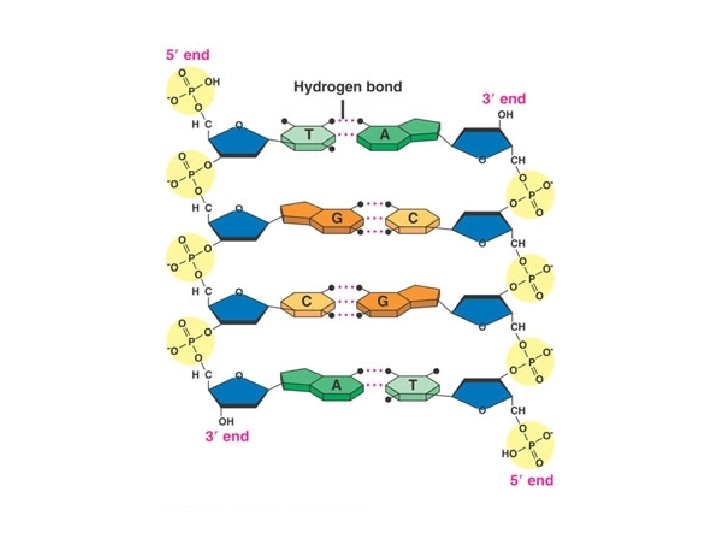
LE 16 -5 Sugar–phosphate backbone Nitrogenous bases 5 end Thymine (T) Adenine (A) Cytosine (C) Phosphate Sugar (deoxyribose) 3 end DNA nucleotide Guanine (G)
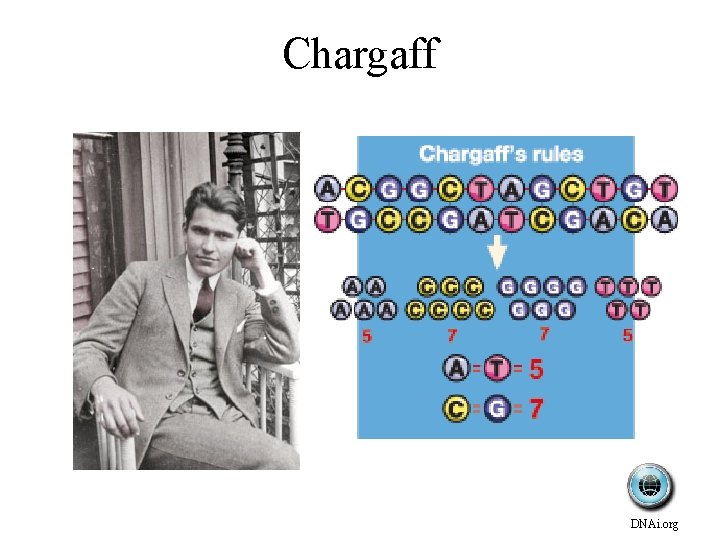
Chargaff DNAi. org

LE 16 -6 Rosalind Franklin’s X-ray diffraction photograph of DNAi. org

Figure 16 -01 DNAi. org
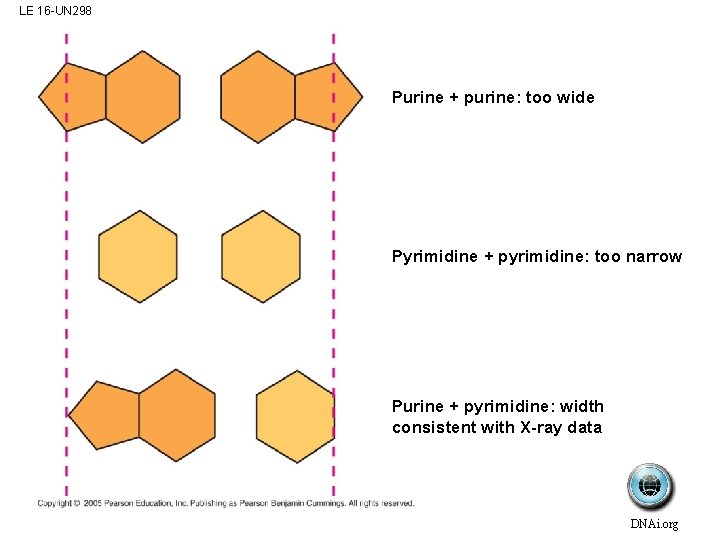
LE 16 -UN 298 Purine + purine: too wide Pyrimidine + pyrimidine: too narrow Purine + pyrimidine: width consistent with X-ray data DNAi. org
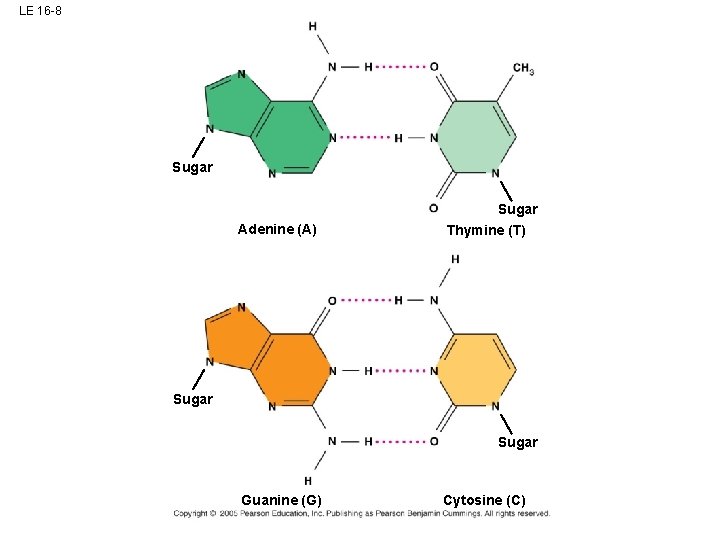
LE 16 -8 Sugar Adenine (A) Sugar Thymine (T) Sugar Guanine (G) Cytosine (C)
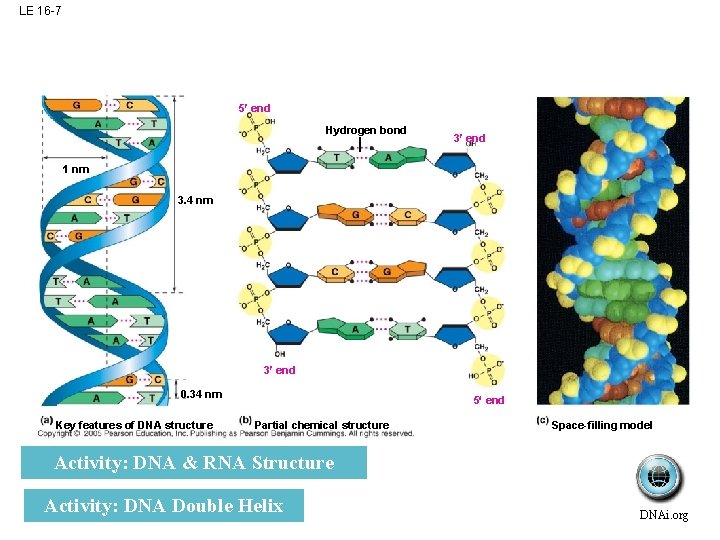
