Cellfree expression of integral membrane proteins Christian Klammt

Cell-free expression of integral membrane proteins Christian Klammt, Innokentiy Maslennikov, Witek Kwiatkowski and Senyon Choe The Salk Institute, Structural Biology Laboratory, La Jolla, CA Center for Structures of Membrane Proteins © 2006

Outline Cell-free (CF) expression systems – Overview (extract sources, reaction set-ups, commercial systems) Protocols for set-up of an E. coli based CF expression system – – – S 30 cell extract preparation T 7 -RNA polymerase preparation List of reaction components Stock solutions of CF components Pipetting protocol Reaction compartments for dialysis mode CF expression of integral membrane proteins (IMPs) – Possible modes for CF IMP expression – – • P-CF (insoluble precipitate) • D-CF (detergent) • L-CF (lipids) List of detergents/ lipids used with CF system Trouble shooting N-CF – a new mode for CF expression of IMPs Center for Structures of Membrane Proteins © 2006
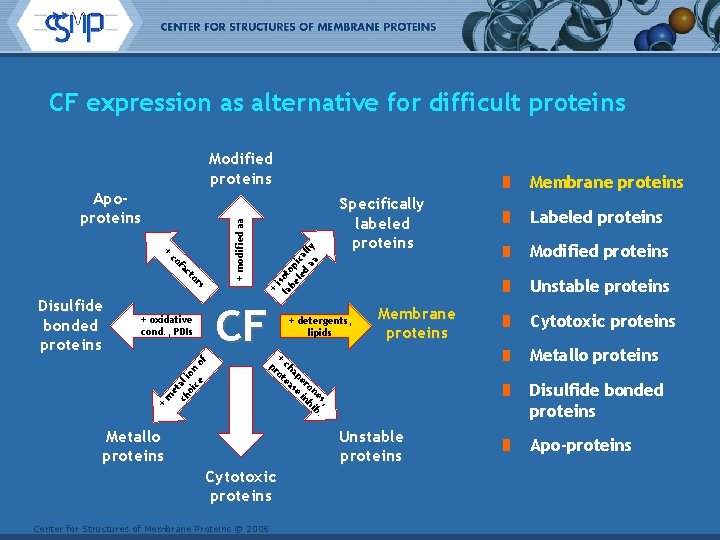
CF expression as alternative for difficult proteins Modified proteins co fa c to rs CF + oxidative cond. , PDIs + m et ch al oi ion ce o f Disulfide bonded proteins + is la oto be p le ica d aa lly + + modified aa Apoproteins Membrane proteins Specifically labeled proteins + detergents, lipids Membrane proteins Center for Structures of Membrane Proteins © 2006 Cytotoxic proteins Metallo proteins Disulfide bonded proteins Unstable proteins Cytotoxic proteins Modified proteins Unstable proteins + pr ch ot ap ea er se on in es, hi b. Metallo proteins Labeled proteins Apo-proteins

CF extract sources and systems Wheat germ – low nuclease activities, long life-times (days), preparative system – difficult extract preparation, complex system E. coli – high efficiency, easy extract preparation, optimized protocols for various applications available, preparative system – high variations, shorter life-times (hours) Rabbit reticulocyte – eukaryotic system – difficult extract preparation, complex non preparative system Insect cells – eukaryotic system, post-translational modifications – complex non preparative system Center for Structures of Membrane Proteins © 2006

