Ch 10 Protein Synthesis Gene Regulation Mutations Genetics















- Slides: 17

Ch. 10: Protein Synthesis, Gene Regulation, & Mutations Genetics

§ DNA controls protein synthesis. § Proteins control most all chemical reactions in living organisms. § DNA is located in the nucleus and does not leave. § Protein synthesis occurs in the ribosome in the cytoplasm. If DNA controls protein synthesis, How does the info get to the ribosome when DNA can’t leave the nucleus?

RNA § RNA acts as a messenger b/t DNA and ribosomes. RNA differs from DNA in 4 ways. 1) RNA contains ribose sugar. 2) Uracil replaces thymine as a nitrogen base. 3) RNA is single strand molecule, DNA is double stranded. 4) RNA utilizes protein-encoding info and DNA maintains the protein-encoding info.

3 Kinds of RNA 1) m. RNA (messenger RNA) carries sequence of nucleotides that code for protein from nucleus to ribosome 2) t. RNA (transfer RNA) picks up individual aa in the cytoplasm & carries them to ribosome (where they are joined together in proper order) 3) r. RNA (ribosomal RNA) plays a structural role in ribosomes

Protein Synthesis Transcription factors (proteins) come together to form a transcription apparatus that binds DNA and starts transcription at specific sites on DNA. The 1 st transcription factor to bind to the promoter is usually a TATA binding protein that attracted to a sequence of the promoter called a TATA box. This factor attracts a complex series of other transcription factors that contort the area so that RNA polymerase joins the complex. This complex guides the RNA polymerase so it can bind just before the gene sequence starts.

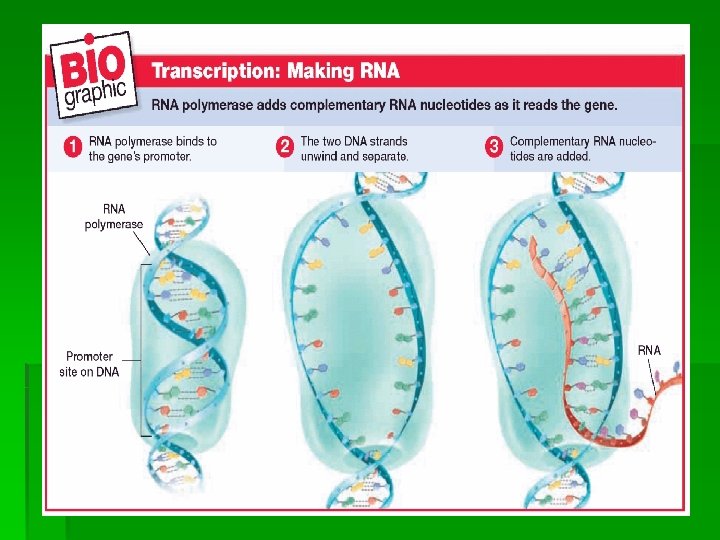
1) Transcription-process where the hereditary info in nucleotide sequence of DNA is transferred and encoded in nucleotide sequence of RNA. The RNA molecule that is formed is called m. RNA polymerase binds to a promotor (start signal) and unwinds & separates the 2 strands of DNA molecule, exposing the nitrogen bases. Only one of the 2 separated DNA strands serves as the pattern for making m. RNA. This DNA strand is called the “coding (sense) strand” and the RNA transcribed from it is called “sense RNA”. Transcription continues until a terminator (stop signal) is reached. The DNA strand that is not transcribed is called the noncoding or antisense strand.

The sequence of nucleotides forming m. RNA is determined by nucleotide sequence of DNA strand on which it is formed. EX: DNA: AGCT m. RNA: UCGA

3 Things that happen before m. RNA can leave the nucleus 1) Cap (sequence of nucleotides) is added to the 5’ end. 2) Poly-A-tail (200 adenines) is added to the 3’ end. Tell the cell which messages should be exported from the nucleus and read. 3) Introns-(noncoding sequences) are cut out 4) Exons-(coding sequences) are joined together to form single m. RNA molecule which travels to cytoplasm


