Chapter 10 The Mutability and Repair of DNA

Chapter 10 The Mutability and Repair of DNA

DNA has a special need for metabolic stability • Ideally, DNA information must be transmitted virtually intact during cell division or reproduction of an organism. • Mutations may disturb the intact of DNA information • Inaccuracy in DNA replication • Chemical damage to the DNA • Insertions by transposons. • To maintain the intact of DNA • Keep replication process in high accuracy. • “DNA repair” mechanisms in correcting genetic information.

In this chapter: Following questions to be addressed • How is DNA mended rapidly to prevent errors? • How cell distinguish parental / daughter strand in repairing replication errors? • How cells restore break or lesion DNA sequences even the original sequence can no longer be read? • How cells cope with lesions that block replication?
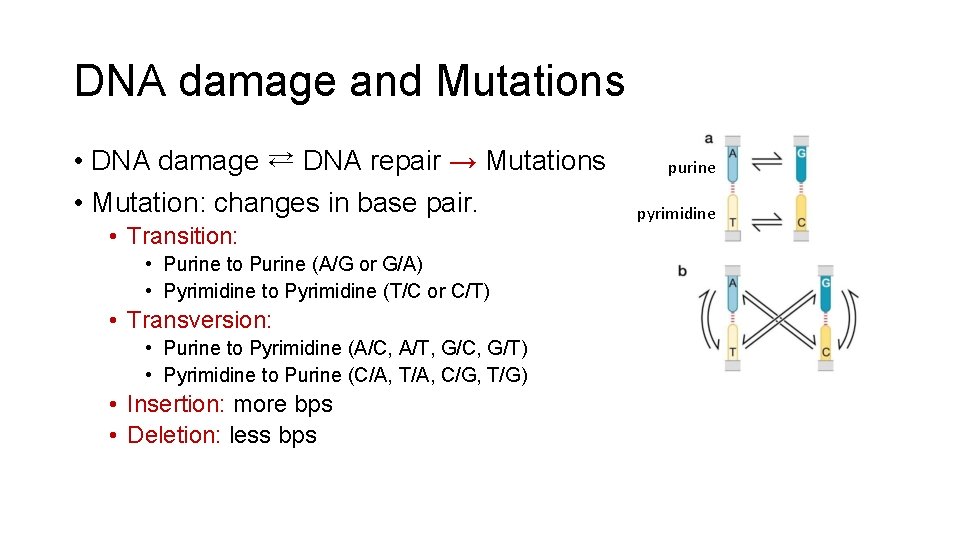
DNA damage and Mutations • DNA damage ⇄ DNA repair → Mutations • Mutation: changes in base pair. • Transition: • Purine to Purine (A/G or G/A) • Pyrimidine to Pyrimidine (T/C or C/T) • Transversion: • Purine to Pyrimidine (A/C, A/T, G/C, G/T) • Pyrimidine to Purine (C/A, T/A, C/G, T/G) • Insertion: more bps • Deletion: less bps purine pyrimidine

