CELL CYCLE AND DOGMA BIOLOGY Cell Cycle The

CELL CYCLE AND DOGMA BIOLOGY
Cell Cycle • The dividing and non-dividing stages in the life of a cell. • Phases: 1. Interphase: Interphase growth and DNA replication 2. Prophase 3. Metaphase Mitotic division 4. Anaphase 5. Telophase

Eukaryotic Cell Cycle 1. Cell growth 2. Chromosome replication 3. Cell division (2 types) • Mitosis • Meiosis
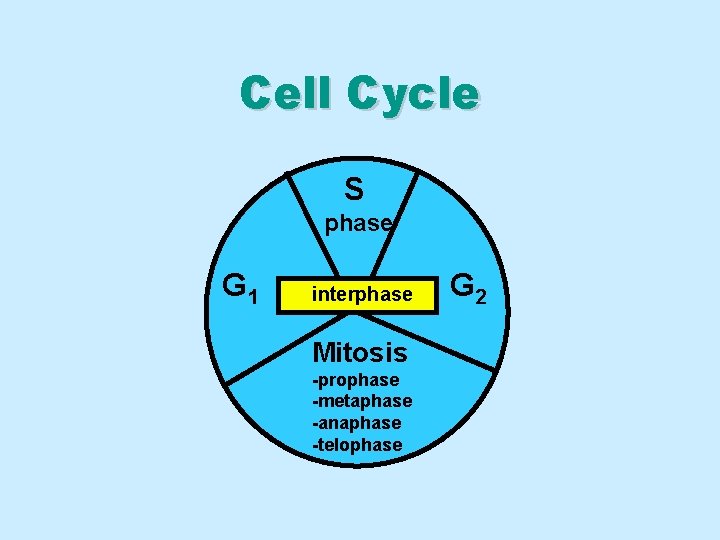
Cell Cycle S phase G 1 interphase Mitosis -prophase -metaphase -anaphase -telophase G 2

Cell Cycle Phases • Interphase: cell growth & DNA replication • Mitosis : nuclear & cell Division

Interphase • Comprises about 90% of the cell cycle • Cellular growth: growth a. protein synthesis b. metabolic activities c. DNA synthesis • Made up of three phases: phases 1. G 1 phase 2. S phase 3. G 2 phase

Interphase 1. G 1 (gap) phase: a. protein synthesis and metabolic activities. b. most cell are arrested in this phase 2. S phase: DNA replication takes place. 3. G 2 (gap) phase: Cellular growth and preparation for M phase
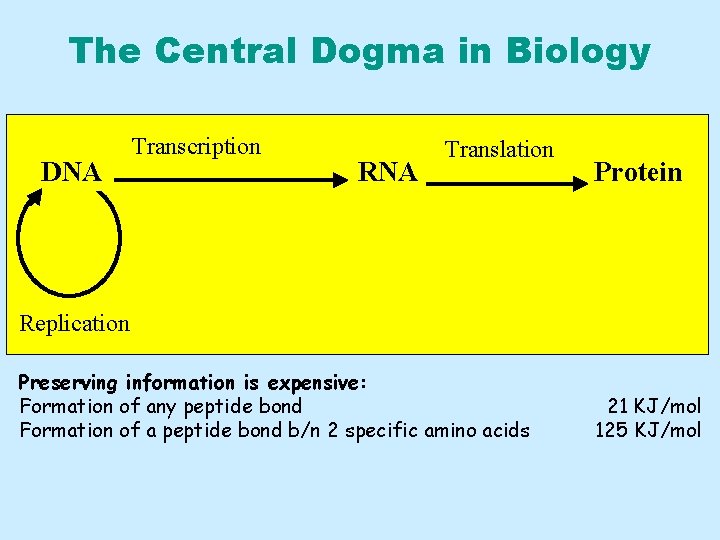
The Central Dogma in Biology DNA Transcription RNA Translation Protein Replication Preserving information is expensive: Formation of any peptide bond Formation of a peptide bond b/n 2 specific amino acids 21 KJ/mol 125 KJ/mol
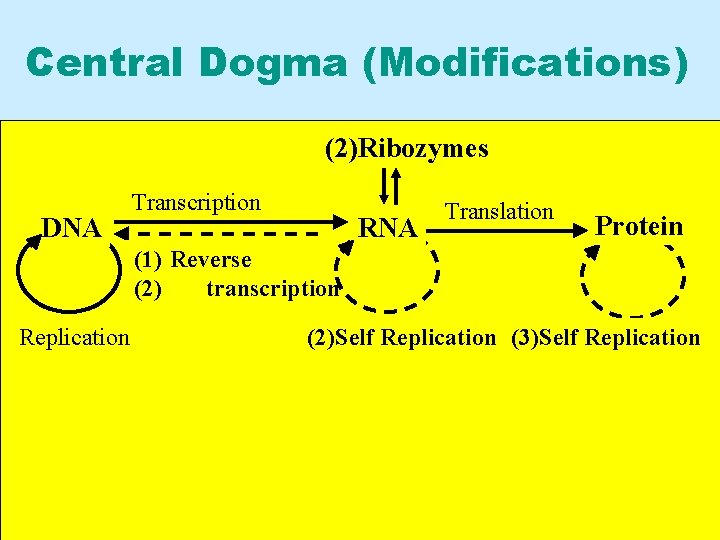
Central Dogma (Modifications) (2)Ribozymes DNA Transcription RNA Translation Protein (1) Reverse (2) transcription Replication (2)Self Replication (3)Self Replication

DNA Replication
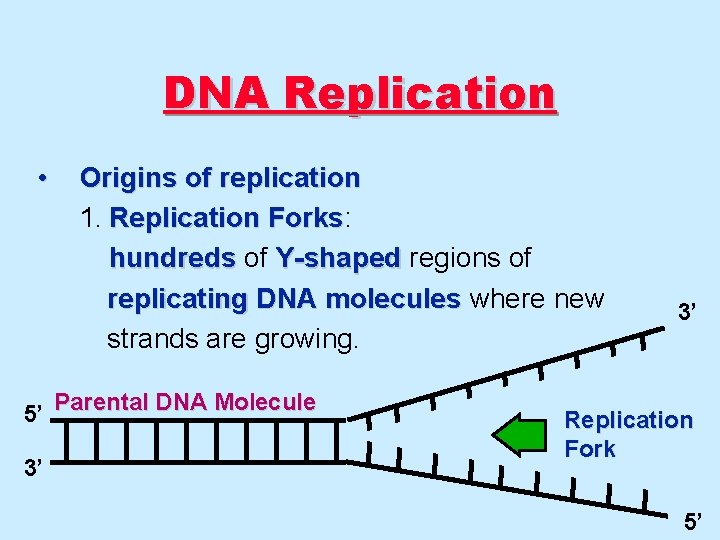
DNA Replication • Origins of replication 1. Replication Forks: Forks hundreds of Y-shaped regions of replicating DNA molecules where new strands are growing. 5’ Parental DNA Molecule 3’ 3’ Replication Fork 5’
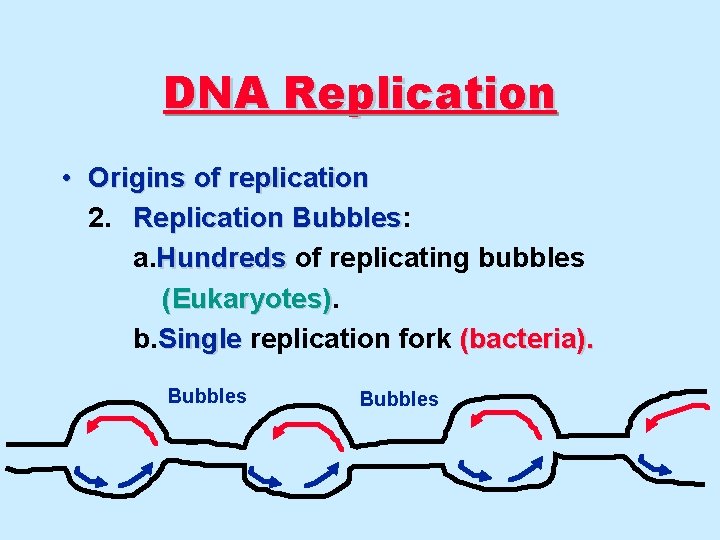
DNA Replication • Origins of replication 2. Replication Bubbles: Bubbles a. Hundreds of replicating bubbles (Eukaryotes) b. Single replication fork (bacteria). Bubbles

