Chaperones CA Garca Seplveda MD Ph D Laboratorio

Chaperones. CA García Sepúlveda MD Ph. D Laboratorio de Genómica Viral y Humana Facultad de Medicina, Universidad Autónoma de San Luis Potosí 1
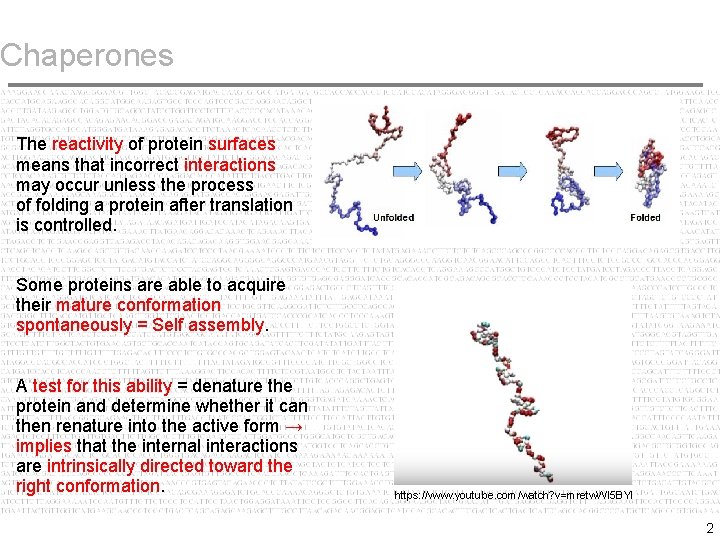
Chaperones The reactivity of protein surfaces means that incorrect interactions may occur unless the process of folding a protein after translation is controlled. Some proteins are able to acquire their mature conformation spontaneously = Self assembly. A test for this ability = denature the protein and determine whether it can then renature into the active form → implies that the internal interactions are intrinsically directed toward the right conformation. https: //www. youtube. com/watch? v=mretw. Wl 5 BYI 2
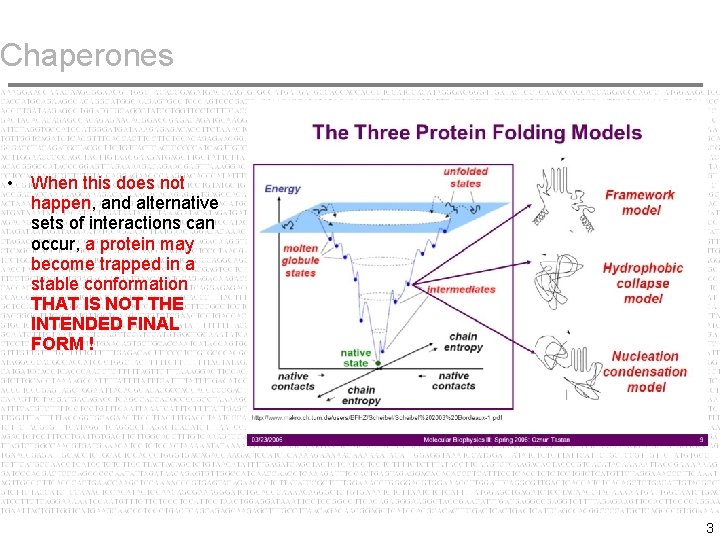
Chaperones • When this does not happen, and alternative sets of interactions can occur, a protein may become trapped in a stable conformation THAT IS NOT THE INTENDED FINAL FORM ! 3