The cause of Mutation • Spontaneous reaction: damages caused by endogenous reactions in organic compound. • Genotoxicity: the property of chemical agents that damages the genetic information within a cell causing mutations. • Base modifications by alkylating agents • Pyrimidine dimers by UV radiation • DNA microsatellites slippage during replication DNA microsatellites slippage
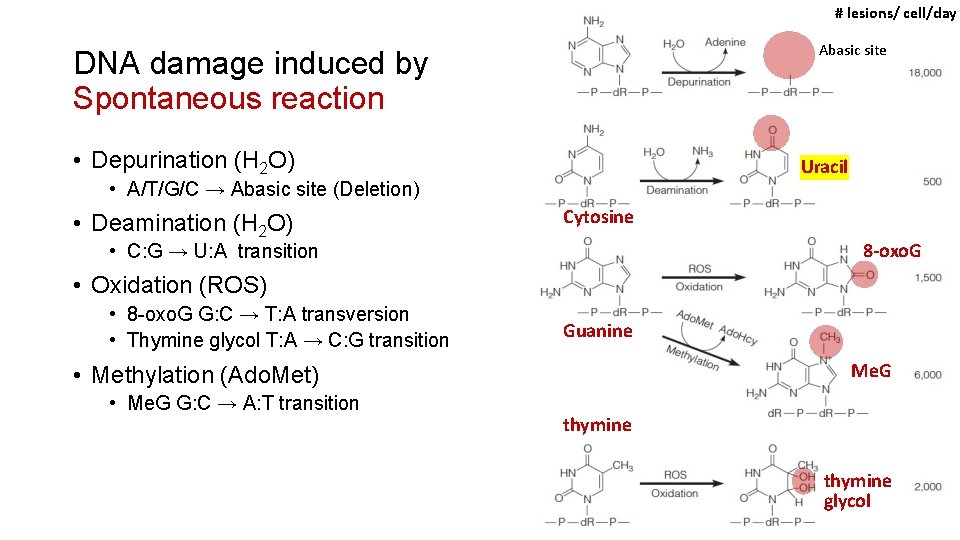
# lesions/ cell/day Abasic site DNA damage induced by Spontaneous reaction • Depurination (H 2 O) Uracil • A/T/G/C → Abasic site (Deletion) • Deamination (H 2 O) Cytosine 8 -oxo. G • C: G → U: A transition • Oxidation (ROS) • 8 -oxo. G G: C → T: A transversion • Thymine glycol T: A → C: G transition Guanine Me. G • Methylation (Ado. Met) • Me. G G: C → A: T transition thymine glycol
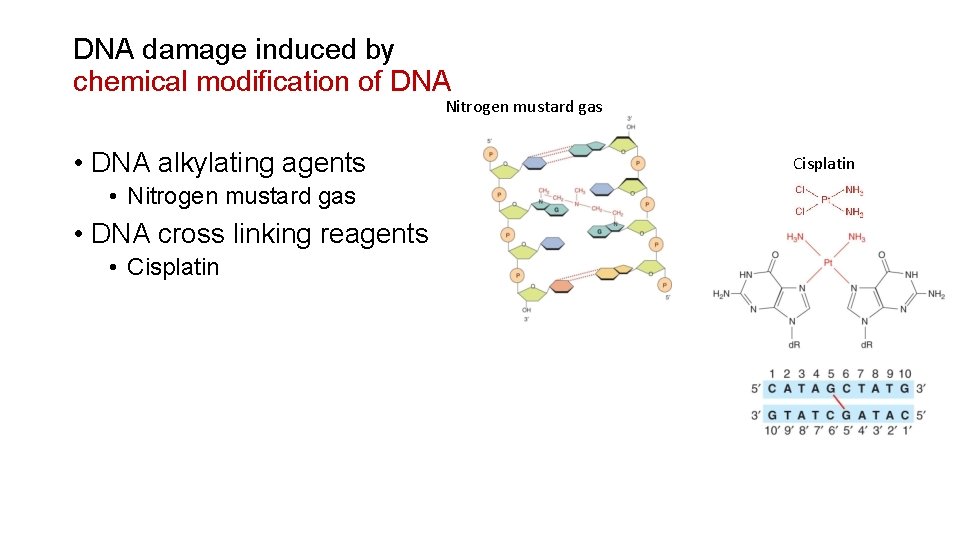
DNA damage induced by chemical modification of DNA Nitrogen mustard gas • DNA alkylating agents • Nitrogen mustard gas • DNA cross linking reagents • Cisplatin
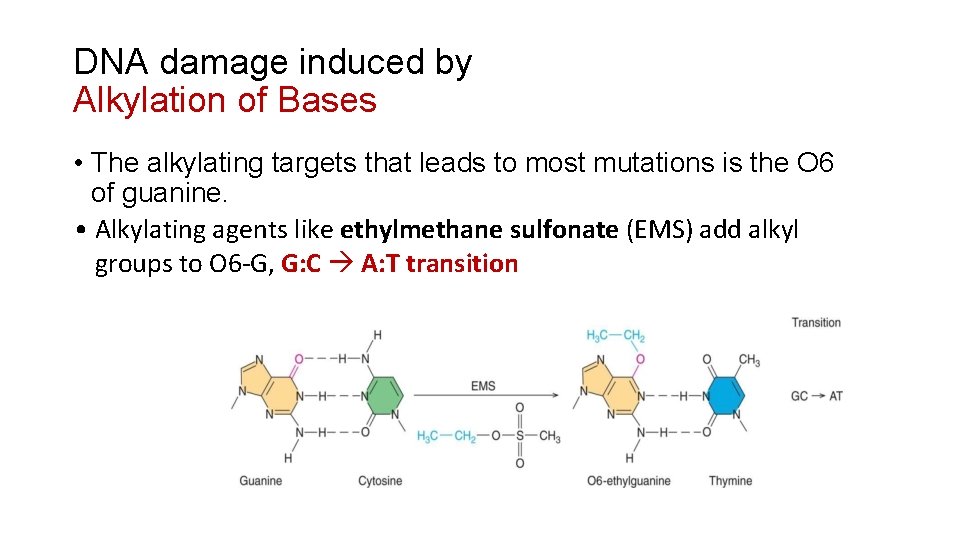
DNA damage induced by Alkylation of Bases • The alkylating targets that leads to most mutations is the O 6 of guanine. • Alkylating agents like ethylmethane sulfonate (EMS) add alkyl groups to O 6 -G, G: C A: T transition

