Chapter 12 Bio and estimation technology Estimation technology

Chapter 12: Bio- and estimation technology • Estimation technology for breeding value estimation • The significance of artificial insemination (AI) for breeding value estimation • Transgenesis and transgenic animals • Application of DNA-markers for disease genes • Finding disease genes or QTL‘s

The most important calculation methods for breeding value estimation • • Selection index (SI) Best Linear Prediction (BLP) Best Linear Unbiased Prediction (BLUP) Animal Model (AM)

The significance of artificial insemination for breeding value estimation Preconditions for breeding value estimations • Standardised environment • Test station or correction for environment • AI, the precondition for equality, as a bull or a boar are sire of offspring in many different herds

The significance of artificial insemination for breeding value estimation • • Accurate estimation (large families) Independent of ownership Comparable Increased selection intensity and accuracy r. IA by AI and super ovulation

Transgenese and transgenic animals • • Gene constructs Methods for gene transfer Production motives for transgenesis. Other motives for transgenesis

Gene constructs • Promoter (regulator-sequence) • Structure gene (DNA-code for a protein)
Methods of gene transfer • Micro injection of DNA in male pronucleus • Embryonic stemcells and homologue recombination • Micro injection or other forms of transfer into foetal cells gene

Motives for transgenese? • • Additive effect 4 A New metabolic traits Pharmaceutics Organ donation
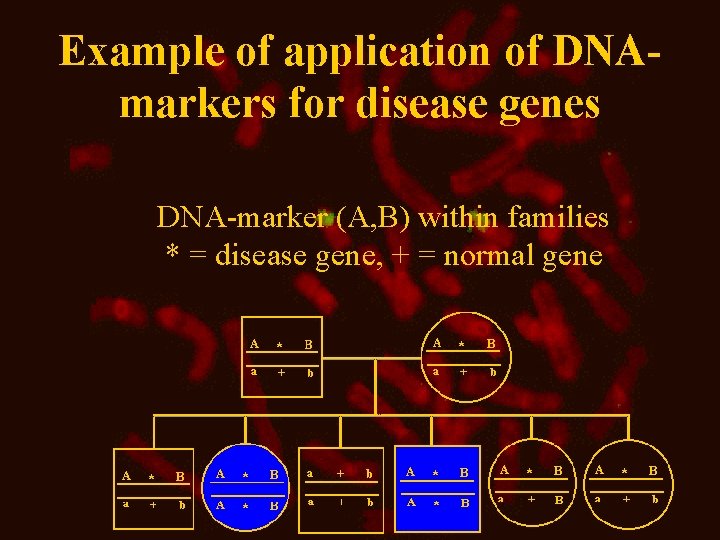
Example of application of DNAmarkers for disease genes DNA-marker (A, B) within families * = disease gene, + = normal gene

Identification of DNA-marker linked to disease genes or QTL‘s • The genome is ca. 3000 centi Morgan (c. M) • A marker covers 20 c. M • 150 DNA-markers are needed to analyse for a given segregation • Ca. half of the markers are informative, so ca. 300 all together have to be applied

Family material suited for identification of a disease gene • • Family size - ca. 50 for an analyse At least 10 to 15 affected The best ratio between affected and normal is 1: 1 The remaining normal are excluded randomly Genotype aa not aa Total affected not affected 20 3 4 23 24 26 Total 23 27 50

Family material suited for identification of QTL’s • • Family size - more than 1000 The existence of QTL’s is unknown Sire or grand-sire design can be applied F 2 animals in relation to exotic crosses

