Chapter 16 DNA Replication Slides with blue borders

Chapter 16 DNA Replication Slides with blue borders come from a slide show by Kim Foglia (http: //explorebiology. com) AP Biology 2007 -2008
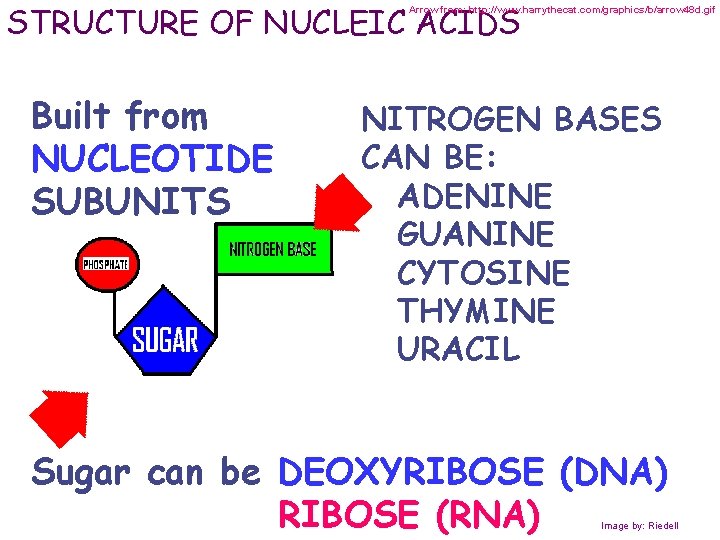
STRUCTURE OF NUCLEIC ACIDS Arrow from: http: //www. harrythecat. com/graphics/b/arrow 48 d. gif Built from NUCLEOTIDE SUBUNITS NITROGEN BASES CAN BE: ADENINE GUANINE CYTOSINE THYMINE URACIL Sugar can be DEOXYRIBOSE (DNA) RIBOSE (RNA) Image by: Riedell
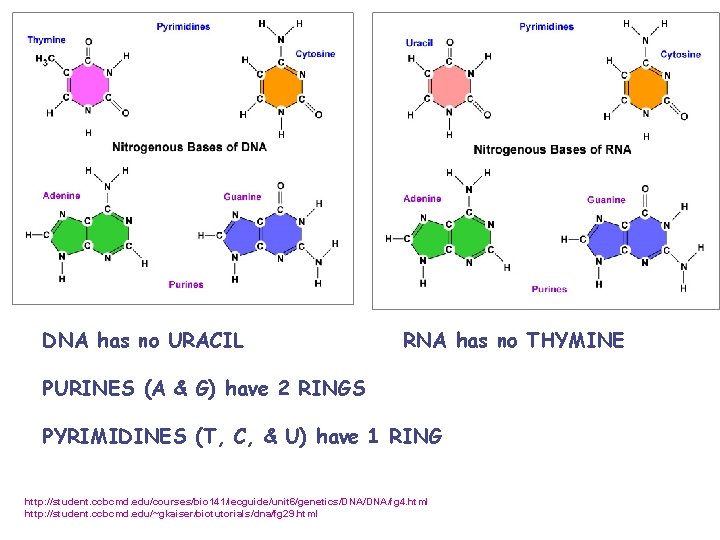
DNA has no URACIL RNA has no THYMINE PURINES (A & G) have 2 RINGS PYRIMIDINES (T, C, & U) have 1 RING http: //student. ccbcmd. edu/courses/bio 141/lecguide/unit 6/genetics/DNA/fg 4. html http: //student. ccbcmd. edu/~gkaiser/biotutorials/dna/fg 29. html
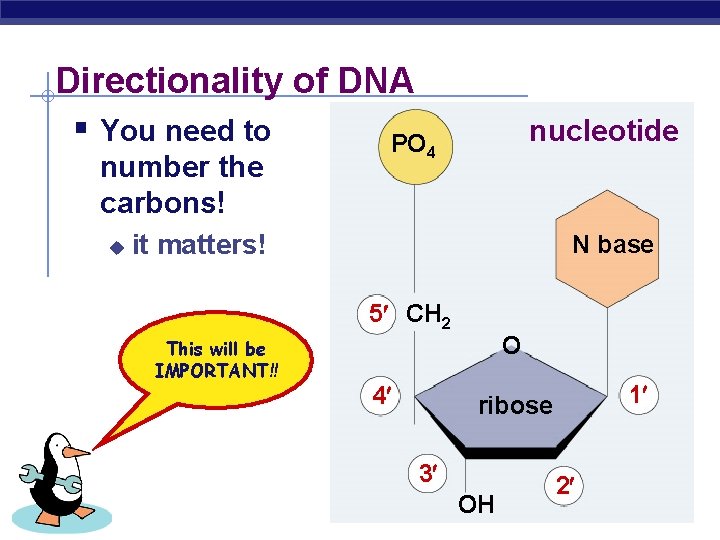
Directionality of DNA § You need to PO 4 nucleotide number the carbons! u it matters! N base 5 CH 2 This will be IMPORTANT!! 4 O 3 AP Biology 1 ribose OH 2
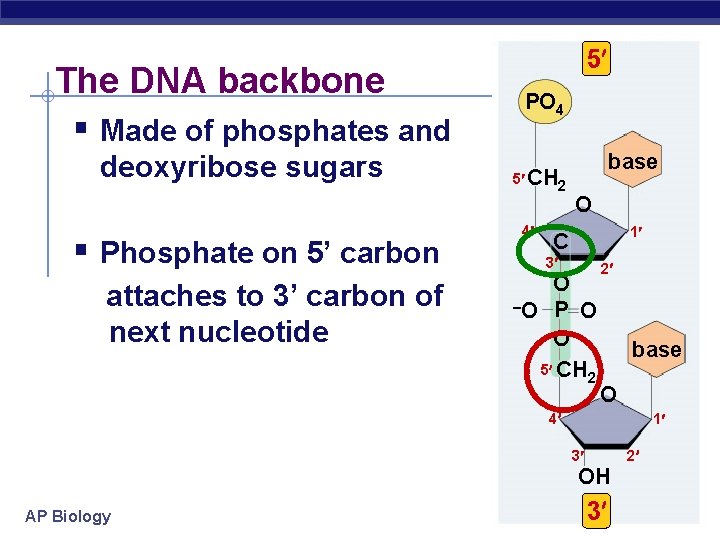
The DNA backbone § Made of phosphates and deoxyribose sugars § Phosphate on 5’ carbon attaches to 3’ carbon of next nucleotide 5 PO 4 5 CH 2 4 base O 1 C 3 O –O P O O 5 CH 2 2 base O 4 1 2 3 OH AP Biology 3
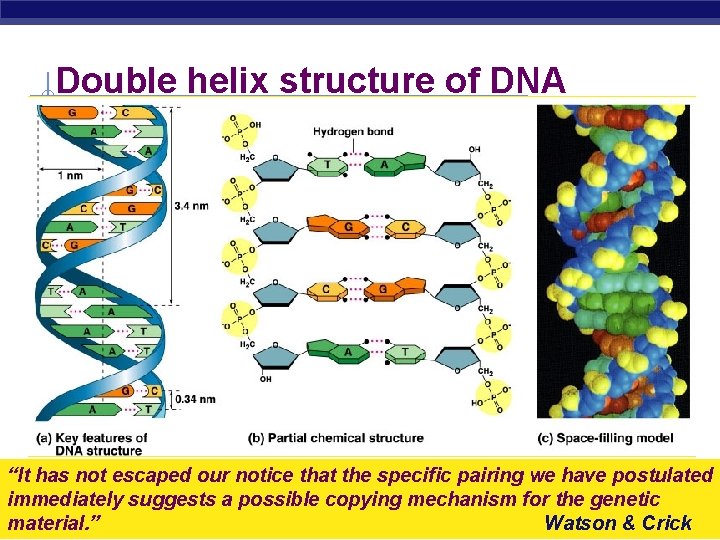
