Cell Structure and Biology Advanced Placement Biology Chapter

Cell Structure and Biology Advanced Placement Biology Chapter 6
Robert Hooke, 1665

Hooke’s First Microscope

History of the Cell • 1665 - Robert Hooke described cork as composed of cellulae (cell). • A few years later-Anton van Leeuwenhoek described live cells. • 1838 and 1839 - Schleiden and Schwann developed the cell theory.

Schleiden and Schwann- Cell Theory • All organisms are composed of cells or at least one. • Cells are the smallest unit of life (a collection of metabolic processes + heredity). • All cells come from other cells. None spontaneously arise.
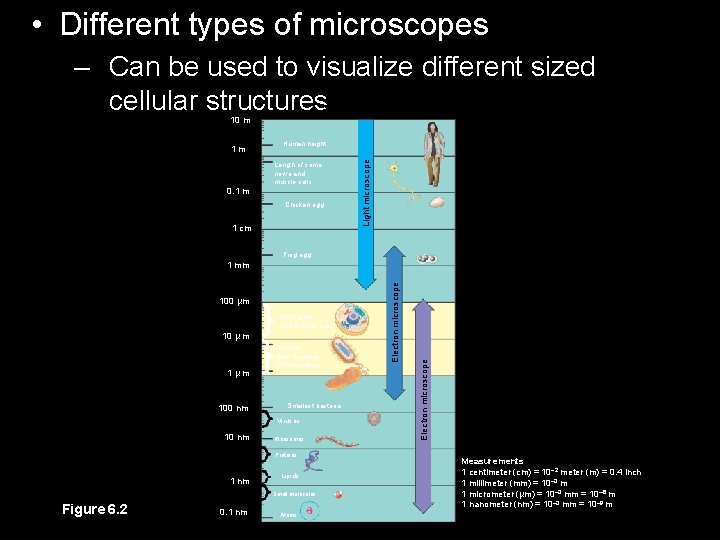
• Different types of microscopes Unaided eye – Can be used to visualize different sized cellular structures 10 m Human height Length of some nerve and muscle cells 0. 1 m Chicken egg 1 cm Light microscope 1 m Frog egg Most plant and Animal cells 10 µ m 1µm 100 nm Nucleus Most bacteria Mitochondrion Smallest bacteria Viruses 10 nm Ribosomes Proteins 1 nm Lipids Small molecules Figure 6. 2 0. 1 nm Atoms Electron microscope 100 µm Electron microscope 1 mm Measurements 1 centimeter (cm) = 10 2 meter (m) = 0. 4 inch 1 millimeter (mm) = 10– 3 m 1 micrometer (µm) = 10– 3 mm = 10– 6 m 1 nanometer (nm) = 10– 3 mm = 10– 9 m

Use different methods for enhancing visualization of cellular structures TECHNIQUE (a) RESULT Brightfield (unstained specimen). Passes light directly through specimen. Unless cell is naturally pigmented or artificially stained, image has little contrast. [Parts (a)–(d) show a human cheek epithelial cell. ] 50 µm (b) (c) Figure 6. 3 Brightfield (stained specimen). Staining with various dyes enhances contrast, but most staining procedures require that cells be fixed (preserved). Phase-contrast. Enhances contrast in unstained cells by amplifying variations in density within specimen; especially useful for examining living, unpigmented cells.
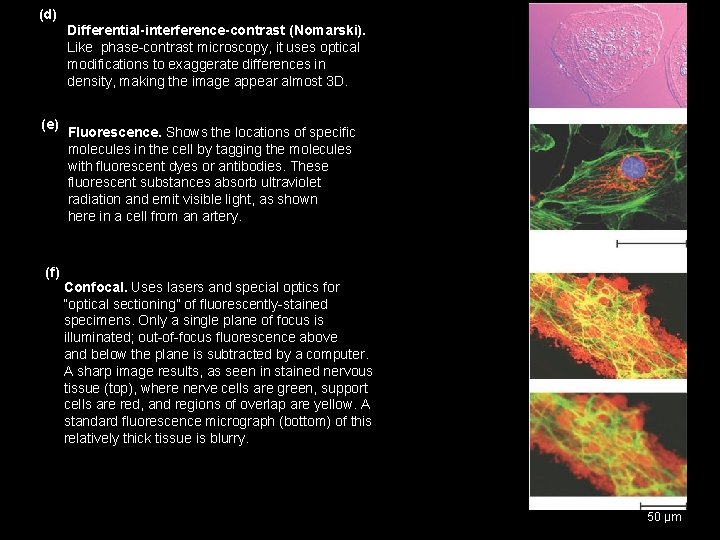
(d) Differential-interference-contrast (Nomarski). Like phase-contrast microscopy, it uses optical modifications to exaggerate differences in density, making the image appear almost 3 D. (e) Fluorescence. Shows the locations of specific molecules in the cell by tagging the molecules with fluorescent dyes or antibodies. These fluorescent substances absorb ultraviolet radiation and emit visible light, as shown here in a cell from an artery. 50 µm (f) Confocal. Uses lasers and special optics for “optical sectioning” of fluorescently-stained specimens. Only a single plane of focus is illuminated; out-of-focus fluorescence above and below the plane is subtracted by a computer. A sharp image results, as seen in stained nervous tissue (top), where nerve cells are green, support cells are red, and regions of overlap are yellow. A standard fluorescence micrograph (bottom) of this relatively thick tissue is blurry. 50 µm

• The scanning electron microscope (SEM) – Provides for detailed study of the surface of a specimen TECHNIQUE RESULTS 1 µm Cilia (a) Scanning electron microscopy (SEM). Micrographs taken with a scanning electron microscope show a 3 D image of the surface of a specimen. This SEM shows the surface of a cell from a rabbit trachea (windpipe) covered with motile organelles called cilia. Beating of the cilia helps move inhaled debris upward toward the throat. Figure 6. 4 (a)

• The transmission electron microscope (TEM) – Provides for detailed study of the internal ultrastructure of cells Cross section Longitudinal section of cilium (b) Transmission electron microscopy (TEM). A transmission electron microscope profiles a thin section of a specimen. Here we see a section through a tracheal cell, revealing its ultrastructure. In preparing the TEM, some cilia were cut along their lengths, creating longitudinal sections, while other cilia were cut straight across, creating cross sections. Figure 6. 4 (b) of cilium 1 µm

• The process of cell fractionation APPLICATION Cell fractionation is used to isolate (fractionate) cell components, based on size and density. TECHNIQUE First, cells are homogenized in a blender to break them up. The resulting mixture (cell homogenate) is then centrifuged at various speeds and durations to fractionate the cell components, forming a series of pellets. Figure 6. 5
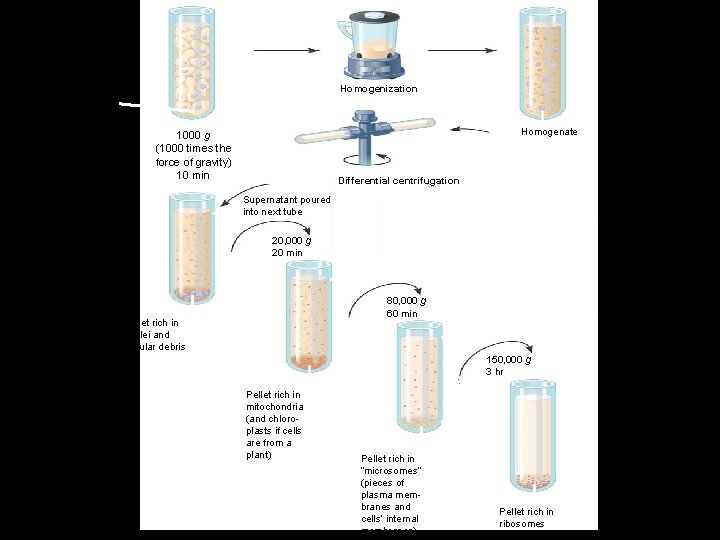
Homogenization Tissue cells Homogenate 1000 g (1000 times the force of gravity) 10 min Differential centrifugation Supernatant poured into next tube 20, 000 g 20 min 80, 000 g 60 min Pellet rich in nuclei and cellular debris 150, 000 g 3 hr Pellet rich in mitochondria (and chloroplasts if cells are from a plant) Figure 6. 5 Pellet rich in “microsomes” (pieces of plasma membranes and cells’ internal membranes) Pellet rich in ribosomes
What’s the world’s largest living cell? Surface to Volume Ratio
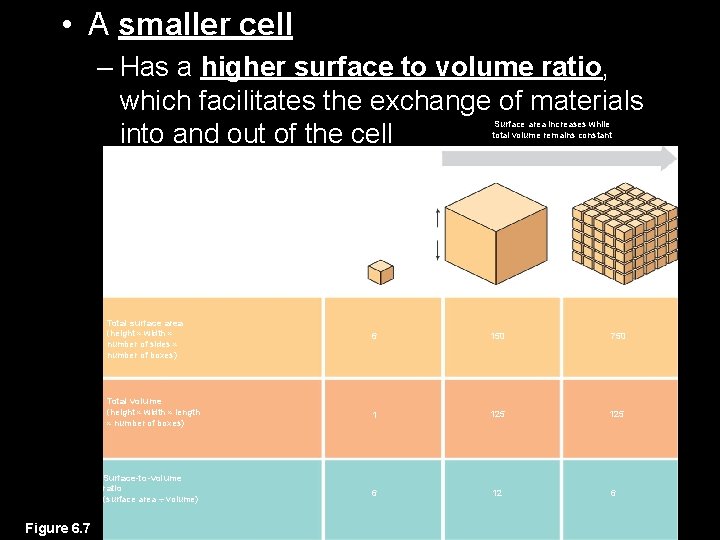
• A smaller cell – Has a higher surface to volume ratio, which facilitates the exchange of materials into and out of the cell Surface area increases while total volume remains constant 5 1 1 Total surface area (height width number of sides number of boxes) 6 150 750 Total volume (height width length number of boxes) 1 125 6 12 6 Surface-to-volume ratio (surface area volume) Figure 6. 7
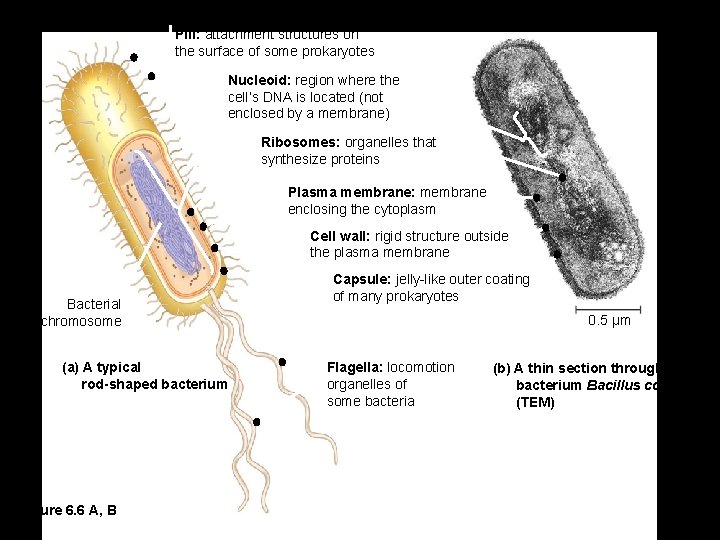
Pili: attachment structures on the surface of some prokaryotes Nucleoid: region where the cell’s DNA is located (not enclosed by a membrane) Ribosomes: organelles that synthesize proteins Plasma membrane: membrane enclosing the cytoplasm Cell wall: rigid structure outside the plasma membrane Bacterial chromosome (a) A typical rod-shaped bacterium Figure 6. 6 A, B Capsule: jelly-like outer coating of many prokaryotes 0. 5 µm Flagella: locomotion organelles of some bacteria (b) A thin section through the bacterium Bacillus coagulans (TEM)