DNA Replication • Strand Separation: Separation 1. Helicase: Helicase enzyme which catalyze the unwinding and separation (breaking H-Bonds) of the parental double helix. 2. Single-Strand Binding Proteins: Proteins proteins which attach and help keep the separated strands apart.
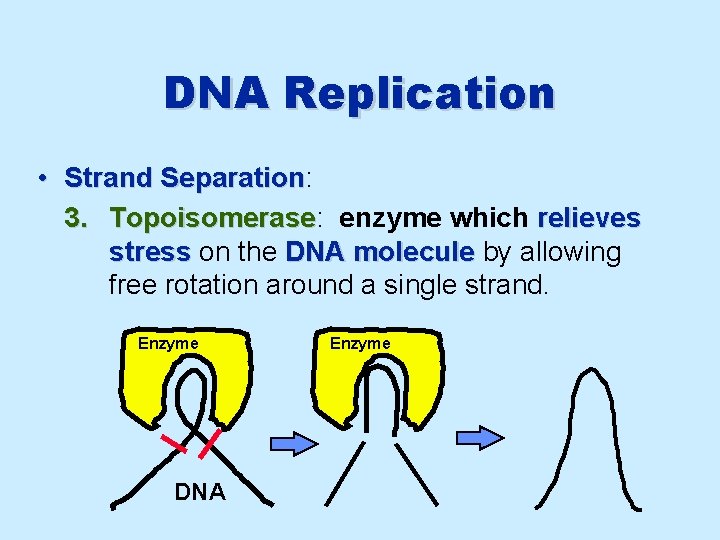
DNA Replication • Strand Separation: Separation 3. Topoisomerase: Topoisomerase enzyme which relieves stress on the DNA molecule by allowing free rotation around a single strand. Enzyme DNA Enzyme

DNA Replication • Priming: 1. RNA primers: primers before new DNA strands can form, there must be small pre-existing primers (RNA) present to start the addition of new nucleotides (DNA Polymerase) 2. Primase: Primase enzyme that polymerizes (synthesizes) the RNA Primer.
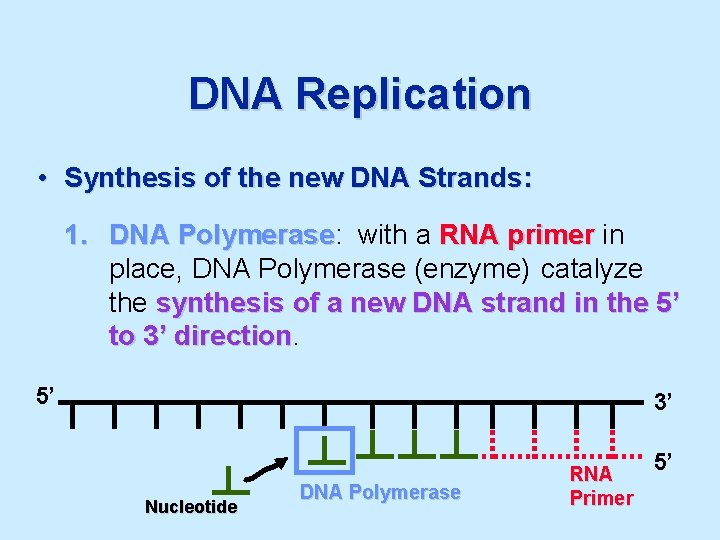
DNA Replication • Synthesis of the new DNA Strands: 1. DNA Polymerase: Polymerase with a RNA primer in place, DNA Polymerase (enzyme) catalyze the synthesis of a new DNA strand in the 5’ to 3’ direction 5’ 3’ Nucleotide DNA Polymerase RNA Primer 5’
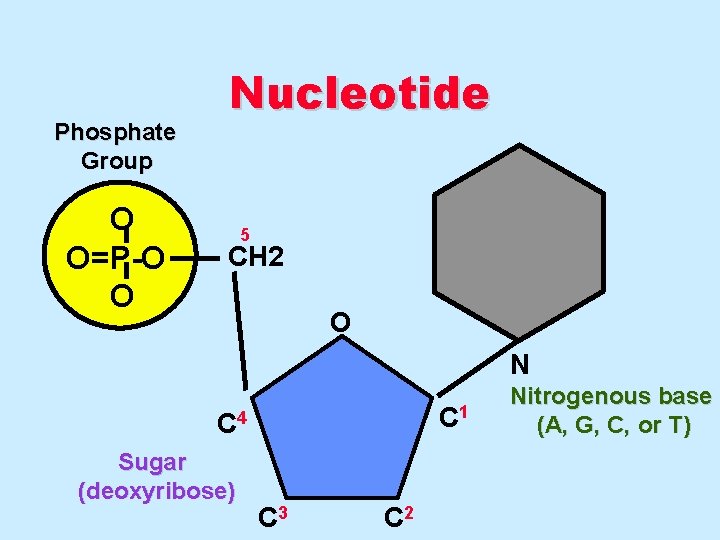
Phosphate Group O O=P-O O Nucleotide 5 CH 2 O N C 1 C 4 Sugar (deoxyribose) C 3 C 2 Nitrogenous base (A, G, C, or T)
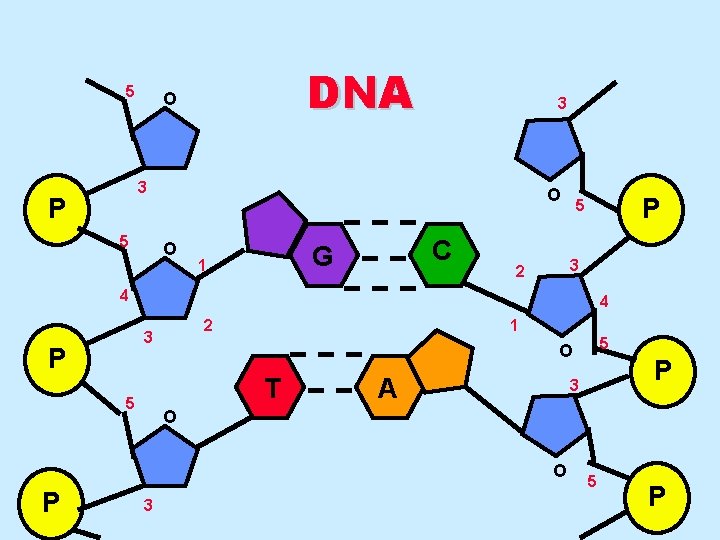
5 DNA O 3 3 P 5 O O C G 1 P 5 3 2 4 4 2 3 P 1 T 5 A P 3 O O P 5 O 3 5 P
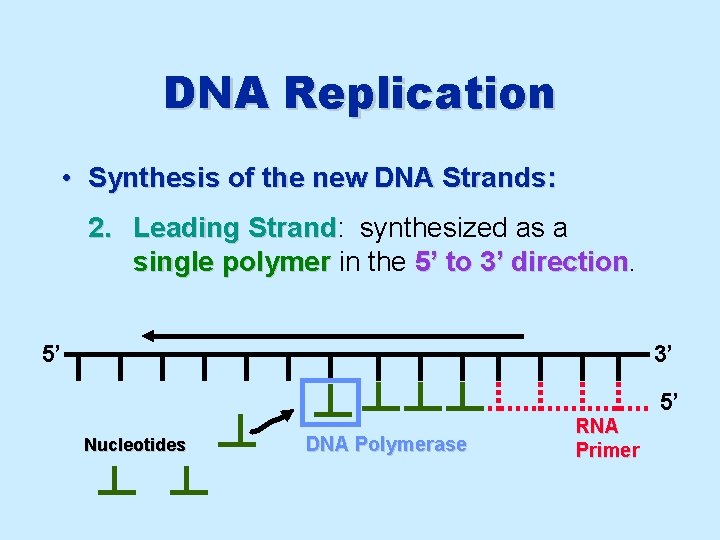
DNA Replication • Synthesis of the new DNA Strands: 2. Leading Strand: Strand synthesized as a single polymer in the 5’ to 3’ direction 5’ 3’ 5’ Nucleotides DNA Polymerase RNA Primer
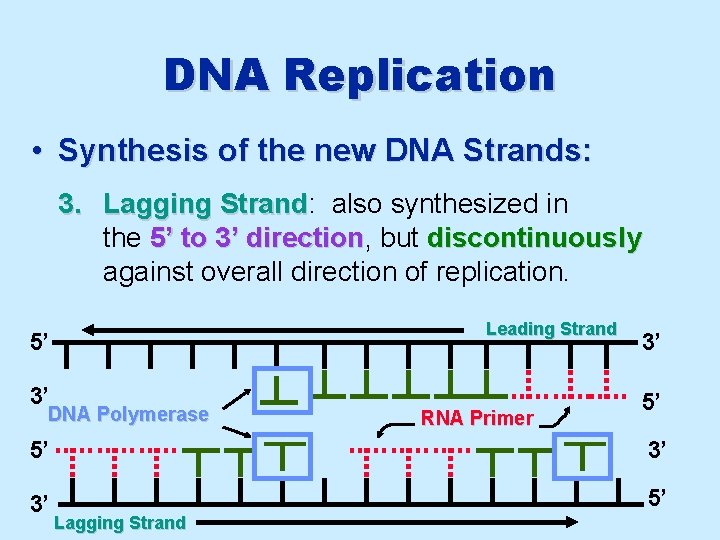
DNA Replication • Synthesis of the new DNA Strands: 3. Lagging Strand: Strand also synthesized in the 5’ to 3’ direction, direction but discontinuously against overall direction of replication. Leading Strand 5’ 3’ DNA Polymerase RNA Primer 3’ 5’ 5’ 3’ 3’ 5’ Lagging Strand
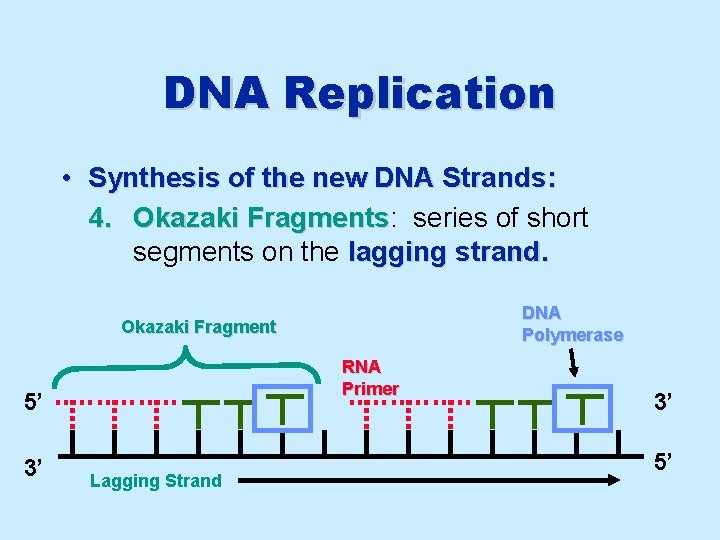
DNA Replication • Synthesis of the new DNA Strands: 4. Okazaki Fragments: Fragments series of short segments on the lagging strand. DNA Polymerase Okazaki Fragment RNA Primer 5’ 3’ Lagging Strand 3’ 5’
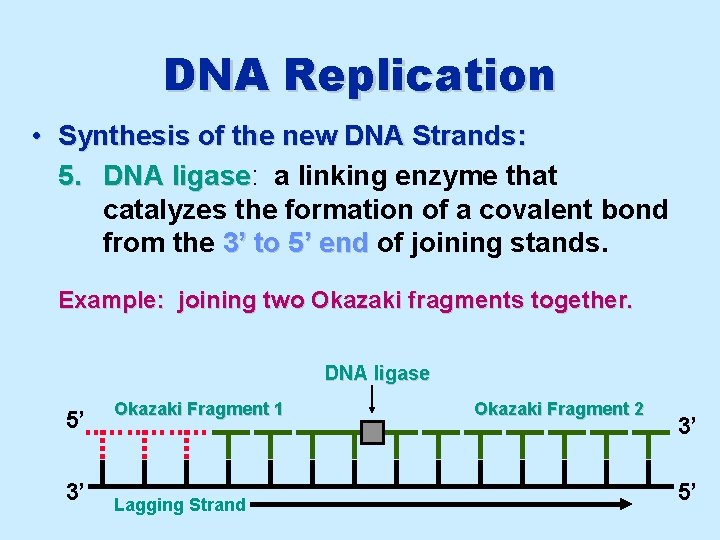
DNA Replication • Synthesis of the new DNA Strands: 5. DNA ligase: ligase a linking enzyme that catalyzes the formation of a covalent bond from the 3’ to 5’ end of joining stands. Example: joining two Okazaki fragments together. DNA ligase 5’ 3’ Okazaki Fragment 1 Lagging Strand Okazaki Fragment 2 3’ 5’