DNA damage induced by Environmental toxin • Bulky electrophilic agents • Polycyclic aromatic hydrocarbons (PAH) such as Benzo[a]pyrene and Aflatoxins (黃麴毒素) must be activated by P 450 system to damage DNA
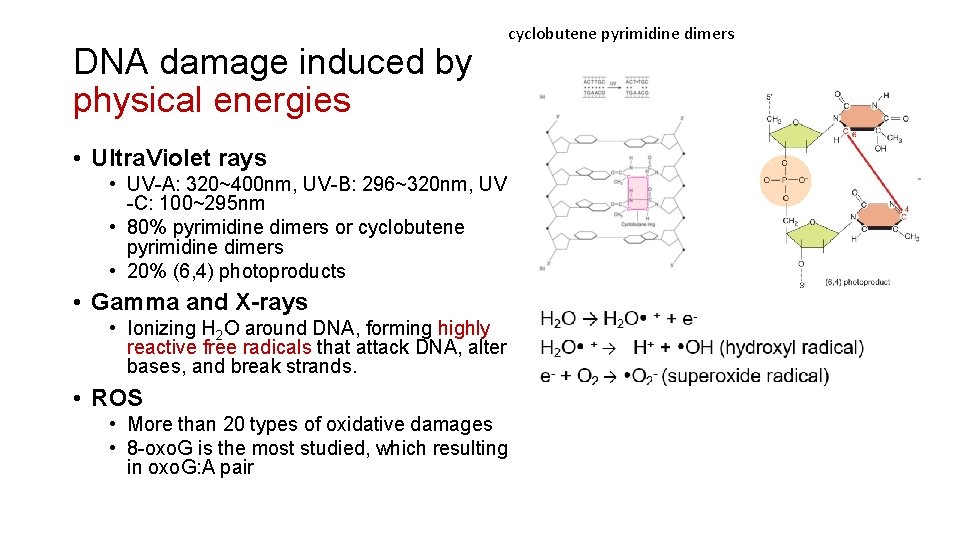
DNA damage induced by physical energies cyclobutene pyrimidine dimers • Ultra. Violet rays • UV-A: 320~400 nm, UV-B: 296~320 nm, UV -C: 100~295 nm • 80% pyrimidine dimers or cyclobutene pyrimidine dimers • 20% (6, 4) photoproducts • Gamma and X-rays • Ionizing H 2 O around DNA, forming highly reactive free radicals that attack DNA, alter bases, and break strands. • ROS • More than 20 types of oxidative damages • 8 -oxo. G is the most studied, which resulting in oxo. G: A pair

Mechanisms to cope with DNA damage • Precise repair • Direct repair: restore the reaction. • Base excision repair: remove and renew the damaged base • Nucleotide excision repair: remove a section of damaged DNA, replaced with normal one. • Mismatch-repair: recognize and correct mismatched DNA created by replication errors. • Homologous recombination • Coping with DNA in emergency • Daughter-strand gap repair: newly replicated DNA duplexes undergo genetic recombination • Translesion synthesis / Error-prone repair: roughly bypass DNA damages that blocks DNA polymerase replication. • Double-strand break repair: rejoin the break ds. DNA randomly.
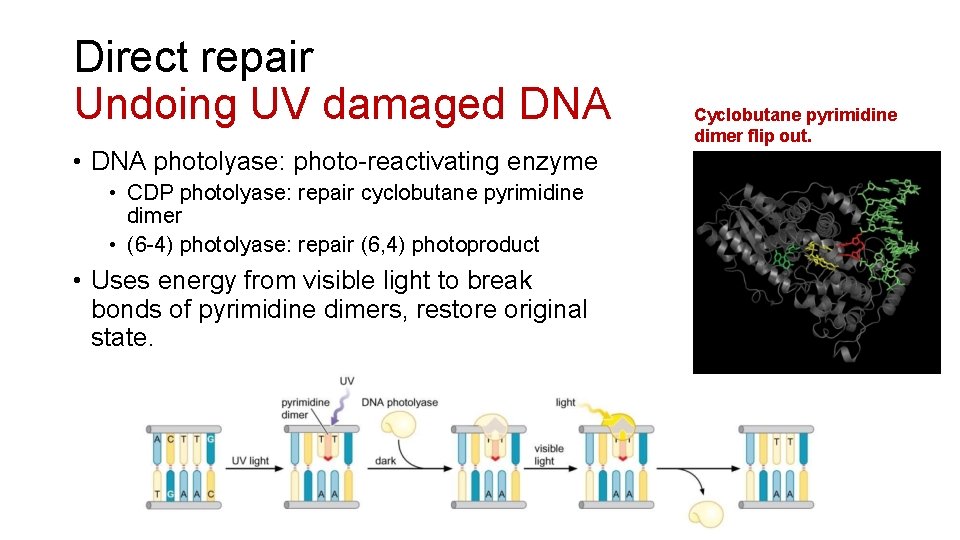
Direct repair Undoing UV damaged DNA • DNA photolyase: photo-reactivating enzyme • CDP photolyase: repair cyclobutane pyrimidine dimer • (6 -4) photolyase: repair (6, 4) photoproduct • Uses energy from visible light to break bonds of pyrimidine dimers, restore original state. Cyclobutane pyrimidine dimer flip out.