Prokaryote vs. Eukaryote Archaebacteria and Animalia, Plantae, Protista, Eubacteria. Fungi. Lack membrane-bound Have true membraneorganelles. bound organelles. DNA in a nucleoid DNA in a nucleus. region. Have plasma membrane. Plants and some protists Cell wall of have a cell wall of peptidoglycan. cellulose. Use 70 S ribosome. Use different ribosomes. Unique flagella-flagellin protein.
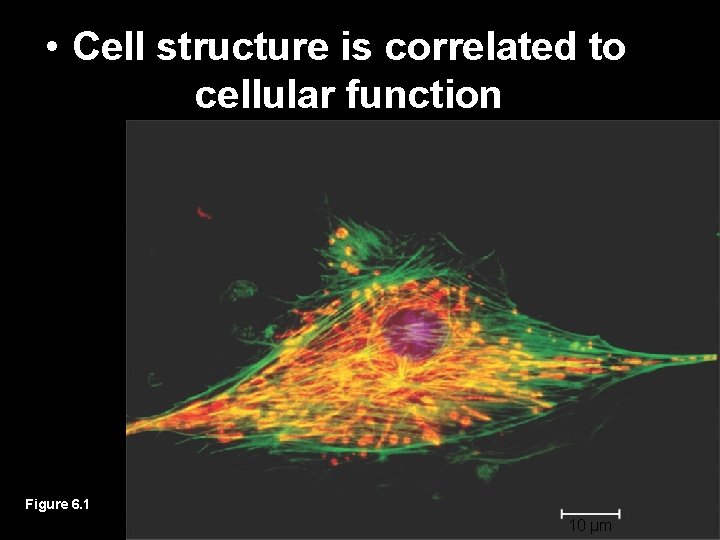
• Cell structure is correlated to cellular function Figure 6. 1 10 µm
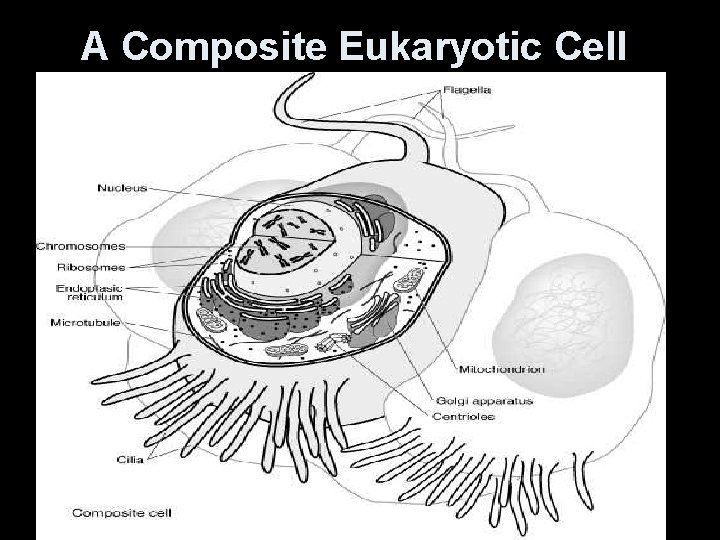
A Composite Eukaryotic Cell
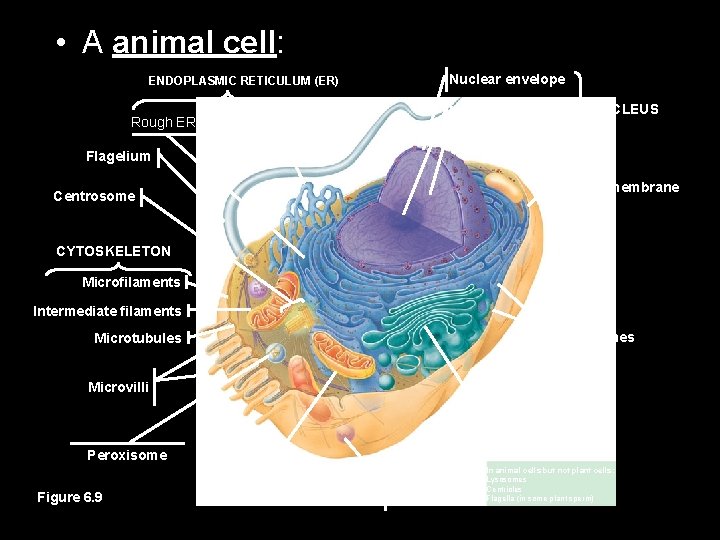
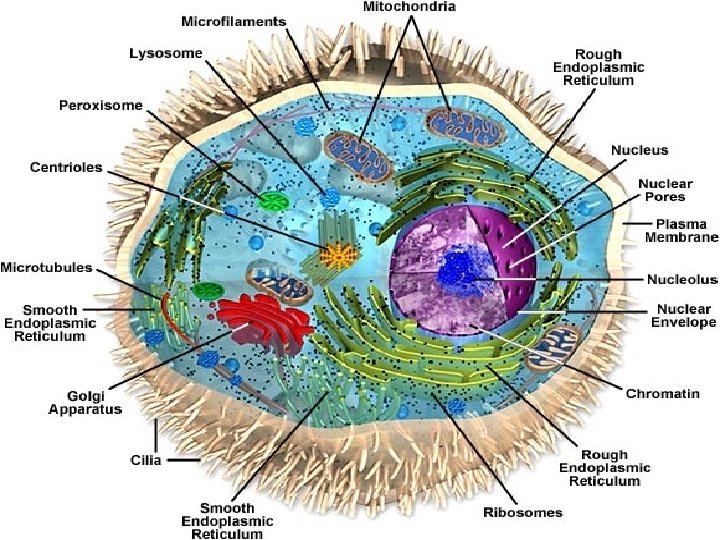
• A animal cell: ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Nuclear envelope Nucleolus NUCLEUS Chromatin Flagelium Plasma membrane Centrosome CYTOSKELETON Microfilaments Intermediate filaments Ribosomes Microtubules Microvilli Golgi apparatus Peroxisome Figure 6. 9 Mitochondrion Lysosome In animal cells but not plant cells: Lysosomes Centrioles Flagella (in some plant sperm)
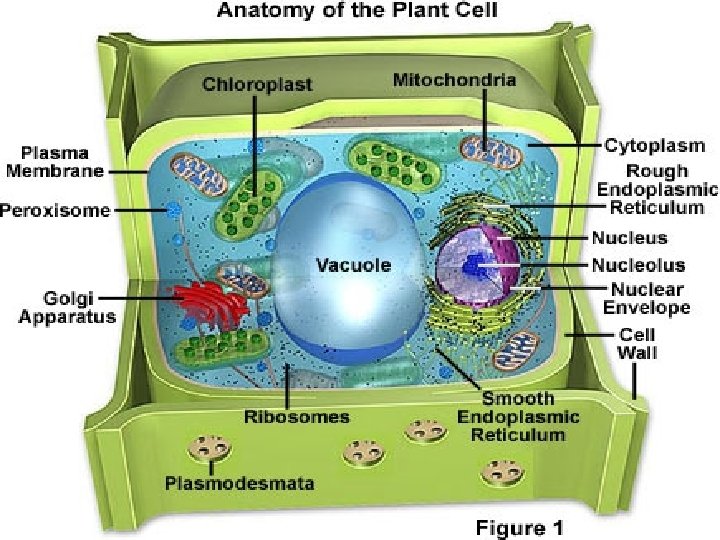
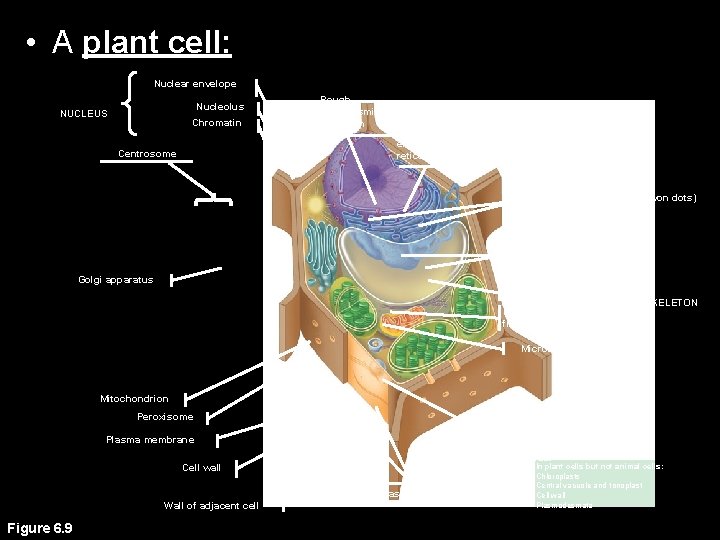
• A plant cell: Nuclear envelope Nucleolus NUCLEUS Chromatin Centrosome Rough endoplasmic reticulum Smooth endoplasmic reticulum Ribosomes (small brwon dots) Central vacuole Tonoplast Golgi apparatus Microfilaments Intermediate filaments CYTOSKELETON Microtubules Mitochondrion Peroxisome Plasma membrane Chloroplast Cell wall Plasmodesmata Wall of adjacent cell Figure 6. 9 In plant cells but not animal cells: Chloroplasts Central vacuole and tonoplast Cell wall Plasmodesmata
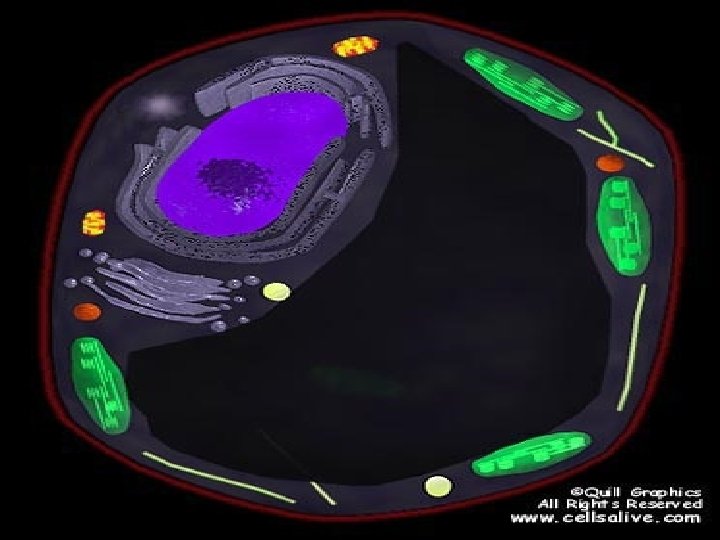
Corn Plant Cell
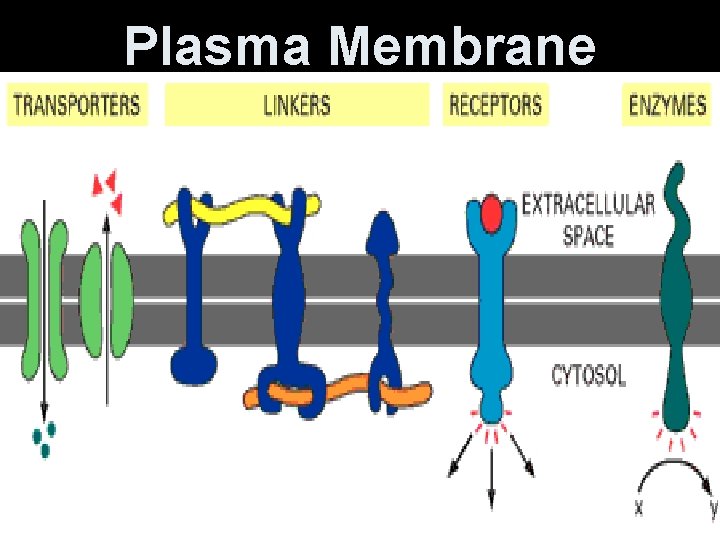
Plasma Membrane
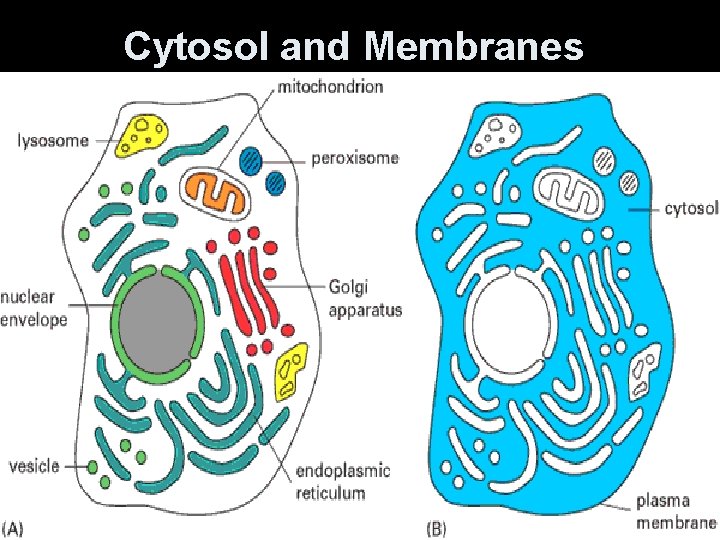
Cytosol and Membranes

