Cellular oncogenes Molecular Biology of Cancer 1 Transfection
Cellular oncogenes Molecular Biology of Cancer 1
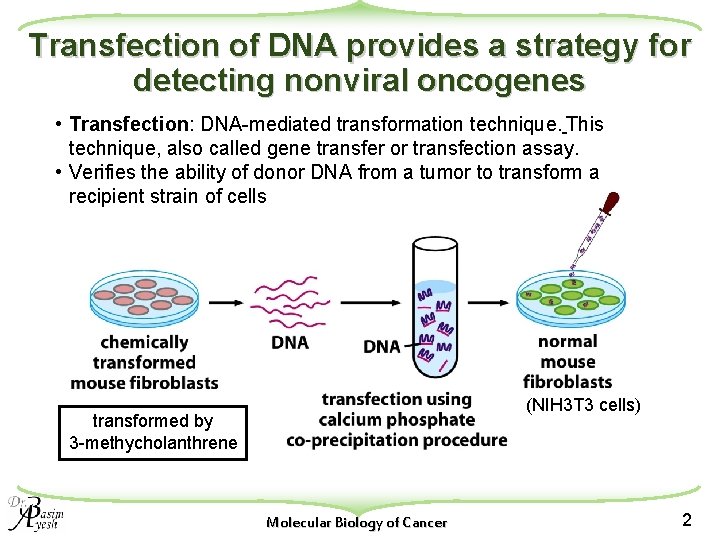
Transfection of DNA provides a strategy for detecting nonviral oncogenes • Transfection: DNA-mediated transformation technique. This technique, also called gene transfer or transfection assay. • Verifies the ability of donor DNA from a tumor to transform a recipient strain of cells (NIH 3 T 3 cells) transformed by 3 -methycholanthrene Molecular Biology of Cancer 2
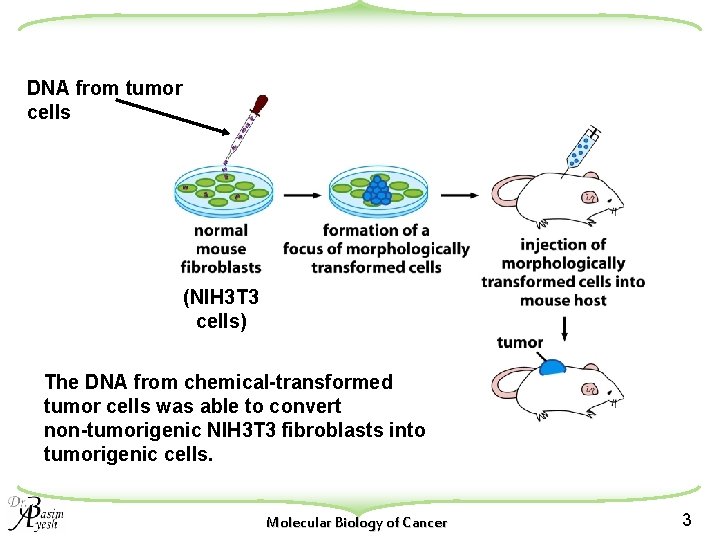
DNA from tumor cells (NIH 3 T 3 cells) The DNA from chemical-transformed tumor cells was able to convert non-tumorigenic NIH 3 T 3 fibroblasts into tumorigenic cells. Molecular Biology of Cancer 3