Double helix structure of DNA “It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic AP Biology material. ” Watson & Crick
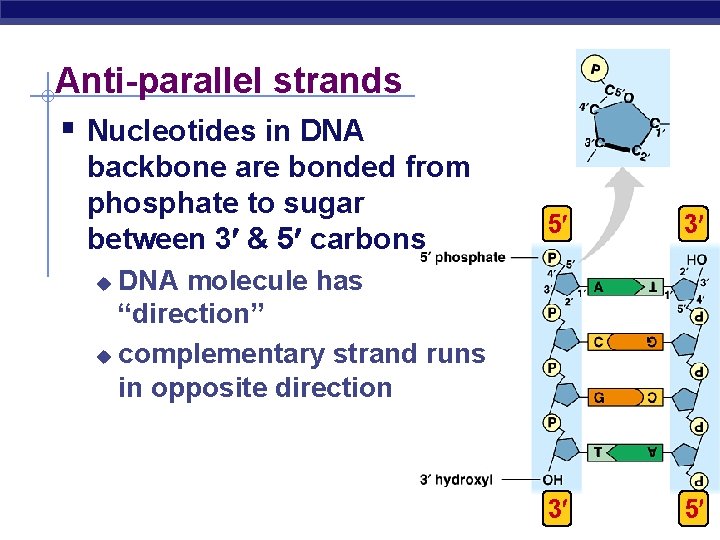
Anti-parallel strands § Nucleotides in DNA backbone are bonded from phosphate to sugar between 3 & 5 carbons 5 3 3 5 DNA molecule has “direction” u complementary strand runs in opposite direction u AP Biology
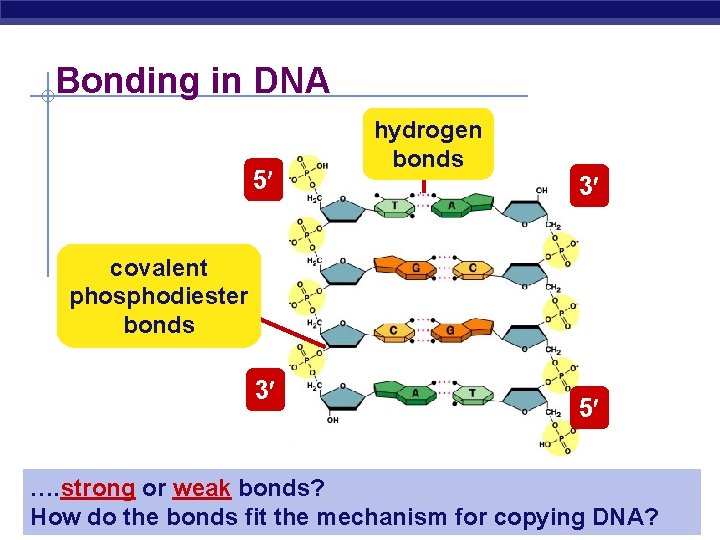
Bonding in DNA 5 hydrogen bonds 3 covalent phosphodiester bonds 3 5 …. strong or weak bonds? AP Biology How do the bonds fit the mechanism for copying DNA?
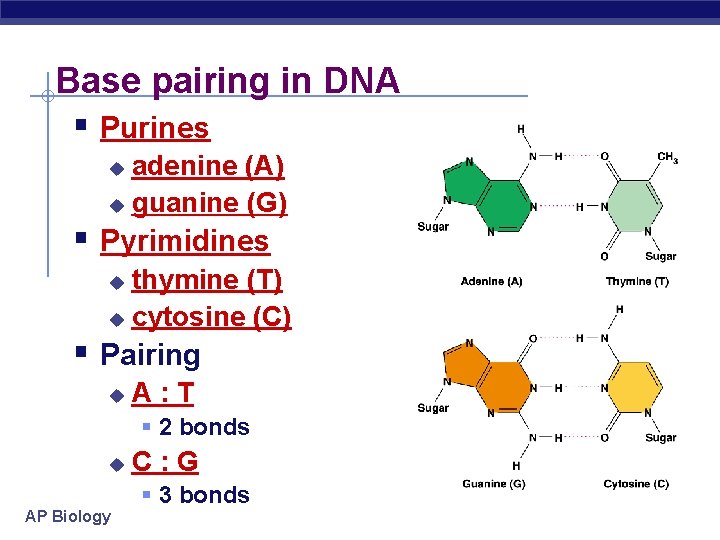
Base pairing in DNA § Purines adenine (A) u guanine (G) u § Pyrimidines thymine (T) u cytosine (C) u § Pairing u A: T § 2 bonds u AP Biology C: G § 3 bonds
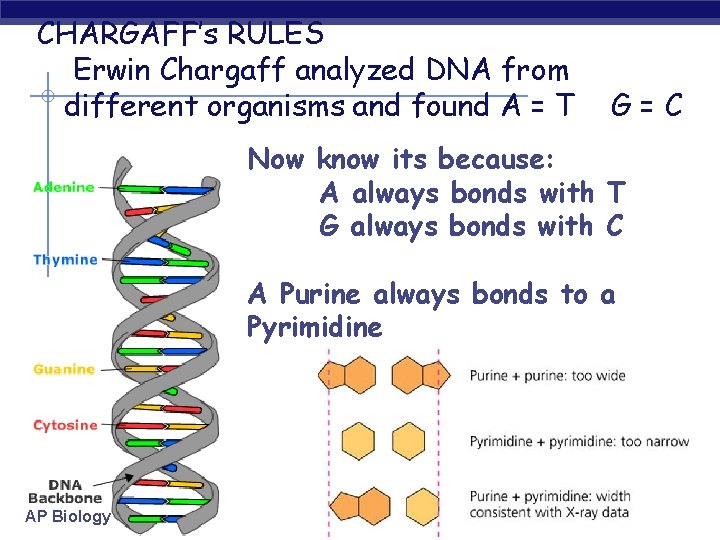
CHARGAFF’s RULES Erwin Chargaff analyzed DNA from different organisms and found A = T G=C Now know its because: A always bonds with T G always bonds with C A Purine always bonds to a Pyrimidine AP Biology
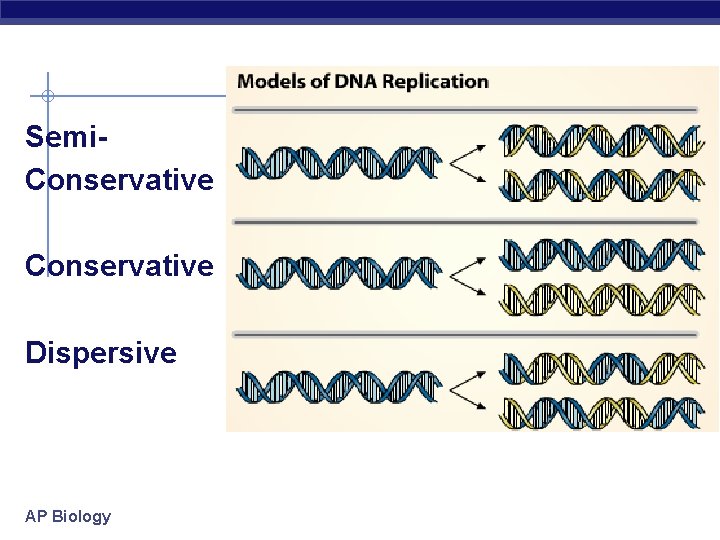
Semi. Conservative Dispersive AP Biology
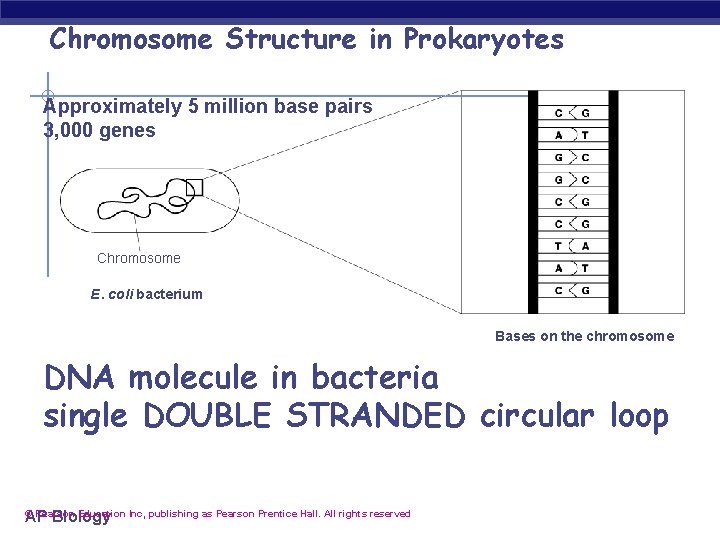
