CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Distance Matrix Methods

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Distance Matrix Methods Anders Gorm Pedersen Molecular Evolution Group Center for Biological Sequence Analysis Technical University of Denmark gorm@cbs. dtu. dk

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Distance Matrix Methods Gorilla : Human : Chimpanzee: Go ACGTCGTA ACGTTCCT ACGTTTCG Go Hu Ch - 4 4 - 2 Hu Ch 1. Construct multiple alignment of sequences 2. Construct table listing all pairwise differences (distance matrix) 3. Construct tree from pairwise distances - 1 1 1 2 Ch Hu Go
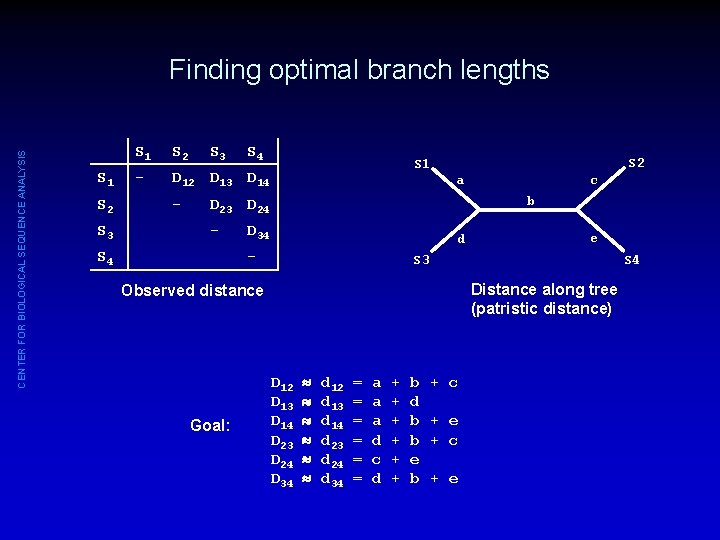
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding optimal branch lengths S 1 S 2 S 3 S 1 S 2 - D 12 D 13 D 14 - S 3 S 4 S 2 S 1 a b D 23 D 24 - S 4 D 34 d - e S 3 S 4 Distance along tree (patristic distance) Observed distance Goal: c D 12 D 13 D 14 D 23 D 24 D 34 d 12 d 13 d 14 d 23 d 24 d 34 = = = a a a d c d + + + b d b b e b + c + e

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Exercise (handout) • Construct distance matrix (count different positions) • Reconstruct tree and find best set of branch lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Optimal Branch Lengths for a Given Tree: Least Squares S 2 S 1 a c • b e d S 3 Fit between given tree and observed distances can be expressed as “sum of squared differences”: S 4 Distance along tree Q = (Dij - dij)2 j>i Goal: D 12 D 13 D 14 D 23 D 24 D 34 d 12 d 13 d 14 d 23 d 24 d 34 = = = a a a d c d + + + b d b b e b + c + e • Find branch lengths that minimize Q - this is the optimal set of branch lengths for this tree.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Optimal Branch Lengths: Least Squares S 2 S 1 a c • Longer distances associated with larger errors e • Squared deviation may be weighted so longer branches contribute less to Q: b d S 3 S 4 Distance along tree Goal: D 12 D 13 D 14 D 23 D 24 D 34 d 12 d 13 d 14 d 23 d 24 d 34 = = = a a a d c d + + + b d b b e b Q = + c + e (Dij - dij)2 Dijn • Power (n) is typically 1 or 2

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths A B C A B - DAB DAC - C A v 1 v 3 DBC - Observed distance DAB d. AB = v 1 + v 2 Goal: DAC d. AC = v 1 + v 3 DBC d. BC = v 2 + v 3 C v 2 B Distance along tree

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Finding Optimal Branch Lengths • System of n linear equations with n unknowns • Can be solved using substitution method or matrix-based methods