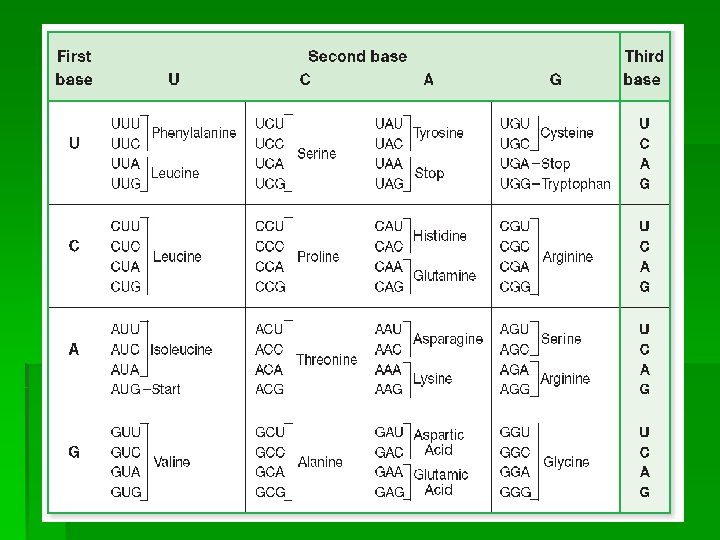
2) Translation-the formation of a protein under the control of m. RNA in the cytoplasm; translate RNA base sequence into the aa sequence of a protein, takes place at the ribosome. The t. RNA carries specific aa to ribosomes where protein is assembled. Codon-sequence of nucleotide bases as combination of 3; each codon is coding for a specific aa; carried by m. RNA


Degenerate codons-different codons that specify the same aa. (usually differ from one another by the base in the third position); this doesn’t affect the protein’s aa sequence. Anticodon-complementary to the m. RNA codon; carried by t. RNA EX: m. RNA: CCU (codon) t. RNA: GGA (anticodon)

2 Stages to Translation 1) Translation initiation Initiation complex begins when m. RNA forms a hydrogen bond with a short sequence of r. RNA in a small ribosomal subunit and with the t. RNA carrying the amino acid. Translation begins w/ the 1 st codon of m. RNA (AUG). 1) Translation elongation A large ribosomal subunit attaches to the initiation complex. The front end of the m. RNA moves across ribosome. Ribosome decodes message of codons. Molecules of t. RNA pick up aa and put them opposite appropriate codons of m. RNA. Incoming aa are linked and bonded to neighboring aa by peptide bond. t. RNA is released back to the cytoplasm to collect another aa. Eventually each ribosome reaches one of the 3 m. RNA “stop” or terminator codons (UAA, UAG, and UCA).

Protein Folding 1. Primary structure (1º)-determined by the sequence of amino acids. 2. Secondary structure (2º)-determined by chemical attractions b/t aa that are close together in the 1º structure. 3. Tertiary structure (3º)-determined by aa interaction with water. 4. Quarternary structure (4º)-proteins that consist of more than one polypeptide chain.

Genes and Mutations 1) Mutations-changes in the DNA of a gene or chromosome 2) Point mutations-chemical changes in just 1 nucleotide or a few nucleotides in a single gene. In gamete, may be passed to offspring. In somatic cell, will not be passed to offspring. 3) Base-pair substitutions-replacement of 1 nucleotide & its partner from the DNA strand with another pair of nucleotides (may or may not be harmful). 4) Frame-shift mutation-insertion of deletion of a nucleotide which causes reading to be out of phase resulting in the production of an abnormal protein.

Gene Regulation in Eukaryotes § Operons are not found in eukaryotes. § More DNA found in eukaryotes than prokaryotes. § Eukaryotic cells are specialized for specific functions and therefore produce different proteins.

Gene Regulation in Bacteria § 1) 2) 3) Scientists first studied gene expression in bacteria (specifically E. coli and lactose sugar). In bacteria (prokaryotes), the basic mechanism that controls gene regulation is the “operon”. Operon-cluster of genes that code for proteins with related functions. (Ex: lac operon code for enzyme beta-galactosidase). Regions of Lac operon Promoter-binding site that signal start of gene for RNA polymerase Operator-acts like on/off switch for operon Structural genes- genes that code for structurally making the proteins. A repressor switches off the lac operon preventing RNA polymerase from reading the code on the structural gene. Is present with no lactose is present. An inducer binds to the repressor molecule causing the repressor to change shape and fall off the operator, switching on the lac operon.