Direct repair Undoing methylated or alkylated DNA • O 6 -methylguanine alkyltransferase: • Removes alkyl group from the methylated site using Cys-SH residue. • Alkylated alkyltransferase: • Gene expression be induced by alkylation. • Specific transcriptional activator of its own gene • Allows adaption, cells could be more resistant to DNA alkylation damages.
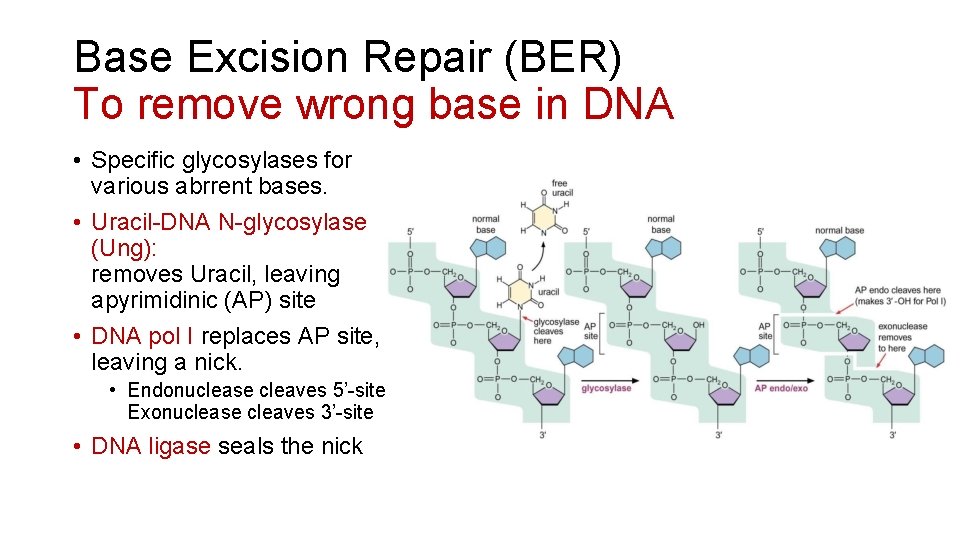
Base Excision Repair (BER) To remove wrong base in DNA • Specific glycosylases for various abrrent bases. • Uracil-DNA N-glycosylase (Ung): removes Uracil, leaving apyrimidinic (AP) site • DNA pol I replaces AP site, leaving a nick. • Endonuclease cleaves 5’-site Exonuclease cleaves 3’-site • DNA ligase seals the nick
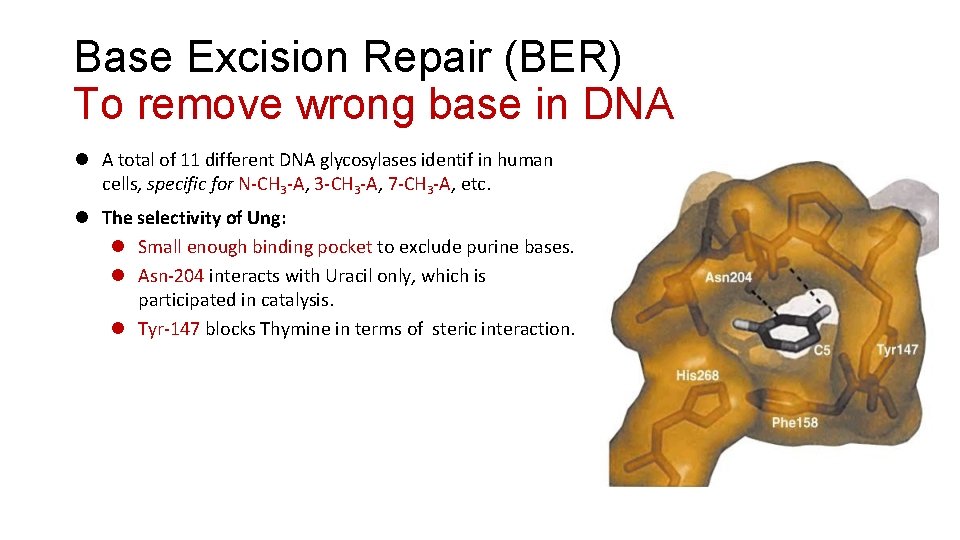
Base Excision Repair (BER) To remove wrong base in DNA A total of 11 different DNA glycosylases identif in human cells, specific for N-CH 3 -A, 3 -CH 3 -A, 7 -CH 3 -A, etc. The selectivity of Ung: Small enough binding pocket to exclude purine bases. Asn-204 interacts with Uracil only, which is participated in catalysis. Tyr-147 blocks Thymine in terms of steric interaction.