Results of recent transfection assay u. Ectopic expression of telomerase catalytic subunit (h. TERT) + SV 40 large T product + mutant H-ras oncogenic direct tumorigenic conversion of normal human epithelial and fibroblast cells. u. Many of the oncogenes identified by genetransfer studies are identical or closely related to those oncogenes transduced by retroviruses. w Members of the ras family have been repeatedly isolated from various human tumors by gene transfer. w Recently, neu, met, and trk Molecular Biology of Cancer 4
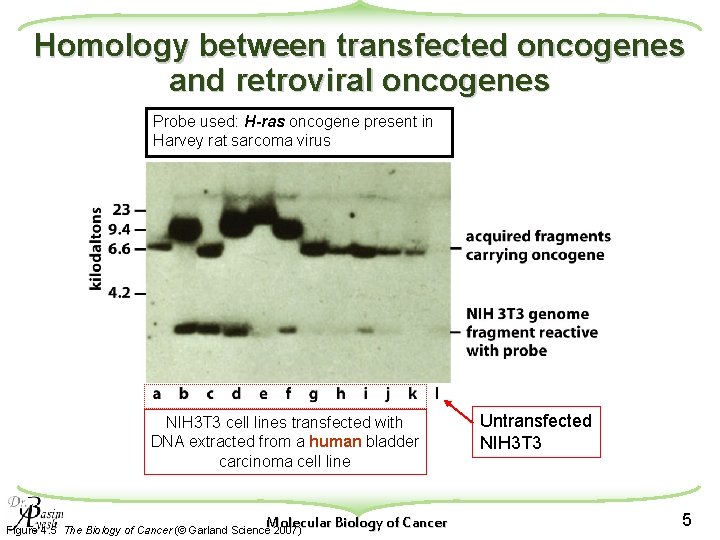
Homology between transfected oncogenes and retroviral oncogenes Probe used: H-ras oncogene present in Harvey rat sarcoma virus NIH 3 T 3 cell lines transfected with DNA extracted from a human bladder carcinoma cell line Molecular Biology of Cancer Figure 4. 5 The Biology of Cancer (© Garland Science 2007) Untransfected NIH 3 T 3 5

ONCOGENES, PROTOONCOGENES, AND THEIR FUNCTIONS u. Oncogenes can be classified into five groups based on functional and biochemical properties of their normal counterparts (proto-oncogenes). 1. growth factors 2. growth factor receptors 3. signal transducers 4. transcription factors 5. others, including programmed cell death regulators. Molecular Biology of Cancer 6

GROWTH FACTORS u Example: sis oncogene of simian sarcoma virus, a retrovirus first isolated from a monkey fibrosarcoma. w Sequence analysis: showed that sis encodes the beta chain of PDGF. w inappropriately expressed growth factors may function as oncogenes. w Constitutive expression of the sis gene product PDGF-ß } neoplastic transformation of fibroblasts containing PDGF receptor } No transformation of cells lacking the PDGF receptor. u Thus, transformation by sis requires interaction of the sis gene product with the PDGF receptor by autocrine stimulation. Molecular Biology of Cancer 7
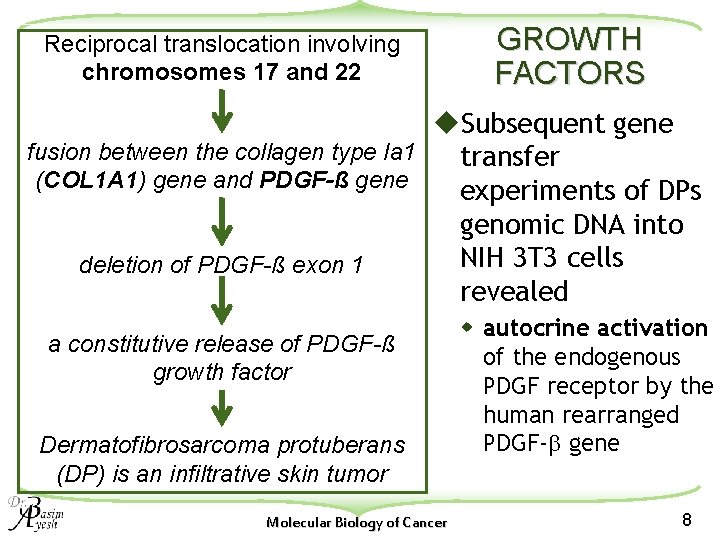
Reciprocal translocation involving chromosomes 17 and 22 GROWTH FACTORS u. Subsequent gene fusion between the collagen type Ia 1 transfer (COL 1 A 1) gene and PDGF-ß gene experiments of DPs genomic DNA into NIH 3 T 3 cells deletion of PDGF-ß exon 1 revealed a constitutive release of PDGF-ß growth factor Dermatofibrosarcoma protuberans (DP) is an infiltrative skin tumor Molecular Biology of Cancer w autocrine activation of the endogenous PDGF receptor by the human rearranged PDGF-b gene 8