Chromosome Structure in Prokaryotes Approximately 5 million base pairs 3, 000 genes Chromosome E. coli bacterium Bases on the chromosome DNA molecule in bacteria single DOUBLE STRANDED circular loop AP Biology © Pearson Education Inc, publishing as Pearson Prentice Hall. All rights reserved
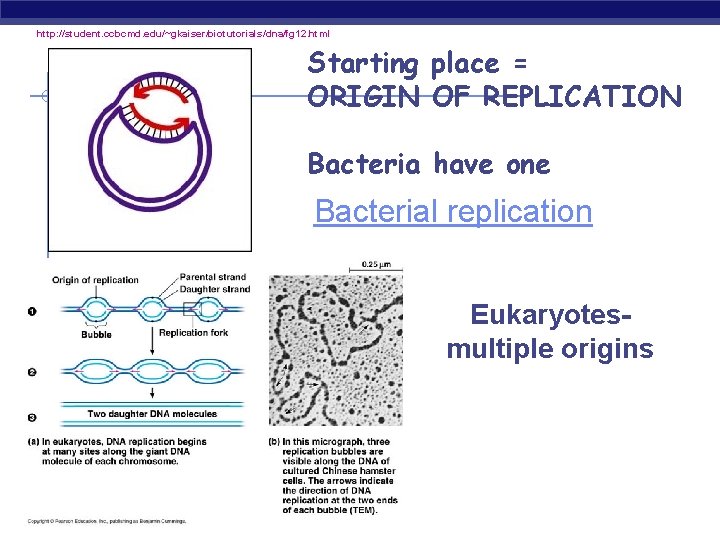
http: //student. ccbcmd. edu/~gkaiser/biotutorials/dna/fg 12. html Starting place = ORIGIN OF REPLICATION Bacteria have one Bacterial replication Eukaryotesmultiple origins AP Biology
HOW NUCLEOTIDES ARE ADDED DNA REPLICATION FORK DNA replication Triphosphate addition DNA replication 2 DNA replication/quiz AP Biology http: //bio. usuhs. mil/biochem 4. html
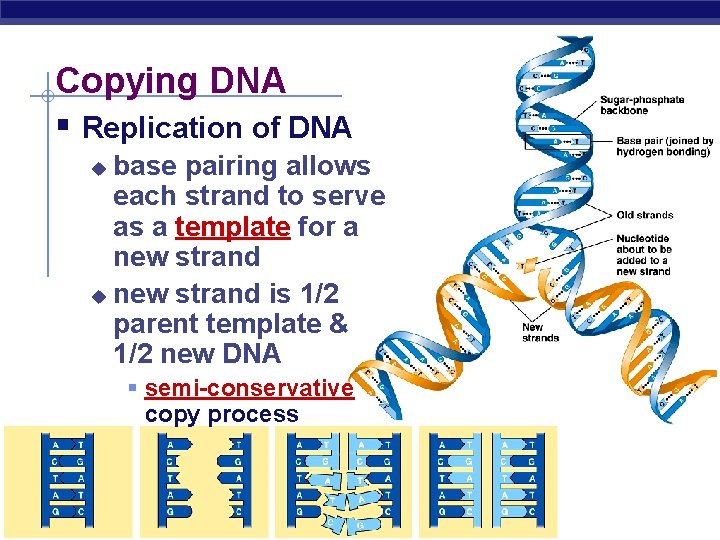
Copying DNA § Replication of DNA base pairing allows each strand to serve as a template for a new strand u new strand is 1/2 parent template & 1/2 new DNA u § semi-conservative copy process AP Biology
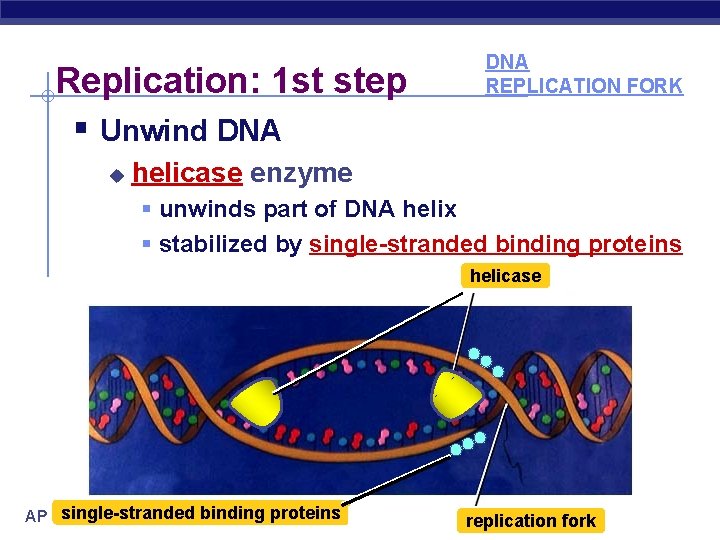
Replication: 1 st step § Unwind DNA u DNA REPLICATION FORK helicase enzyme § unwinds part of DNA helix § stabilized by single-stranded binding proteins helicase single-stranded binding proteins AP Biology replication fork
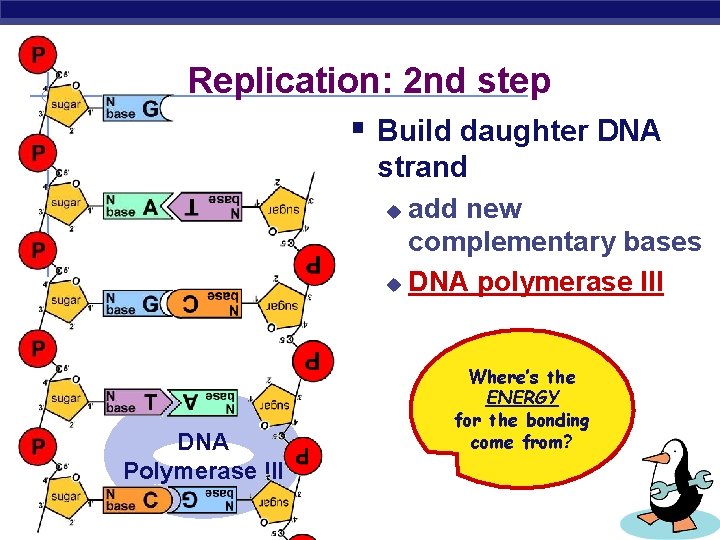
Replication: 2 nd step § Build daughter DNA strand add new complementary bases u DNA polymerase III u DNA Polymerase III AP Biology Where’s the ENERGY for the bonding come from?