Base Excision Repair (BER) To remove wrong base in DNA • The most abundant DNA oxidation product: 8 -oxo. G: A • Three specific BER enzyme for 8 oxo. G: • Level 1 • Mut. T: Hydrolyzes 8 -oxo. GTP • Level 2 (The fail-safe glycosylase system) • Mut. Y: Excises A from the wrong paired 8 -oxo. G: A pair. • Level 3 • Mut. M: excise oxo. G from 8 -oxo. G: C pair

Nucleotide Excision Repair (NER) To remove drastic helix distorting • Specialized nucleases for versatile purposes • In E. coli • uvr. A-B complex: • ATP hydrolysis promotes formation of Uvr. A dimer, complexed with Uvr. B dimer, scans DNA to identify a distortion • uvr. B-C complex: (Excinuclease) • Once the distortion be scanned, uvr. B dimer left on the distortion, one uvr. B leave and recruit uvr. C, uvr. C creates nicks at 3’ and 5’ to the lesion (12 -13 bases) • uvr. D DNA Helicase: • releases ss-fragment from duplex • DNA pol II or I / DNA ligase repair and seal

Nucleotide Excision Repair (NER) To remove drastic helix distorting • In Eukaryotics: far more complicated system • Global genome-NER (GG-NER) • Distortion detected by XPC, • recruit XPE-complex (UV-DDB) • Transcriptional coupled NER (TC-NER) • Distortion encountered by RNA pol, • recruit CSA/B complex • XPA: verification • XPF: endonuclease for 5’ end • XPG: endonuclease for 3’ end • DNA pol ε/κ/δ and DNA ligase: repair and seal