GROWTH FACTOR RECEPTORS u. Some viral oncogenes are altered versions of normal growth factor (receptor tyrosine kinases). u. Growth factor receptors constitute an important class of protooncogenes involved in the regulation of normal cell growth. u. Examples include: erb B, erb B-2, fms, kit, met, ros, and trk. u. Mutation or abnormal expression of growth factor receptors can convert them into oncogenes. Molecular Biology of Cancer 9
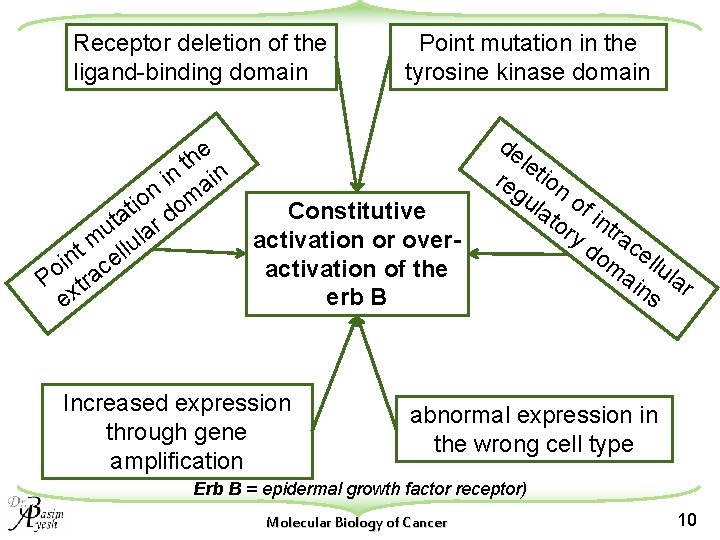
Receptor deletion of the ligand-binding domain Point mutation in the tyrosine kinase domain e th n in ai it on dom Constitutive a r t u la activation or overm u l t l in ce activation of the o P tra erb B ex Increased expression through gene amplification de let re ion gu lat of i or ntr y d ac om ellu ain lar s abnormal expression in the wrong cell type Erb B = epidermal growth factor receptor) Molecular Biology of Cancer 10

SIGNAL TRANSDUCERS u Many protooncogenes are members of signal transduction pathways. Consist of two main groups: 1. Nonreceptor protein kinases } tyrosine kinases (abl, lck, and src) } Serine/threonine kinases (raf-1, mos, and pim-1) 2. Guanosine triphosphate (GTP)-binding proteins } Monomeric GTP-binding proteins: ras family of protooncogenes (H-ras, K-ras, and N-ras) } Heterotrimeric GTP-binding proteins (G proteins) implicated as protooncogenes: currently include gsp and gip. u Signal transducers are often converted to oncogenes by mutations that lead to their unregulated activity, which in turn leads to uncontrolled cellular proliferation. Molecular Biology of Cancer 11

TRANSCRIPTION FACTORS u. Many protooncogenes are transcription factors that were discovered through their retroviral homologs u. Examples include erb A, ets, fos, jun, myb, and cmyc. w Fos + jun AP-1 transcription factor positively regulates a number of cell division activating genes w Erb A is the receptor for the T 3 thyroid hormone, triiodothyronine. Molecular Biology of Cancer 12

TRANSCRIPTION FACTORS u. Protooncogenes that function as transcription factors are often activated by chromosomal translocations in hematologic and solid neoplasms. u. Example: c-myc gene: w helps to control the expression of genes leading to cell proliferation. w c-myc gene is frequently activated by chromosomal translocations in human leukemia and lymphoma. Molecular Biology of Cancer 13