DNA Replication • Synthesis of the new DNA Strands: 6. Proofreading: Proofreading initial base-pairing errors are usually corrected by DNA polymerase
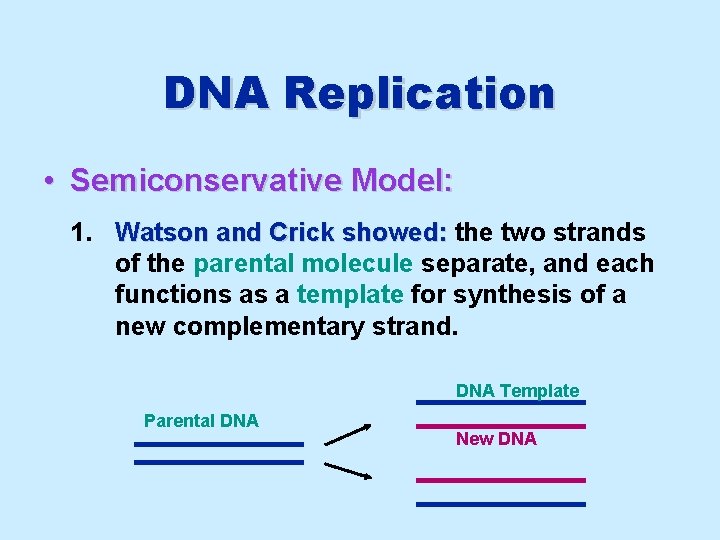
DNA Replication • Semiconservative Model: 1. Watson and Crick showed: the two strands of the parental molecule separate, and each functions as a template for synthesis of a new complementary strand. DNA Template Parental DNA New DNA

DNA Repair • Excision repair: 1. Damaged segment is excised by a repair enzyme (there are over 50 repair enzymes). 2. DNA polymerase and DNA ligase replace and bond the new nucleotides together.

Gene Expression üTranscription üTranslation

What is gene expression? • Biological processes, such as transcription, and in case of proteins, also translation, that yield a gene product. • A gene is expressed when its biological product is present and active. • Gene expression is regulated at multiple levels.

Expression of Genetic Information Beadle and Tatum (1941) showed in the fungus Neurospora crassa that there is a relationship between a gene and each enzyme needed in a biochemical pathway, resulting in the one geneone enzyme hypothesis (now modified to one gene-one polypeptide, since not all proteins are enzymes and some require more than one polypeptide).