What is the function of organelles? • To compartmentalize chemical reactions that may proceed simultaneously. • To provide membranes on which to catalyze reactions.
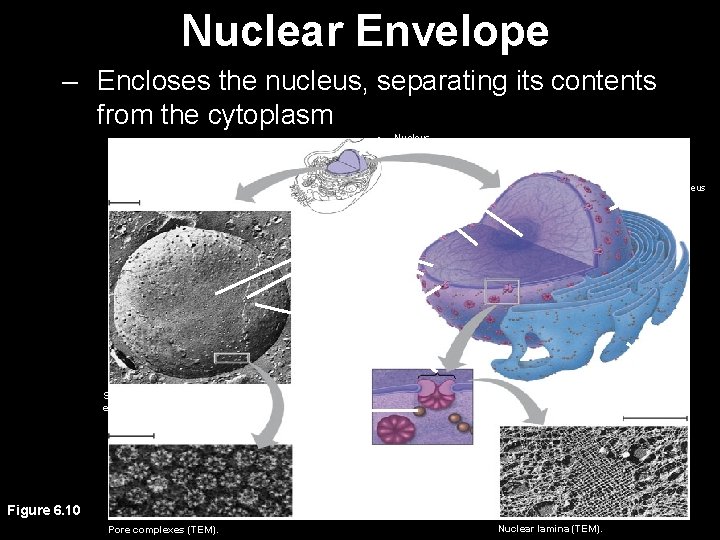
Nuclear Envelope – Encloses the nucleus, separating its contents from the cytoplasm Nucleus 1 µm Nucleus Nucleolus Chromatin Nuclear envelope: Inner membrane Outer membrane Nuclear pore Pore complex Rough ER Surface of nuclear envelope. 1 µm Ribosome 0. 25 µm Close-up of nuclear envelope Figure 6. 10 Pore complexes (TEM). Nuclear lamina (TEM).
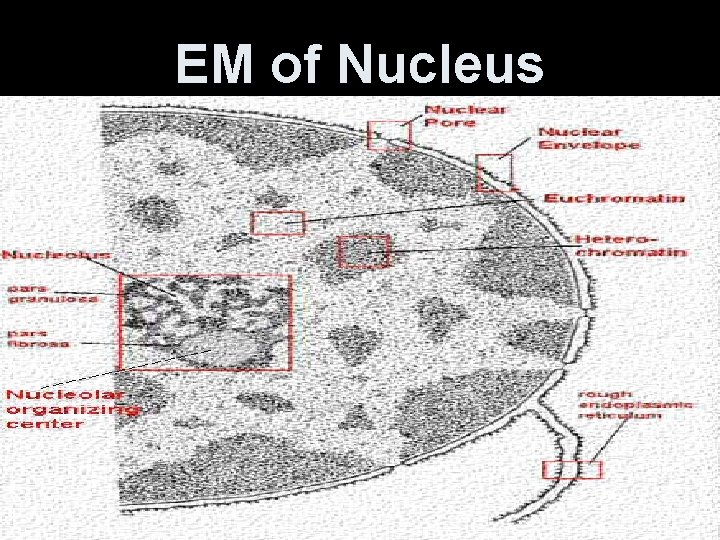
EM of Nucleus

1. The Nucleus • Largest organelle, centralized in animal cells. • Stores and protects the cell’s genetic information. • Surrounded by two phospholipid bilayer membranes-nuclear envelope. • Where both layers are fused - nuclear pores + transport protein.