PROGRAMMED CELL DEATH REGULATION u. The only protooncogene thus far shown to regulate programmed cell death is bcl-2. w Bcl-2 was discovered by the study of chromosomal translocations in human lymphoma. ubcl-2 activation inhibits programmed cell death in lymphoid cell populations (survival). ubcl-2 is not the only apoptosis gene involved in neoplasia, additional protooncogenes await identification Molecular Biology of Cancer 14

MECHANISMS OF ONCOGENE ACTIVATION u. Three genetic mechanisms activate oncogenes in human neoplasms: 1. Mutation 2. gene amplification 3. chromosome rearrangements u. These mechanisms result in either an alteration of protooncogene structure or an increase in protooncogene expression u. Because neoplasia is a multistep process w more than one of these mechanisms often contribute to the genesis of human tumors by altering a number of cancer-associated genes. Molecular Biology of Cancer 15

MUTATION Mutations activate proto-oncogenes through structural alterations in their encoded proteins Usually involve critical protein regulatory regions Often lead to the uncontrolled, continuous activity of the mutated protein Molecular Biology of Cancer 16

MUTATION u. Retroviral oncogenes, often have deletions that contribute to their activation. w deletions in the amino- terminal ligand-binding domains of the erb B, kit, ros, met, and trk oncogenes. u. In human tumors, most characterized oncogene mutations are base substitutions (point mutations) that change a single amino acid within the protein. Molecular Biology of Cancer 17
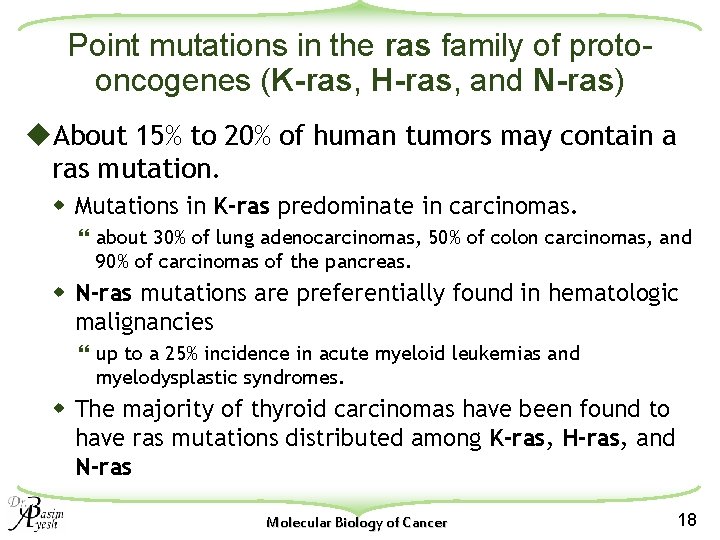