LE 16 -7 5 end Hydrogen bond 3 end 1 nm 3. 4 nm 3 end 0. 34 nm Key features of DNA structure 5 end Partial chemical structure Space-filling model Activity: DNA & RNA Structure Activity: DNA Double Helix DNAi. org
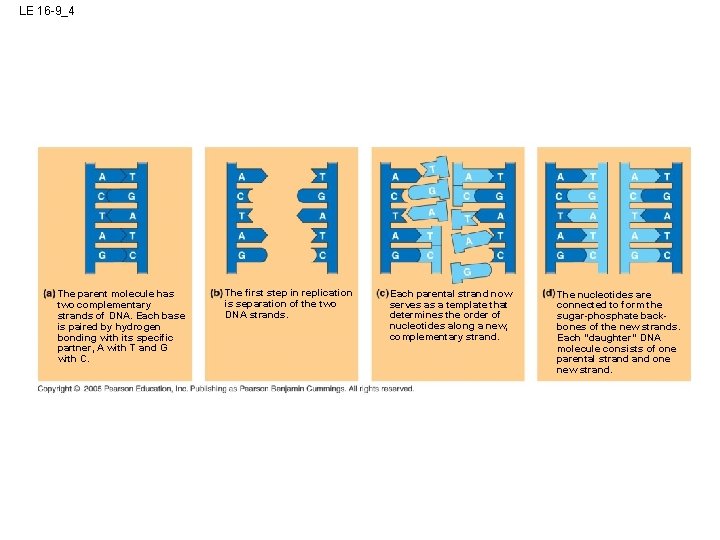
LE 16 -9_4 The parent molecule has two complementary strands of DNA. Each base is paired by hydrogen bonding with its specific partner, A with T and G with C. The first step in replication is separation of the two DNA strands. Each parental strand now serves as a template that determines the order of nucleotides along a new, complementary strand. The nucleotides are connected to form the sugar-phosphate backbones of the new strands. Each “daughter” DNA molecule consists of one parental strand one new strand.