Nucleotide Excision Repair (NER) To remove drastic helix distorting • Mutations in XPA~XPG: • Xeroderman pigmentosum (XP) • > 1000 fold incidence of skin cancer upon sun exposure • Mutations in CS • TC-NER affected • Cockayne’s syndrome (CS) • Retarded growth, neurological dysfunction, and photosensitivity.
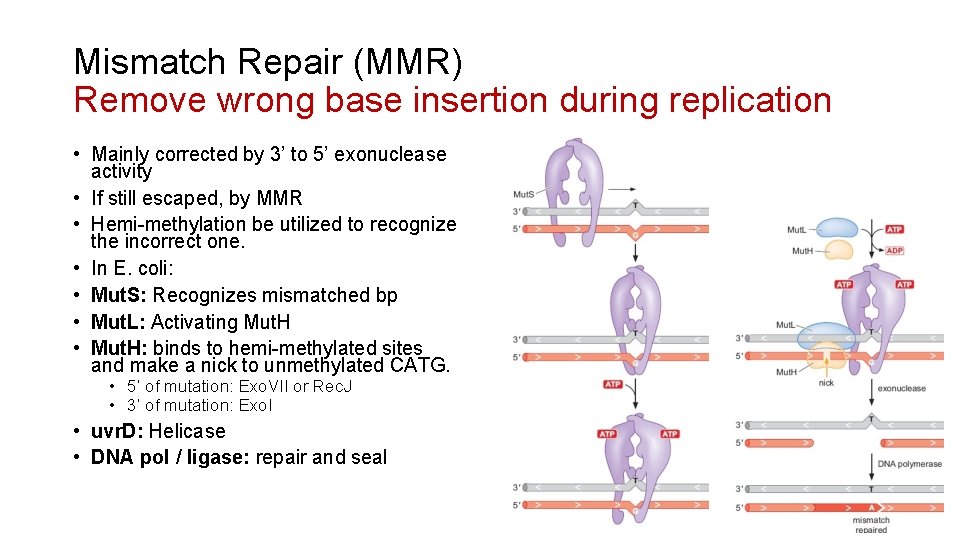
Mismatch Repair (MMR) Remove wrong base insertion during replication • Mainly corrected by 3’ to 5’ exonuclease activity • If still escaped, by MMR • Hemi-methylation be utilized to recognize the incorrect one. • In E. coli: • Mut. S: Recognizes mismatched bp • Mut. L: Activating Mut. H • Mut. H: binds to hemi-methylated sites and make a nick to unmethylated CATG. • 5’ of mutation: Exo. VII or Rec. J • 3’ of mutation: Exo. I • uvr. D: Helicase • DNA pol / ligase: repair and seal
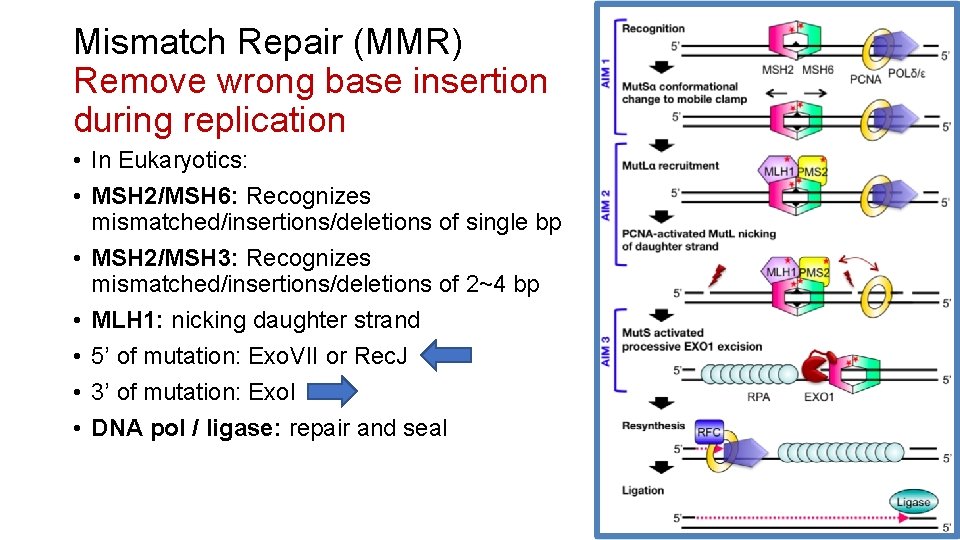
Mismatch Repair (MMR) Remove wrong base insertion during replication • In Eukaryotics: • MSH 2/MSH 6: Recognizes mismatched/insertions/deletions of single bp • MSH 2/MSH 3: Recognizes mismatched/insertions/deletions of 2~4 bp • MLH 1: nicking daughter strand • 5’ of mutation: Exo. VII or Rec. J • 3’ of mutation: Exo. I • DNA pol / ligase: repair and seal

Mismatch Repair (MMR) Remove wrong base insertion during replication • Mutations in MMR proteins: • HNPCC (Heritable nonpolyposis colon cancer), or Lynch syndrome • 1/200 American affected, counts for 15% of colon cancers • Mostly, h. MLH 1 and h. MSH 2 mutation • Microsatellite instability: Large mutations in regions of repeating units of 1~4 nts, increased in number of repeats during patients’ lifetime. • Caused by unrepaired mismatch bubbles created by slippage in DNA replication.

