Cell biology of carbohydrate metabolism Glycogen Galactose Glycogen

Cell biology of carbohydrate metabolism
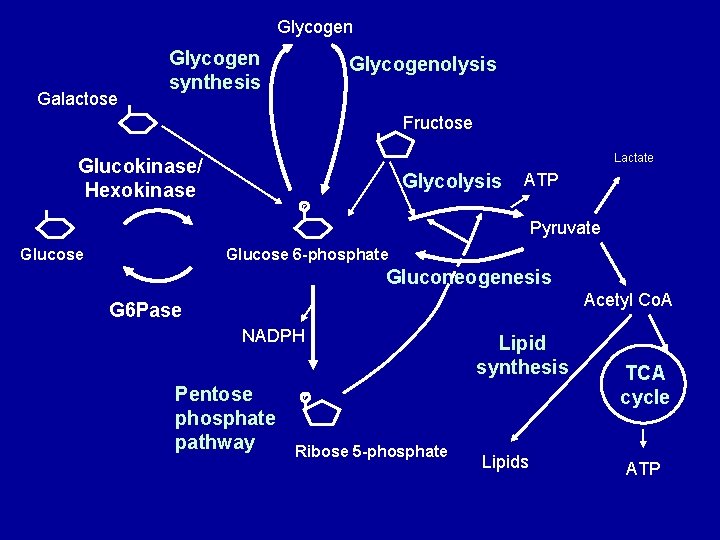
Glycogen Galactose Glycogen synthesis Glycogenolysis Fructose Lactate Glucokinase/ Hexokinase Glycolysis ATP P Pyruvate Glucose 6 -phosphate Gluconeogenesis Acetyl Co. A G 6 Pase NADPH Pentose phosphate pathway Lipid synthesis P Ribose 5 -phosphate Lipids TCA cycle ATP