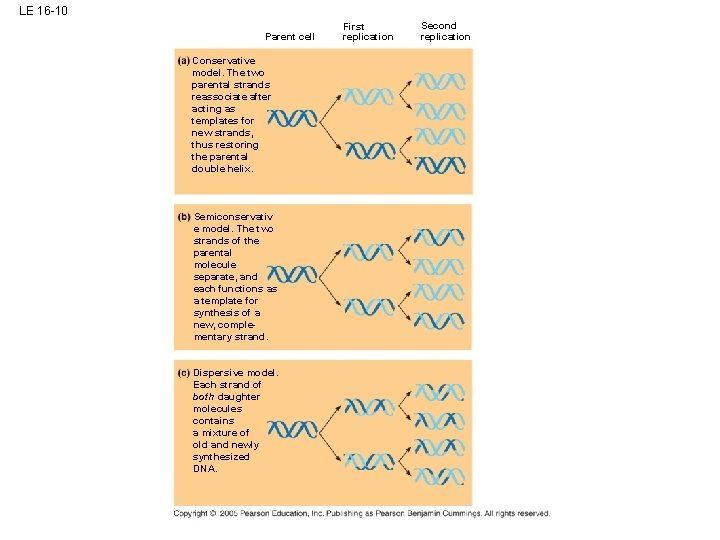
LE 16 -10 Parent cell Conservative model. The two parental strands reassociate after acting as templates for new strands, thus restoring the parental double helix. Semiconservativ e model. The two strands of the parental molecule separate, and each functions as a template for synthesis of a new, complementary strand. Dispersive model. Each strand of both daughter molecules contains a mixture of old and newly synthesized DNA. First replication Second replication
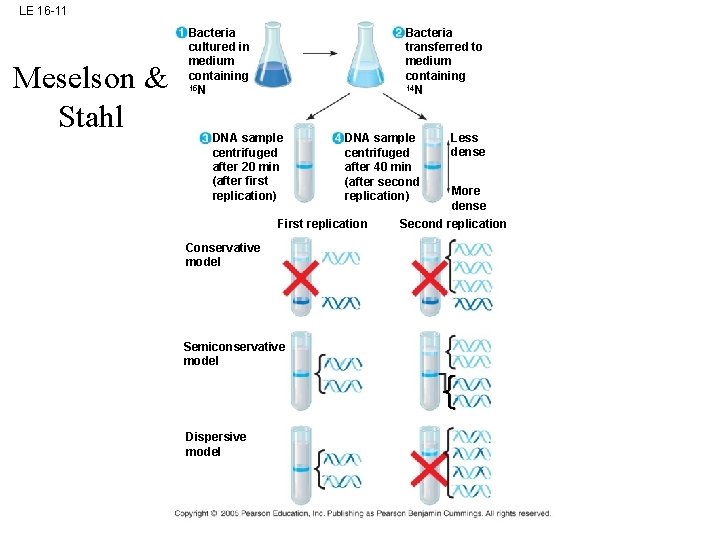
LE 16 -11 Meselson & Stahl Bacteria cultured in medium containing 15 N Bacteria transferred to medium containing 14 N DNA sample centrifuged after 20 min (after first replication) DNA sample centrifuged after 40 min (after second replication) First replication Conservative model Semiconservative model Dispersive model Less dense More dense Second replication
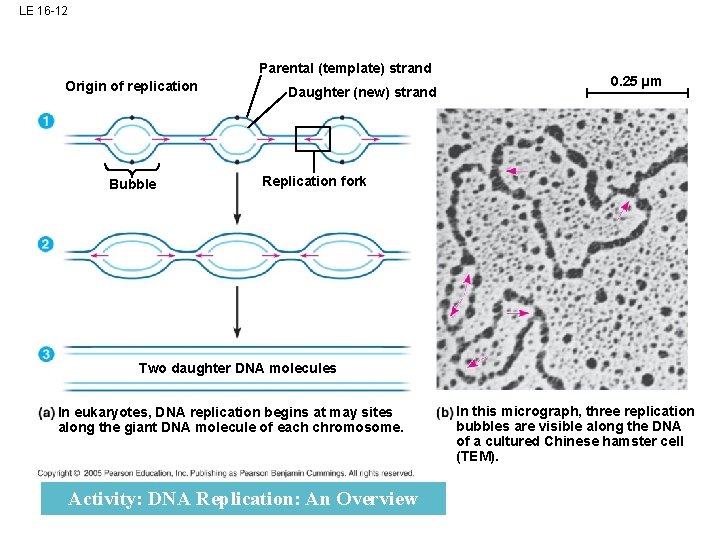