The Nucleolus • Site within the nucleus of ribosomal subunits are manufactured- r. RNA + ribosomal proteins. • Ribosomes leave the nucleus as subunits through the nuclear pore and are later reassembled. • May be free (in the cytoplasm) or attached to the ER (rough ER).
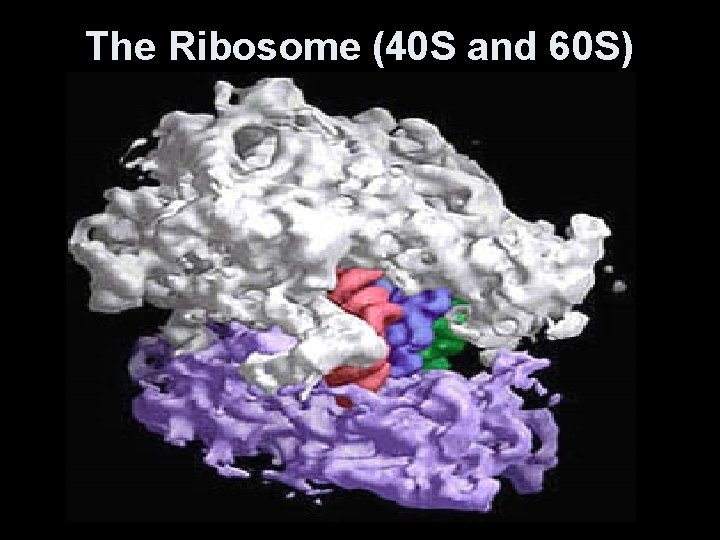
The Ribosome (40 S and 60 S)
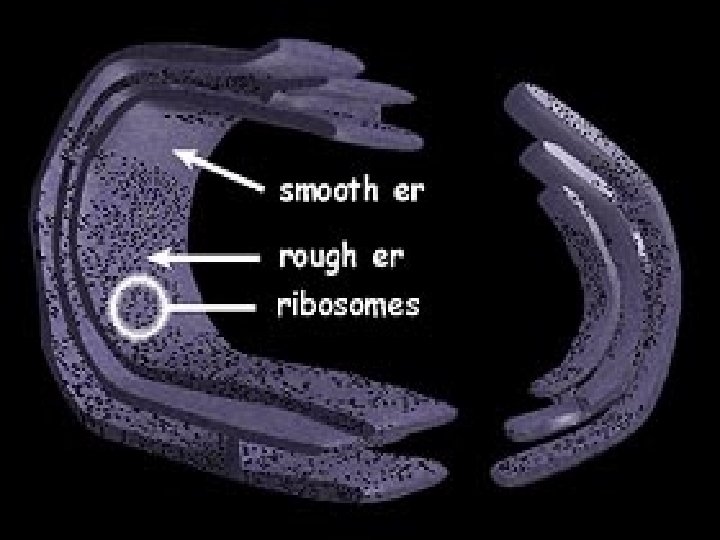
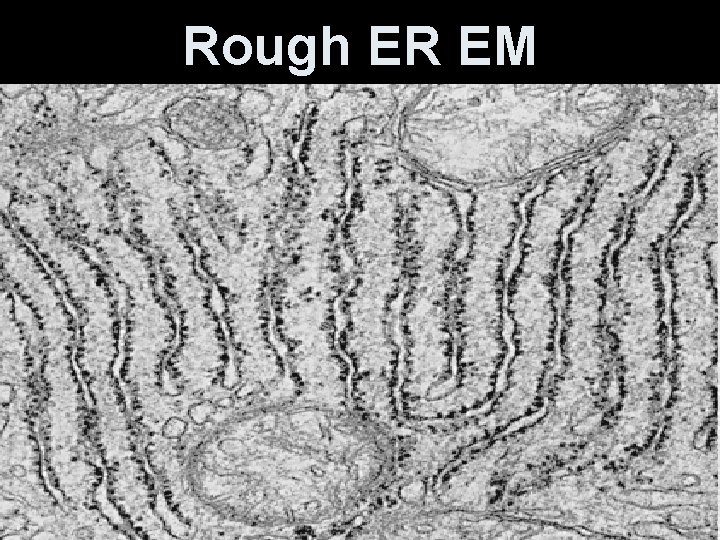
Rough ER EM

Endoplasmic Reticulum (ER) • Means “little net within the cytoplasm” • Internal membrane system with a lipid bilayer + proteins. • Weaved in sheets- forming channels. • Outer membrane of the nuclear envelope is continuous with the ER membrane. • Some regions have embedded ribosomes.
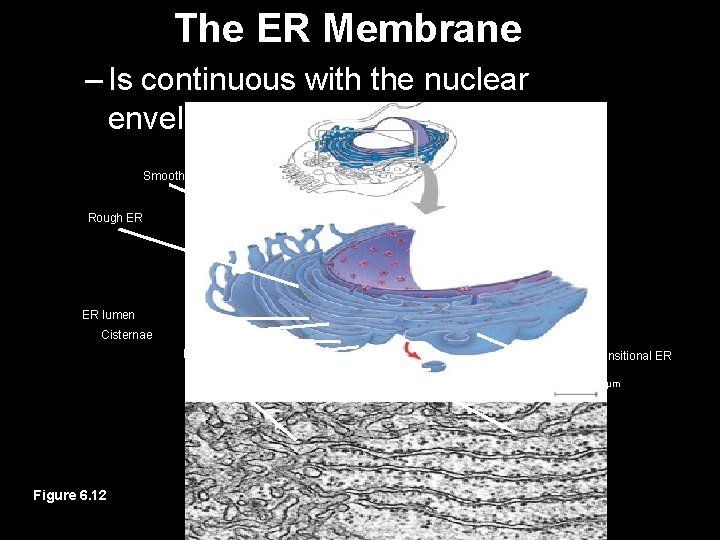
The ER Membrane – Is continuous with the nuclear envelope Smooth ER Nuclear envelope Rough ER ER lumen Cisternae Ribosomes Transport vesicle Smooth ER Figure 6. 12 Transitional ER Rough ER 200 µm

Two Types of (ER) 1. Rough ER: heavily studded with ribosomes- protein synthesis. Proteins have signal sequences which direct to a docking site on the surface of the ER. 2. Smooth ER: lack ribosomes; have enzymes embedded in membrane for carbohydrate and lipid synthesis. 3. Both secrete finished products in transport vesicles.

Functions of Smooth ER The smooth ER: –Synthesizes lipids –Metabolizes carbohydrates –Stores calcium –Detoxifies poison

The Golgi Complex • Flattened stacks of membranes in the cytoplasm-cisternae. • Collection, packaging and distribution of proteins and lipids. • Transport vesicles from RER and SER fuse with the Golgi membrane.