Reaction set-up - batch vs. dialysis batch COUPLED TRANSCRIPTION/ TRANSLATION Ribosomes, ARSases, t. RNA, Polymerases, DNA, m. RNA, IFs, EFs, RFs, NTPs, amino acids, Energy system stirrer SUBSTRATES: amino acids, NTPs, energy LMW substrates PRODUCTS: NMPs, PPi, Pi Reaction mixture (RM) Dialysis membrane Feeding mixture (FM) dialysis Easy set-up Need for suitable dialysis set-up Short reaction times (<3 hours) Long reaction times (> 12 hours) Agglomeration of harmful LMW products, fast energy consumption Continuous exchange of harmful LMW products and supply of energy High throughput screening Up to 6 mg/ml of protein in RM Up to 1 mg/ml Limited high throughput screening Center for Structures of Membrane Proteins © 2006

Commercially available systems E. coli: – – Expressway. TM (Invitrogen) RTS E. coli (Roche) Easy. Express E. coli (Qiagen) S 30 T 7 High-Yield (Promega) Wheat germ – – – ENDEXT® (Cell-Free Sciences) RTS wheat germ (Roche) TNT® wheat germ (Promega) Rabbit reticulocyte – TNT® rabbit reticulocyte (Promega) Insect – – Easy. Express insect (Qiagen) TNT® insect (Promega) Center for Structures of Membrane Proteins © 2006 Roche is using a dialysis set-up (patent), other systems are based on batch mode. Convenient, easy to use kits High costs compared to individual systems “Black box” systems with limited room for optimizations
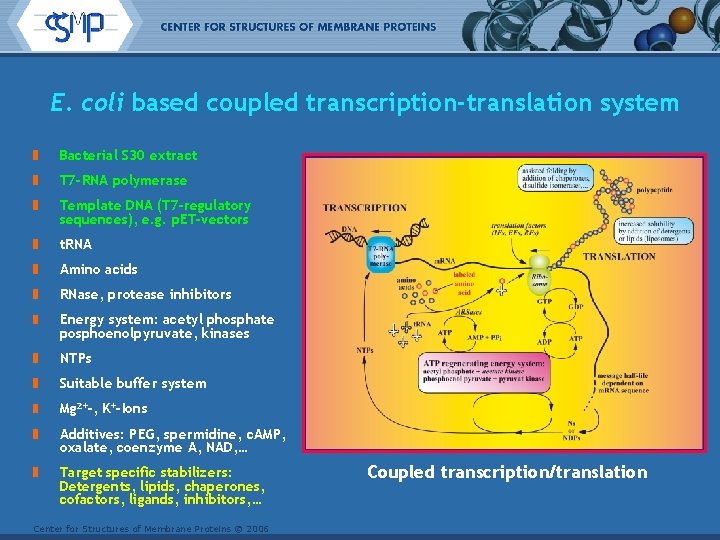
E. coli based coupled transcription-translation system Bacterial S 30 extract T 7 -RNA polymerase Template DNA (T 7 -regulatory sequences), e. g. p. ET-vectors t. RNA Amino acids RNase, protease inhibitors Energy system: acetyl phosphate posphoenolpyruvate, kinases NTPs Suitable buffer system Mg 2+-, K+-Ions Additives: PEG, spermidine, c. AMP, oxalate, coenzyme A, NAD, … Target specific stabilizers: Detergents, lipids, chaperones, cofactors, ligands, inhibitors, … Center for Structures of Membrane Proteins © 2006 Coupled transcription/translation