LE 16 -12 Parental (template) strand Origin of replication Bubble Daughter (new) strand 0. 25 µm Replication fork Two daughter DNA molecules In eukaryotes, DNA replication begins at may sites along the giant DNA molecule of each chromosome. Activity: DNA Replication: An Overview In this micrograph, three replication bubbles are visible along the DNA of a cultured Chinese hamster cell (TEM).
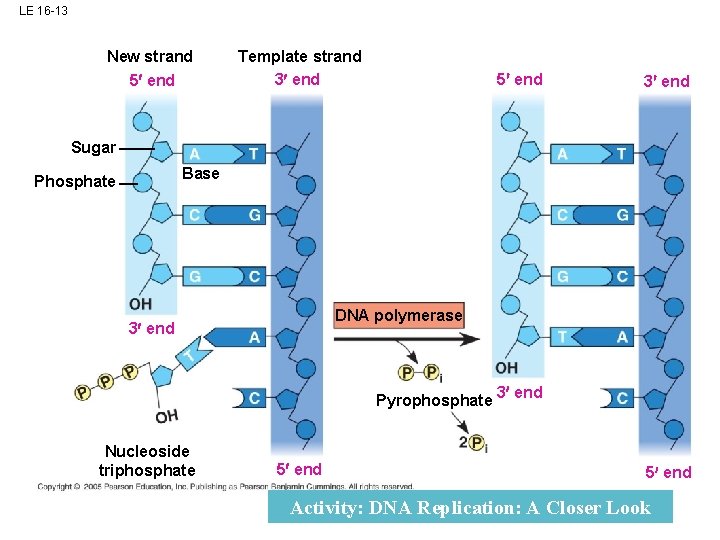
LE 16 -13 New strand 5 end Template strand 3 end 5 end 3 end Sugar Base Phosphate DNA polymerase 3 end Pyrophosphate Nucleoside triphosphate 5 end 3 end 5 end Activity: DNA Replication: A Closer Look
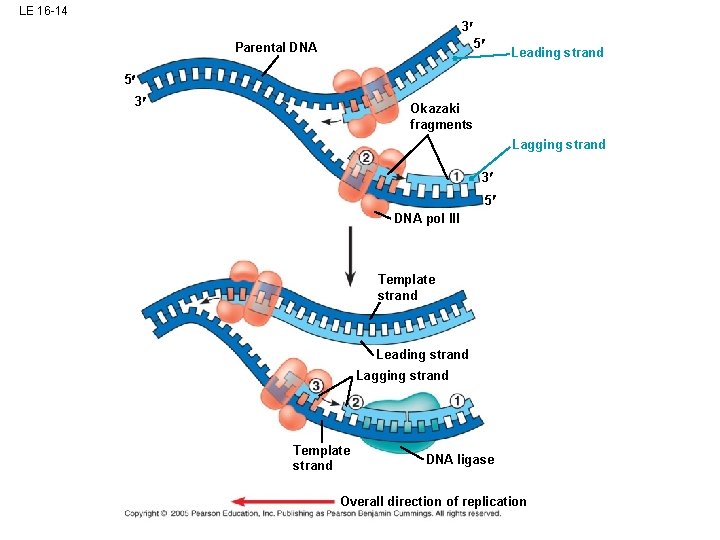
LE 16 -14 3 5 Parental DNA Leading strand 5 3 Okazaki fragments Lagging strand 3 5 DNA pol III Template strand Leading strand Lagging strand Template strand DNA ligase Overall direction of replication
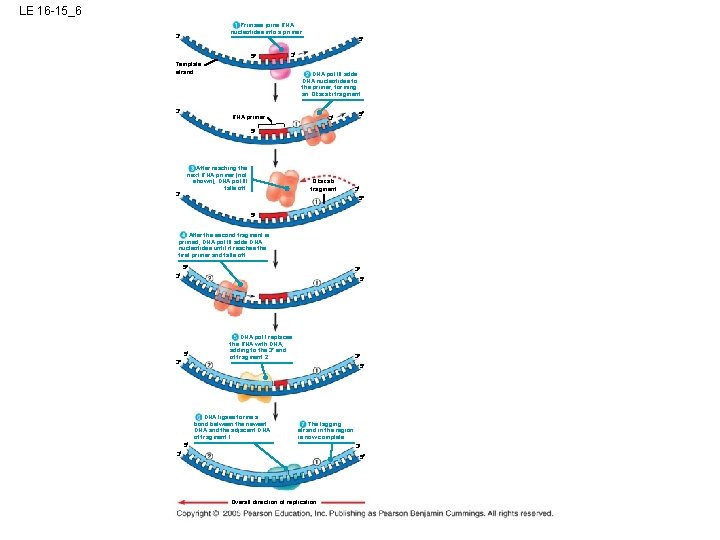