Point mutations in the ras family of protooncogenes (K-ras, H-ras, and N-ras) u. About 15% to 20% of human tumors may contain a ras mutation. w Mutations in K-ras predominate in carcinomas. } about 30% of lung adenocarcinomas, 50% of colon carcinomas, and 90% of carcinomas of the pancreas. w N-ras mutations are preferentially found in hematologic malignancies } up to a 25% incidence in acute myeloid leukemias and myelodysplastic syndromes. w The majority of thyroid carcinomas have been found to have ras mutations distributed among K-ras, H-ras, and N-ras Molecular Biology of Cancer 18
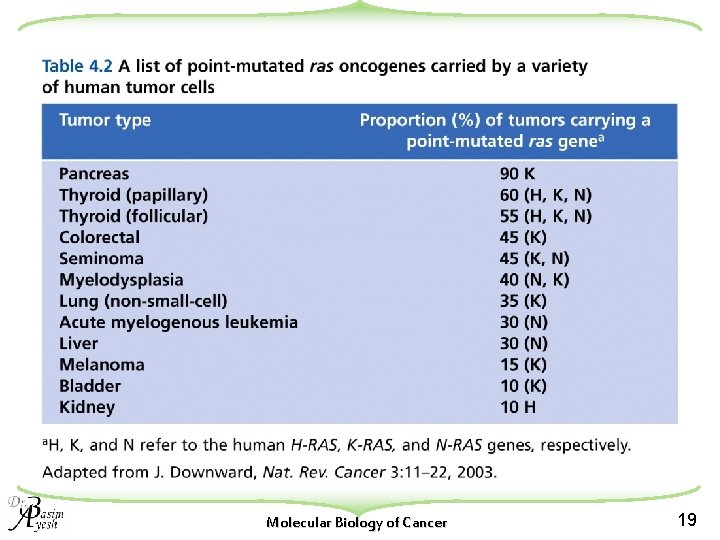
Molecular Biology of Cancer 19
Point mutations in the ras family of protooncogenes (K-ras, H-ras, and N-ras) Carcinogen exposure. Ras mutations The majority involve codon 12 of the gene, a smaller number involving other regions such as codons 13 Constitutive activation of the signal-transduction Molecular Biology of Cancer 20
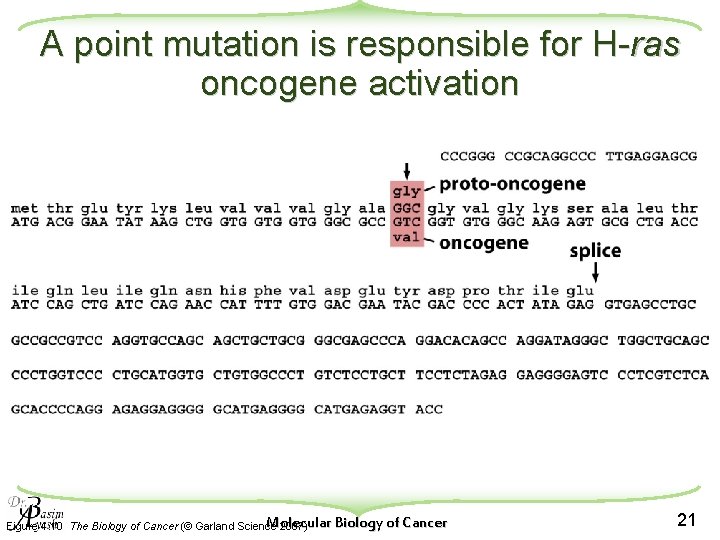
A point mutation is responsible for H-ras oncogene activation Molecular Biology of Cancer Figure 4. 10 The Biology of Cancer (© Garland Science 2007) 21

GENE AMPLIFICATION u The process of gene amplification occurs through redundant replication of genomic DNA, often giving rise to karyotypic abnormalities: w double-minute chromosomes (DMs) } are characteristic minichromosome structures without centromeres w homogeneous staining regions (HSRs) } HSRs are segments of chromosomes that lack the normal alternating pattern of light- and dark-staining bands u up to several hundred copies of a gene. u Amplification leads to the increased expression of genes, which in turn can confer a selective advantage for cell growth. Molecular Biology of Cancer 22