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Least Squares Optimality Criterion • Search through all (or many) tree topologies • For each investigated tree, find best branch lengths using least squares criterion • Among all investigated trees, the best tree is the one with the smallest sum of squared errors. • Least squares criterion used both for finding branch lengths on individual trees, and for finding best tree.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Minimum Evolution Optimality Criterion • Search through all (or many) tree topologies • For each investigated tree, find best branch lengths using least squares criterion • Among all investigated trees, the best tree is the one with the smallest sum of branch lengths (the shortest tree). • Least squares criterion used for finding branch lengths on individual trees, minimum tree length used for finding best tree.
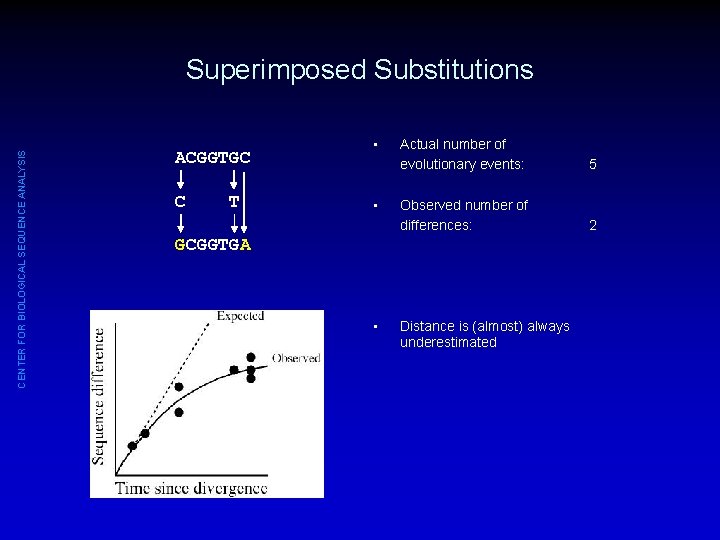
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Superimposed Substitutions ACGGTGC C T • • Actual number of evolutionary events: 5 Observed number of differences: 2 GCGGTGA • Distance is (almost) always underestimated

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Model-based correction for superimposed substitutions • Goal: try to infer the real number of evolutionary events (the real distance) based on 1. Observed data (sequence alignment) 2. A model of how evolution occurs

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Jukes and Cantor Model • Four nucleotides assumed to be equally frequent (f=0. 25) • All 12 substitution rates assumed to be equal • Under this model the corrected distance is: DJC = -0. 75 x ln(1 -1. 33 x DOBS) A C G T • A -3 C -3 G -3 T -3 For instance: DOBS=0. 43 => DJC=0. 64

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Clustering Algorithms • Starting point: Distance matrix • Cluster least different pair of nodes: – Tree: connect pair of nodes to common ancestral node, compute branch lengths from ancestral node to both descendants – Distance matrix: combine two entries into one. Compute new distance matrix, by finding distance from new node to all other nodes • Repeat until all nodes are linked • Results in only one tree, there is no measure of tree-goodness.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm • For each tip compute ui = j Dij/(n-2) (this is essentially the average distance to all other tips, except the denominator is n-2 instead of n) • Find the pair of tips, i and j, where Dij-ui-uj is smallest • Connect the tips i and j, forming a new ancestral node. The branch lengths from the ancestral node to i and j are: vi = 0. 5 Dij + 0. 5 (u (ui-uj) vj = 0. 5 Dij + 0. 5 (u (uj-ui) • Update the distance matrix: Compute distance between new node and each remaining tip as follows: Dij, k (Dik+Djk-Dij)/2 ij, k = (D • Replace tips i and j by the new node which is now treated as a tip • Repeat until only two nodes remain.

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B C D - 17 21 27 - 12 18 - 14 -

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B C D i ui - 17 21 27 A (17+21+27)/2=32. 5 - 12 18 B (17+12+18)/2=23. 5 - 14 C (21+12+14)/2=23. 5 - D (27+18+14)/2=29. 5

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B C D i ui - 17 21 27 A (17+21+27)/2=32. 5 - 12 18 B (17+12+18)/2=23. 5 - 14 C (21+12+14)/2=23. 5 - D (27+18+14)/2=29. 5 B C D A B C D - -39 -35 -35 - -39 D Dij-ui-uj

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B C D i ui - 17 21 27 A (17+21+27)/2=32. 5 - 12 18 B (17+12+18)/2=23. 5 - 14 C (21+12+14)/2=23. 5 - D (27+18+14)/2=29. 5 B C D A B C D - -39 -35 -35 - -39 D Dij-ui-uj