E. coli S 30 extract preparation (3 days) Cell growth (day 1): – Inoculate 5 liters of TB media at a ratio of 1: 100 with fresh overnight culture of E. coli strain (A 19, BL 21, D 10) in a fermentor and start rapid cooling when cells reach log-phase (OD 600 = 5 -6). All following steps on ice! Cell wash: – – Harvest cells by centrifugation (7, 000 x g, 4°C, 10 min) and resuspend in Buffer A (10 m. M Tris-Acetat, p. H 8. 2, 14 m. M Mg(o. Ac)2, 0. 6 m. M KCl, 6 m. M BME) using a glass rod. Centrifuge cells (8, 000 x g, 4°C, 30 min) and repeat washing, [STORE ON ICE OVER NIGHT], wash 3 rd time. Cell lysis (day 2): – Weight cells and resuspend in 110% w/v buffer B (10 m. M Tris-Acetat, p. H 8. 2, 14 m. M Mg(o. Ac)2, 0. 6 m. M KCl, 1 m. M DTT, 0. 1 m. M PMSF). Disrupt cells by French press (20, 000 psi) or cell disrupter at 4°C. Isolation of cell extract: – Centrifuge disrupted cells (30, 000 x g, 4°C, 30 min), keep first 2/3 of supernatant and centrifuge again. keep 2/3 of second supernatant. Run-off procedure: – Supplement extract with 400 m. M Na. Cl (5 M stock) and incubate for 45 min at 42°C. Dialysis: – Dialyse against 2 x 40 volumes over night with first buffer exchange after 2 h into S 30 buffer (10 m. M Tris-Acetat, p. H 8. 2, 14 m. M Mg(o. Ac)2, 0. 6 m. M Ko. Ac, 0. 5 m. M DTT at 4°C (25 k. Da MWCO membrane). Store extract (day 3) – Clear extract by centrifugation (30, 000 x g, 4°C, 30 min), aliquot and freeze in liquid N 2, store at -80°C. Center for Structures of Membrane Proteins © 2006