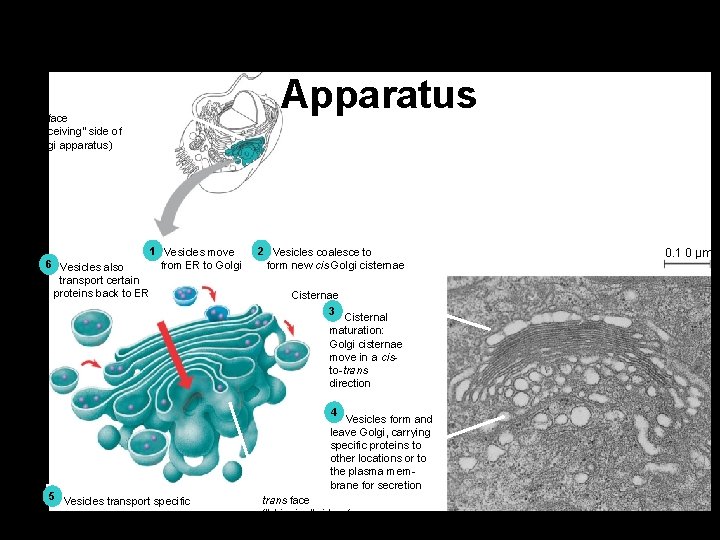
Functions of the Golgi Apparatus Golgi cis face (“receiving” side apparatus of Golgi apparatus) 1 Vesicles move from ER to Golgi 6 Vesicles also transport certain proteins back to ER 2 Vesicles coalesce to form new cis Golgi cisternae 0. 1 0 µm Cisternae 3 Cisternal maturation: Golgi cisternae move in a cisto-trans direction 4 5 Vesicles transport specific proteins backward to newer Golgi cisternae Vesicles form and leave Golgi, carrying specific proteins to other locations or to the plasma membrane for secretion trans face (“shipping” side of Golgi apparatus) TEM of Golgi apparatus Figure 6. 13
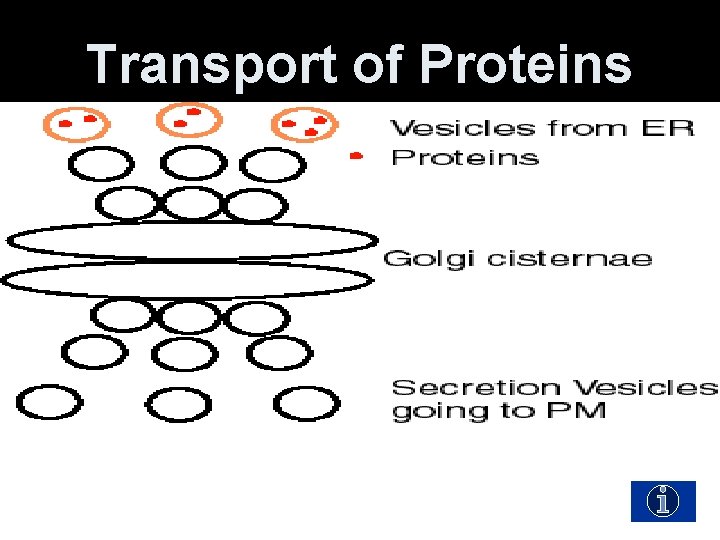
Transport of Proteins
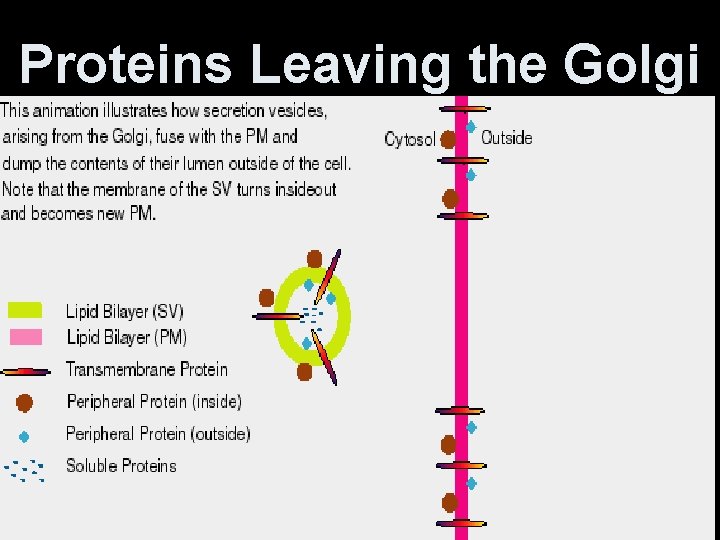
Proteins Leaving the Golgi

The Golgi Complex • Proteins (from RER) may have short sugar chains added--> glycoproteins. • Lipids (from SER) may have short sugar chains added-->glycolipids. • Both collect at flattened ends-cisternae. • Cisternae membranes pinch off the glycoproteins and glycolipids into secretory vesicles (liposomes). • Liposomes may fuse with plasma membane or organelle membranes.

Lysosomes • Membrane-bound organelle with digestive enzymes. • Breakdown protein, nucleic acid, carbos, lipids. • Digest old organelles and invading bacterial cells. • Digestive enzymes only active at low p. H.
Lysosomes

Lysosomes • Inactive lysosomes-Primary Lysosomes, high p. H, enzymes are inactive. • Once fused with food vacuole- pump H+ into compartment- active, Secondary Lysosomes. • Involved in normal cell death and programmed cell death (apoptosis). • Ex. Tadpole tail tissue; webbing between human fingers.

Peroxisomes: Oxidation • Peroxisomes: – Produce hydrogen peroxide and convert it to water. Chloroplast Peroxisome Mitochondrion Figure 6. 19 1 µm
EM of Peroxisome
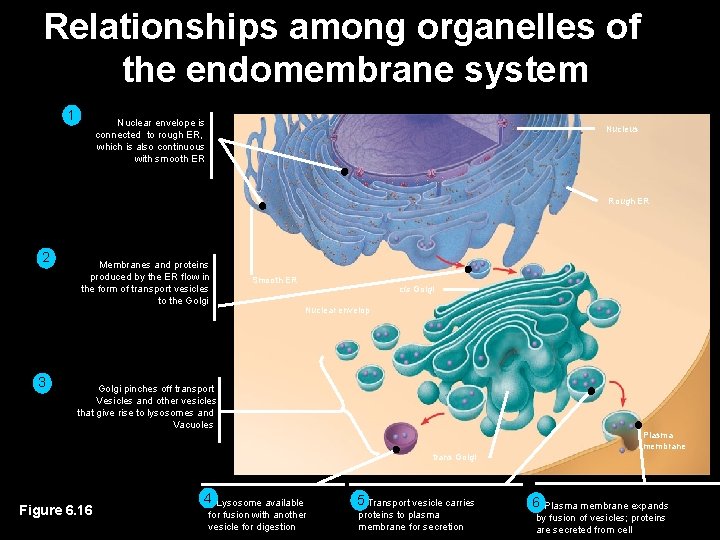
Relationships among organelles of the endomembrane system 1 Nuclear envelope is connected to rough ER, which is also continuous with smooth ER Nucleus Rough ER 2 3 Membranes and proteins produced by the ER flow in the form of transport vesicles to the Golgi Smooth ER cis Golgi Nuclear envelop Golgi pinches off transport Vesicles and other vesicles that give rise to lysosomes and Vacuoles Plasma membrane trans Golgi Figure 6. 16 4 Lysosome available for fusion with another vesicle for digestion 5 Transport vesicle carries proteins to plasma membrane for secretion 6 Plasma membrane expands by fusion of vesicles; proteins are secreted from cell
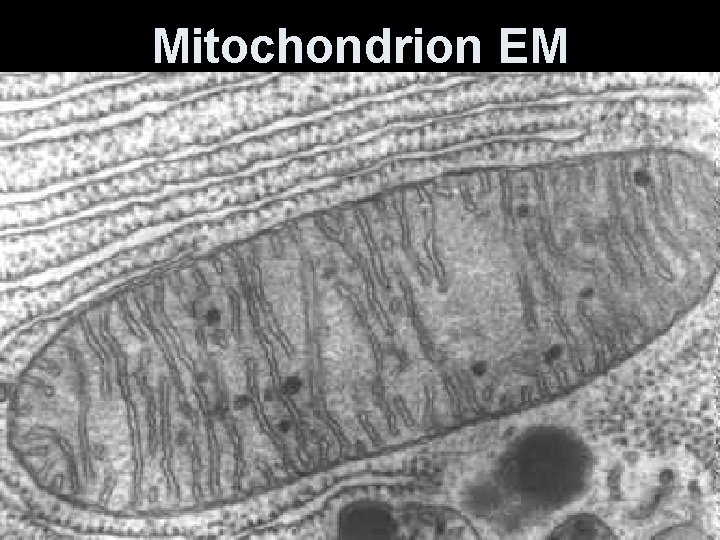
Mitochondrion EM
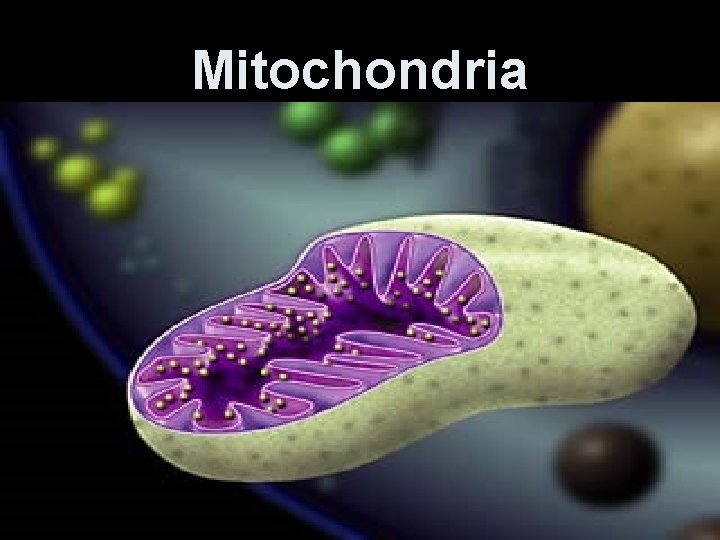
Mitochondria