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B C D i ui - 17 21 27 A (17+21+27)/2=32. 5 - 12 18 B (17+12+18)/2=23. 5 - 14 C (21+12+14)/2=23. 5 - D (27+18+14)/2=29. 5 B C D A B C D - -39 -35 -35 - -39 D Dij-ui-uj C D X

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B C D i ui - 17 21 27 A (17+21+27)/2=32. 5 - 12 18 B (17+12+18)/2=23. 5 - 14 C (21+12+14)/2=23. 5 - D (27+18+14)/2=29. 5 B C D A B C D - -39 -35 -35 - -39 D Dij-ui-uj C D 4 X 10 v. C = 0. 5 x 14 + 0. 5 x (23. 5 -29. 5) = 4 v. D = 0. 5 x 14 + 0. 5 x (29. 5 -23. 5) = 10
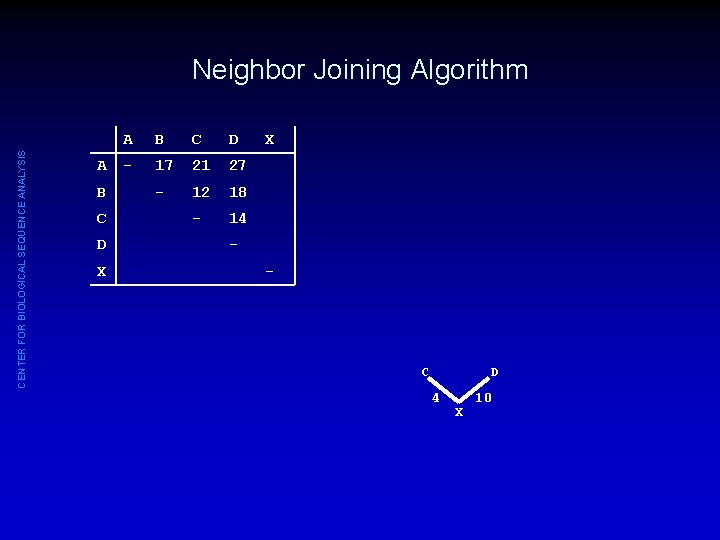
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B C D X A B C D - 17 21 27 - 12 18 - 14 X - C D 4 X 10

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B C D X A B C D - 17 21 27 - 12 18 - 14 X DXA = (DCA + DDA - DCD)/2 = (21 + 27 - 14)/2 = 17 - DXB = (DCB + DDB - DCD)/2 = (12 + 18 - 14)/2 = 8 C D 4 X 10

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B C D X - 17 21 27 17 - 12 18 8 - 14 - DXA = (DCA + DDA - DCD)/2 = (21 + 27 - 14)/2 = 17 DXB = (DCB + DDB - DCD)/2 = (12 + 18 - 14)/2 = 8 C D 4 X 10

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B X - 17 17 - 8 - DXA = (DCA + DDA - DCD)/2 = (21 + 27 - 14)/2 = 17 DXB = (DCB + DDB - DCD)/2 = (12 + 18 - 14)/2 = 8 C D 4 X 10

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B X i ui - 17 17 A (17+17)/1 = 34 - 8 B (17+8)/1 = 25 - X (17+8)/1 = 25 C D 4 X 10

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B X i - 17 17 A (17+17)/1 = 34 - 8 B (17+8)/1 = 25 - X (17+8)/1 = 25 B X A B X - -42 -28 - -28 X Dij-ui-uj ui C D 4 X 10