Chaperones • Proteins in this category cannot self-assemble. Their acquisition of proper structure requires the assistance of a chaperone. • Chaperones are proteins that mediate correct assembly by causing a target protein to acquire one possible conformation instead of others. • Accomplished by binding to reactive surfaces in the target protein that are exposed during the assembly process and preventing them from interacting. • Chaperones function by preventing formation of incorrect structures rather than by promoting formation of correct structures. 4
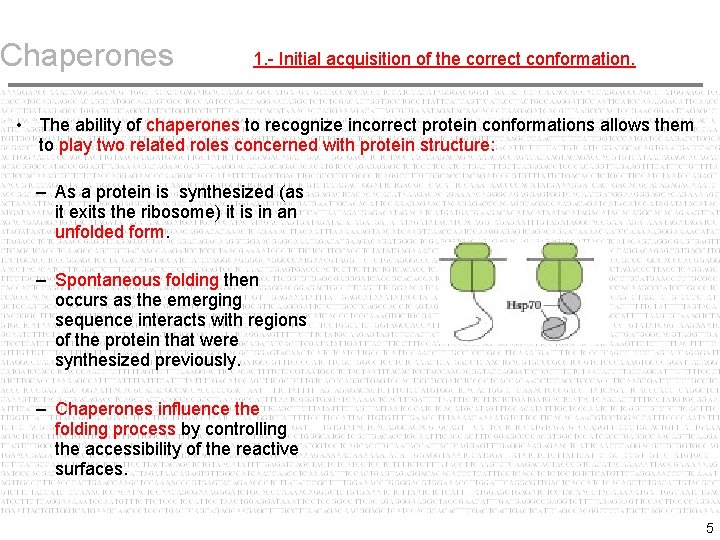
Chaperones 1. - Initial acquisition of the correct conformation. • The ability of chaperones to recognize incorrect protein conformations allows them to play two related roles concerned with protein structure: – As a protein is synthesized (as it exits the ribosome) it is in an unfolded form. – Spontaneous folding then occurs as the emerging sequence interacts with regions of the protein that were synthesized previously. – Chaperones influence the folding process by controlling the accessibility of the reactive surfaces. 5
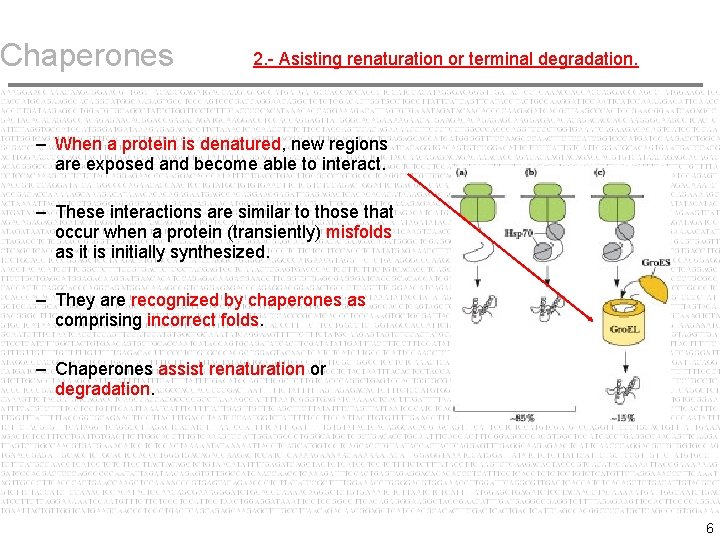
