Ch IPSeq Analysis Simon Andrews simon andrewsbabraham ac

Ch. IP-Seq Analysis Simon Andrews simon. andrews@babraham. ac. uk @simon_andrews v 2018 -07

What this course covers • • • The theory of Ch. IP-Seq library properties Sequencing, Data processing and QC Data visualisation and exploration Types of analysis – Peak Calling – Differential Binding

What is Ch. IP-Seq is a technology which uses high-throughput sequencing to infer the positions of any mark associated with DNA which can be captured by an antibody.

Types of antibody • Transcription factors / repressors – nanog, CTCF • Histones and histone modifications – H 3, H 3 K 4 me 3 • DNA modifications – Methyl-Cytosine, Formyl cytosine • Chromatin remodelling proteins – BMI 1, EZH 2 • Transcription machinery – Pol 2

Related Techniques • ATAC-Seq – Uses transposases to digest exposed DNA to enrich for accessible DNA. • Cut and Run – Uses transposes fused to antibodies to find marked, accessible chromatin • Dam. ID/Dam. IP – Fuses a methyltransferase to a protein then measures methyl. Adenine by bisulphite seq (Dam. ID) or m. A Ch. IP (Dam. IP)
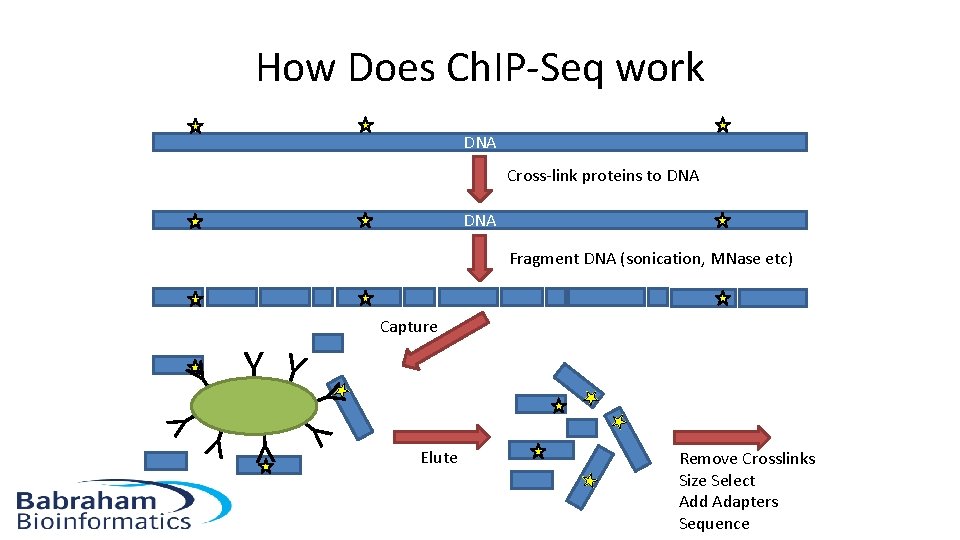