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B X i - 17 17 A (17+17)/1 = 34 - 8 B (17+8)/1 = 25 - X (17+8)/1 = 25 B X A B X - -42 -28 - -28 X Dij-ui-uj ui C D 4 X 10
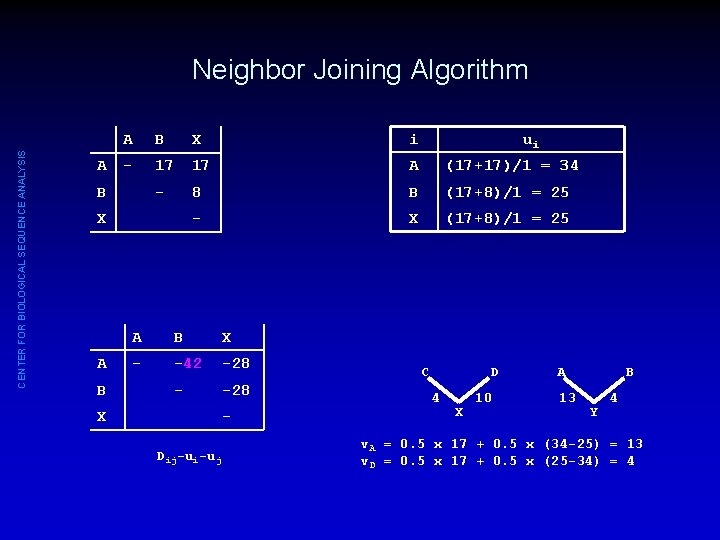
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A A B X i - 17 17 A (17+17)/1 = 34 - 8 B (17+8)/1 = 25 - X (17+8)/1 = 25 B X A B X - -42 -28 - -28 X Dij-ui-uj ui C D 4 X 10 A 13 B Y 4 v. A = 0. 5 x 17 + 0. 5 x (34 -25) = 13 v. D = 0. 5 x 17 + 0. 5 x (25 -34) = 4
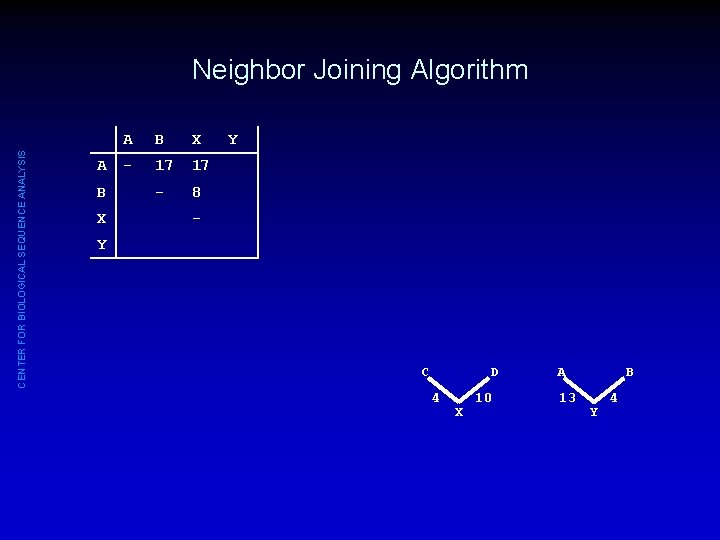
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B X - 17 17 - 8 Y - Y C D 4 X 10 A 13 B Y 4
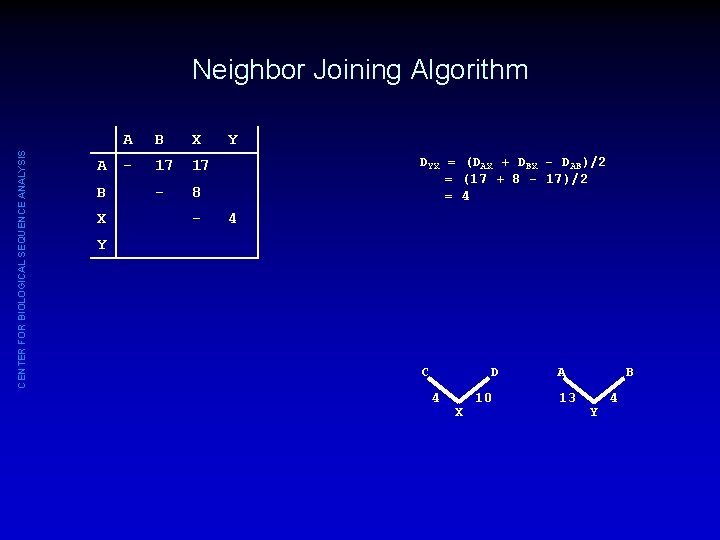