Mitochondrion (ia) • Rod-shaped organelle, 1 -3 micrometers long. • Bounded by two membranes- outer is smooth; inner is folded into continuous layers-cristae. • Two compartments- matrix-inside the inner membrane and intermembrane space between the two membranes. • Enzymes for oxidative metabolism are embedded in the inner membrane.
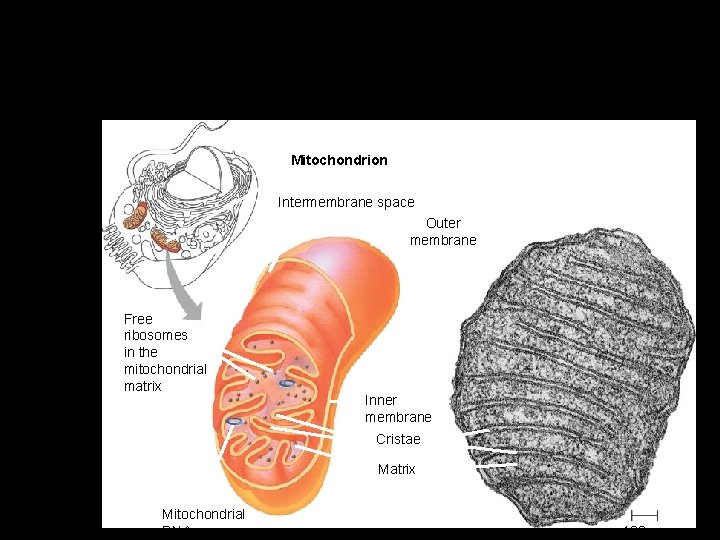
Mitochondria are enclosed by two membranes – A smooth outer membrane – An inner membrane folded into cristae Mitochondrion Intermembrane space Outer membrane Free ribosomes in the mitochondrial matrix Inner membrane Cristae Matrix Figure 6. 17 Mitochondrial DNA 100 µm

Mitochondria • Contain a circular piece of DNA for many of the proteins in oxidative metabolism. • Also has its own r. RNA and ribosomal proteins--> own protein synthesis. • Involved in its own replication. • Circular DNA? Two membranes? Own Genes? Own replication? • What does that sound like?

The Plastids • Chloroplasts • Leucoplasts • Amyloplasts • Chromoplasts
EM of Chloroplast
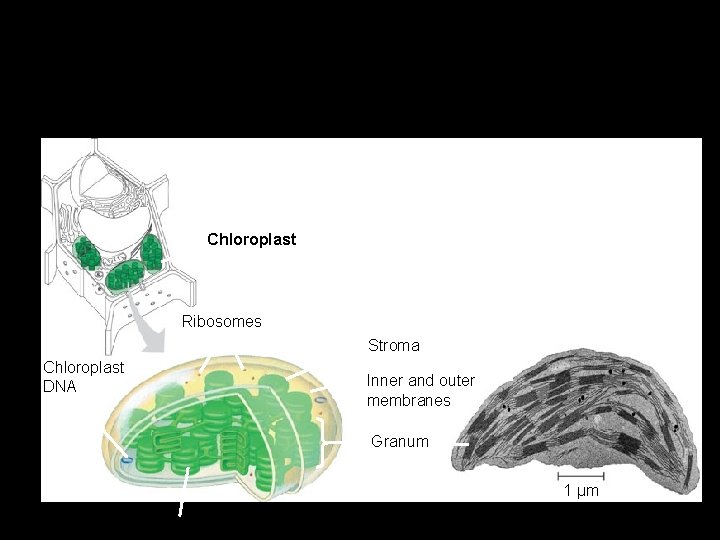
Chloroplasts – Are found in leaves and other green organs of plants and in algae. Chloroplast Ribosomes Stroma Chloroplast DNA Inner and outer membranes Granum 1 µm Figure 6. 18 Thylakoid

Chloroplasts • Algae and plants have organelles for photosynthesis. • Two membranes- outer and inner membranes. • A closed, stacked network of membranesgranum (a). • Fluid-filled space around grana-stroma. • Disc-shaped structures-thylakoids. • Light-capturing enzymes are embedded on thylakoids.

Chloroplasts • Have DNA which encode many enzymes necessary for photosynthesis. • Do all plant cells have chloroplasts? • May lose internal structure-leucoplasts. • A leucoplast that stores starchamyloplast. Found in root cells. • A leucoplast that stores other pigmentschromoplasts.

Centriole • Barrel-shaped organelles in animals and protists, NOT plants. • Usually found in pairs around the nuclear membrane. • Hollow cylinders made of microtubules (protein). Have their own DNA. • Help move chromosome during cell division.
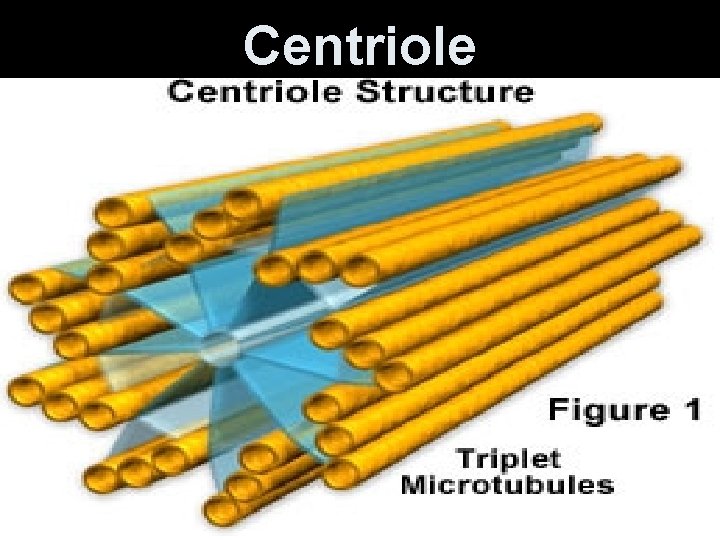
Centriole

Other Organelles • Central Vacuole or Tonoplast: in plants, for protein, water, and waste storage. • Vesicles: in animals, usually smaller sacs used for storage and transport of materials.
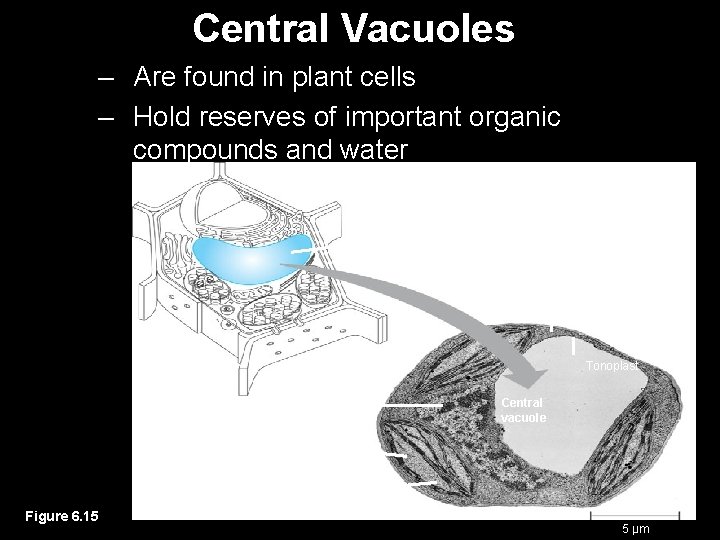
Central Vacuoles – Are found in plant cells – Hold reserves of important organic compounds and water Central vacuole Cytosol Tonoplast Nucleus Central vacuole Cell wall Chloroplast Figure 6. 15 5 µm

The Cytoskeleton!
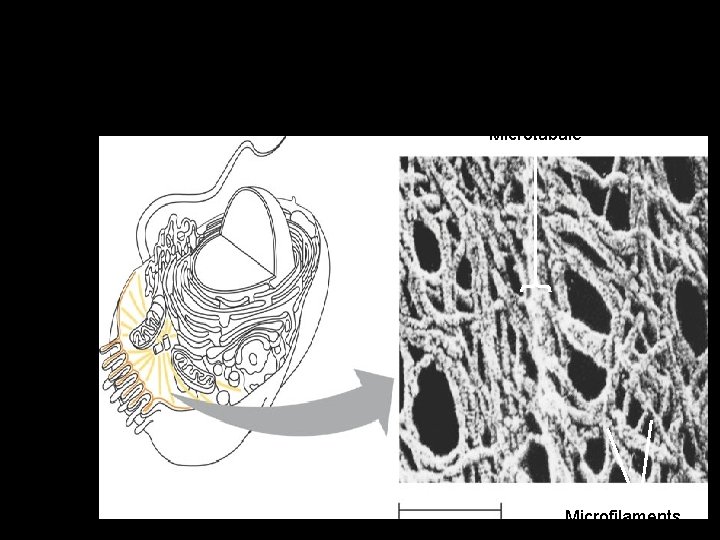
Figure 6. 20 Cytoskeleton – Is a network of fibers extending throughout the cytoplasm Microtubule 0. 25 µm Microfilaments