Double strand break repair • Double strand break: the most dangerous DNA damage, breaking chromosomes. • About 50 times per cell cycle, cell death or cancer if not repaired. • Eukaryotes deal with DSB in 2 ways • Homologous recombination (HR): repair precisely, happened in S/G 2 phase, use unbroken sister chromatid as partner to repair. • Non-Homologous End-Joining (NHEJ): repair efficiently but result in errors, happened in G 1 phase, no homologous chromosome for reference.
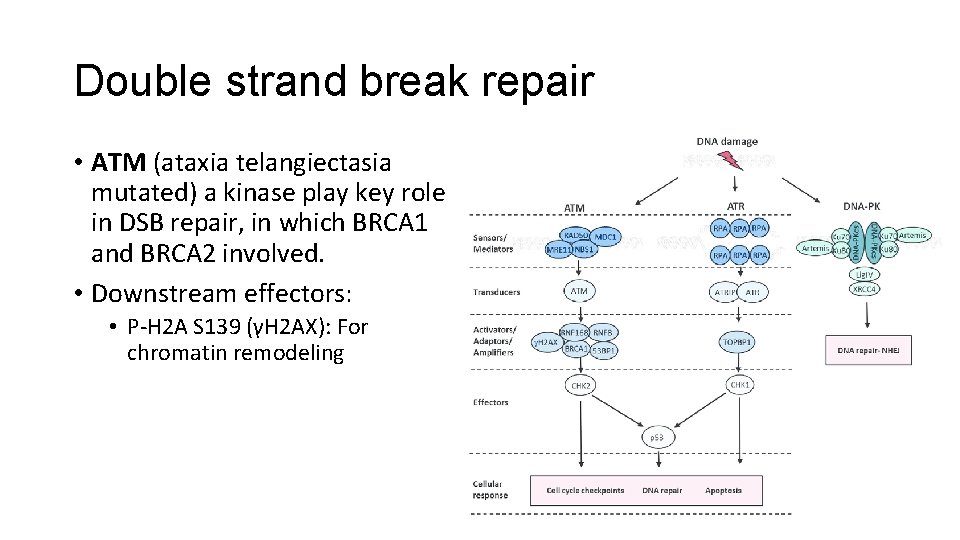
Double strand break repair • ATM (ataxia telangiectasia mutated) a kinase play key role in DSB repair, in which BRCA 1 and BRCA 2 involved. • Downstream effectors: • P-H 2 A S 139 (γH 2 AX): For chromatin remodeling

Homologous Recombination Repair DSB precisely using template • ATM kinase signaling • Phosphorylation activated Nuclease in MRN complex (Mre 11 -Rad 50 -Nbs 1) and downstream effectors H 2 AX, BRCA 1 • MRN nuclease trim away 5’-end, leaving 3’ single-strand ends, coated by RPA. • 3’ ss end activated ATR to initiate more signaling • Rad 51 replaces RPA, aided by BRCA 2. • Strand invasion of undamaged homologous chromosome. • DNA synthesis and cutting • Yield either recombinant or non-recombinant chromosomes.

Non. Homologous End-Joining (NHEJ) Repair DSB without template • Ku 70/80 dimer: form ring-like circle around DNA ends. • 2 DNA-PKCS: be recruited and phosphorylate each other. • Artemis: recruited by ATM signaling, then activated by DNA-PK • DNA polymerase and ligase: gap filled and ligated. DNA ligase

Coping with DNA Damage Without Repairing • Daughter-Strand Gap Repair • Regression of Replication Fork • Translesion Synthesis (Error-prone bypass)

Daughter-Strand Gap Repair Damage bypass mechanism for errors on lagging strand • Direct reversal and excision repair: true repair processes, short-term metabolic responses. • However, if photoreactivation and excision repair systems defective, or saturated by DNA damage too extensive, longer- term systems come into play. • Replicative polymerase encounter a lesion, leaving a gap. • The undamaged parental strand transferred to the gap, mediated by Rec. A. • The gapped daughter strand transferred to the position where parental strand across from. • Recombination mediated by Ruv. ABC • Refilled by DNA polymerase, ligase. • However, the original damage still unrepaired.