How Does Ch. IP-Seq work DNA Cross-link proteins to DNA Fragment DNA (sonication, MNase etc) Capture Y YY Y Elute Remove Crosslinks Size Select Add Adapters Sequence Y Y
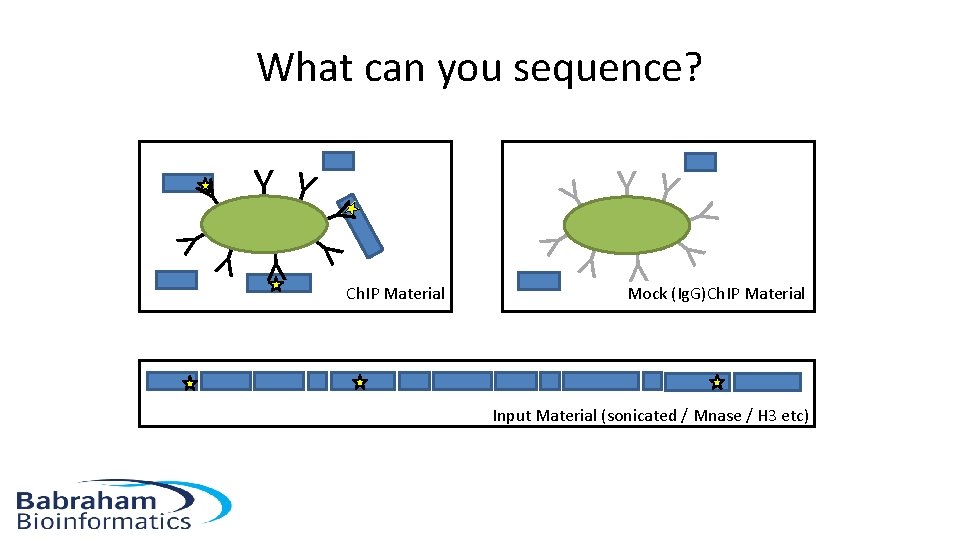
What can you sequence? Y Y Ch. IP Material Y YY YY Y Mock (Ig. G)Ch. IP Material Input Material (sonicated / Mnase / H 3 etc) Y Y
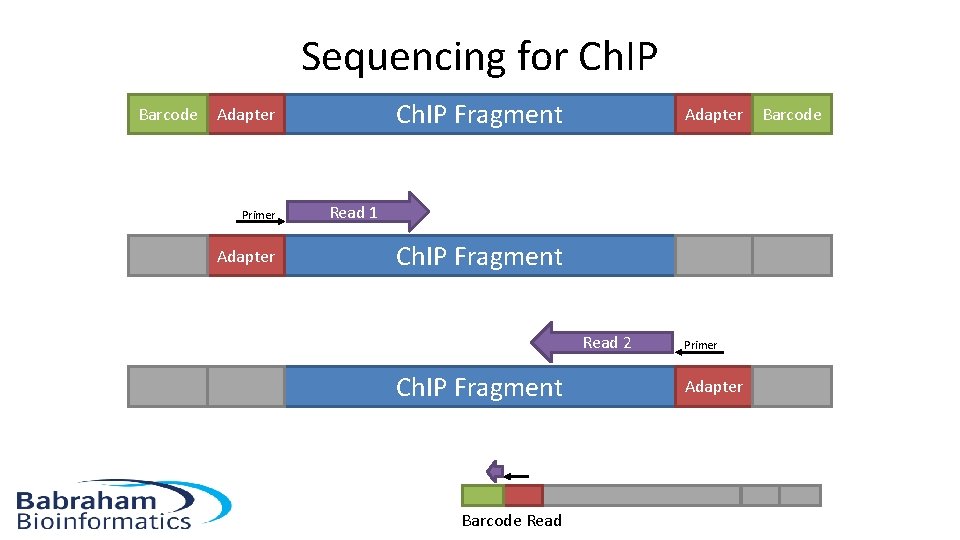
Sequencing for Ch. IP Barcode Ch. IP Fragment Adapter Primer Adapter Read 1 Ch. IP Fragment Read 2 Ch. IP Fragment Barcode Read Primer Adapter Barcode
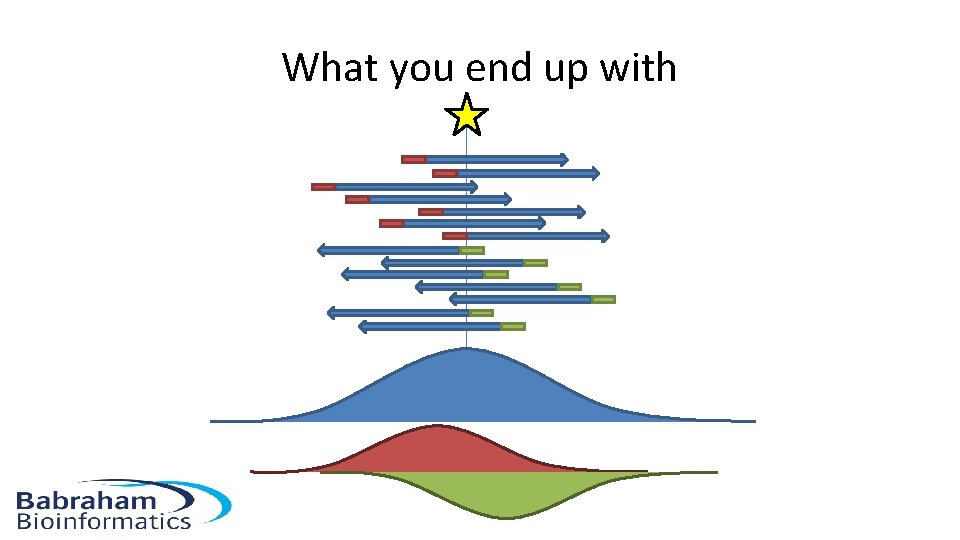
What you end up with
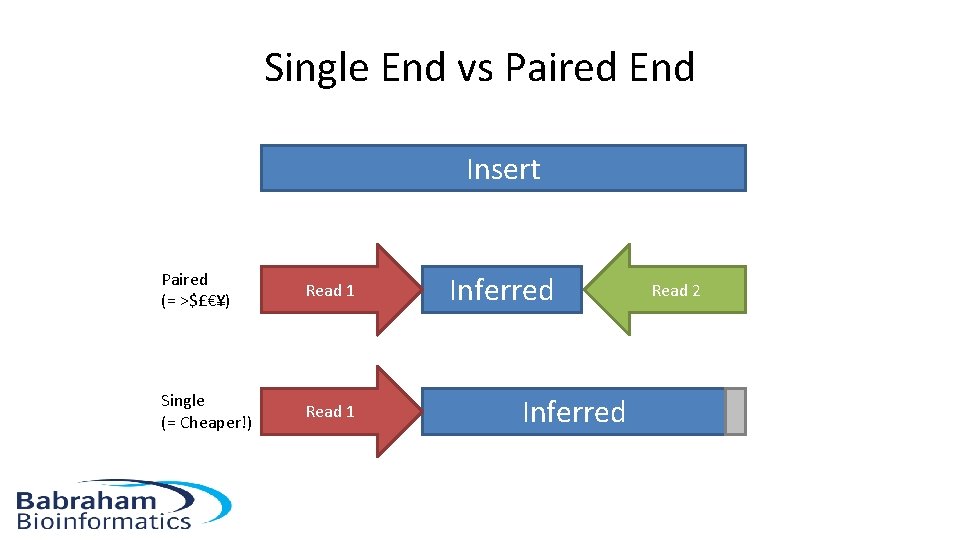