LE 16 -15_6 Primase joins RNA nucleotides into a primer. 3 5 5 3 Template strand 3 DNA pol III adds DNA nucleotides to the primer, forming an Okazaki fragment. RNA primer 3 5 5 3 After reaching the next RNA primer (not shown), DNA pol III falls off. Okazaki fragment 3 5 5 After the second fragment is primed, DNA pol III adds DNA nucleotides until it reaches the first primer and falls off. 5 3 3 5 5 3 DNA pol I replaces the RNA with DNA, adding to the 3 end of fragment 2. 3 5 DNA ligase forms a bond between the newest DNA and the adjacent DNA of fragment 1. The lagging strand in the region is now complete. 5 3 3 5 Overall direction of replication
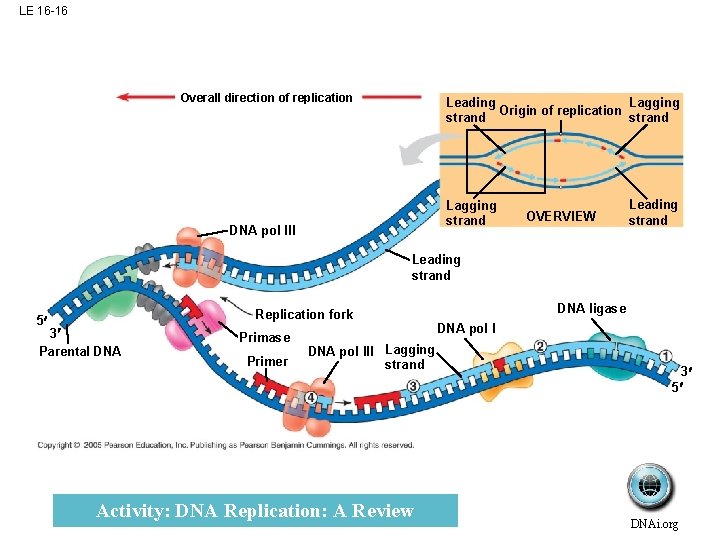
LE 16 -16 Overall direction of replication Lagging Leading Origin of replication strand Lagging strand DNA pol III OVERVIEW Leading strand 5 Replication fork 3 Parental DNA Primase Primer DNA pol III Lagging strand Activity: DNA Replication: A Review DNA ligase DNA pol I 3 5 DNAi. org