Replication fork regression Damage bypass mechanism for errors on leading strand • (a) Replication fork encounter a lesion in leading strand • (b) Regression of replication fork pairs the damaged site with undamaged template (green). • (c) The daughter–strand complement of the damaged strand (black) can then be copied from the other daughter strand (green). • (d) Bypass the lesion site lesion in leading template regression
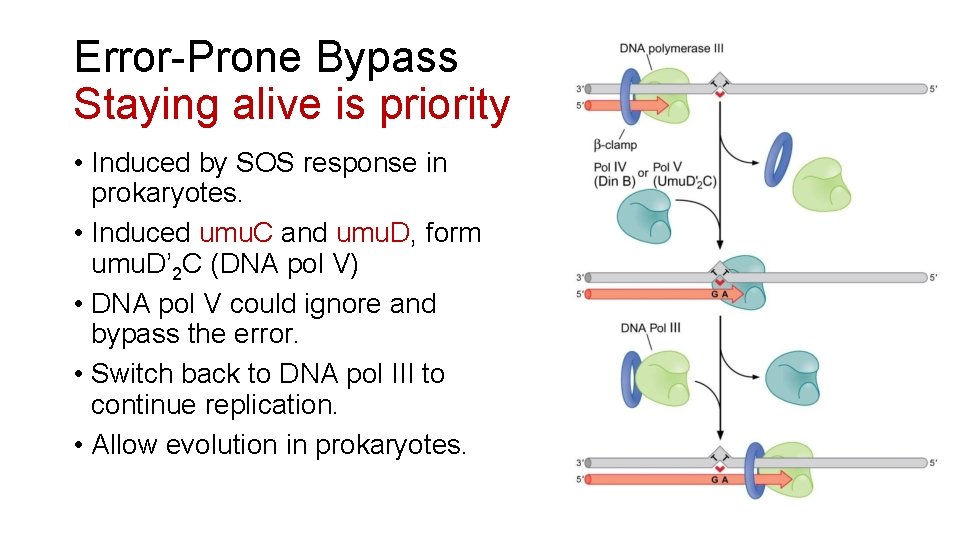
Error-Prone Bypass Staying alive is priority • Induced by SOS response in prokaryotes. • Induced umu. C and umu. D, form umu. D’ 2 C (DNA pol V) • DNA pol V could ignore and bypass the error. • Switch back to DNA pol III to continue replication. • Allow evolution in prokaryotes.
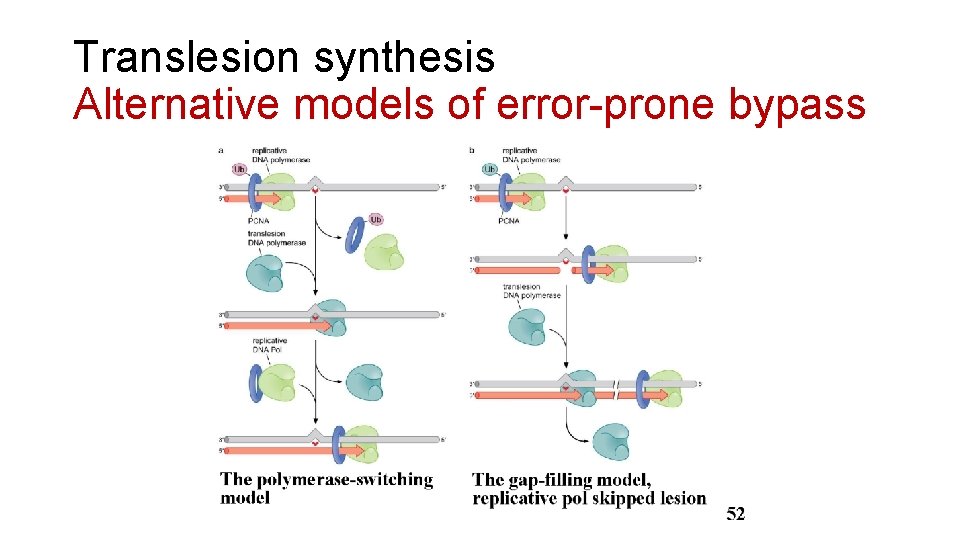
Translesion synthesis Alternative models of error-prone bypass
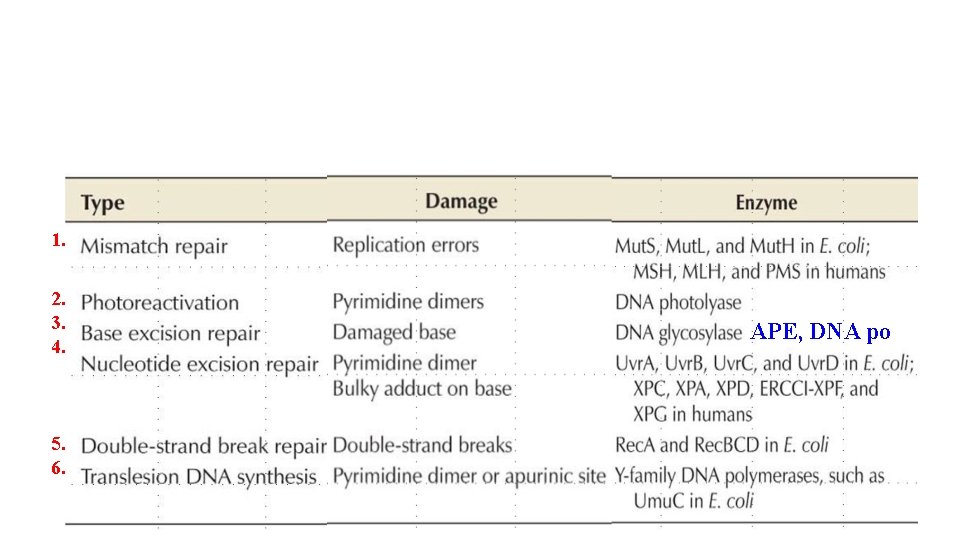
1. 2. 3. 4. 5. 6. APE, DNA po
- Slides: 32