T 7 -RNA polymerase (T 7 RNAP) preparation (2 days) Cell growth, harvest (day 1): – – Inoculate 4 liters of LB in Erlenmeyer culture flasks at a ratio of 1: 100 with fresh over night culture of BL 21 Star x p. AR 1219. Incubate at 37°C with vigorous shaking until OD 600 of 0. 6 -0. 8. Induce T 7 RNAP with 1 m. M IPTG and incubate at 37°C with vigorous shaking for 5 h. Harvest cells by centrifugation (4, 500 x g, 4°C, 15 min) and resuspend cell pellet in 120 ml T 7 buffer (30 m. M Tris, p. H 8, 10 m. M EDTA, 50 m. M Na. Cl, 5% Glycerol, 10 m. M BME). Cell lysis: – – Disrupt cells with French press (20, 000 psi) or cell disrupter. Adjust the supernatant to final concentration of 2% streptomycin sulfate by gentle stirring and drop wise addition of 10% stock. Remove precipitated DNA from supernatant by centrifugation (30, 000 x g, 4°C, 30 min). Purification: – – Purify T 7 RNAP by anion exchange chromatography. Equilibrate Q-Sepharose column (1. 6 x 10 cm) with T 7 buffer, load supernatant with 1 ml/min flow rate. Elute T 7 RNAP in gradient from 50 m. M to 500 m. M Na. Cl in 15 column volumes at a 3 ml/min flow rate. T 7 RNAP starts to elute at ~150 m. M Na. Cl. Dialysis: – T 7 RNAP fractions are pooled and dialyzed against 2 x 100 volumes of dialysis buffer (10 m. M Tris, p. H 8, 1 m. M EDTA, 10 m. M Na. Cl, 10% glycerol, 1 m. M DTT) over night. Activity test (day 2): – T 7 RNAP activity is tested by in vitro transcription and units are determined by comparison with a commercial control. Store T 7 RNAP: – Store T 7 RNAP in aliquots in dialysis buffer adjusted to 50% Glycerol at -20°C. Center for Structures of Membrane Proteins © 2006
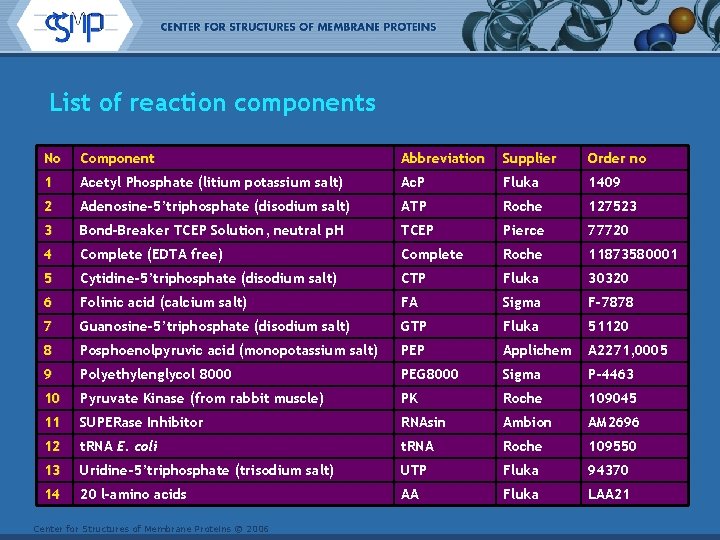
List of reaction components No Component Abbreviation Supplier Order no 1 Acetyl Phosphate (litium potassium salt) Ac. P Fluka 1409 2 Adenosine-5’triphosphate (disodium salt) ATP Roche 127523 3 Bond-Breaker TCEP Solution, neutral p. H TCEP Pierce 77720 4 Complete (EDTA free) Complete Roche 11873580001 5 Cytidine-5’triphosphate (disodium salt) CTP Fluka 30320 6 Folinic acid (calcium