CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Multiple Alignment Anders

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Multiple Alignment Anders Gorm Pedersen / Henrik Nielsen Molecular Evolution Group Center for Biological Sequence Analysis gorm@cbs. dtu. dk / hnielsen@cbs. dtu. dk

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Outline • Refresher: pairwise alignments • Multiple alignments: what use are they? • Multiple alignment by dynamic programming – Why it is not done • Multiple alignment by progressive cluster and profile alignment – How it is done • Multiple alignment programs, features and comparisons • Take-home messages

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Refresher: pairwise alignments 43. 2% identity; alpha beta Global alignment score: 374 10 20 30 40 50 V-LSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLS-----HGSA : : . : . : : : : . VHLTPEEKSAVTALWGKV--NVDEVGGEALGRLLVVYPWTQRFFESFGDLSTPDAVMGNP 10 20 30 40 50 60 70 80 90 100 110 QVKGHGKKVADALTNAVAHVDDMPNALSALSDLHAHKLRVDPVNFKLLSHCLLVTLAAHL. : : : : . . . . : : : : : . KVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKLHVDPENFRLLGNVLVCVLAHHF 60 70 80 90 100 110 120 130 140 alpha PAEFTPAVHASLDKFLASVSTVLTSKYR : : : . : . . . : : . beta GKEFTPPVQAAYQKVVAGVANALAHKYH 120 130 140

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Refresher: pairwise alignments A R N D C Q E G 5 -2 -1 -1 -1 0 7 -1 7 -2 2 8 -4 -2 -4 1 0 0 2 -3 0 -1 • • 13 -3 7 -3 2 6 -3 -2 -3 • 8 . . . A R N D C Q E • G. . . • K L A A S V I L S D A L K L A A - - S D A L -10 + 3 x (-1)=-13 Alignment score is calculated from substitution matrix Identities on diagonal have high scores Similar amino acids have high scores Dissimilar amino acids have low (negative) scores Gaps penalized by gap-opening + gap elongation

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Refresher: pairwise alignments The number of possible pairwise alignments increases explosively with the length of the sequences: Two protein sequences of length 100 amino acids can be aligned in approximately 1060 different ways 1060 bottles of beer would fill up our entire galaxy

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Refresher: pairwise alignments T C G C • Solution: dynamic programming • Essentially: the best path through any grid point in the alignment matrix must originate from one of three previous points • Far fewer computations • Optimal alignment guaranteed to be found A T C x C A

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Refresher: pairwise alignments • Most used substitution matrices are themselves derived empirically from simple multiple alignments Multiple alignment Score(A/C) = log Calculate substitution frequencies Freq(A/C), observed Freq(A/C), expected A/A 2. 15% A/C 0. 03% A/D 0. 07%. . . Convert to scores