Chaperones 2. - Asisting renaturation or terminal degradation. – When a protein is denatured, new regions are exposed and become able to interact. – These interactions are similar to those that occur when a protein (transiently) misfolds as it is initially synthesized. – They are recognized by chaperones as comprising incorrect folds. – Chaperones assist renaturation or degradation. 6
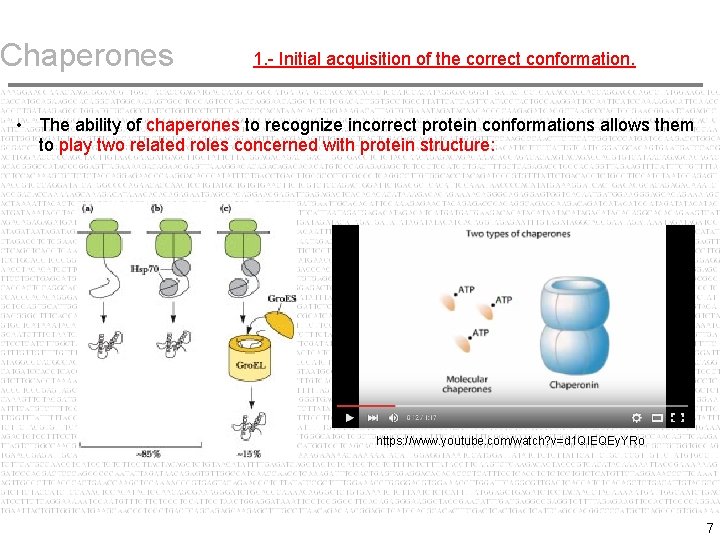
Chaperones 1. - Initial acquisition of the correct conformation. • The ability of chaperones to recognize incorrect protein conformations allows them to play two related roles concerned with protein structure: https: //www. youtube. com/watch? v=d 1 QIEQEy. YRo 7
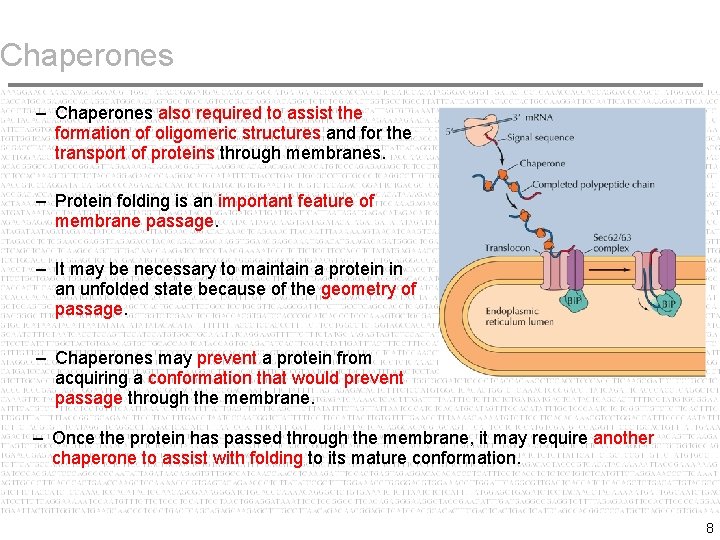
Chaperones – Chaperones also required to assist the formation of oligomeric structures and for the transport of proteins through membranes. – Protein folding is an important feature of membrane passage. – It may be necessary to maintain a protein in an unfolded state because of the geometry of passage. – Chaperones may prevent a protein from acquiring a conformation that would prevent passage through the membrane. – Once the protein has passed through the membrane, it may require another chaperone to assist with folding to its mature conformation. 8