salt) FA Sigma F-7878 7 Guanosine-5’triphosphate (disodium salt) GTP Fluka 51120 8 Posphoenolpyruvic acid (monopotassium salt) PEP Applichem A 2271, 0005 9 Polyethylenglycol 8000 PEG 8000 Sigma P-4463 10 Pyruvate Kinase (from rabbit muscle) PK Roche 109045 11 SUPERase Inhibitor RNAsin Ambion AM 2696 12 t. RNA E. coli t. RNA Roche 109550 13 Uridine-5’triphosphate (trisodium salt) UTP Fluka 94370 14 20 l-amino acids AA Fluka LAA 21 Center for Structures of Membrane Proteins © 2006
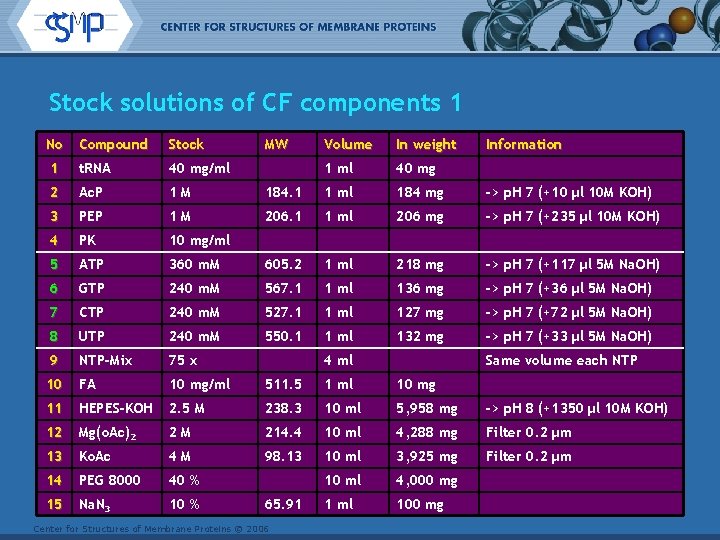
Stock solutions of CF components 1 No Compound Stock 1 t. RNA 40 mg/ml 2 Ac. P 1 M 3 PEP 1 M 4 PK 10 mg/ml 5 ATP 6 MW Volume In weight 1 ml 40 mg 184. 1 1 ml 184 mg -> p. H 7 (+10 µl 10 M KOH) 206. 1 1 ml 206 mg -> p. H 7 (+235 µl 10 M KOH) 360 m. M 605. 2 1 ml 218 mg -> p. H 7 (+117 µl 5 M Na. OH) GTP 240 m. M 567. 1 1 ml 136 mg -> p. H 7 (+36 µl 5 M Na. OH) 7 CTP 240 m. M 527. 1 1 ml 127 mg -> p. H 7 (+72 µl 5 M Na. OH) 8 UTP 240 m. M 550. 1 1 ml 132 mg -> p. H 7 (+33 µl 5 M Na. OH) 9 NTP-Mix 75 x 10 FA 10 mg/ml 511. 5 1 ml 10 mg 11 HEPES-KOH 2. 5 M 238. 3 10 ml 5, 958 mg -> p. H 8 (+1350 µl 10 M KOH) 12 Mg(o. Ac)2 2 M 214. 4 10 ml 4, 288 mg Filter 0. 2 µm 13 Ko. Ac 4 M 98. 13 10 ml 3, 925 mg Filter 0. 2 µm 14 PEG 8000 40 % 10 ml 4, 000 mg 15 Na. N 3 10 % 1 ml 100 mg 4 ml 65. 91 Center for Structures of Membrane Proteins © 2006 Information Same volume each NTP
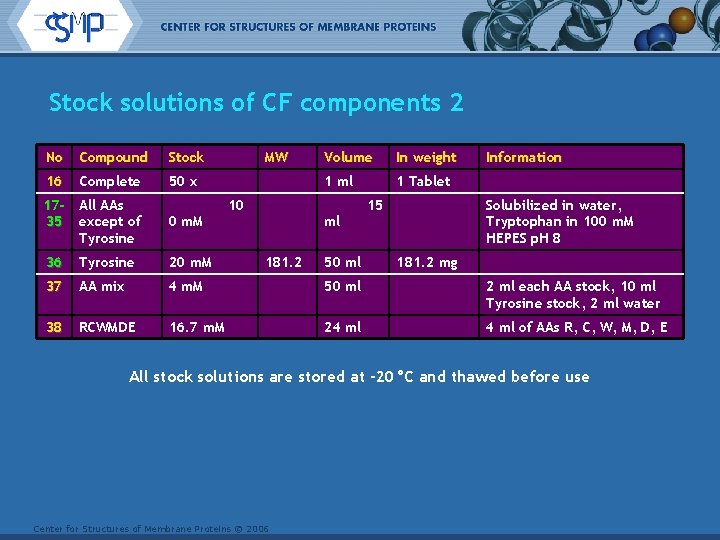
Stock solutions of CF components 2 No Compound Stock 16 Complete 50 x MW Volume In weight 1 ml 1 Tablet 10 15 Information 17 - All AAs 35 except of Tyrosine Solubilized in water, Tryptophan in 100 m. M HEPES p. H 8 0 m. M 36 Tyrosine 20 m. M 37 AA mix 4 m. M 50 ml 2 ml each AA stock, 10 ml Tyrosine stock, 2 ml water 38 RCWMDE 16. 7 m. M 24 ml of AAs R, C, W, M, D, E ml 181. 2 50 ml 181. 2 mg All stock solutions are stored at -20°C and thawed before use Center for Structures of Membrane Proteins © 2006