• There are three main types of fibers that make up the cytoskeleton: Table 6. 1
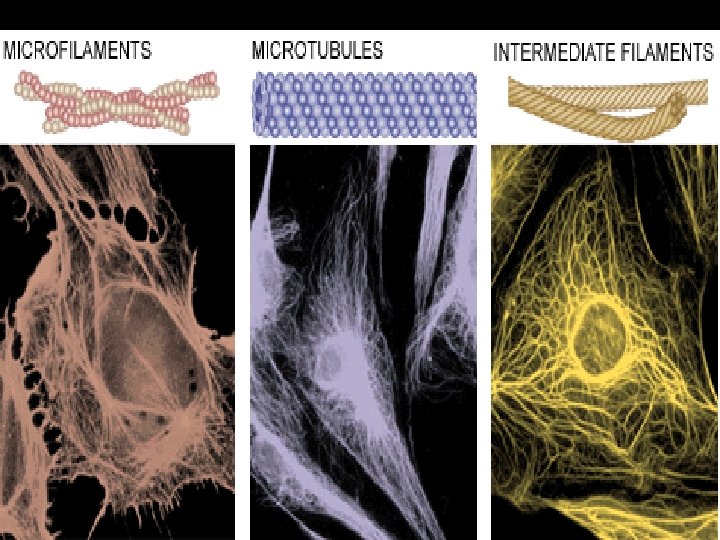

Actin Filaments • Made of globular protein monomers - actin • Actin monomers polymerize to form actin filaments • Filaments are connected to proteins within the plasma membrane.
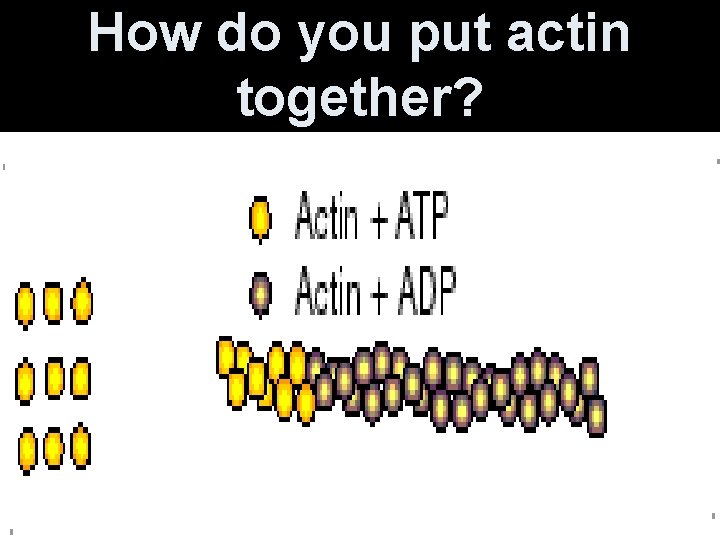
How do you put actin together?

Actin Filaments • Actin filaments are thinner, cause cellular movements like ameboid movements, cell pinching during division. • Provide shape for the cell.
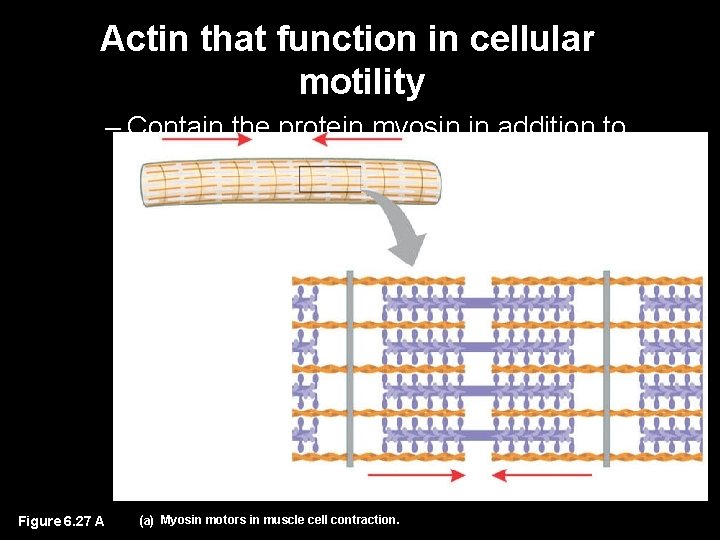
Actin that function in cellular motility – Contain the protein myosin in addition to actin Muscle cell Actin filament Myosin arm Figure 6. 27 A (a) Myosin motors in muscle cell contraction.

Microtubules • 2 globular monomers- tubulin and tubulin polymerize to form 13 protofilaments • Filaments form wide, hollow tubesmicrotubules. • Form from nucleation centers (near nucleus) and radiate out.
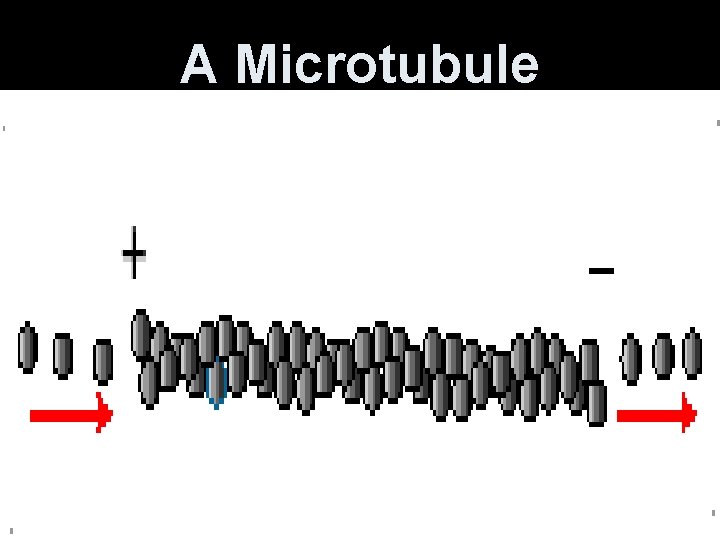
A Microtubule
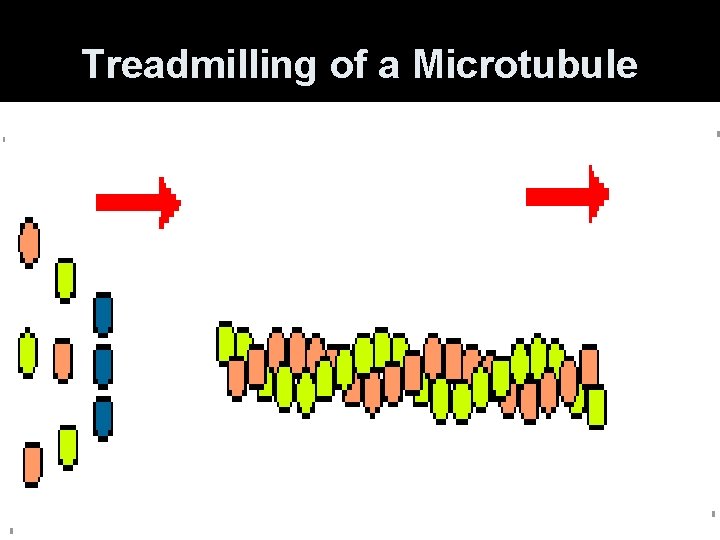
Treadmilling of a Microtubule

Microtubules • Constantly polymerize and depolymerize. GTP-binding at ends. • Ends are + (away from center) or (toward center). • Cellular movements and intracellular movement of materials and organelles.