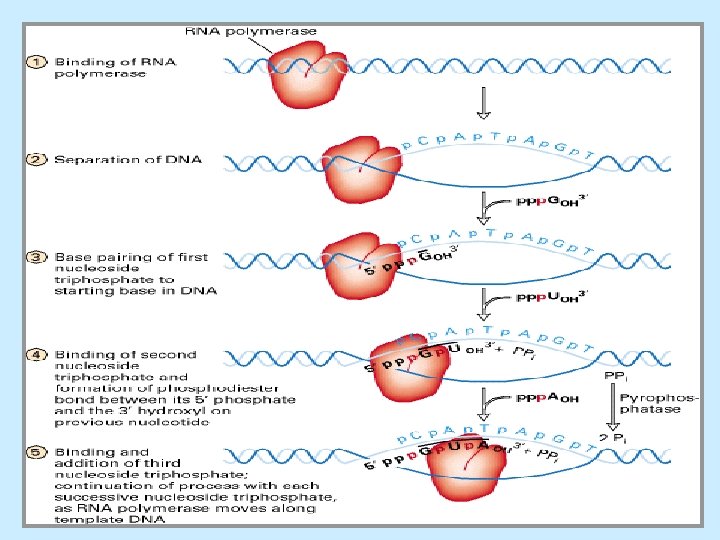
Expression of Genetic Information Production of proteins requires two steps: – Transcription involves an enzyme (RNA polymerase) making an RNA copy of part of one DNA strand. There are four main classes of RNA: i. Messenger RNAs (m. RNA), which specify the amino acid sequence of a protein by using codons of the genetic code. ii. Transfer RNAs (t. RNA). iii. Ribosomal RNAs (r. RNA). iv. Small nuclear RNAs (sn. RNA), found only in eukaryotes. – Translation converts the information in m. RNA into the amino acid sequence of a protein using ribosomes, large complexes of r. RNAs and proteins.
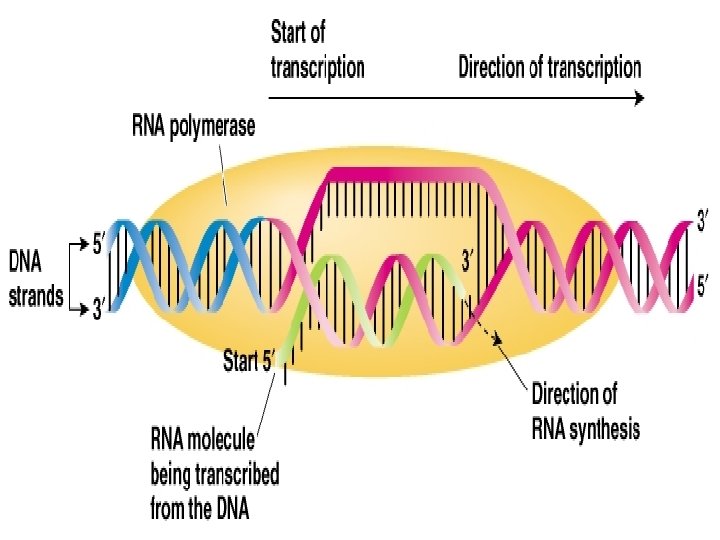
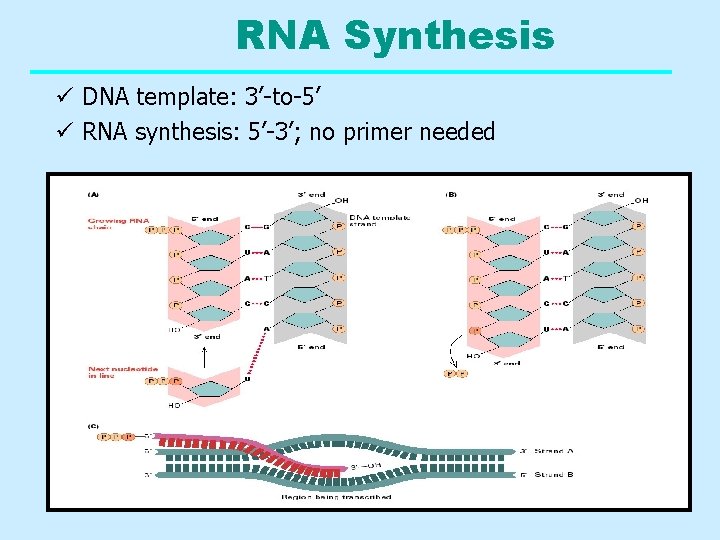
RNA Synthesis ü DNA template: 3’-to-5’ ü RNA synthesis: 5’-3’; no primer needed

Expression of Genetic Information ü Only some of the genes in a cell are active at any given time, and activity also varies by tissue type and developmental stage. ü Regulation of gene expression is not completely understood, but it has been shown to involve an array of controlling signals. a. Jacob and Monod (1961) proposed the operon model to explain prokaryotic gene regulation, showing that a genetic switch is used to control production of the enzymes needed to metabolize lactose. Similar systems control many genes in bacteria and their viruses. b. Genetic switches used in eukaryotes are different and more complex, with much remaining to be learned about their function.