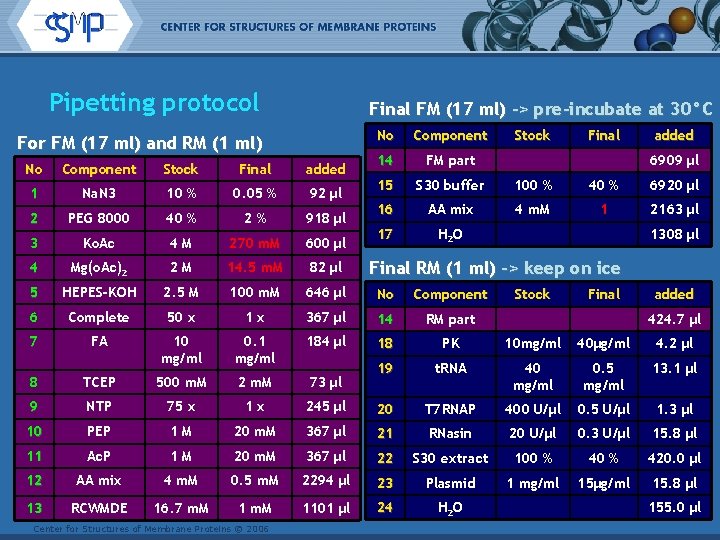
Pipetting protocol Final FM (17 ml) -> pre-incubate at 30°C For FM (17 ml) and RM (1 ml) No Component Stock Final added 14 FM part 15 S 30 buffer 100 % 40 % 6920 µl 16 AA mix 4 m. M 1 2163 µl 17 H 2 O 6909 µl No Component Stock Final added 1 Na. N 3 10 % 0. 05 % 92 µl 2 PEG 8000 40 % 2% 918 µl 3 Ko. Ac 4 M 270 m. M 600 µl 4 Mg(o. Ac)2 2 M 14. 5 m. M 82 µl 5 HEPES-KOH 2. 5 M 100 m. M 646 µl No Component 6 Complete 50 x 1 x 367 µl 14 RM part 7 FA 10 mg/ml 0. 1 mg/ml 184 µl 18 PK 10 mg/ml 40µg/ml 4. 2 µl 8 TCEP 500 m. M 2 m. M 73 µl 19 t. RNA 40 mg/ml 0. 5 mg/ml 13. 1 µl 9 NTP 75 x 1 x 245 µl 20 T 7 RNAP 400 U/µl 0. 5 U/µl 1. 3 µl 10 PEP 1 M 20 m. M 367 µl 21 RNasin 20 U/µl 0. 3 U/µl 15. 8 µl 11 Ac. P 1 M 20 m. M 367 µl 22 S 30 extract 100 % 420. 0 µl 12 AA mix 4 m. M 0. 5 m. M 2294 µl 23 Plasmid 1 mg/ml 15µg/ml 15. 8 µl 13 RCWMDE 16. 7 m. M 1101 µl 24 H 2 O Center for Structures of Membrane Proteins © 2006 1308 µl Final RM (1 ml) -> keep on ice Stock Final added 424. 7 µl 155. 0 µl

Reaction compartments for dialysis mode Analytical scale (50 -100 ul): Preparative scale (1 -3 ml): Slide-ALyzer MINI incubate at 30°C, 200 rpm 100 ul RM 1400 ul FM 75 ul RM 1050 ul FM Slide-A-Lyzer 3 ml RM 51 ml FM incubate at 30°C, 200 rpm 50 ul RM 700 ul FM No Volume MWCO Name Supplier Order no 1 50 -100 µl 20 k. Da Slide-A-Lyzer MINI Pierce 69590 2 0. 5 – 3 ml 20 k. Da Slide-A-Lyzer Pierce 66003 Center for Structures of Membrane Proteins © 2006
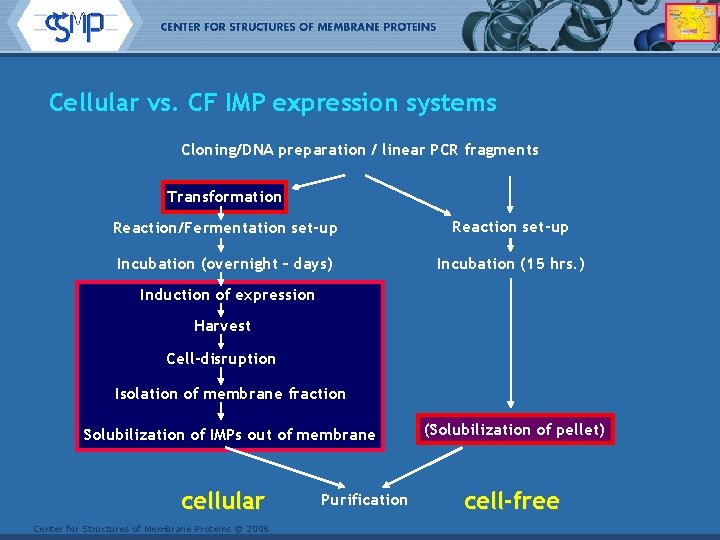
Cellular vs. CF IMP expression systems Cloning/DNA preparation / linear PCR fragments Transformation Reaction/Fermentation set-up Reaction set-up Incubation (overnight – days) Incubation (15 hrs. ) Induction of expression Harvest Cell-disruption Isolation of membrane fraction Solubilization of IMPs out of membrane cellular Center for Structures of Membrane Proteins © 2006 Purification (Solubilization of pellet) cell-free