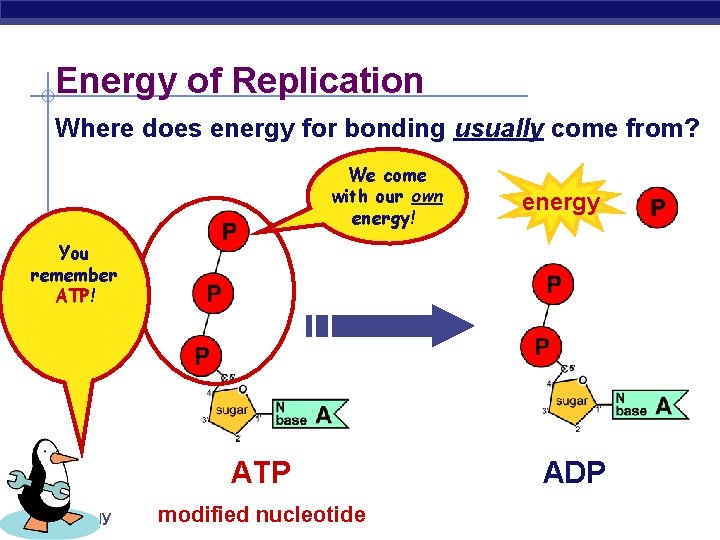
Energy of Replication Where does energy for bonding usually come from? We come with our own energy! energy You remember ATP! ATP AP Biology modified nucleotide ADP
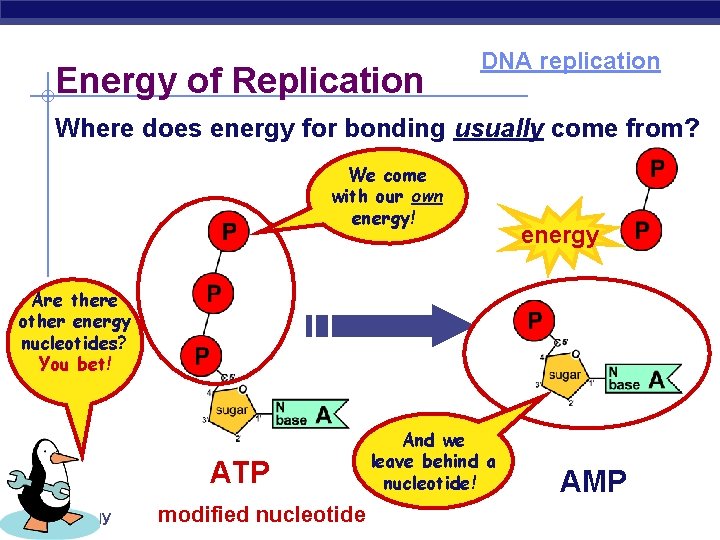
Energy of Replication DNA replication Where does energy for bonding usually come from? We come with our own energy! energy Are there other energy nucleotides? You bet! ATP AP Biology modified nucleotide And we leave behind a nucleotide! AMP
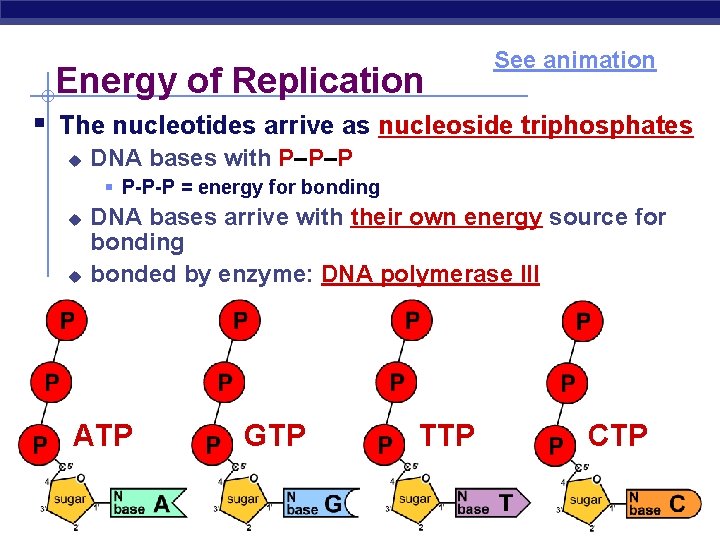
Energy of Replication See animation § The nucleotides arrive as nucleoside triphosphates u DNA bases with P–P–P § P-P-P = energy for bonding u u DNA bases arrive with their own energy source for bonding bonded by enzyme: DNA polymerase III ATP AP Biology GTP TTP CTP
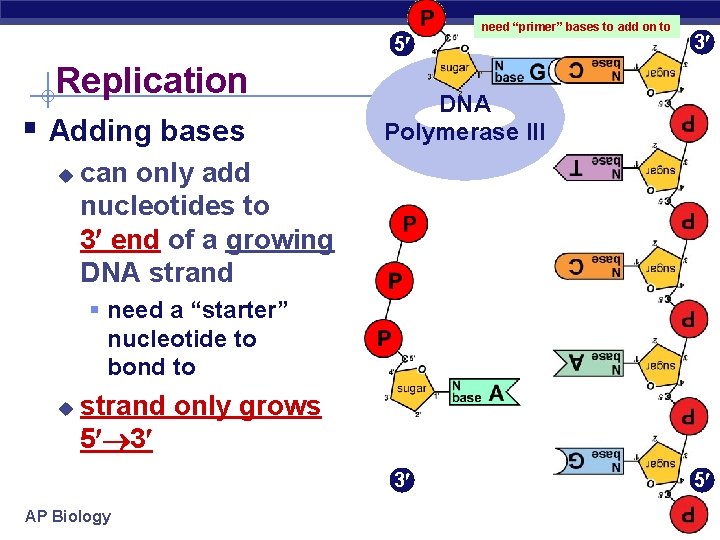
5 Replication § Adding bases u need “primer” bases to add on to 3 DNA Polymerase III can only add nucleotides to 3 end of a growing DNA strand § need a “starter” nucleotide to bond to u strand only grows 5 3 3 AP Biology 5
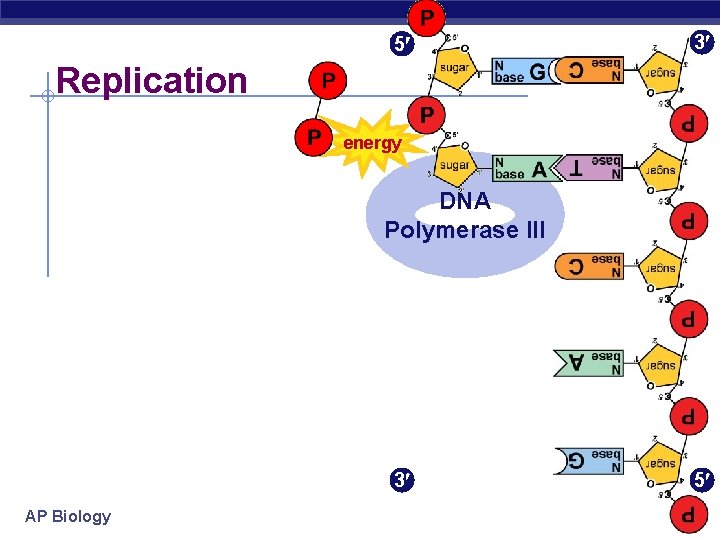
5 3 Replication energy DNA Polymerase III 3 AP Biology 5
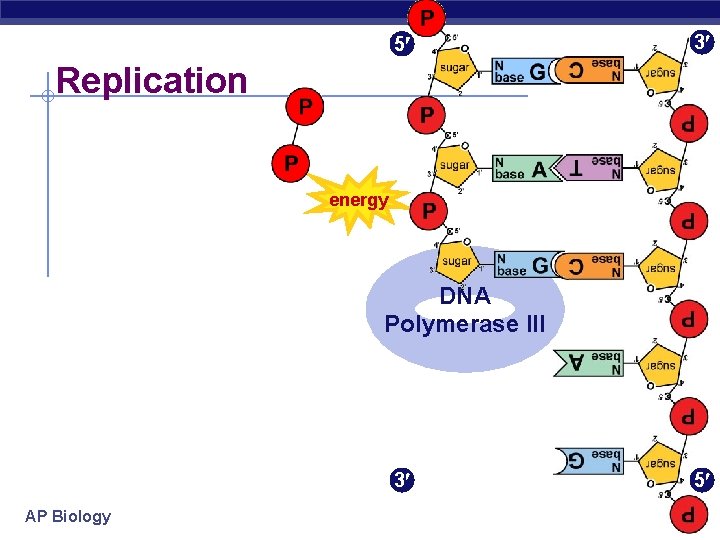
5 3 Replication energy DNA Polymerase III 3 AP Biology 5
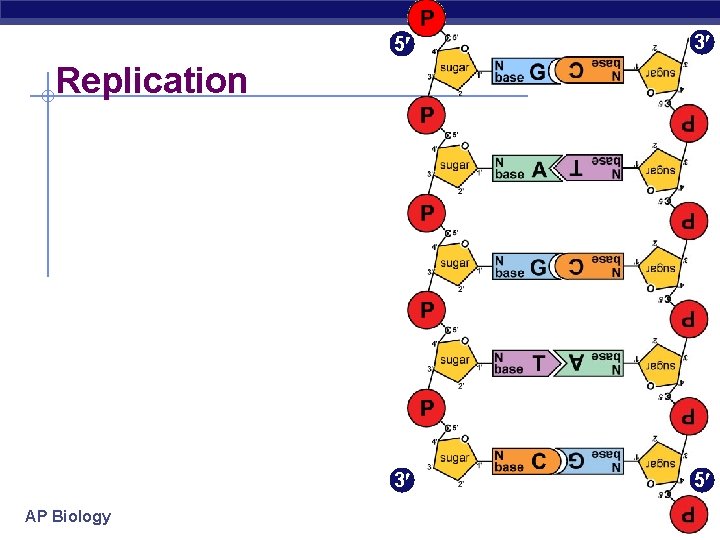
5 3 3 5 Replication AP Biology