Refresher: pairwise alignments CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS • “Optimal alignment” means “having the highest possible score, given substitution matrix and set of gap penalties”. • This is NOT necessarily the biologically most meaningful alignment. • Specifically, the underlying assumptions are often wrong: substitutions are not equally frequent at all positions, affine gap penalties do not model insertion/deletion well, etc. • Pairwise alignment programs always produce an alignment even when it does not make sense to align sequences.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Database searching • Using pairwise alignments to search databases for similar sequences Query sequence Database
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Multiple alignment
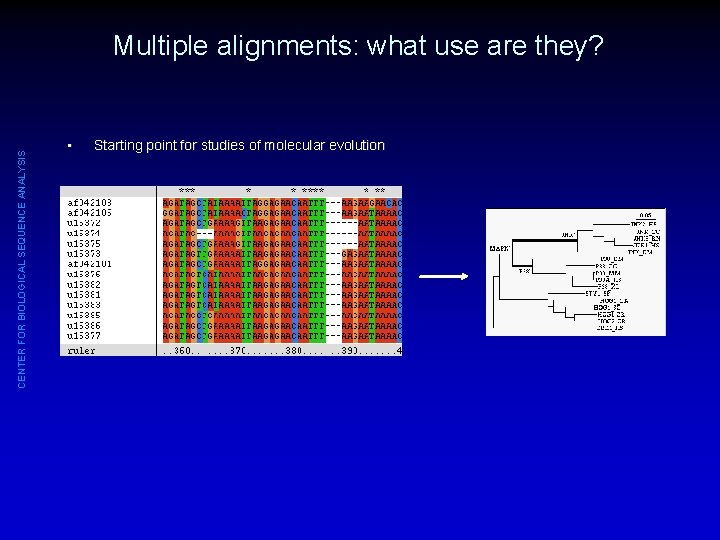
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Multiple alignments: what use are they? • Starting point for studies of molecular evolution
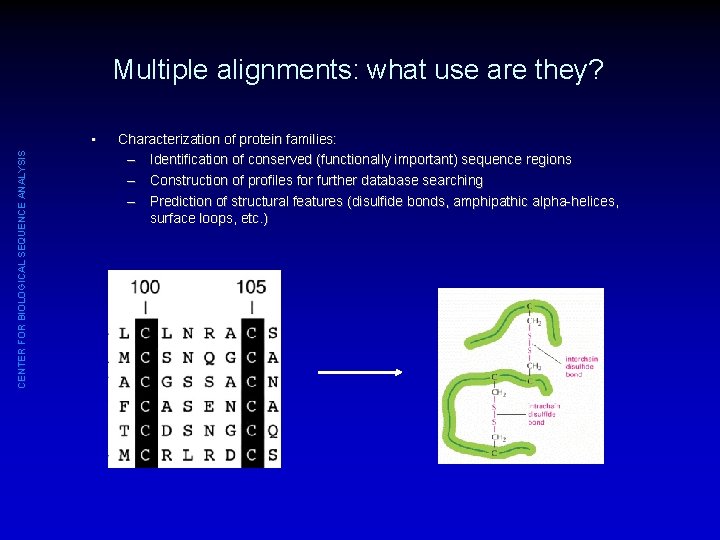
Multiple alignments: what use are they? CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS • Characterization of protein families: – Identification of conserved (functionally important) sequence regions – Construction of profiles for further database searching – Prediction of structural features (disulfide bonds, amphipathic alpha-helices, surface loops, etc. )

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Scoring a multiple alignment: the “sum of pairs” score . . . A. . . S. . . T. . . One column from alignment AA: 4, AS: 1, AT: 0 ST: 1 SP-score: 4+1+0+1 = 7 Weighted sum of pairs: each SP-score is multiplied by a weight reflecting the evolutionary distance (avoids undue influence on score by sets of very similar sequences) => In theory, it is possible to define an alignment score for multiple alignments (there are several alternative scoring systems)
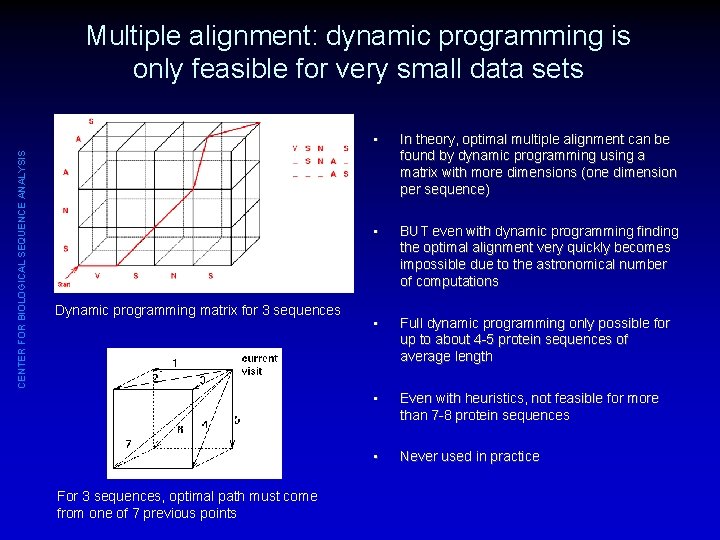
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Multiple alignment: dynamic programming is only feasible for very small data sets Dynamic programming matrix for 3 sequences For 3 sequences, optimal path must come from one of 7 previous points • In theory, optimal multiple alignment can be found by dynamic programming using a matrix with more dimensions (one dimension per sequence) • BUT even with dynamic programming finding the optimal alignment very quickly becomes impossible due to the astronomical number of computations • Full dynamic programming only possible for up to about 4 -5 protein sequences of average length • Even with heuristics, not feasible for more than 7 -8 protein sequences • Never used in practice