Protein Coding Genes ü ORF – long (usually >100 aa) – “known” proteins likely ü Basal signals – Transcription, translation ü Regulatory signals – Depend on organism • Prokaryotes vs Eukaryotes
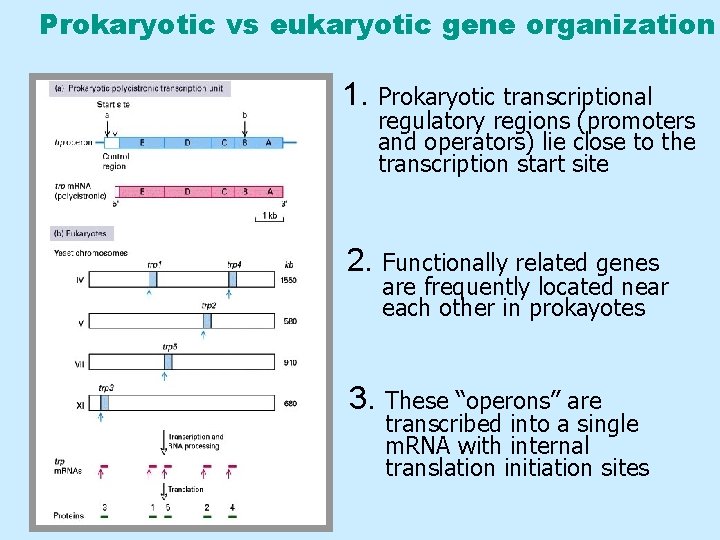
Prokaryotic vs eukaryotic gene organization 1. Prokaryotic transcriptional regulatory regions (promoters and operators) lie close to the transcription start site 2. Functionally related genes are frequently located near each other in prokayotes 3. These “operons” are transcribed into a single m. RNA with internal translation initiation sites
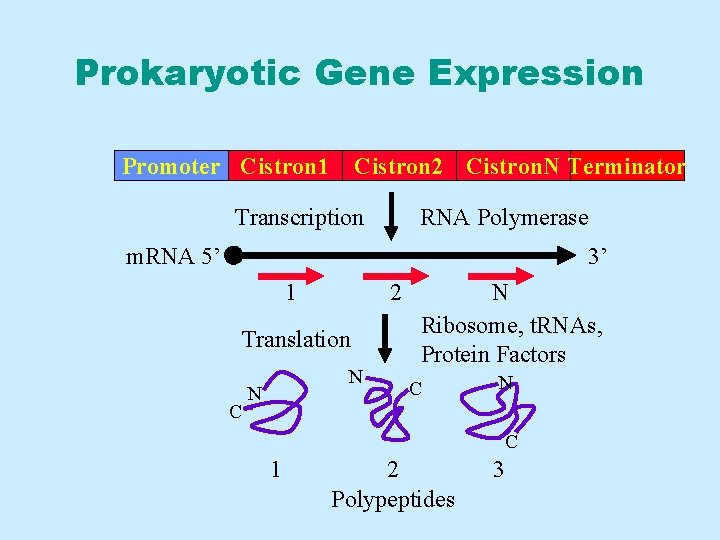
Prokaryotic Gene Expression Promoter Cistron 1 Cistron 2 Cistron. N Terminator Transcription RNA Polymerase m. RNA 5’ 3’ 1 2 Translation C N N N Ribosome, t. RNAs, Protein Factors C N C 1 2 Polypeptides 3
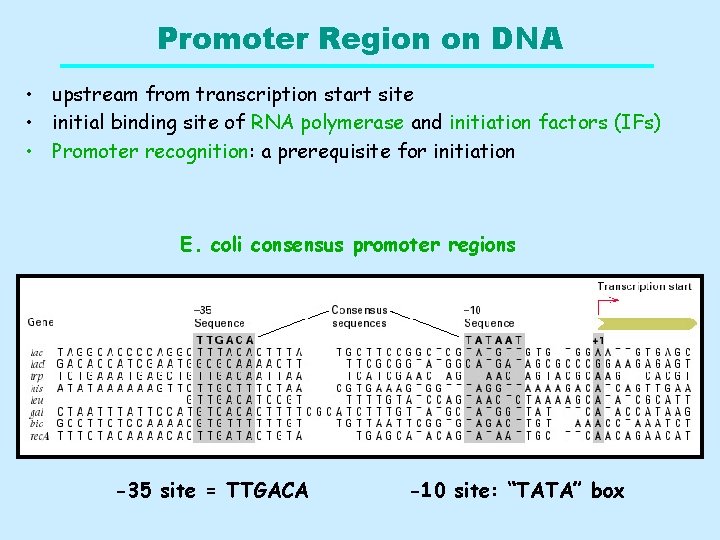
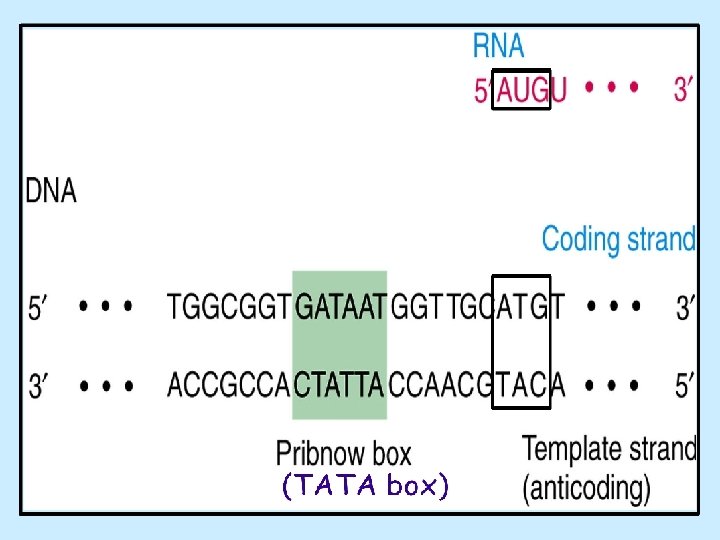
Promoter Region on DNA • upstream from transcription start site • initial binding site of RNA polymerase and initiation factors (IFs) • Promoter recognition: a prerequisite for initiation E. coli consensus promoter regions -35 site = TTGACA -10 site: “TATA” box
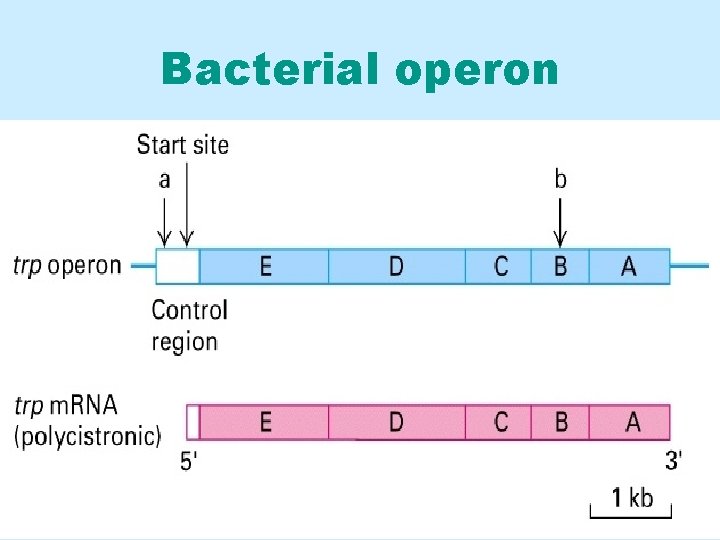
Bacterial operon