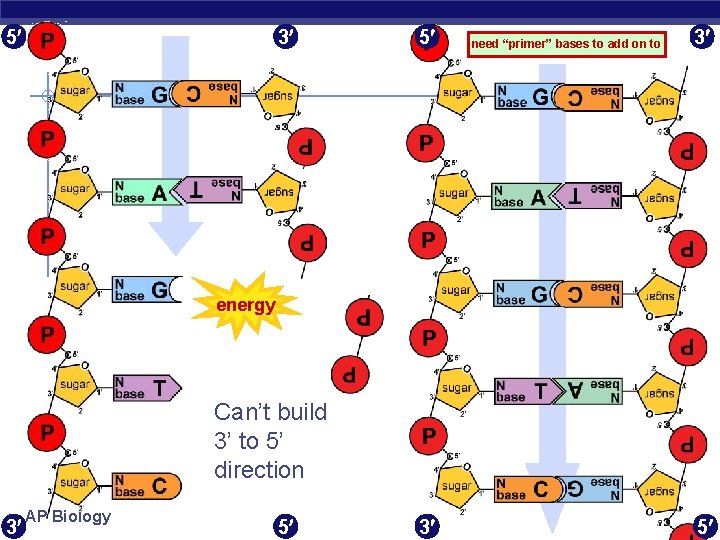
5 3 5 need “primer” bases to add on to 3 energy Can’t build 3’ to 5’ direction 3 AP Biology 5 3 5
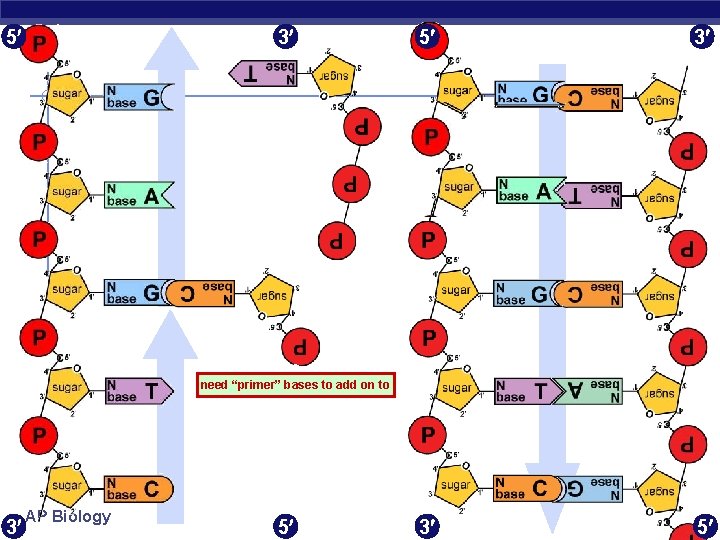
5 3 3 5 need “primer” bases to add on to 3 AP Biology 5
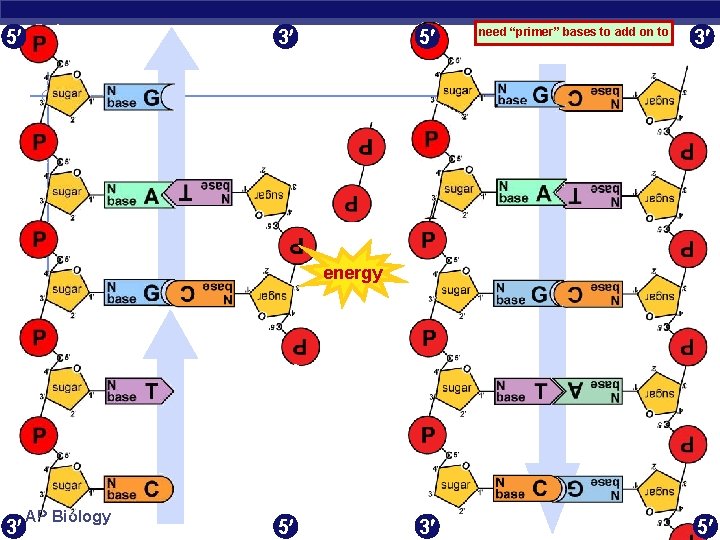
5 3 5 need “primer” bases to add on to 3 energy 3 AP Biology 5 3 5
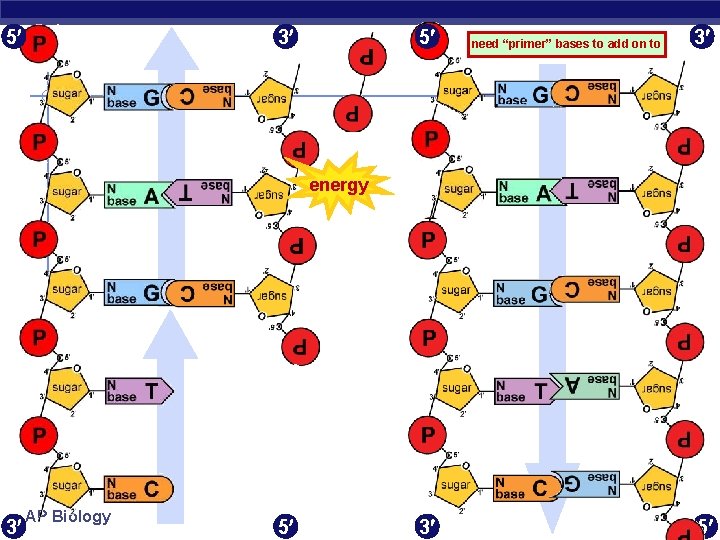
5 3 5 need “primer” bases to add on to 3 energy 3 AP Biology 5 3 5
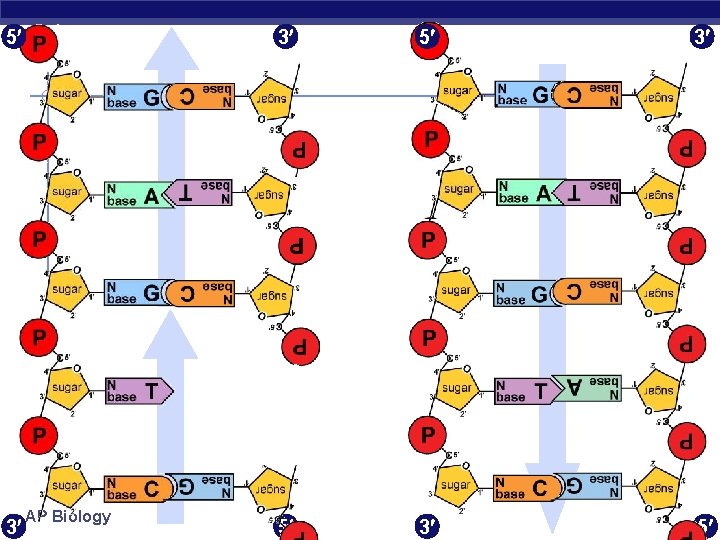
5 3 AP Biology 3 5
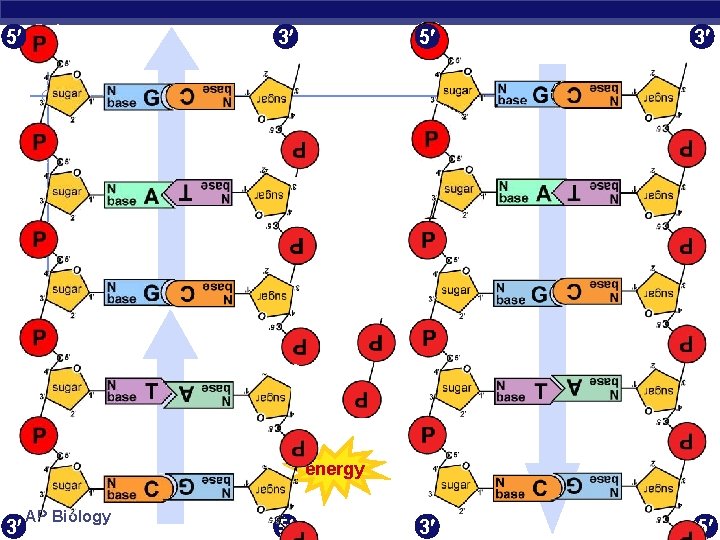
5 3 3 5 energy 3 AP Biology 5
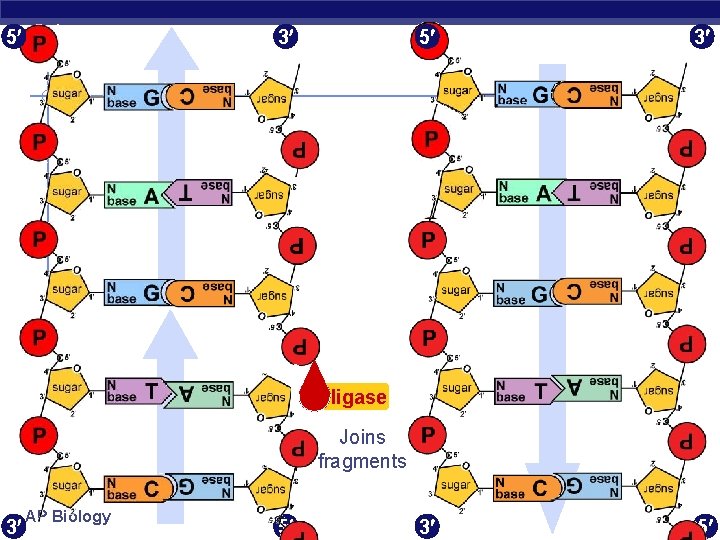
5 3 3 5 ligase Joins fragments 3 AP Biology 5
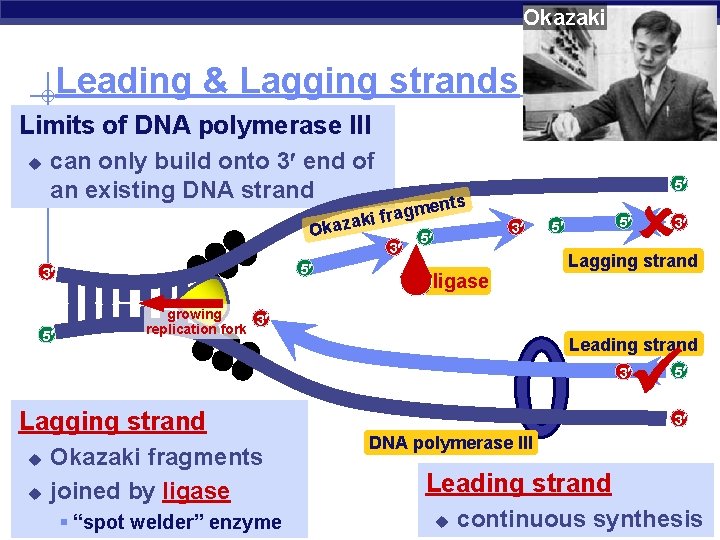
Okazaki Leading & Lagging strands Limits of DNA polymerase III u can only build onto 3 end of an existing DNA strand en m g a r f ki Okaza 3 5 5 ts 3 5 ligase growing 3 replication fork 5 5 Lagging strand Leading strand 3 Lagging strand u u Okazaki fragments joined by ligase AP Biology § “spot welder” enzyme 3 5 3 DNA polymerase III Leading strand u continuous synthesis
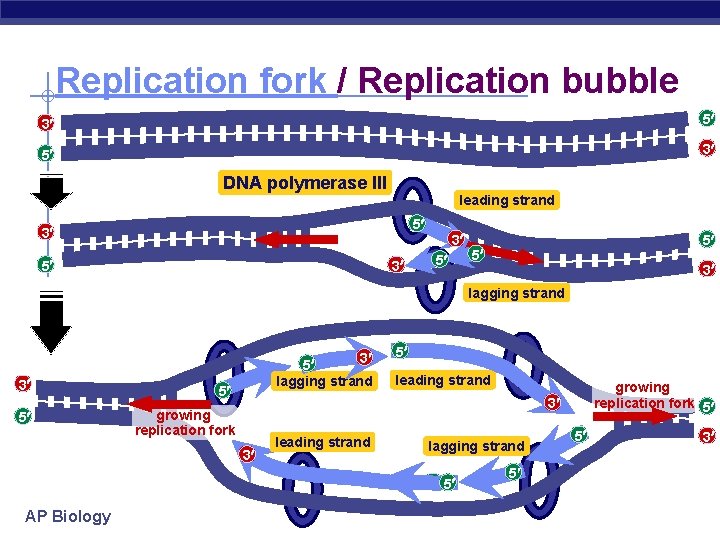
Replication fork / Replication bubble 3 5 5 3 DNA polymerase III leading strand 5 3 3 5 5 5 3 lagging strand 3 5 lagging strand 5 5 leading strand 3 growing replication fork 3 leading strand lagging strand 5 5 AP Biology growing replication fork 5 5 5 3