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Multiple alignment: an approximate solution • Progressive alignment: 1. Perform all pairwise alignments; keep track of sequence similarities between all pairs of sequences (construct “distance matrix”) 2. Align the most similar pair of sequences 3. Progressively add sequences to the (constantly growing) multiple alignment in order of decreasing similarity.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Progressive alignment: details 1) Perform all pairwise alignments, note pairwise distances (construct “distance matrix”) S 1 S 2 S 3 S 4 6 pairwise alignments S 1 S 2 S 3 S 4 3 1 3 3 2) Construct pseudo-phylogenetic tree from pairwise distances S 1 S 2 S 3 S 4 3 1 3 3 2 3 S 1 S 3 S 4 S 2 “Guide tree”

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Progressive alignment: details 3) a) Use tree as guide for multiple alignment: Align most similar pair of sequences using dynamic programming S 1 S 3 b) S 1 S 3 S 4 S 2 Align next most similar pair S 2 S 4 c) Align alignments using dynamic programming - preserve gaps S 1 S 3 S 2 S 4 New gap to optimize alignment of (S 2, S 4) with (S 1, S 3)

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Scoring profile alignments The naïve way: Compare each residue in one profile to all residues in second profile. Score is average of all comparisons. . A. . . S. . . AS: 1, AT: 0 + SS: 4, ST: 1 . . . S. . . T. . . One column from alignment Score: 1+0+4+1 = 1. 5 4

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Scoring profile alignments In practice: Use each profile as an estimate of a position-specific scoring matrix. This is combined with the original scoring matrix. More details to follow in session about “weight matrices”. Profile-to-sequence or profile-to-profile alignments are potentially more biologically relevant than sequence-to-sequence alignments, because position-specific patterns of substitution can be taken into account.

Additional heuristics for quality CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS • Sequence weighting (most programs): – scores from similar groups of sequences are down-weighted • Variable substitution matrices (Clustal. W/X): – during alignment Clustal. W/X uses different substitution matrices depending on how similar the sequences/profiles are • Context-dependent gap penalties (Clustal. W/X) – e. g. higher gap penalties in hydrophobic stretches • Reiterate alignment (MUSCLE, MAFFT) – calculate a new guide tree based on first multiple alignment, and use that for the next round of alignment • Consistency (T-Coffee, Prob. Cons) – use all the pairwise alignments to compute the match/mismatch scores in the multiple alignment (slow)

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Additional heuristics for speed • Use fast comparison instead of full dynamic programming when calculating the guide tree (Clustal. W/X, MUSCLE, MAFFT, …) • Reduce dynamic programming time in profile alignments by identifying regions of high similarity with a Fast Fourier Transform (MAFFT) • Build an approximate guide tree without making all pairwise comparisons (ClustalΩ)

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Global methods (e. g. , Clustal. X) get into trouble when data is not globally related!!!
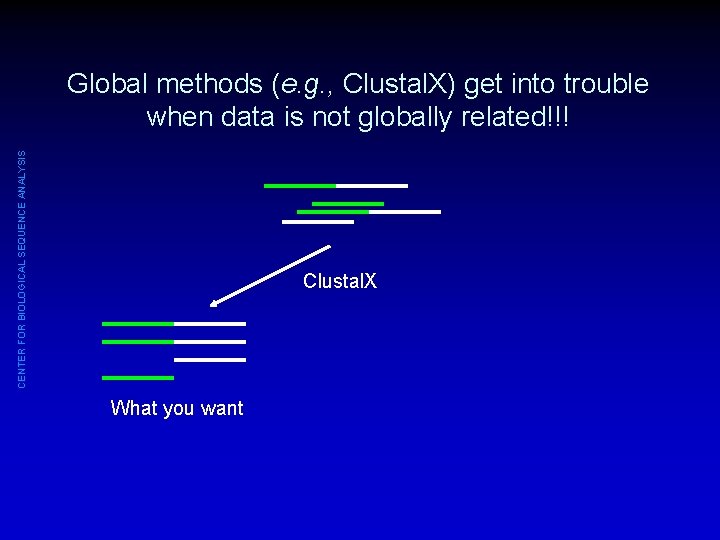
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Global methods (e. g. , Clustal. X) get into trouble when data is not globally related!!! Clustal. X What you want