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B X - 17 17 - 8 - Y DYX = (DAX + DBX - DAB)/2 = (17 + 8 - 17)/2 = 4 4 Y C D 4 X 10 A 13 B Y 4
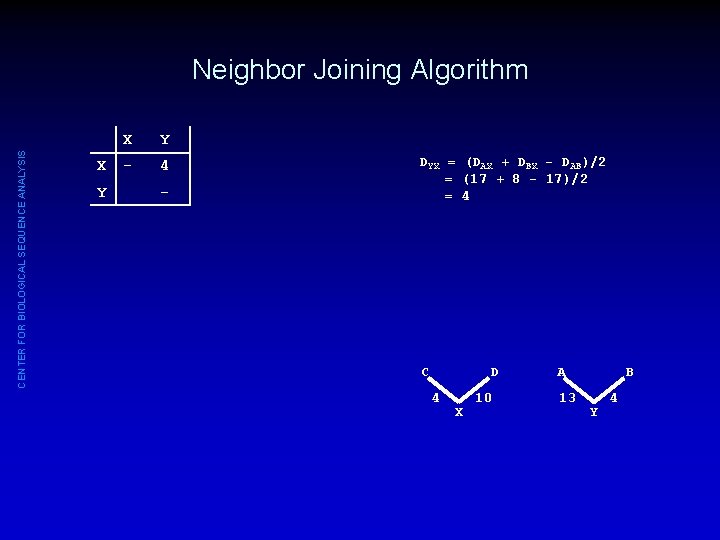
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm X Y - 4 - DYX = (DAX + DBX - DAB)/2 = (17 + 8 - 17)/2 = 4 C D 4 X 10 A 13 B Y 4
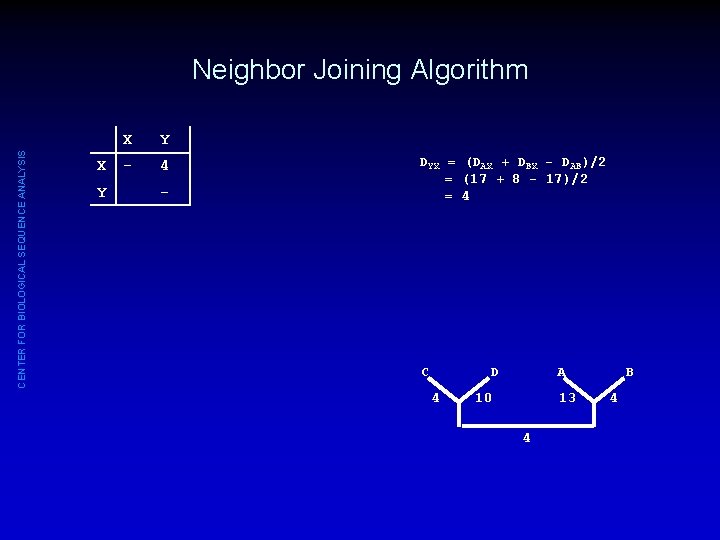
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm X Y - 4 - DYX = (DAX + DBX - DAB)/2 = (17 + 8 - 17)/2 = 4 C A D 4 10 13 4 B 4
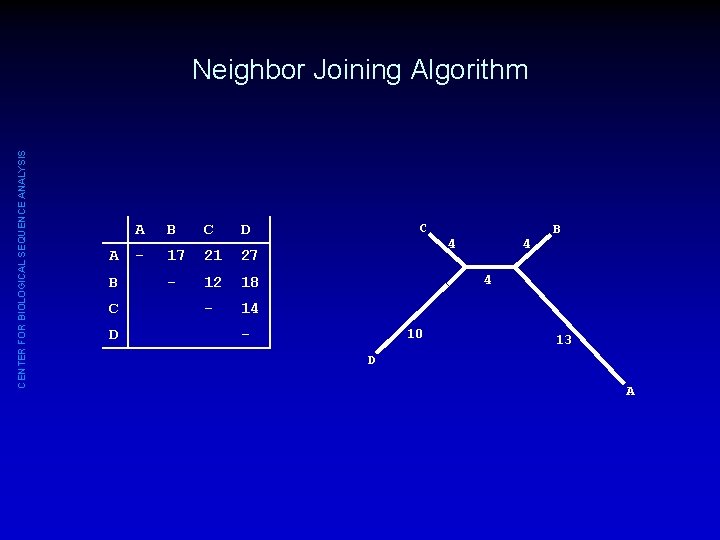
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Neighbor Joining Algorithm A B C D - 17 21 27 - 12 18 - 14 C 4 4 B 4 - 10 13 D A
- Slides: 41