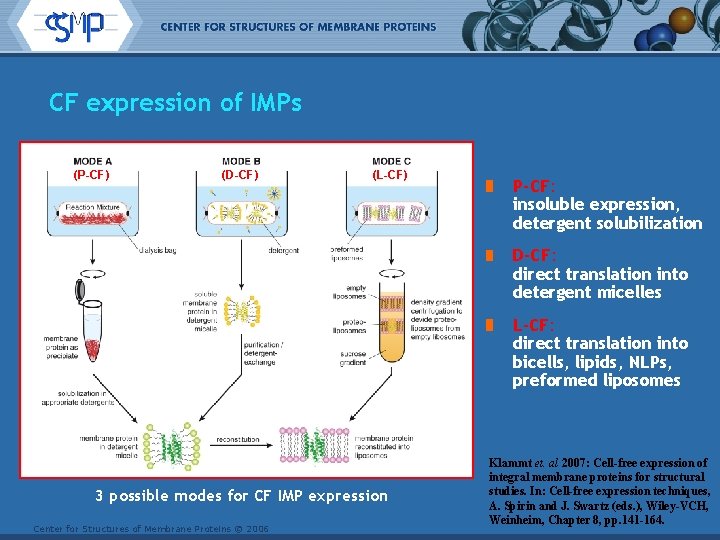
CF expression of IMPs (P-CF) (D-CF) (L-CF) P-CF: insoluble expression, detergent solubilization D-CF: direct translation into detergent micelles L-CF: direct translation into bicells, lipids, NLPs, preformed liposomes 3 possible modes for CF IMP expression Center for Structures of Membrane Proteins © 2006 Klammt et. al 2007: Cell-free expression of integral membrane proteins for structural studies. In: Cell-free expression techniques, A. Spirin and J. Swartz (eds. ), Wiley-VCH, Weinheim, Chapter 8, pp. 141 -164.
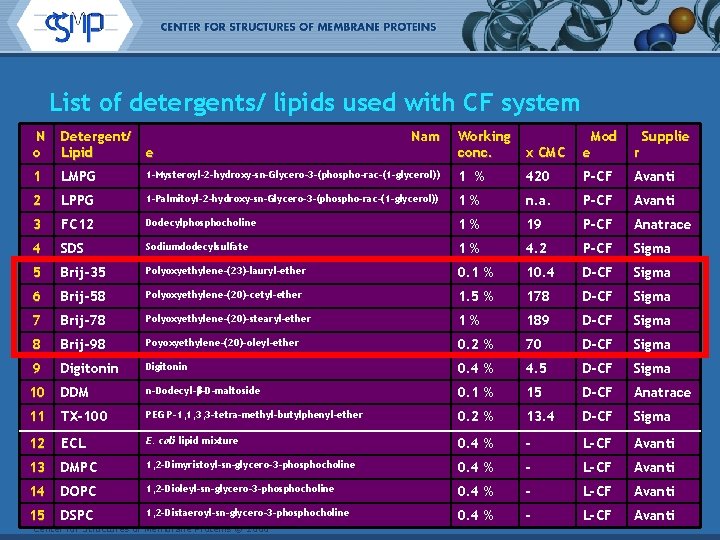
List of detergents/ lipids used with CF system N o Detergent/ Lipid e 1 LMPG 2 Working conc. x CMC Mod e r 1 -Mysteroyl-2 -hydroxy-sn-Glycero-3 -(phospho-rac-(1 -glycerol)) 1 % 420 P-CF Avanti LPPG 1 -Palmitoyl-2 -hydroxy-sn-Glycero-3 -(phospho-rac-(1 -glycerol)) 1% n. a. P-CF Avanti 3 FC 12 Dodecylphosphocholine 1% 19 P-CF Anatrace 4 SDS Sodiumdodecylsulfate 1% 4. 2 P-CF Sigma 5 Brij-35 Polyoxyethylene-(23)-lauryl-ether 0. 1 % 10. 4 D-CF Sigma 6 Brij-58 Polyoxyethylene-(20)-cetyl-ether 1. 5 % 178 D-CF Sigma 7 Brij-78 Polyoxyethylene-(20)-stearyl-ether 1% 189 D-CF Sigma 8 Brij-98 Poyoxyethylene-(20)-oleyl-ether 0. 2 % 70 D-CF Sigma 9 Digitonin 0. 4 % 4. 5 D-CF Sigma 10 DDM n-Dodecyl-b-D-maltoside 0. 1 % 15 D-CF Anatrace 11 TX-100 PEG P-1, 1, 3, 3 -tetra-methyl-butylphenyl-ether 0. 2 % 13. 4 D-CF Sigma 12 ECL E. coli lipid mixture 0. 4 % - L-CF Avanti 13 DMPC 1, 2 -Dimyristoyl-sn-glycero-3 -phosphocholine 0. 4 % - L-CF Avanti 14 DOPC 1, 2 -Dioleyl-sn-glycero-3 -phosphocholine 0. 4 % - L-CF Avanti 15 DSPC 1, 2 -Distaeroyl-sn-glycero-3 -phosphocholine 0. 4 % - L-CF Avanti Center for Structures of Membrane Proteins © 2006 Nam Supplie