u Studies demonstrated that three protooncogene families (myc, erb B, and ras) are amplified in a significant number of human tumors: u Examples: w c-myc amplification: about 20% to 30% of breast and ovarian cancers; and in some types of squamous cell carcinomas w erb B amplification: found in up to 50% of glioblastomas and in 10% to 20% of squamous carcinomas of the head and neck. w erb. B-2 (HER-2/neu) gene amplification: Approximately in 15% to 30% of breast and ovarian cancers. w Members of the ras gene family, including K-ras and N-ras, are sporadically amplified in various carcinomas. Molecular Biology of Cancer 23
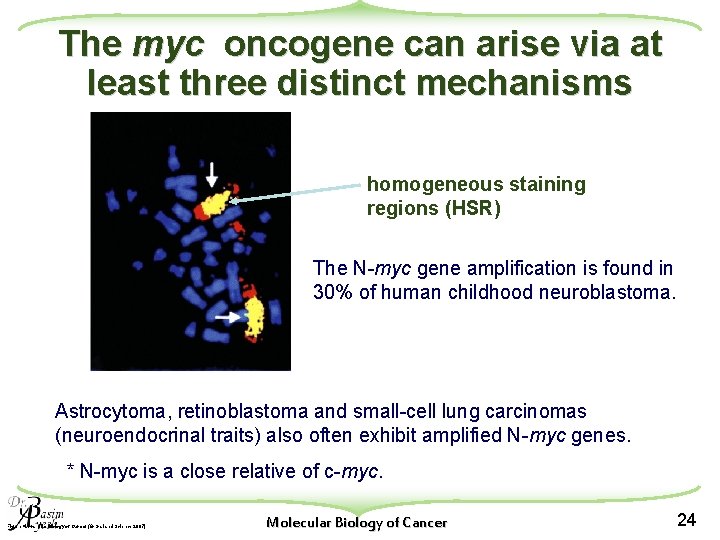
The myc oncogene can arise via at least three distinct mechanisms homogeneous staining regions (HSR) The N-myc gene amplification is found in 30% of human childhood neuroblastoma. Astrocytoma, retinoblastoma and small-cell lung carcinomas (neuroendocrinal traits) also often exhibit amplified N-myc genes. * N-myc is a close relative of c-myc. Figure 4. 11 a The Biology of Cancer (© Garland Science 2007) Molecular Biology of Cancer 24
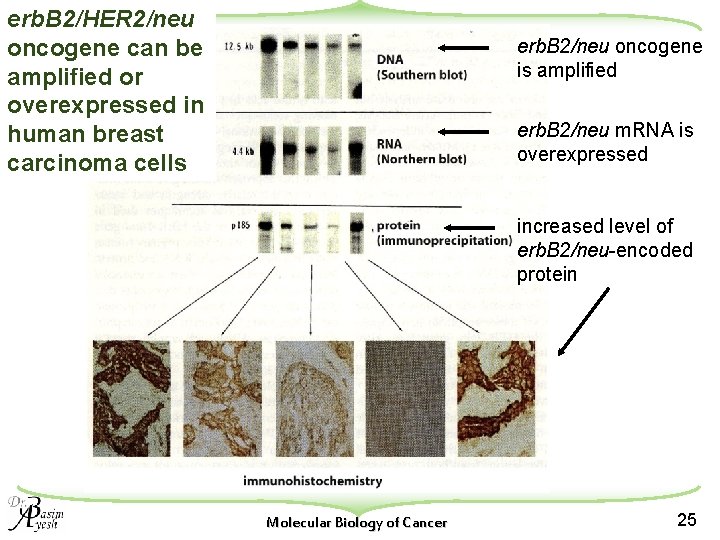
erb. B 2/HER 2/neu oncogene can be amplified or overexpressed in human breast carcinoma cells erb. B 2/neu oncogene is amplified erb. B 2/neu m. RNA is overexpressed increased level of erb. B 2/neu-encoded protein Molecular Biology of Cancer 25
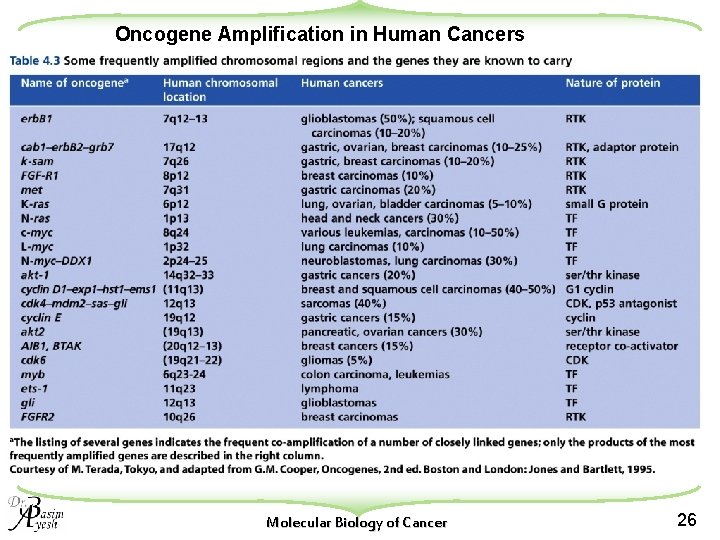
Oncogene Amplification in Human Cancers Molecular Biology of Cancer 26