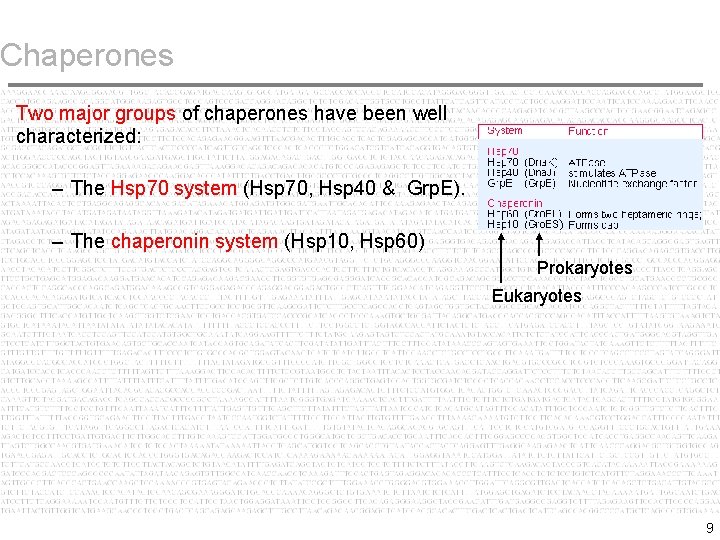
Chaperones Two major groups of chaperones have been well characterized: – The Hsp 70 system (Hsp 70, Hsp 40 & Grp. E). – The chaperonin system (Hsp 10, Hsp 60) Prokaryotes Eukaryotes 9
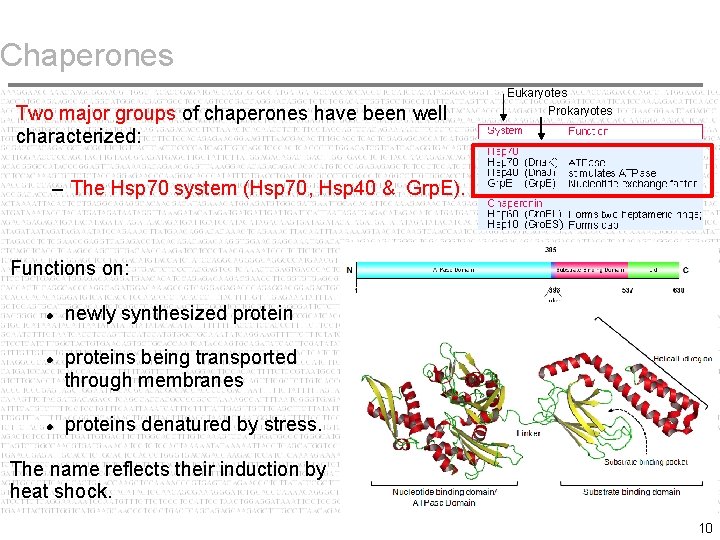
Chaperones Eukaryotes Two major groups of chaperones have been well characterized: Prokaryotes – The Hsp 70 system (Hsp 70, Hsp 40 & Grp. E). Functions on: newly synthesized proteins being transported through membranes proteins denatured by stress. The name reflects their induction by heat shock. 10
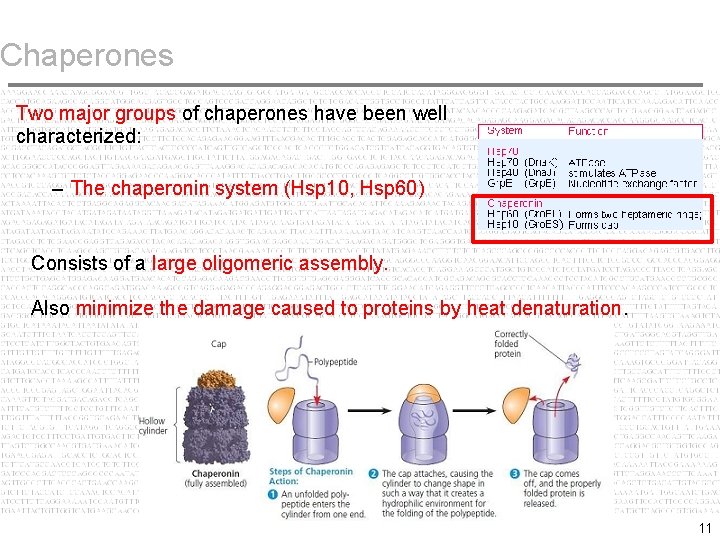
Chaperones Two major groups of chaperones have been well characterized: – The chaperonin system (Hsp 10, Hsp 60) Consists of a large oligomeric assembly. Also minimize the damage caused to proteins by heat denaturation. 11

The Hsp 70 System – The Hsp 70 system (Hsp 70, Hsp 40 & Grp. E). – The Hsp 70 family is found in bacteria, eukaryotic cytosol, in the ER & in chloroplasts/mitochondria. – Members help protein folding by multiple cycles of association