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Global methods (e. g. , Clustal. X) get into trouble when data is not globally related!!! Clustal. X What you want What you might get Possible solutions: (1) Cut out conserved regions of interest and THEN align them (2) Use method that deals with local similarity (e. g. , Dialign)

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Other multiple alignment programs pileup DIALIGN multalign SBpima multal MLpima saga T-Coffee hmmt mafft MUSCLE poa Prob. Cons prank. . .

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Quantifying the Performance of Protein Sequence Multiple Alignment Programs • Compare to alignment that is known (or strongly believed) to be correct • Quantify by counting e. g. fraction of correctly paired residues • Option 1: Compare performance to benchmark data sets for which 3 D structures and structural alignments are available (BALi. BASE, PREfab, SABmark, SMART). – Advantage: real, biological data with real characteristics – Problem: we only have good benchmark data for core regions, no good knowledge of how gappy regions really look • Option 2: Construct synthetic alignments by letting a computer simulate evolution of a sequence along a phylogenetic tree – Advantage: we know the real alignment including where the gaps are – Problem: Simulated data may miss important aspects of real biological data
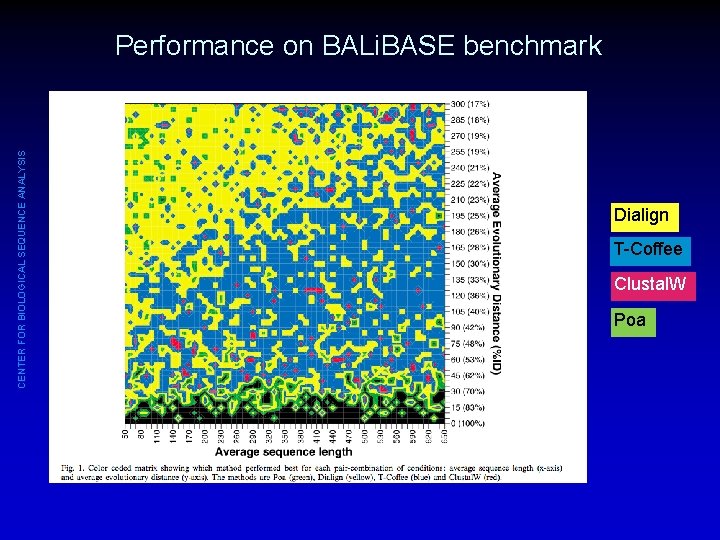
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Performance on BALi. BASE benchmark Dialign T-Coffee Clustal. W Poa
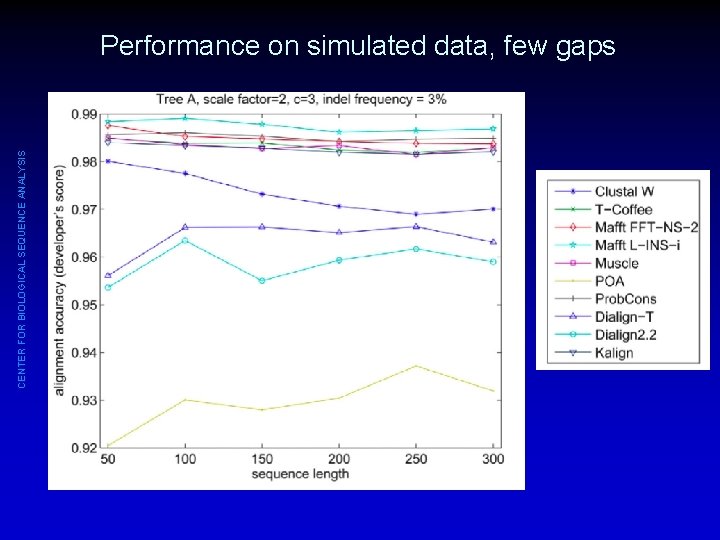
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Performance on simulated data, few gaps