CHROMOSOMAL REARRANGEMENTS u Recurring chromosomal rearrangements are often detected in hematologic as well as in some solid malignancies w mainly of chromosomal translocations w less frequently, chromosomal inversions. u Chromosomal rearrangements can lead to hematologic malignancy via two different mechanisms: 1. The transcriptional activation of proto-oncogenes w w Chromosomal rearrangements move a proto-oncogene close to an immunoglobulin or T-cell receptor gene Transcription of the protooncogene then falls under control of regulatory elements from the immunoglobulin or T-cell receptor locus. Molecular Biology of Cancer 27

CHROMOSOMAL REARRANGEMENTS 2. The creation of fusion genes. w Fusion genes can be created by chromosomal rearrangements when the chromosomal breakpoints fall within the loci of two different genes. w Fusion genes encode chimeric proteins with transforming activity. w In some cases, the same protooncogene is involved in several different translocations (cmyc, ews, and ret). Molecular Biology of Cancer 28

Biology of Cancer Table 4. 5 The Biology of Cancer (© Garland Science. Molecular 2007) 29
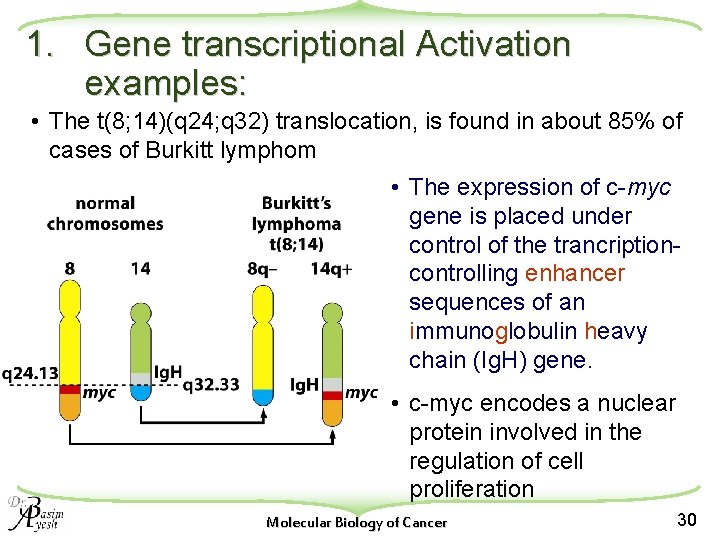
1. Gene transcriptional Activation examples: • The t(8; 14)(q 24; q 32) translocation, is found in about 85% of cases of Burkitt lymphom • The expression of c-myc gene is placed under control of the trancriptioncontrolling enhancer sequences of an immunoglobulin heavy chain (Ig. H) gene. • c-myc encodes a nuclear protein involved in the regulation of cell proliferation Molecular Biology of Cancer 30

1. Gene transcriptional Activation examples: u. In addition to c-myc, several proto-oncogenes that encode nuclear proteins are activated by various chromosomal translocations in T-ALL involving the T-cell receptor a or ß locus. u. These include HOX 11, TAL 2, and RBTN 1/Tgt 1. w The proteins encoded by these genes are thought to function as transcription factors through DNAbinding and protein-protein interactions. Molecular Biology of Cancer 31