and dissociation to nascent peptide. Prokaryotes Eukaryotes – Hsp 70 functions in conjunction with two further components, best characterized in bacteria as Dna. J and Grp. E. 12
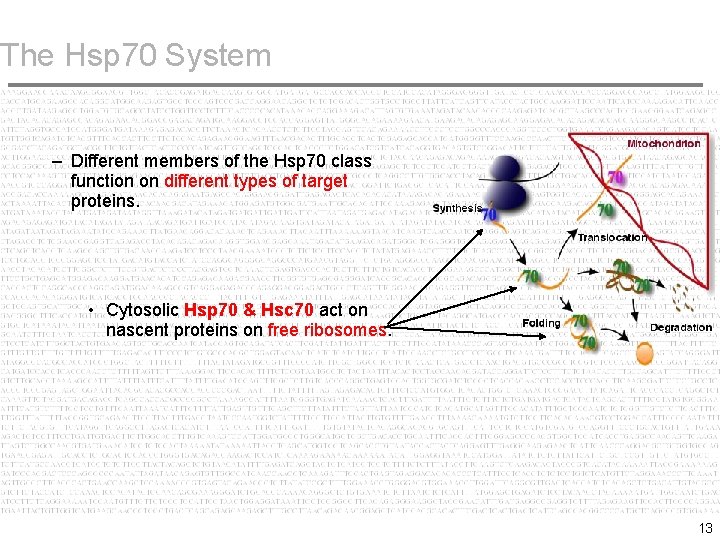
The Hsp 70 System – Different members of the Hsp 70 class function on different types of target proteins. • Cytosolic Hsp 70 & Hsc 70 act on nascent proteins on free ribosomes. 13
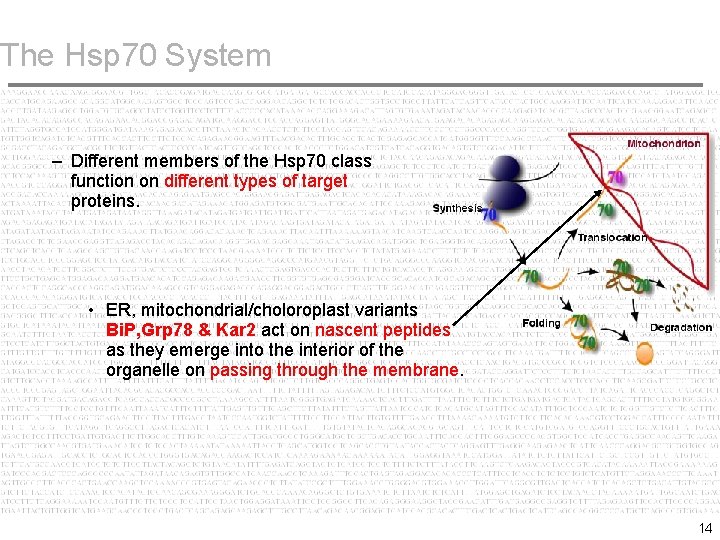
The Hsp 70 System – Different members of the Hsp 70 class function on different types of target proteins. • ER, mitochondrial/choloroplast variants Bi. P, Grp 78 & Kar 2 act on nascent peptides as they emerge into the interior of the organelle on passing through the membrane. 14
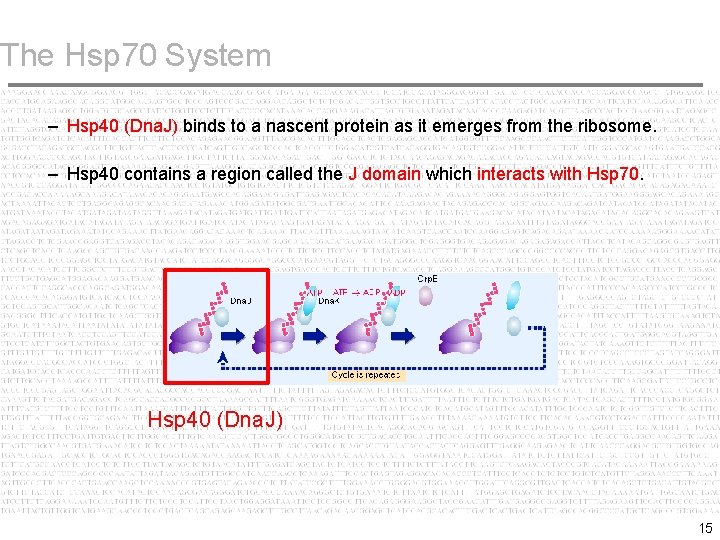
The Hsp 70 System – Hsp 40 (Dna. J) binds to a nascent protein as it emerges from the ribosome. – Hsp 40 contains a region called the J domain which interacts with Hsp 70. Hsp 40 (Dna. J) 15
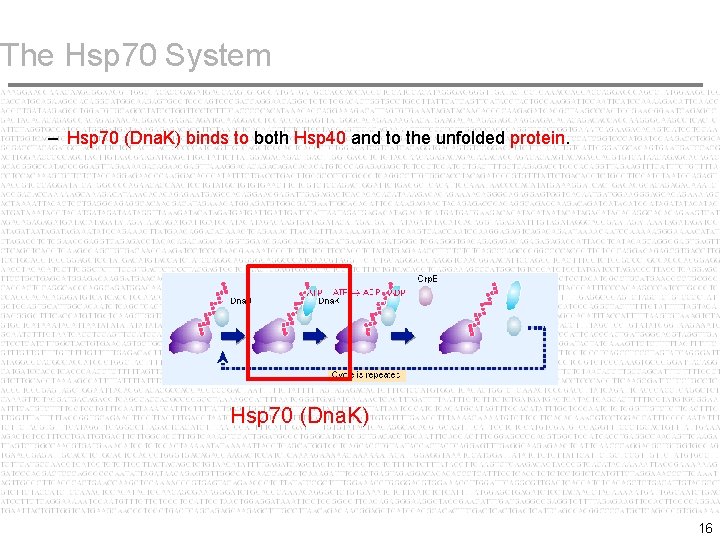
The Hsp 70 System – Hsp 70 (Dna. K) binds to both Hsp 40 and to the unfolded protein. Hsp 70 (Dna. K) 16
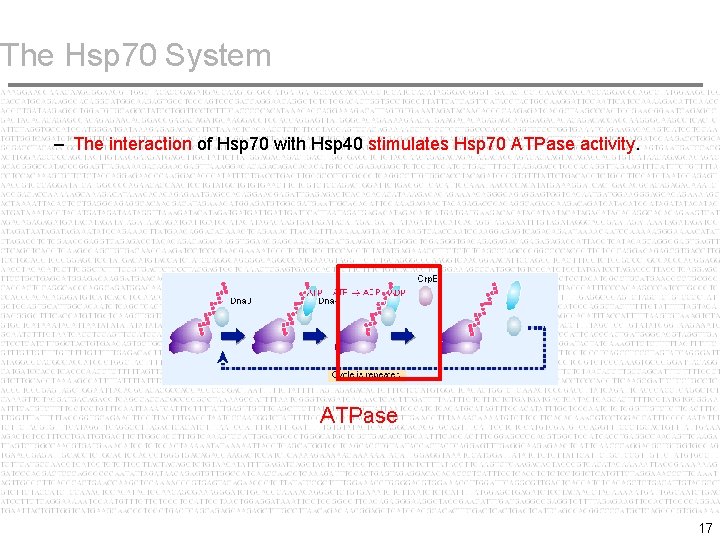