Single End vs Paired End Insert Paired (= >$£€¥) Read 1 Single (= Cheaper!) Read 1 Inferred Read 2
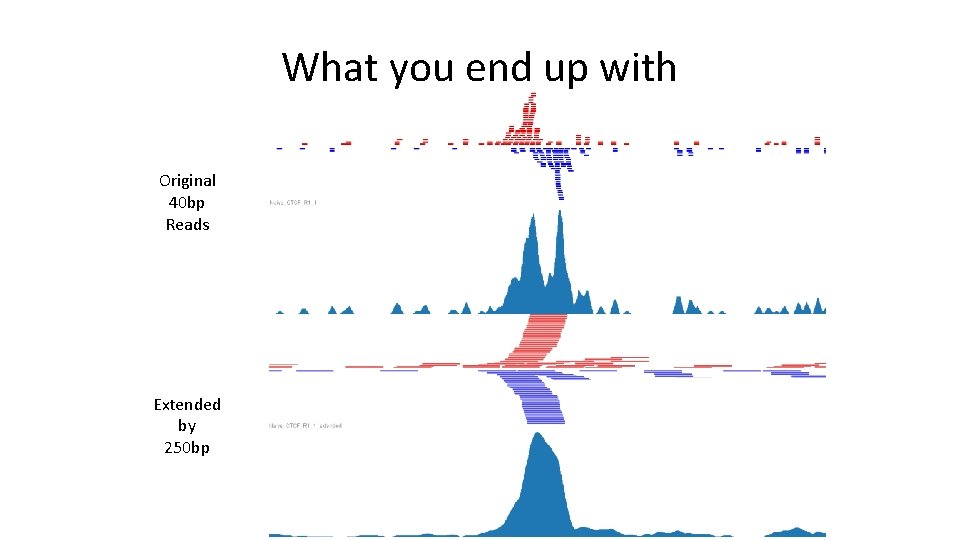
What you end up with Original 40 bp Reads Extended by 250 bp
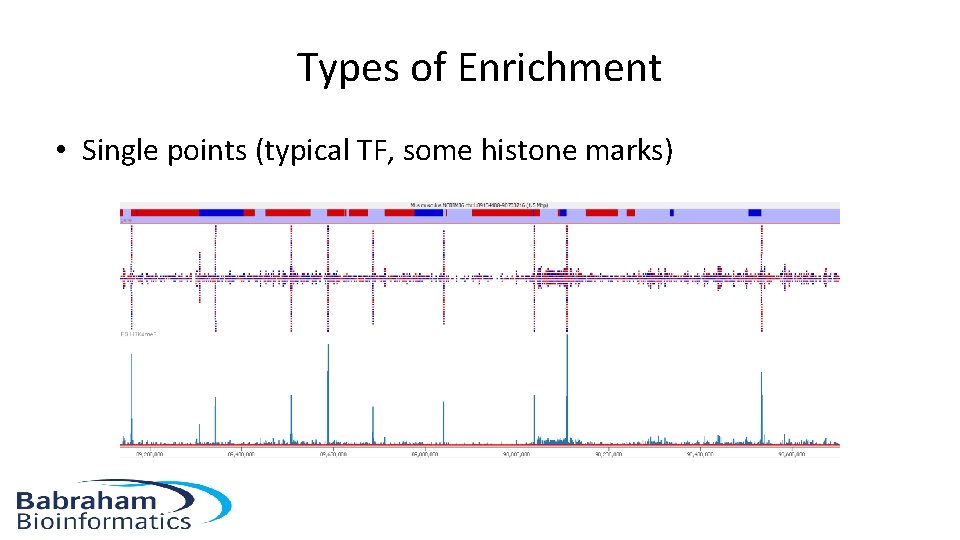
Types of Enrichment • Single points (typical TF, some histone marks)
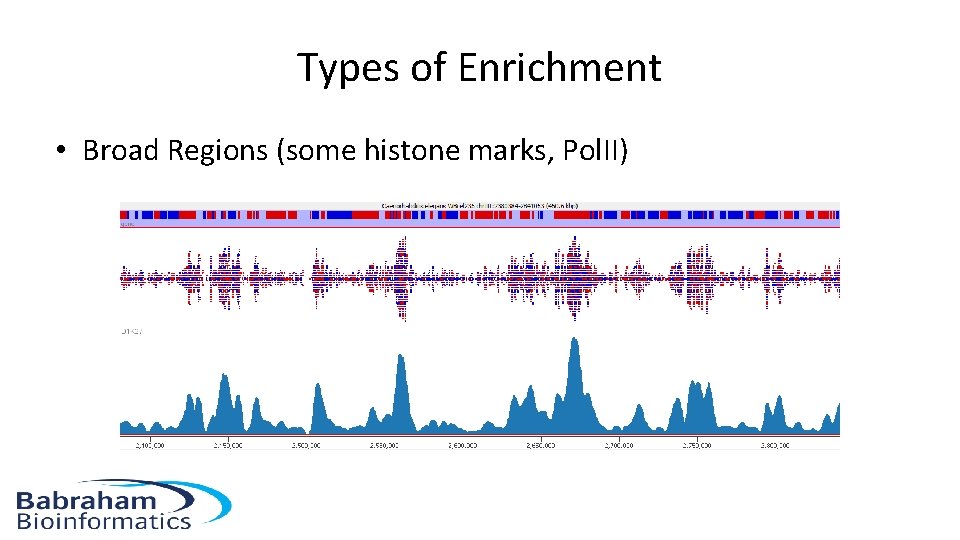
Types of Enrichment • Broad Regions (some histone marks, Pol. II)
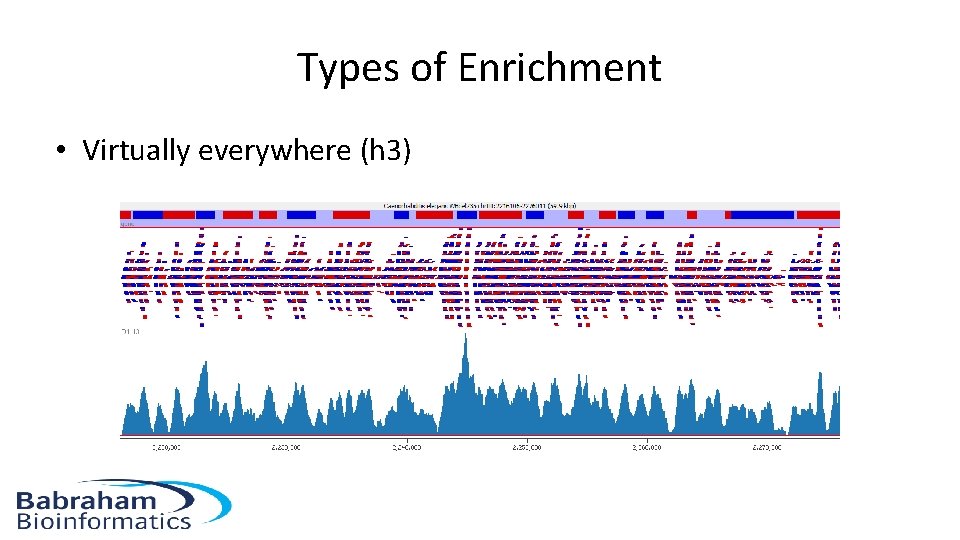
Types of Enrichment • Virtually everywhere (h 3)
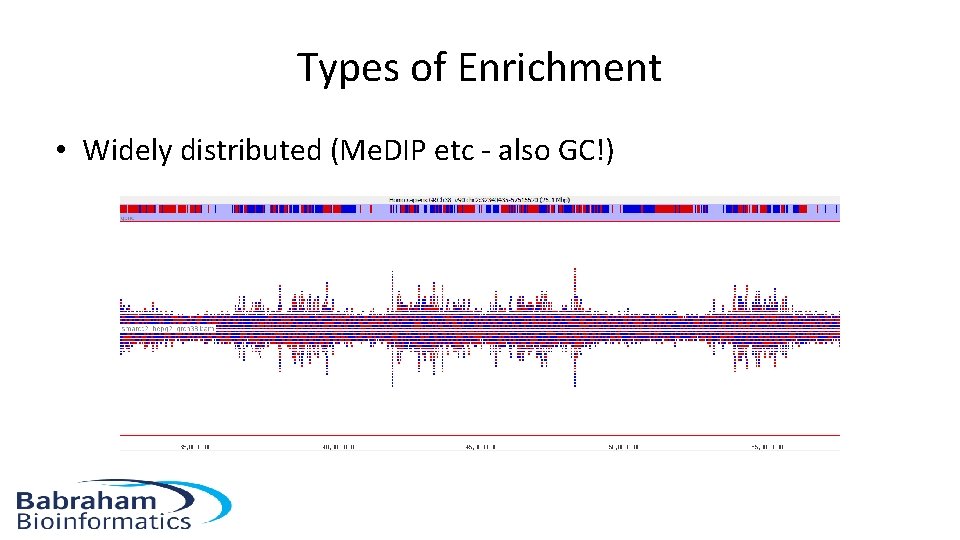
Types of Enrichment • Widely distributed (Me. DIP etc - also GC!)