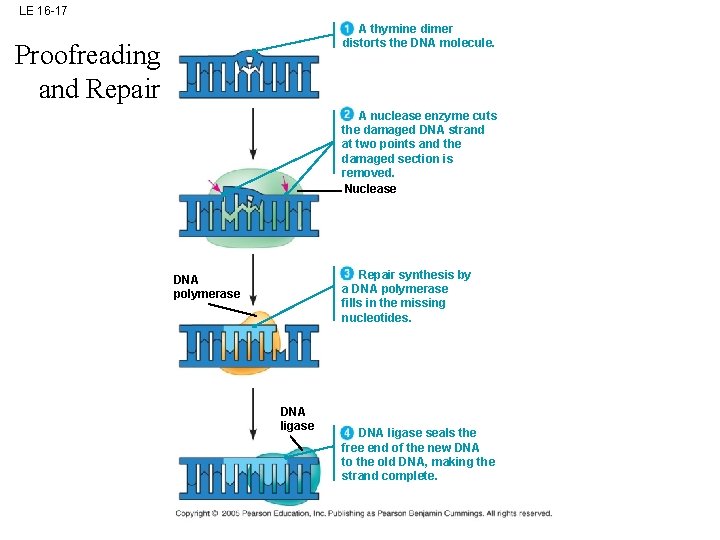
LE 16 -17 A thymine dimer distorts the DNA molecule. Proofreading and Repair A nuclease enzyme cuts the damaged DNA strand at two points and the damaged section is removed. Nuclease Repair synthesis by a DNA polymerase fills in the missing nucleotides. DNA polymerase DNA ligase seals the free end of the new DNA to the old DNA, making the strand complete.
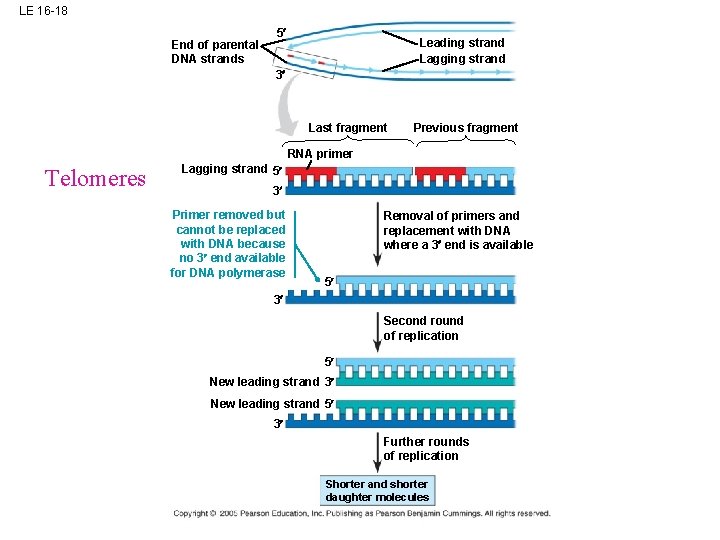
LE 16 -18 End of parental DNA strands 5 Leading strand Lagging strand 3 Last fragment Previous fragment RNA primer Telomeres Lagging strand 5 3 Primer removed but cannot be replaced with DNA because no 3 end available for DNA polymerase Removal of primers and replacement with DNA where a 3 end is available 5 3 Second round of replication 5 New leading strand 3 New leading strand 5 3 Further rounds of replication Shorter and shorter daughter molecules
- Slides: 22