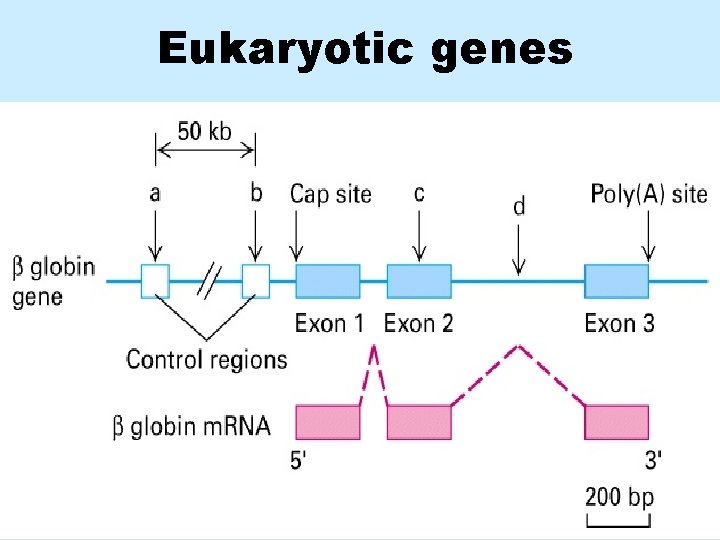
Eukaryotic genes
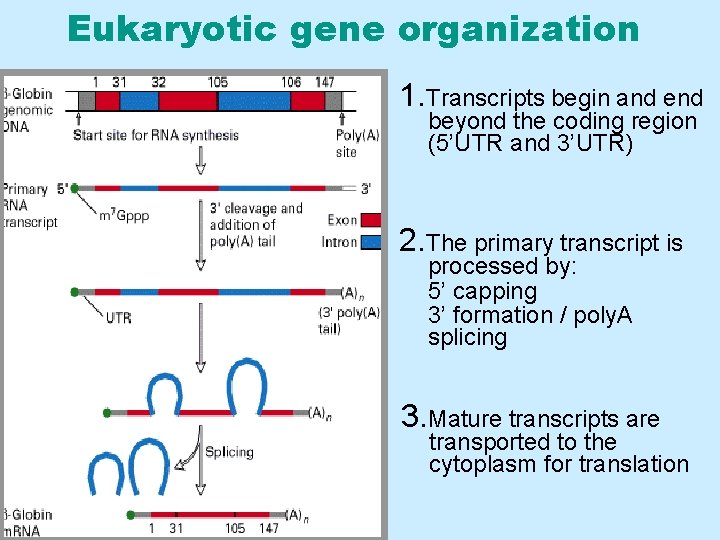
Eukaryotic gene organization 1. Transcripts begin and end beyond the coding region (5’UTR and 3’UTR) 2. The primary transcript is processed by: 5’ capping 3’ formation / poly. A splicing 3. Mature transcripts are transported to the cytoplasm for translation
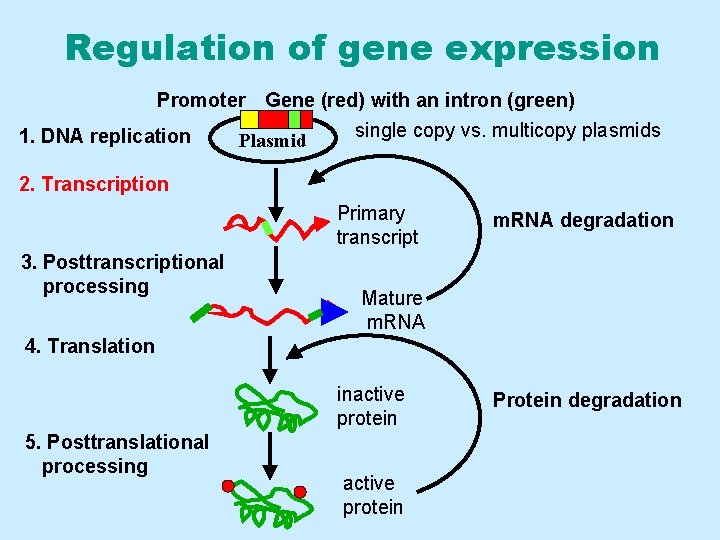
Regulation of gene expression Promoter Gene (red) with an intron (green) single copy vs. multicopy plasmids 1. DNA replication Plasmid 2. Transcription Primary transcript 3. Posttranscriptional processing m. RNA degradation Mature m. RNA 4. Translation inactive protein 5. Posttranslational processing active protein Protein degradation
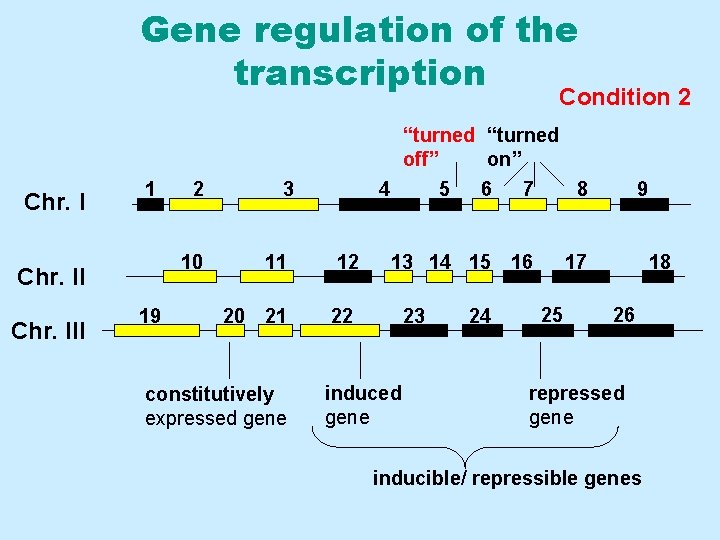
Gene regulation of the transcription Condition 2 1 “turned off” on” Chr. I 1 10 Chr. III 2 19 3 4 11 12 20 21 22 constitutively expressed gene 5 7 8 13 14 15 16 17 23 induced gene 6 24 25 9 18 26 repressed gene inducible/ repressible genes
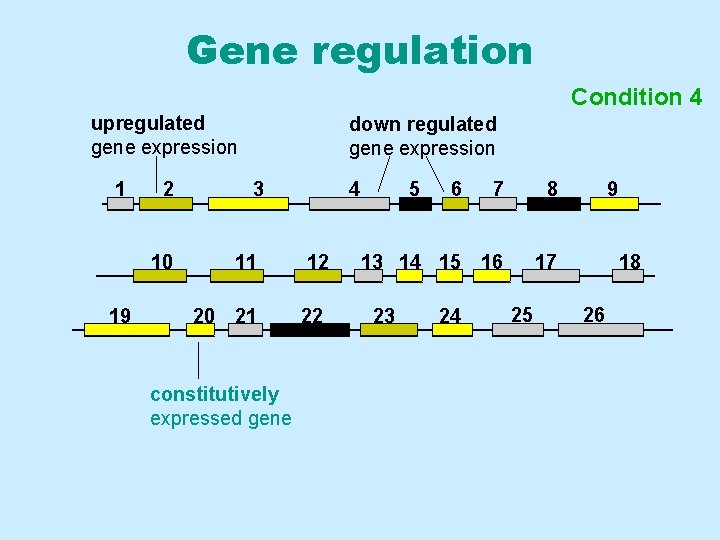
Gene regulation Condition 43 upregulated gene expression 1 2 10 19 down regulated gene expression 3 4 11 12 20 21 22 constitutively expressed gene 5 7 8 13 14 15 16 17 23 6 24 25 9 18 26