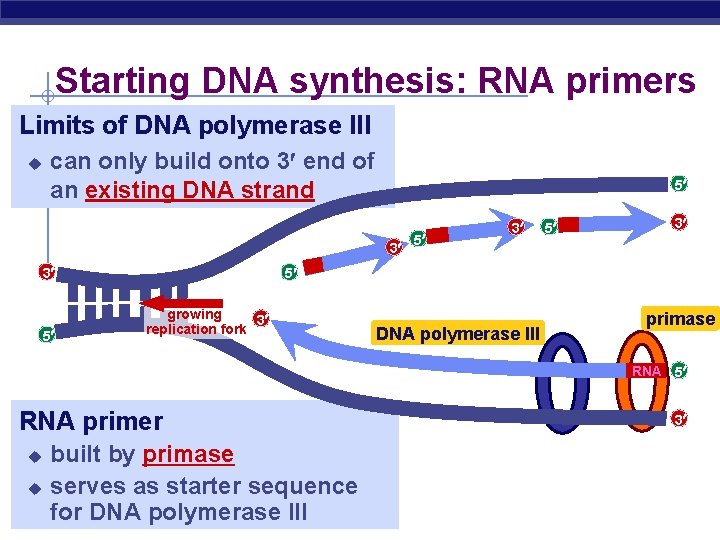
Starting DNA synthesis: RNA primers Limits of DNA polymerase III u can only build onto 3 end of an existing DNA strand 5 3 3 5 5 3 5 growing 3 replication fork DNA polymerase III primase RNA 5 RNA primer built by primase u serves as starter sequence DNA polymerase III AP for Biology u 3
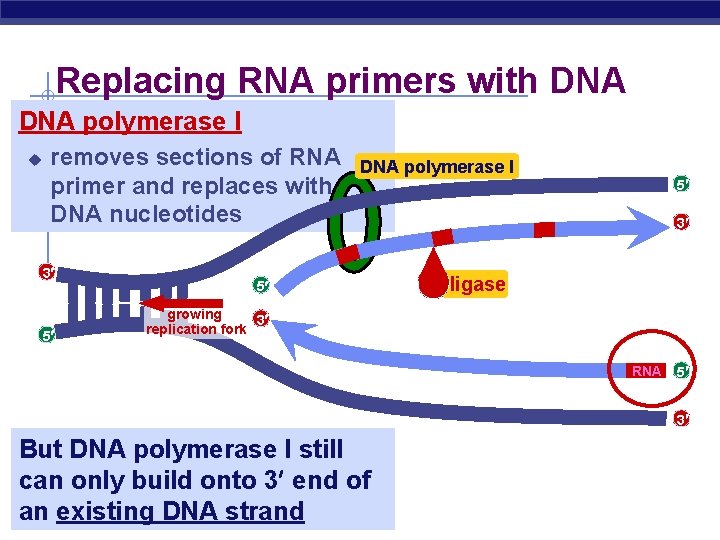
Replacing RNA primers with DNA polymerase I u removes sections of RNA primer and replaces with DNA nucleotides 3 5 DNA polymerase I 5 5 3 ligase growing 3 replication fork RNA 5 3 But DNA polymerase I still can only build onto 3 end of an existing DNA strand AP Biology
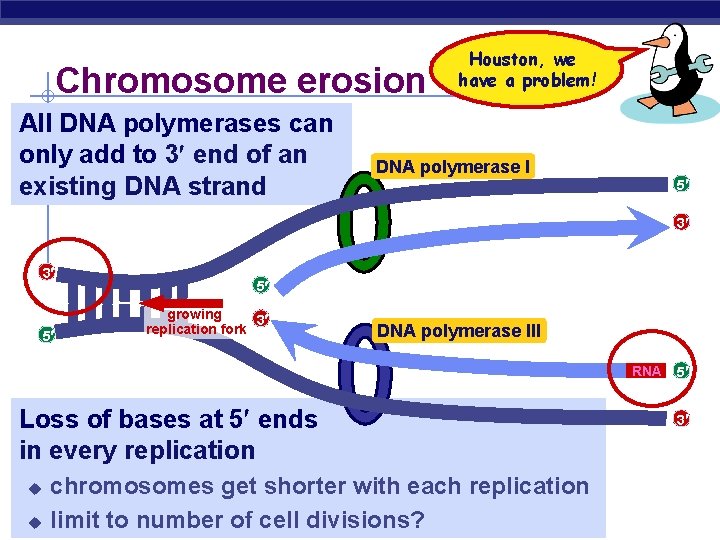
Chromosome erosion All DNA polymerases can only add to 3 end of an existing DNA strand Houston, we have a problem! DNA polymerase I 5 3 3 5 5 growing 3 replication fork DNA polymerase III RNA Loss of bases at 5 ends in every replication chromosomes get shorter with each replication AP Biologyto number of cell divisions? u limit u 5 3
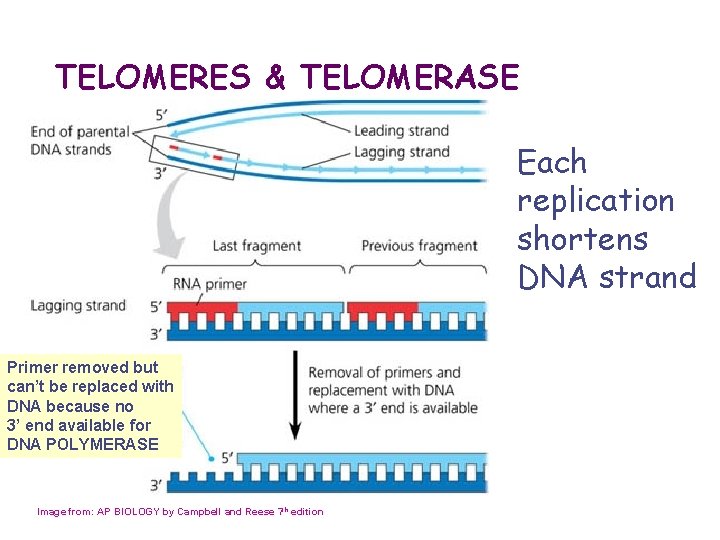
TELOMERES & TELOMERASE Each replication shortens DNA strand Primer removed but can’t be replaced with DNA because no 3’ end available for DNA POLYMERASE Image from: AP BIOLOGY by Campbell and Reese 7 th edition