The Hsp 70 System – The interaction of Hsp 70 with Hsp 40 stimulates Hsp 70 ATPase activity. ATPase 17

The Hsp 70 System – The ADP-bound form of the complex remains associated with the protein substrate until Grp. E displaces the ADP. Grp. E displaces ADP 18
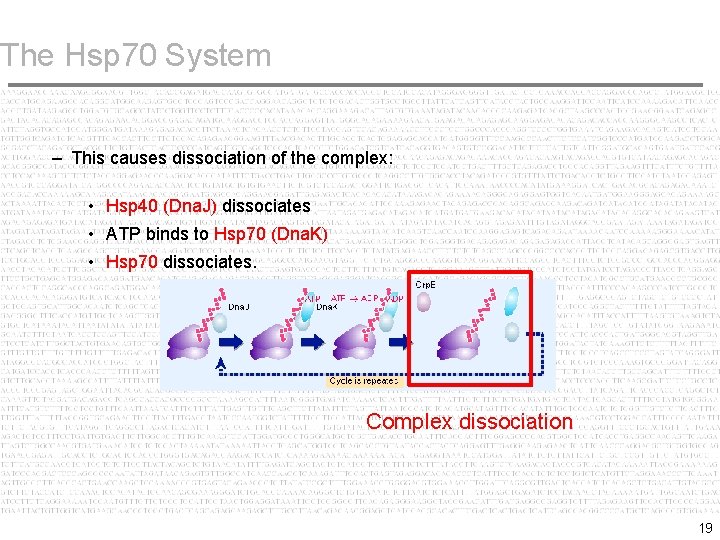
The Hsp 70 System – This causes dissociation of the complex: • Hsp 40 (Dna. J) dissociates • ATP binds to Hsp 70 (Dna. K) • Hsp 70 dissociates. Complex dissociation 19

The Hsp 70 System – As the protein chain lengthens, Hsp 70 (Dna. K) may dissociate from one binding site and reassociate with another. – Thus releasing parts of the substrate protein to fold in an ordered manner. – Finally, the intact protein is released from the ribosome, folded into its mature conformation Protein release 20
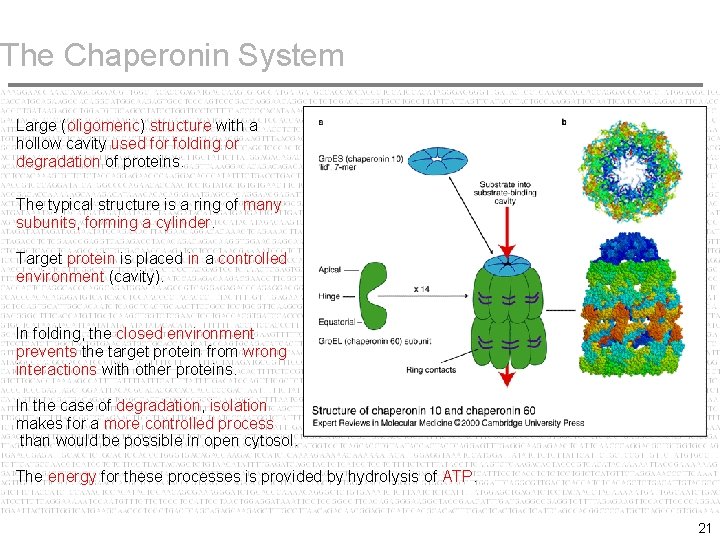
The Chaperonin System Large (oligomeric) structure with a hollow cavity used for folding or degradation of proteins. The typical structure is a ring of many subunits, forming a cylinder. Target protein is placed in a controlled environment (cavity). In folding, the