Definitions • Constitutively expressed genes Genes that are actively transcribed (and translated) under all experimental conditions, at essentially all developmental stages, or in virtually all cells. • Inducible genes Genes that are transcribed and translated at higher levels in response to an inducing factor • Repressible genes Genes whose transcription and translation decreases in response to a repressing signal

Definitions • Housekeeping genes – genes for enzymes of central metabolic pathways (e. g. TCA cycle) – these genes are constitutively expressed – the level of gene expression may vary
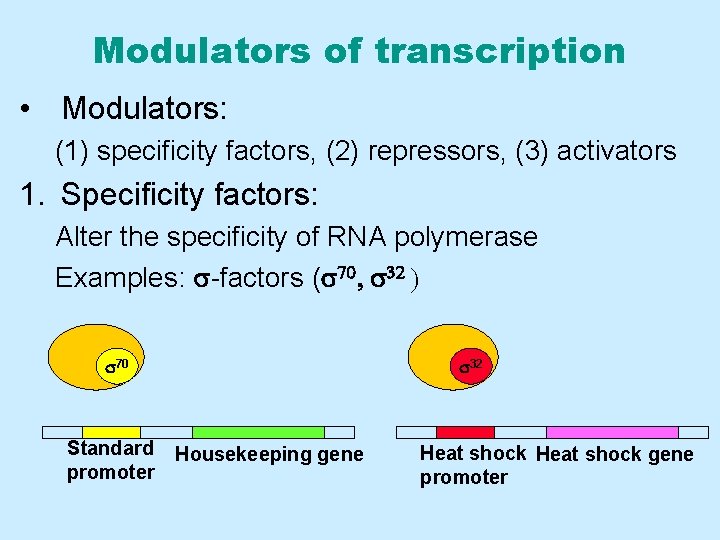
Modulators of transcription • Modulators: (1) specificity factors, (2) repressors, (3) activators 1. Specificity factors: Alter the specificity of RNA polymerase Examples: s-factors (s 70, s 32 ) s 70 Standard promoter s 32 Housekeeping gene Heat shock gene promoter
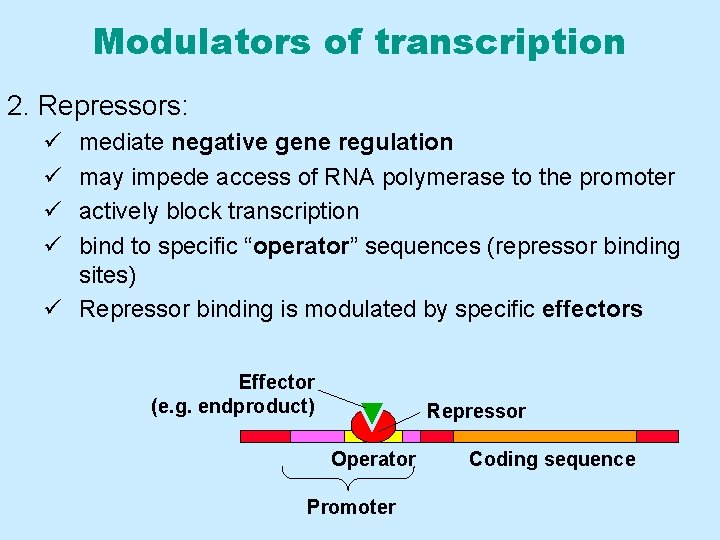
Modulators of transcription 2. Repressors: ü ü mediate negative gene regulation may impede access of RNA polymerase to the promoter actively block transcription bind to specific “operator” sequences (repressor binding sites) ü Repressor binding is modulated by specific effectors Effector (e. g. endproduct) Repressor Operator Promoter Coding sequence
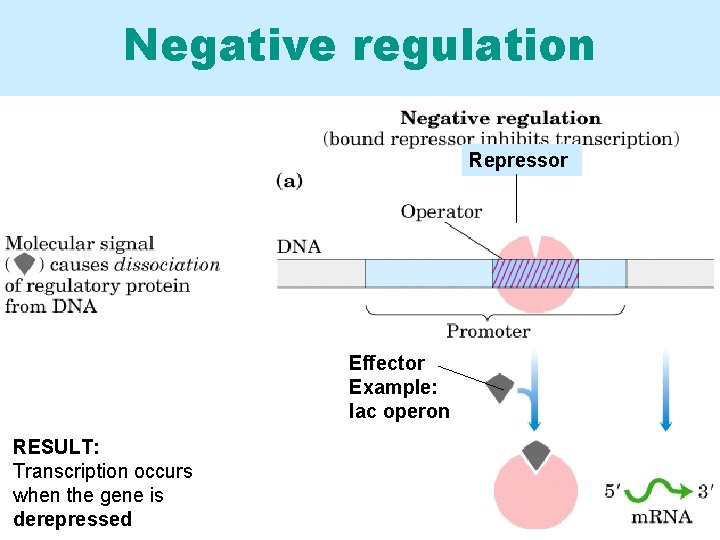
Negative regulation Repressor Effector Example: lac operon RESULT: Transcription occurs when the gene is derepressed
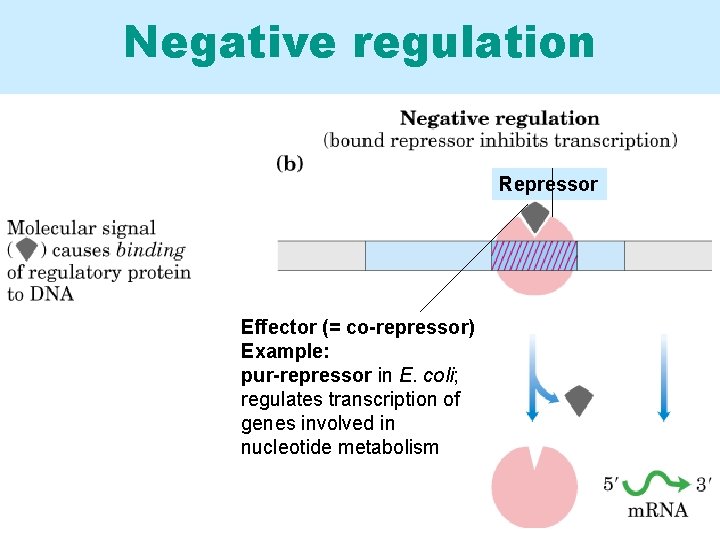
Negative regulation Repressor Effector (= co-repressor) Example: pur-repressor in E. coli; regulates transcription of genes involved in nucleotide metabolism
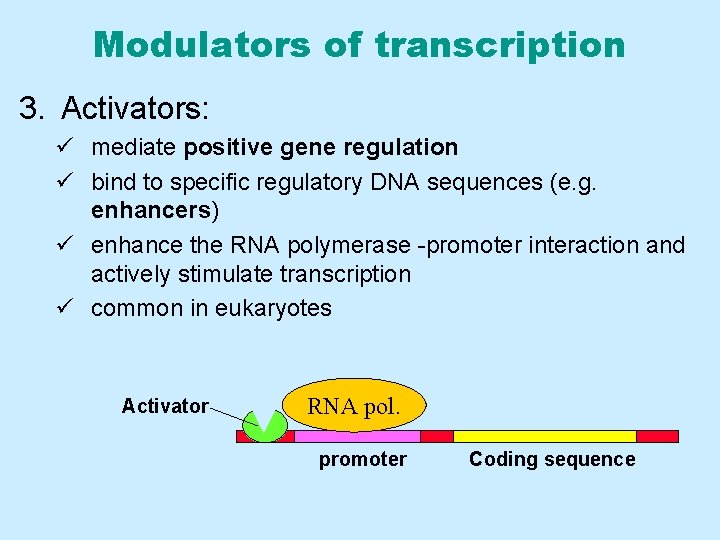
Modulators of transcription 3. Activators: ü mediate positive gene regulation ü bind to specific regulatory DNA sequences (e. g. enhancers) ü enhance the RNA polymerase -promoter interaction and actively stimulate transcription ü common in eukaryotes Activator RNA pol. promoter Coding sequence
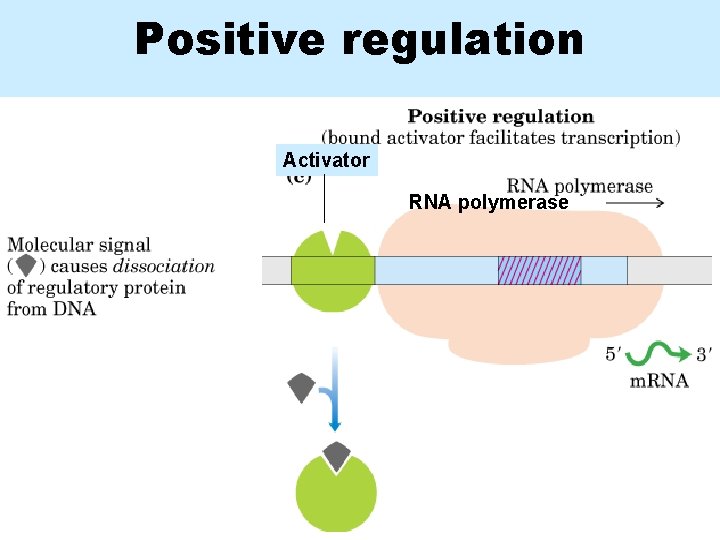
Positive regulation Activator RNA polymerase
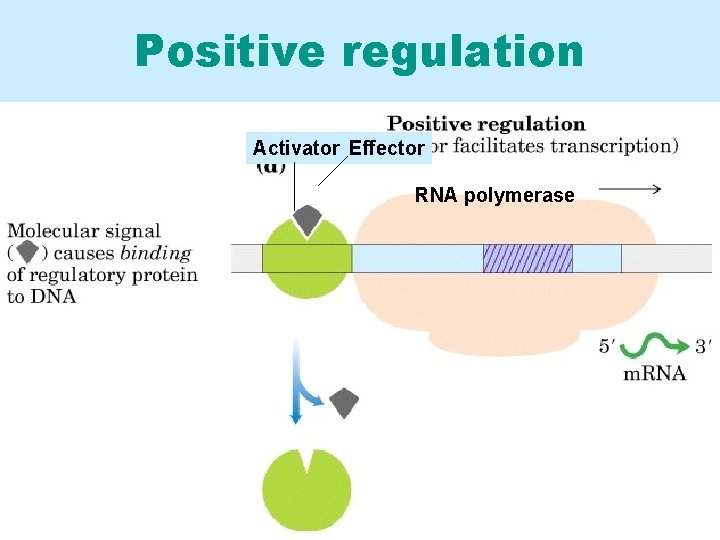
Positive regulation Activator Effector RNA polymerase

Operons üa promoter plus a set of adjacent genes whose gene products function together. üusually contain 2 – 6 genes, (up to 20 genes) üthese genes are transcribed as a polycistronic transcript. ü relatively common in prokaryotes ü rare in eukaryotes