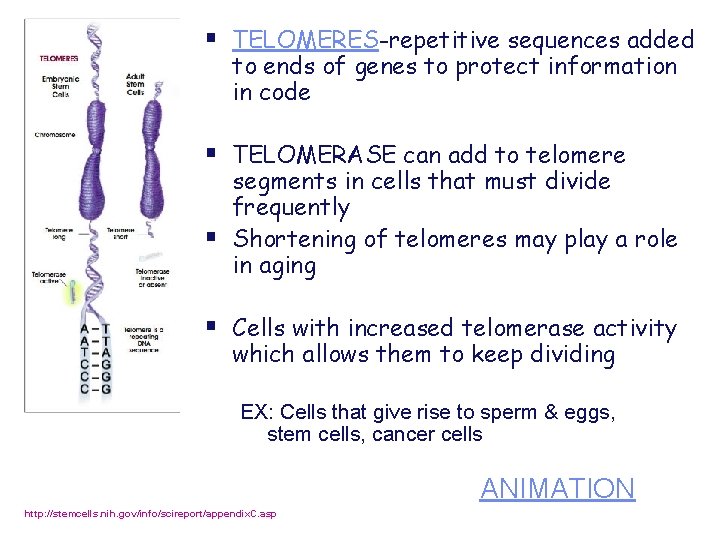
§ TELOMERES-repetitive sequences added to ends of genes to protect information in code § TELOMERASE can add to telomere § segments in cells that must divide frequently Shortening of telomeres may play a role in aging § Cells with increased telomerase activity which allows them to keep dividing EX: Cells that give rise to sperm & eggs, stem cells, cancer cells ANIMATION http: //stemcells. nih. gov/info/scireport/appendix. C. asp
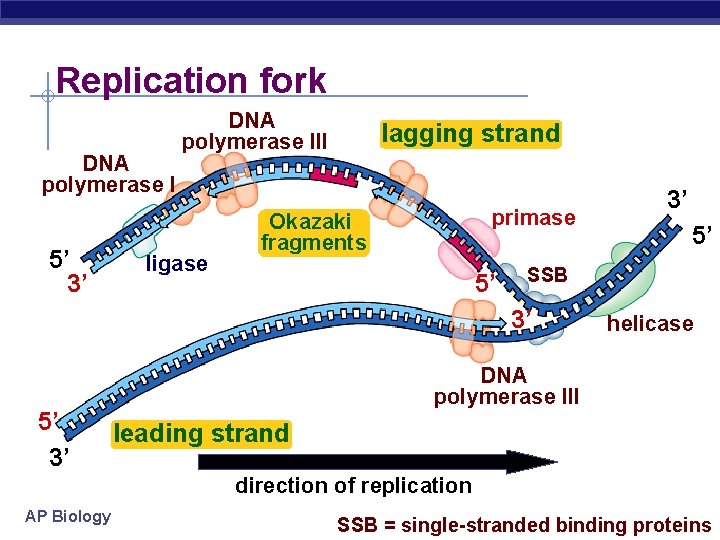
Replication fork DNA polymerase I 5’ 3’ DNA polymerase III ligase lagging strand primase Okazaki fragments 5’ 5’ SSB 3’ 5’ 3’ 3’ helicase DNA polymerase III leading strand direction of replication AP Biology SSB = single-stranded binding proteins
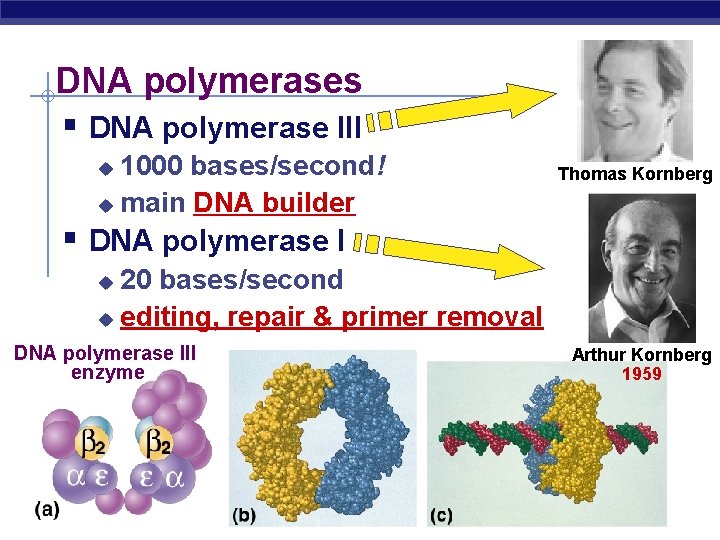
DNA polymerases § DNA polymerase III 1000 bases/second! u main DNA builder u Thomas Kornberg § DNA polymerase I 20 bases/second u editing, repair & primer removal u DNA polymerase III enzyme AP Biology Arthur Kornberg 1959

Fast & accurate! § It takes E. coli <1 hour to copy 5 million base pairs in its single chromosome u divide to form 2 identical daughter cells § Human cell copies its 6 billion bases & divide into daughter cells in only few hours remarkably accurate u only ~1 error per 100 million bases u ~30 errors per cell cycle u AP Biology