2. Gene Fusion Examples: u The first example of gene fusion was discovered through the cloning of the breakpoint of the Philadelphia chromosome in chronic myelogenous leukemia (CML). u The t(9; 22)(q 34; q 11) translocation in CML fuses the c-abl gene, normally located at 9 q 34, with the bcr gene at 22 q 11 w The bcr/abl fusion, created on the der(22) chromosome, encodes a chimeric protein of 210 k. Da increased tyrosine kinase activity and abnormal cellular localization. u The t(9; 22) translocation is also found in up to 20% of cases of acute lymphoblastic leukemia (ALL). w In these cases, the breakpoint in the bcr gene differs somewhat from that found in CML, resulting in a 185 k. Da bcr/abl fusion protein. Molecular Biology of Cancer 32
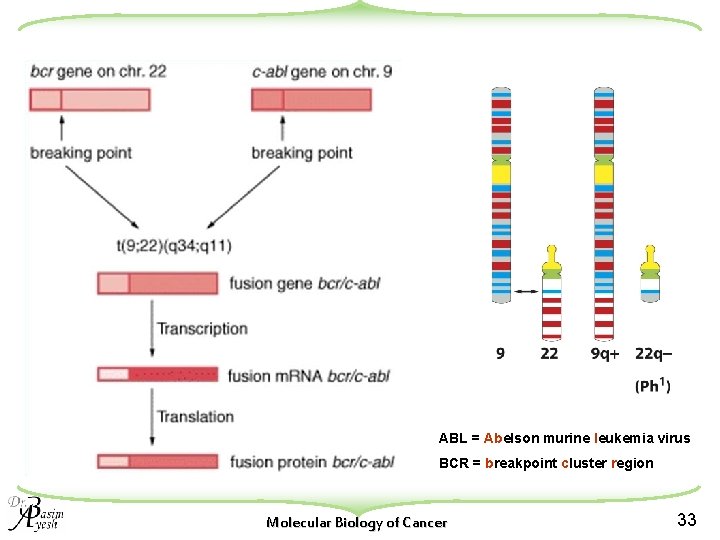
ABL = Abelson murine leukemia virus BCR = breakpoint cluster region Molecular Biology of Cancer 33
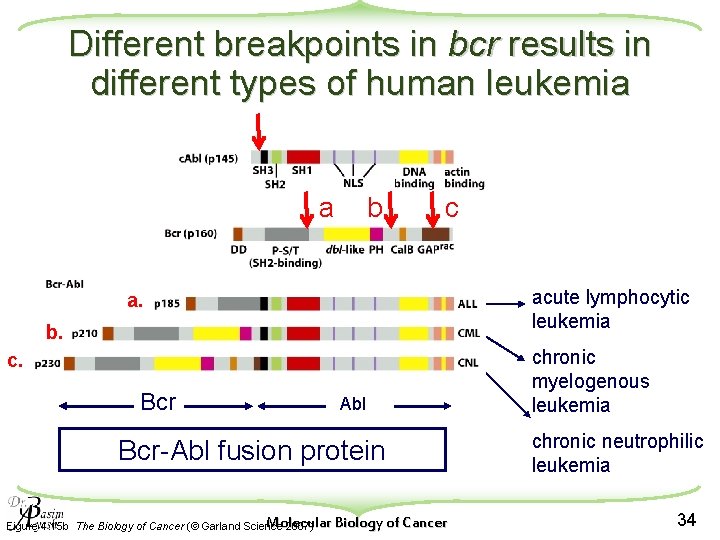
Different breakpoints in bcr results in different types of human leukemia a b c acute lymphocytic leukemia a. b. c. Bcr Abl Bcr-Abl fusion protein Molecular Biology of Cancer Figure 4. 15 b The Biology of Cancer (© Garland Science 2007) chronic myelogenous leukemia chronic neutrophilic leukemia 34
- Slides: 34