closed environment prevents the target protein from wrong interactions with other proteins. In the case of degradation, isolation makes for a more controlled process than would be possible in open cytosol. The energy for these processes is provided by hydrolysis of ATP. 21
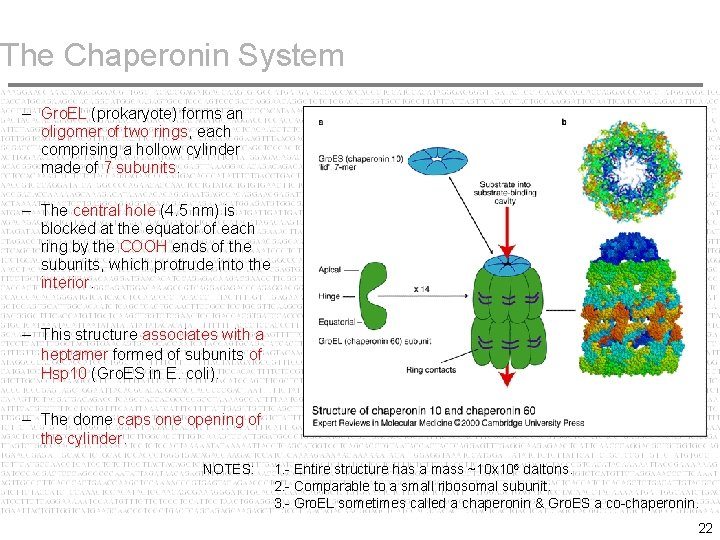
The Chaperonin System – Gro. EL (prokaryote) forms an oligomer of two rings, each comprising a hollow cylinder made of 7 subunits. – The central hole (4. 5 nm) is blocked at the equator of each ring by the COOH ends of the subunits, which protrude into the interior. – This structure associates with a heptamer formed of subunits of Hsp 10 (Gro. ES in E. coli). – The dome caps one opening of the cylinder. NOTES: 1. - Entire structure has a mass ~10 x 106 daltons. 2. - Comparable to a small ribosomal subunit. 3. - Gro. EL sometimes called a chaperonin & Gro. ES a co-chaperonin. 22

The Overview (i. Biology with Arthur Horwich) https: //www. youtube. com/watch? v=j. Oh. Ny. Vjk. Ch. M 23
die Proteinfaltung auf Max Planck Gesselschaft https: //www. youtube. com/watch? v=n. EHe 3 Aie 9 Ek 24
- Slides: 24