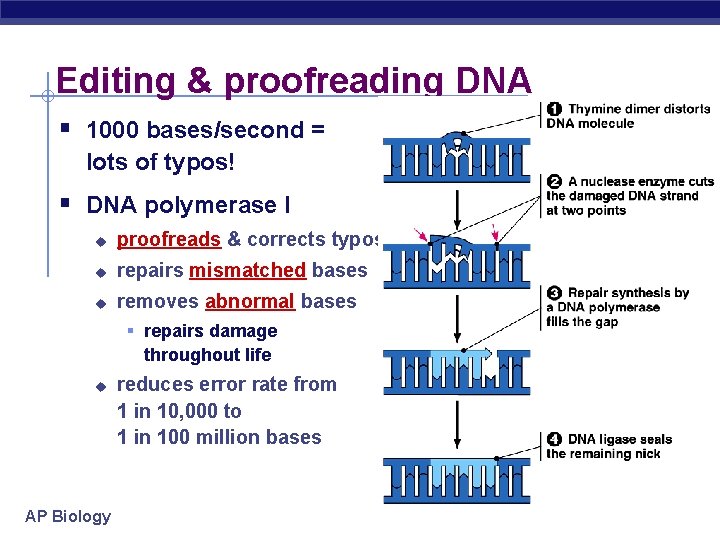
Editing & proofreading DNA § 1000 bases/second = lots of typos! § DNA polymerase I u proofreads & corrects typos u repairs mismatched bases u removes abnormal bases § repairs damage throughout life u AP Biology reduces error rate from 1 in 10, 000 to 1 in 100 million bases
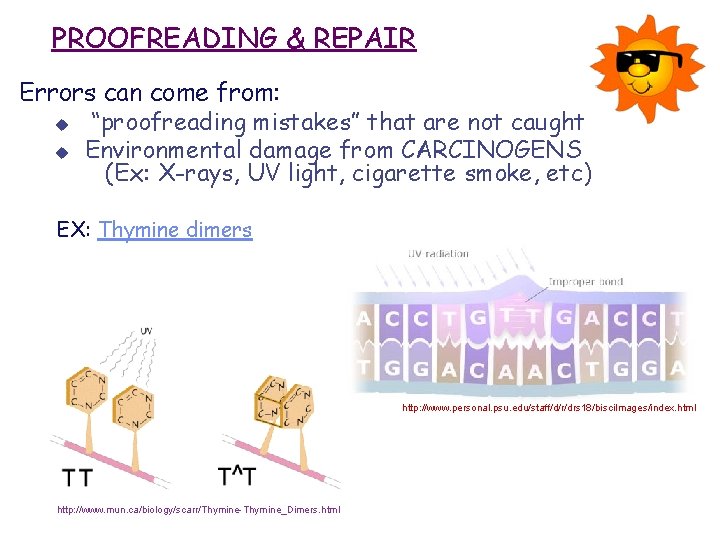
PROOFREADING & REPAIR Errors can come from: u “proofreading mistakes” that are not caught u Environmental damage from CARCINOGENS (Ex: X-rays, UV light, cigarette smoke, etc) EX: Thymine dimers http: //www. personal. psu. edu/staff/d/r/drs 18/bisci. Images/index. html http: //www. mun. ca/biology/scarr/Thymine-Thymine_Dimers. html
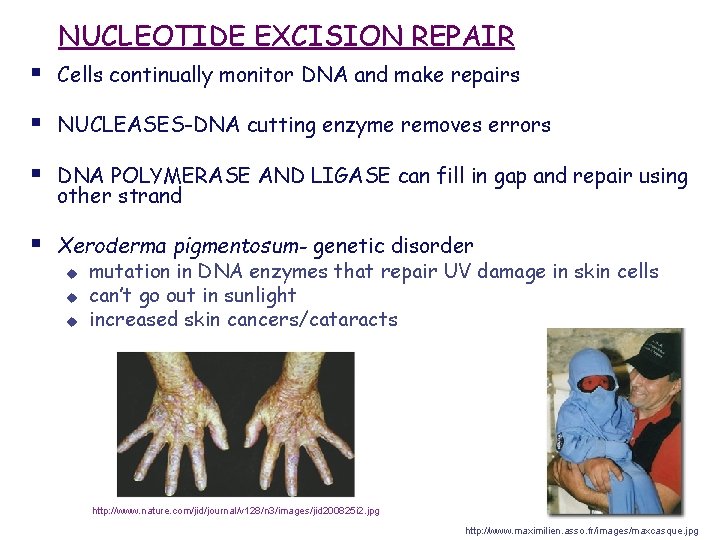
NUCLEOTIDE EXCISION REPAIR § Cells continually monitor DNA and make repairs § NUCLEASES-DNA cutting enzyme removes errors § DNA POLYMERASE AND LIGASE can fill in gap and repair using other strand § Xeroderma pigmentosum- genetic disorder u u u mutation in DNA enzymes that repair UV damage in skin cells can’t go out in sunlight increased skin cancers/cataracts http: //www. nature. com/jid/journal/v 128/n 3/images/jid 200825 i 2. jpg http: //www. maximilien. asso. fr/images/maxcasque. jpg
- Slides: 44