The lactose (lac) operon Pi I Q 3 P Q 1 Z Q 2 Y A • Contains several elements – – lac. Z gene = b-galactosidase lac. Y gene = galactosidase permease lac. A gene = thiogalactoside transacetylase lac. I gene = lac repressor – – Pi = promoter for the lac. I gene P = promoter for lac-operon Q 1 = main operator Q 2 and Q 3 = secondary operator sites (pseudo-operators)
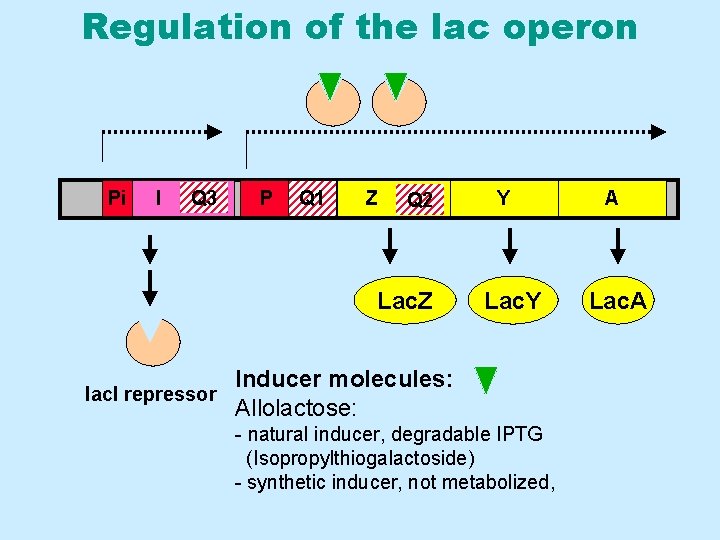
Regulation of the lac operon Pi I Q 3 P Q 1 Z Q 2 Lac. Z Y Lac. Y Inducer molecules: lac. I repressor Allolactose: - natural inducer, degradable IPTG (Isopropylthiogalactoside) - synthetic inducer, not metabolized, A Lac. A
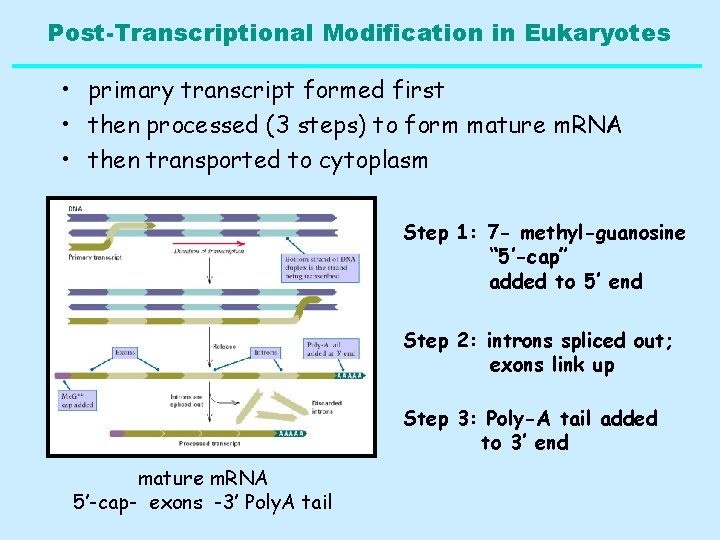
Post-Transcriptional Modification in Eukaryotes • primary transcript formed first • then processed (3 steps) to form mature m. RNA • then transported to cytoplasm Step 1: 7 - methyl-guanosine “ 5’-cap” added to 5’ end Step 2: introns spliced out; exons link up Step 3: Poly-A tail added to 3’ end mature m. RNA 5’-cap- exons -3’ Poly. A tail
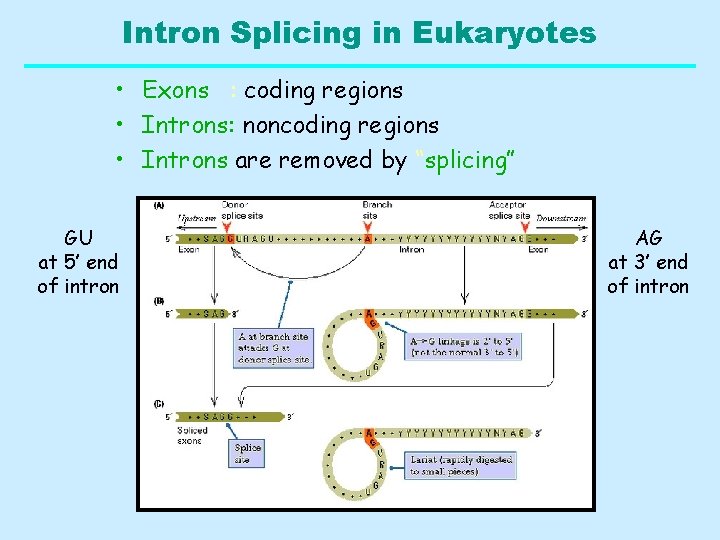
Intron Splicing in Eukaryotes • Exons : coding regions • Introns: noncoding regions • Introns are removed by “splicing” GU at 5’ end of intron AG at 3’ end of intron
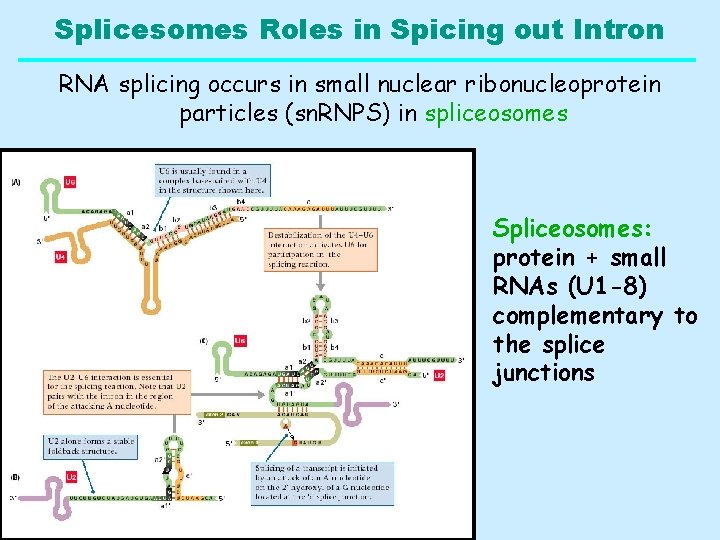
Splicesomes Roles in Spicing out Intron RNA splicing occurs in small nuclear ribonucleoprotein particles (sn. RNPS) in spliceosomes Spliceosomes: protein + small RNAs (U 1 -8) complementary to the splice junctions
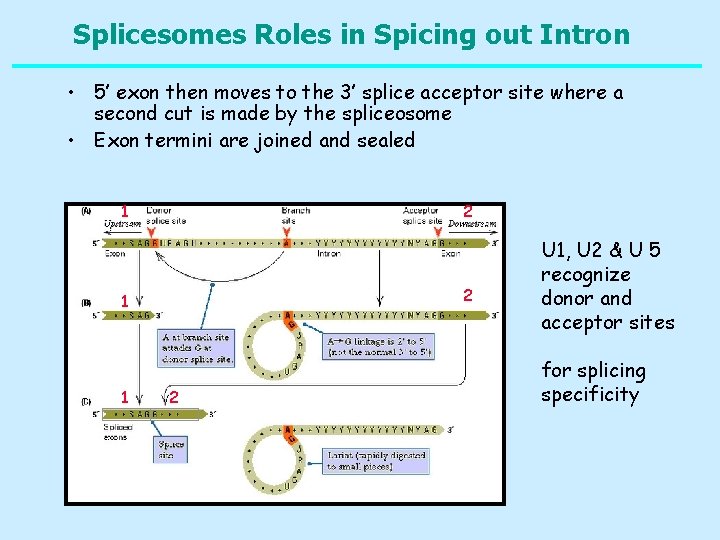
Splicesomes Roles in Spicing out Intron • 5’ exon then moves to the 3’ splice acceptor site where a second cut is made by the spliceosome • Exon termini are joined and sealed 1 2 2 1 1 2 U 1, U 2 & U 5 recognize donor and acceptor sites for splicing specificity
- Slides: 60