Trouble shooting No IMP product formation – – – Verify efficiency of CF system by expression controls (e. g. , with GFP) Ensure high quality of template DNA Modify DNA template design (e. g. , addition of expression tags) Analyze m. RNA secondary structures to ensure efficient initiation and elongation of translation Ensure compatibility of extra compounds supplied with CF system Low IMP expression levels – – Optimize Mg 2+ and K+ ion concentration Adjust amino acid composition Increase t. RNA concentration Try potentially beneficial additives Low IMP solubilization in D-CF – – Evaluate a series of detergent types Optimize the final detergent concentration Provide detergent mixtures or combinations of detergents/lipids Decrease expression temperature Production of IMP fragments – – – Stabilize IMP by protease inhibitors during CF expression Increase t. RNA concentration Use S 30 extracts from protease-negative strains Center for Structures of Membrane Proteins © 2006

N-CF – a new mode for CF expression of IMPs The polymer NVoy (Expedeon) enables soluble CF expression and analysis of functional GPCRs Analyzed GPCRs in the presence of NVoy are homogenous, functional and can be analyzed by NMR ligand binding N-CF expression of diverse GPCRs no aggregation homogenous by SEC-UV/RI/LS Center for Structures of Membrane Proteins © 2006 preliminary NMR data

Recommended references Klammt et al. 2004 (EJB) – P-CF protocol for CF IMP expression Klammt, Schwarz et al. 2005 (FEBS J) – detailed D-CF protocol for different IMPs Klammt et al. 2006 (FEBS J) – detailed review about CF IMP in P-CF, D-CF Klammt et al. 2007 (MMB) – detailed expression protocols for extract preparation and P-CF and D-CF set-up Schwarz et al. 2007 (Nature Prot) – detailed expression protocols for extract and T 7 RNAP preparation and P-CF, D-CF Klammt et al. 2007 (JSB) – CF expression of GPCRs Schwarz et al. 2008 (Proteomics) – recent review about CF IMP in P-CF, D-CF, L-CF Klammt et al. 2009 (submitted) – N-CF expression of functional GPCRs Center for Structures of Membrane Proteins © 2006
- Slides: 20