What are you actually measuring? • Ch. IP Seq measures RELATIVE enrichment – Region A has twice as much signal as Region B • Without some external calibration, NOTHING in Ch. IP-Seq gives an ABSOLUTE measure.
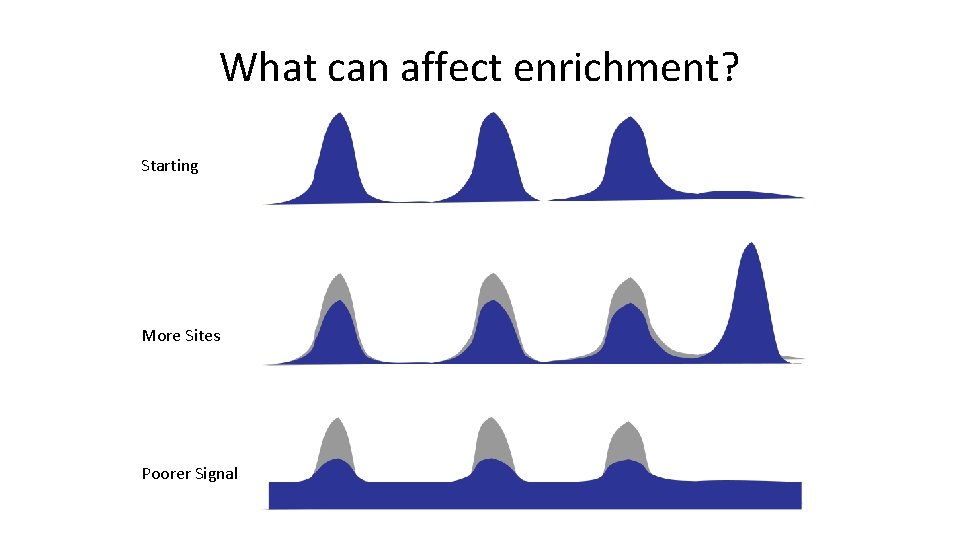
What can affect enrichment? Starting More Sites Poorer Signal

What sort of questions can you answer? • Where is this mark present? – General - it's in promoters, gene bodies etc. – Specific - it's at these loci • How does this mark change when I do XXX? – Categorical: A peak disappears – Quantitative: The enrichment of a locus changes
- Slides: 18