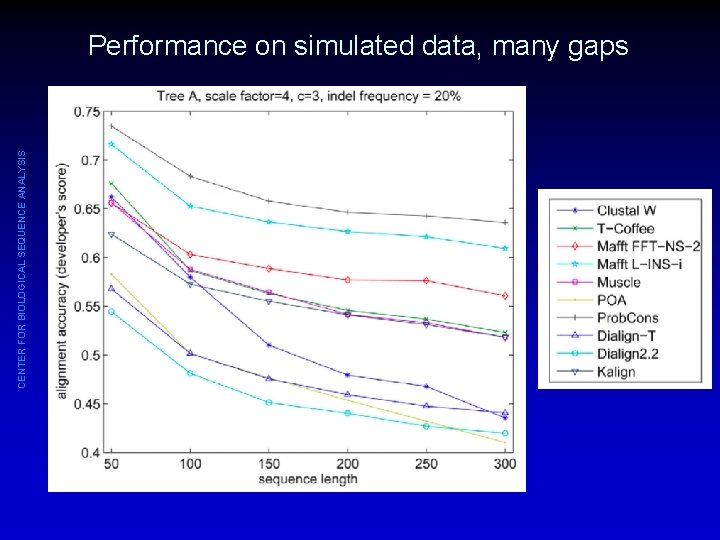
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Performance on simulated data, many gaps
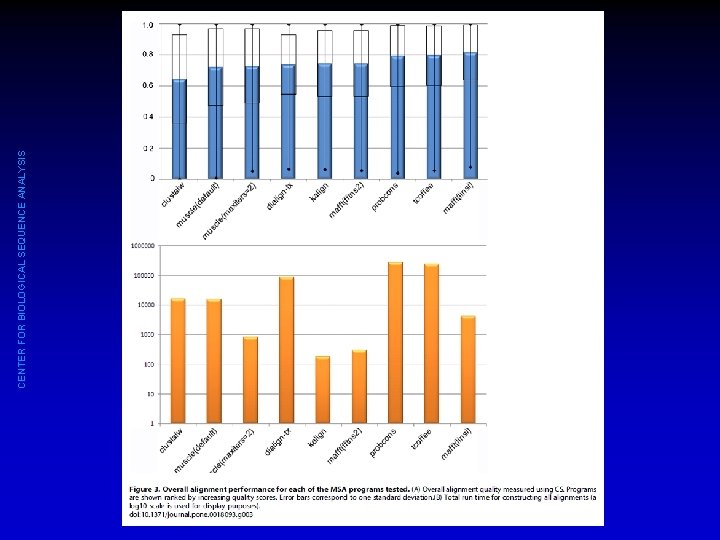
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS So which method should I choose? • Performance depends on way of measuring and on nature of data set • No single method performs best under all conditions • Slower is not necessarily better • Clustal Ω looks quite good with very large alignments • To be on the safe side, you ought to check that results are robust to alignment uncertainty (try a number of methods, check conclusions on each alignment). • T-Coffee can run several methods for you and report degree of agreement

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Special purpose alignment program • Rev. Trans: alignment of coding DNA based on information at protein level • Codon-codon boundaries maintained

Future perspectives CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS • Alignment inferred along with rest of analysis (e. g. phylogeny) • Bayesian techniques, conclusions based on probability distribution over possible alignments.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Take-home messages • Unlike pairwise alignment, the multiple alignment problem cannot be solved exactly • Nevertheless, multiple alignments are often more biologically correct than pairwise alignments • No single program performs best under all conditions, but MAFFT seems like a good compromise in most situations • Although Clustal. W/X is the best known program, it is not among the best or fastest.
- Slides: 34