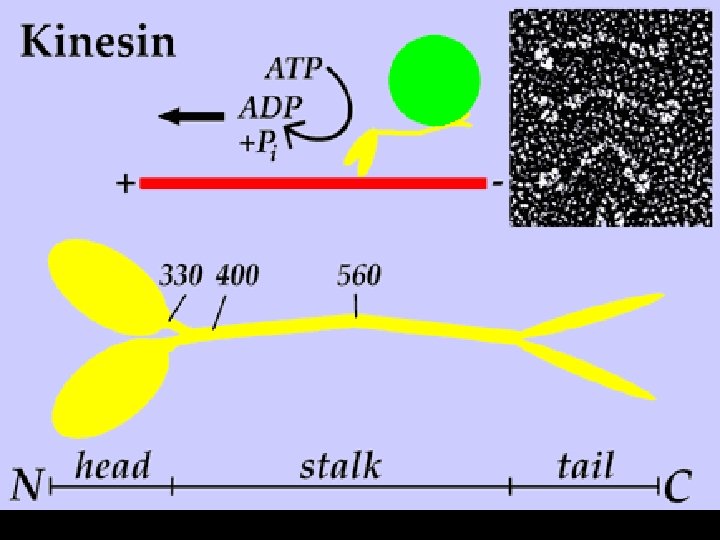
Microtubules • Use specialized motor proteins to move organelles along the microtubule. • Kinesins- move organelles toward the + end (toward cell periphery) • Dyneins- move them toward the - end (toward the center of cell)
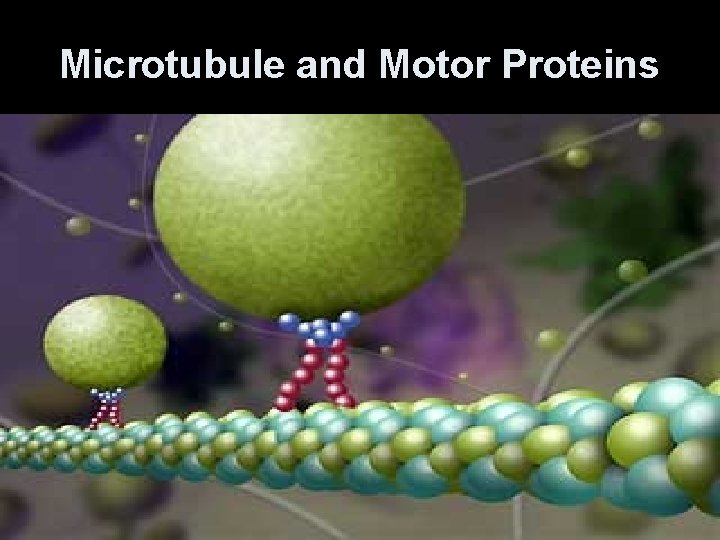
Microtubule and Motor Proteins
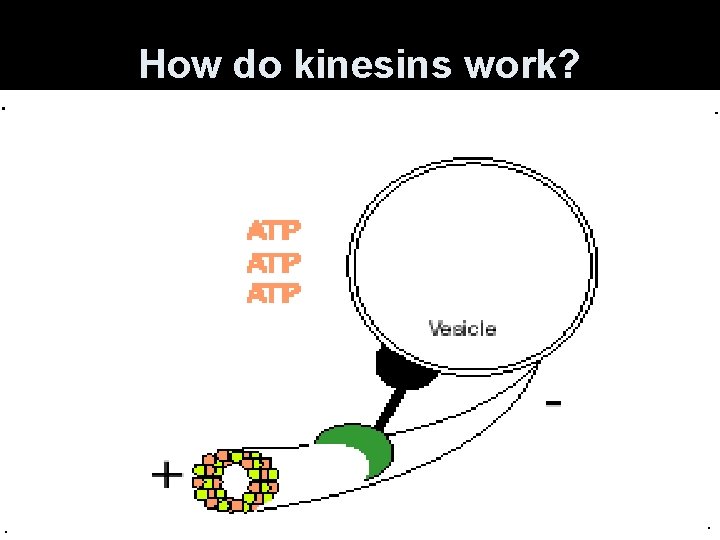
How do kinesins work?
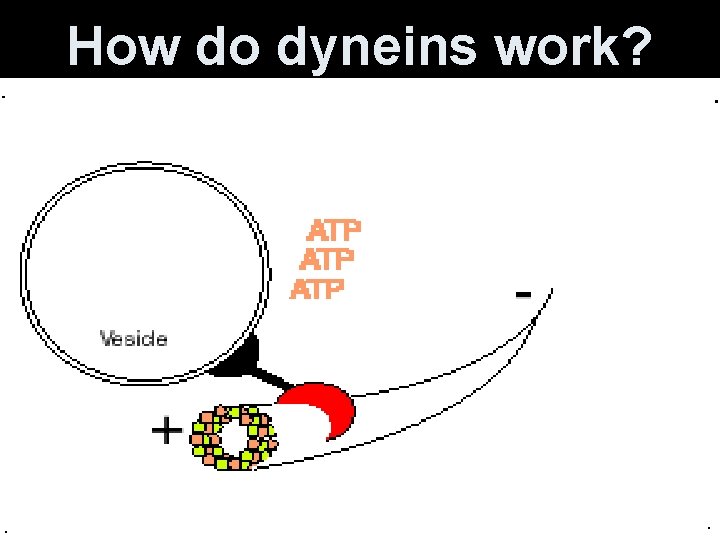
How do dyneins work?

Could a simple defect in a kinesin affect a whole organism?

Wild-type Drosophila larva

Wild-type (Normal) Drosophila Movement

Mutant Kinesins in Drosophila (khc 6 mutant)

Intermediate Filaments • Most durable protein filament- tough fibrous filaments of overlapping tetramers of protein (rope-like). • Between actin and microtubules in size. Stable • Ex. of Fibers- vimentin and keratin
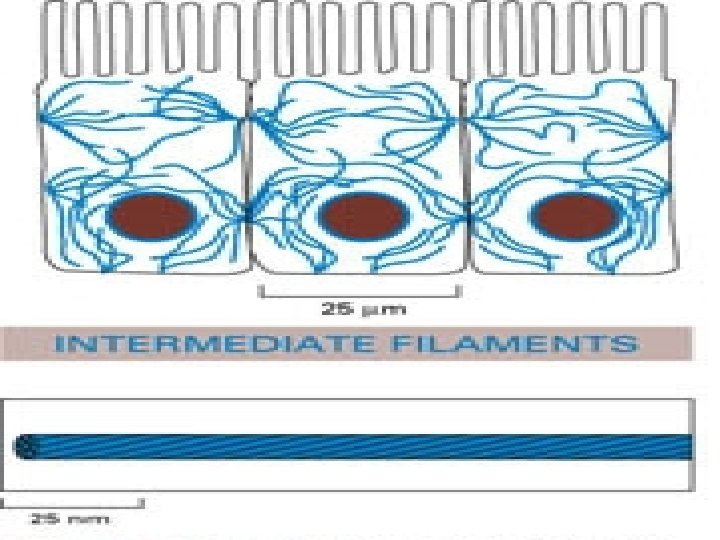

EM of Intermediate Filaments

Intermediate Filaments • Anchored to proteins embedded into plasma membrane. • Provide mechanical support to cell.
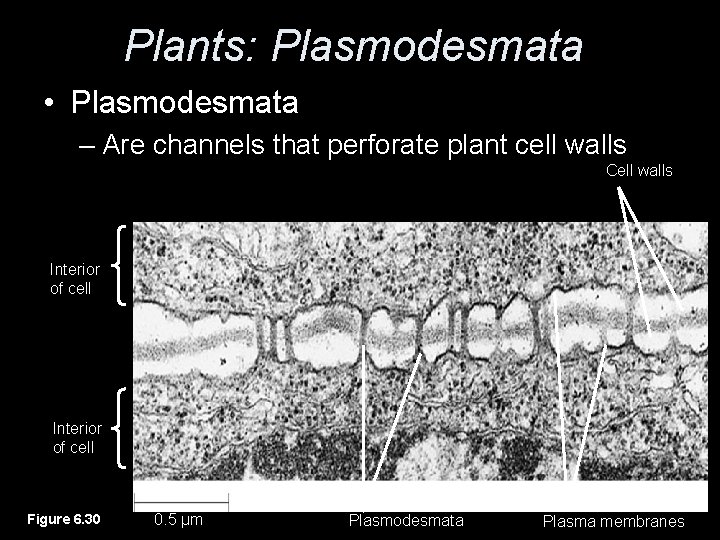
Plants: Plasmodesmata • Plasmodesmata – Are channels that perforate plant cell walls Cell walls Interior of cell Figure 6. 30 0. 5 µm Plasmodesmata Plasma membranes

The Extracellular Matrix (ECM) of Animal Cells • Animal cells – Lack cell walls – Are covered by an elaborate matrix, the ECM
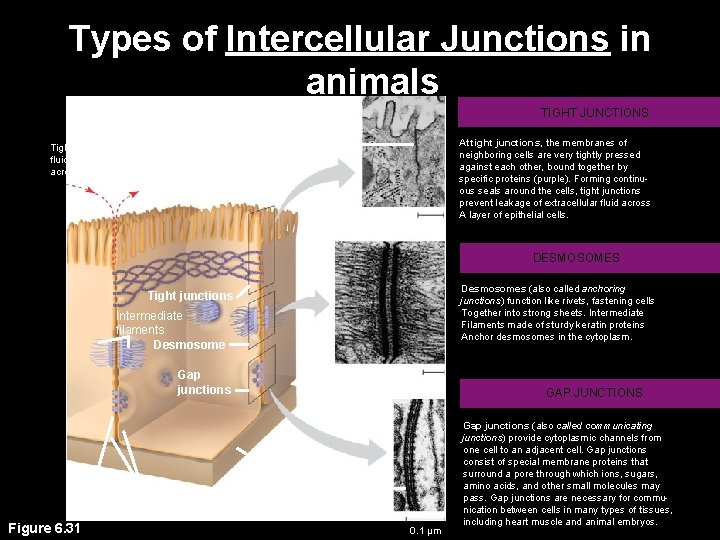
Types of Intercellular Junctions in animals TIGHT JUNCTIONS At tight junctions, the membranes of neighboring cells are very tightly pressed against each other, bound together by specific proteins (purple). Forming continuous seals around the cells, tight junctions prevent leakage of extracellular fluid across A layer of epithelial cells. Tight junctions prevent fluid from moving across a layer of cells 0. 5 µm DESMOSOMES Desmosomes (also called anchoring junctions) function like rivets, fastening cells Together into strong sheets. Intermediate Filaments made of sturdy keratin proteins Anchor desmosomes in the cytoplasm. Tight junctions Intermediate filaments Desmosome Gap junctions Space between cells Figure 6. 31 1 µm Extracellular matrix Plasma membranes of adjacent cells Gap junction 0. 1 µm GAP JUNCTIONS Gap junctions (also called communicating junctions) provide cytoplasmic channels from one cell to an adjacent cell. Gap junctions consist of special membrane proteins that surround a pore through which ions, sugars, amino acids, and other small molecules may pass. Gap junctions are necessary for communication between cells in many types of tissues, including heart muscle and animal embryos.
Cilia
- Slides: 94