CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Bootstrapping Anders Gorm

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Bootstrapping Anders Gorm Pedersen Molecular Evolution Group Center for Biological Sequence Analysis Technical University of Denmark (DTU)

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Bootstrap sampling: assess uncertainty about quantity estimated from data • Starting point: N observations x 1, x 2, x 3, . . . , x. N • Construct a “bootstrap sample”: – Using sampling with replacement, select N data points from the original observations – This means some data points are present more than once, some exactly once, some are not present in the bootstrap sample • Repeat many times (e. g. , 1000) • From each bootstrap sample: estimate quantity of interest • The distribution of estimates indicate uncertainty
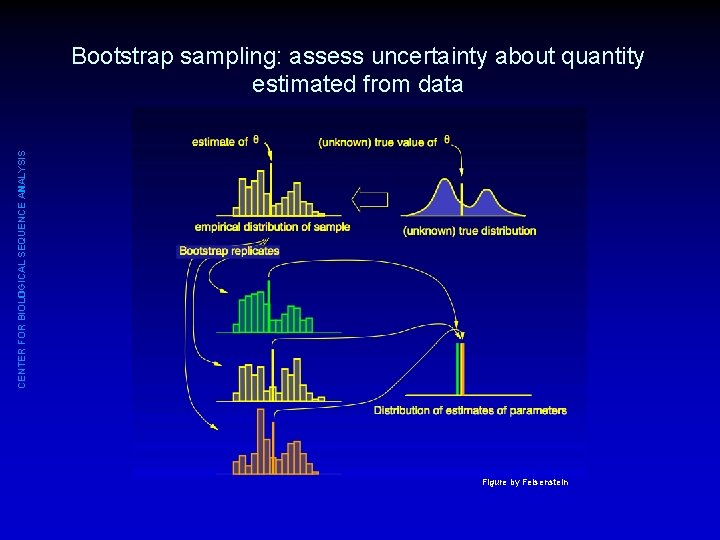
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Bootstrap sampling: assess uncertainty about quantity estimated from data Figure by Felsenstein

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Bootstrapping phylogenetic trees 1. 00 0. 74 Consensus tree Figure by Felsenstein
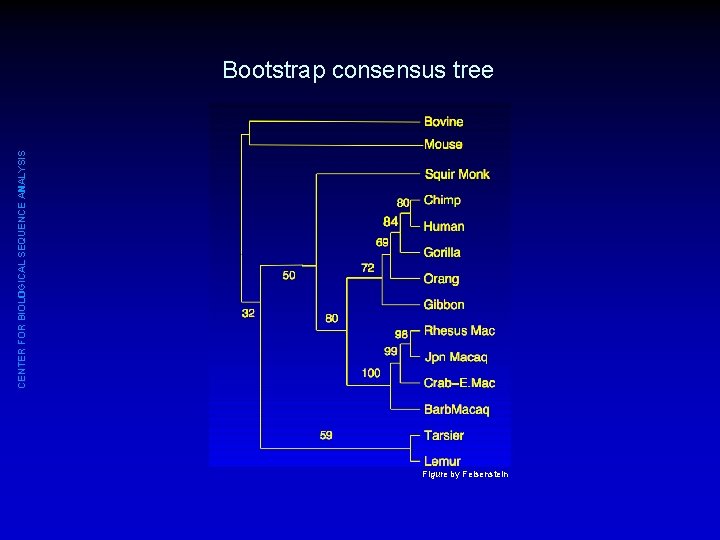
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Bootstrap consensus tree Figure by Felsenstein

CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Parametric bootstrapping • Starting point: data set with observations • Fit model to data, estimate parameters (e. g. , Θ 1, Θ 2, Θ 3) • Using parameter estimates: generate large number of simulated datasets (e. g. , 1000) • From each simulated dataset: estimate parameters • The distribution of estimates indicate uncertainty
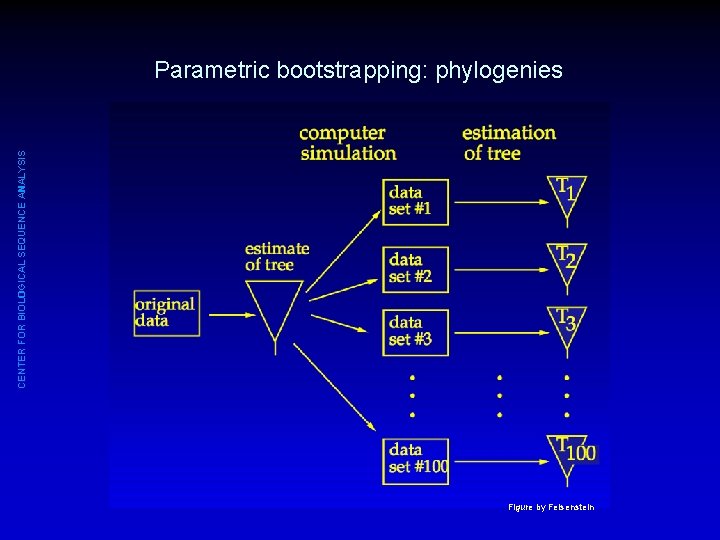
CENTER FOR BIOLOGICAL SEQUENCE ANALYSIS Parametric bootstrapping: phylogenies Figure by Felsenstein
- Slides: 7