Sire design or Grand sire design QTL estimation • Grand sire design Estimated effect on offspring’s breeding values • Sire design Estimated effect on offspring phenotypes • Marker typing of parents and offspring, in both cases
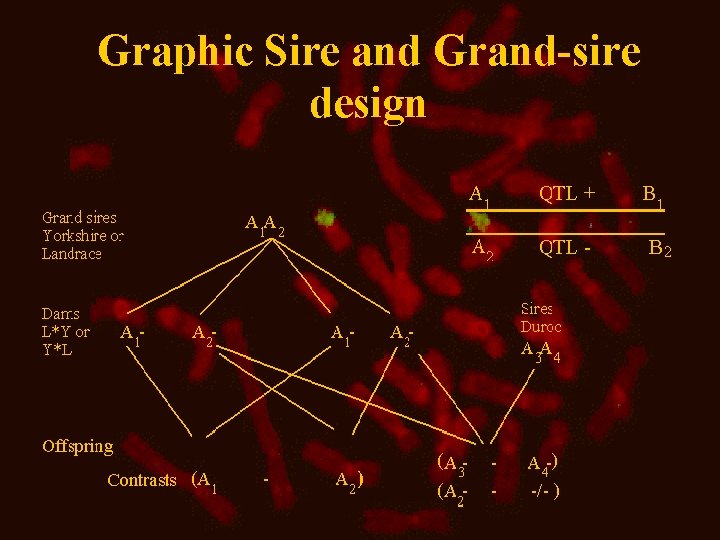
Graphic Sire and Grand-sire design

Sire design or Grand-sire design, test size Least significant difference (LSD) in disease frequency for offspring with A 1 and offspring with A 2 Number A 1 Number A 2 S. D. LSD for disease frequency 5000 500 100 0. 0060 0. 0190 0. 0420 1. 8 % units 5. 7 % units 12. 6 % units 5000 500 100 Number of affected bin(N; 0. 1 0. 9/N)

Results of selection experiments for disease resistance Successfull selection experiments in layers for resistance to neo plasma fatalities, Cole and Hutt (1973) Successfull selection for high and low leukocyte count in mice, Chai (1975)
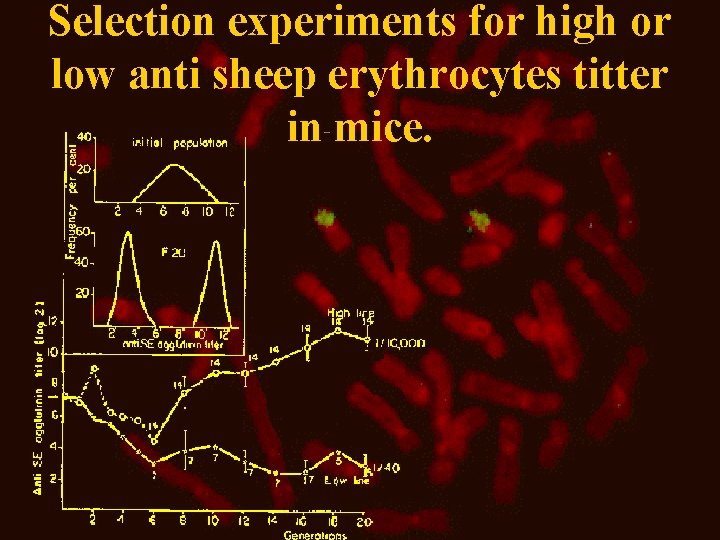
Selection experiments for high or low anti sheep erythrocytes titter in mice.
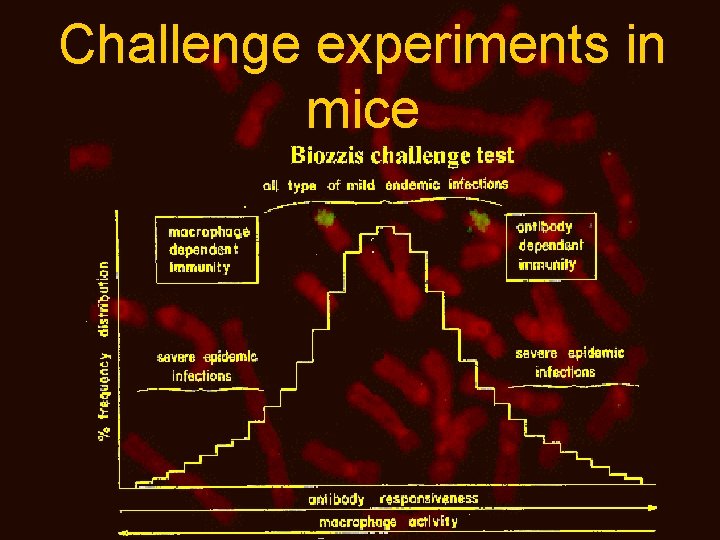
Challenge experiments in mice

Conclusion of experiments for general disease resistance Breeding measures to healthy animals Use crosses but avoid F 2 and four-way High uniformity in production
- Slides: 19