Regulation of metabolic pathways The flow of intermediates through metabolic pathways is controlled by four mechanisms 1. Substrate availability 2. Allosteric activation or inhibition 3. Covalent modification of enzymes 4. Induction/repression of enzyme synthesis + the types of transporters and enzymes that exist in a certain cell type
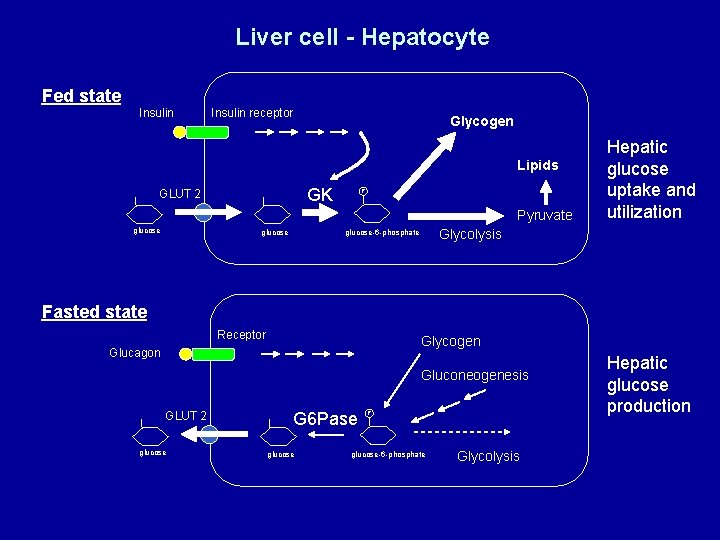
Liver cell - Hepatocyte Fed state Insulin receptor Glycogen Lipids GK GLUT 2 P Pyruvate glucose Glycolysis glucose-6 -phosphate glucose Hepatic glucose uptake and utilization Fasted state Receptor Glycogen Glucagon Gluconeogenesis GLUT 2 glucose G 6 Pase glucose P glucose-6 -phosphate Glycolysis Hepatic glucose production
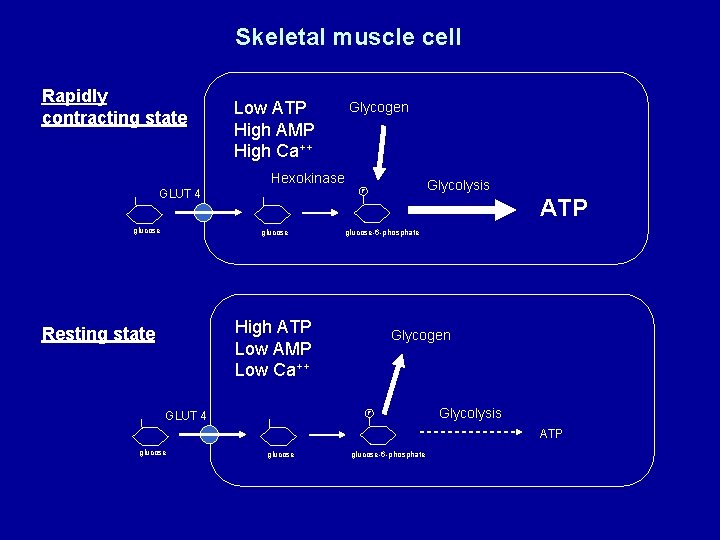
Skeletal muscle cell Rapidly contracting state Low ATP High AMP High Ca++ Glycogen Hexokinase GLUT 4 glucose Glycolysis P glucose ATP glucose-6 -phosphate High ATP Low AMP Low Ca++ Resting state GLUT 4 Glycogen P Glycolysis ATP glucose-6 -phosphate
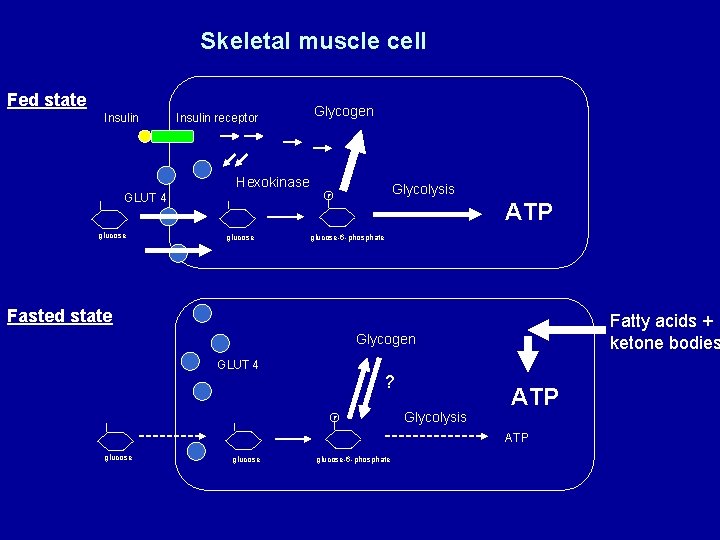
Skeletal muscle cell Fed state Insulin receptor Glycogen Hexokinase GLUT 4 glucose Glycolysis P glucose ATP glucose-6 -phosphate Fasted state Fatty acids + ketone bodies Glycogen GLUT 4 ? P Glycolysis ATP glucose-6 -phosphate
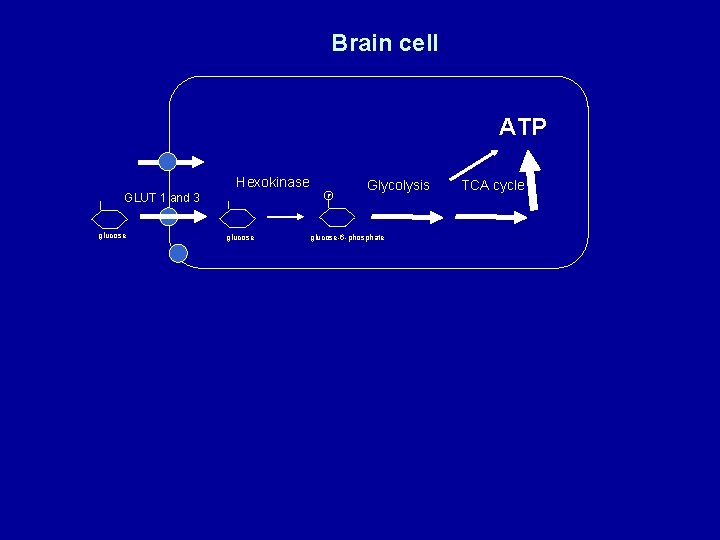
Brain cell ATP Hexokinase GLUT 1 and 3 glucose P glucose Glycolysis glucose-6 -phosphate TCA cycle

Glucose as a regulator of metabolism Concentration effects on enzymatic steps High Km of glucokinase determines rate of glucose catabolism and insulin secretion in pancreatic beta cells High Km of glucokinase allows liver cells to rapidly phosphorylate glucose at high concentrations Allosteric effector Glucose stimulates release of glucokinase from the glucokinase regulatory protein (GKRP) Glucose inhibits liver glycogen phosphorylase Transcriptional effects Glucose stimulates gene transcription through glucose-responsive regions of gene promoters High glucose concentration stimulates the expression of the pyruvate kinase in liver

A carbohydrate response element (Cho. RE) consists of 2 E-box like elements separated by 5 nucleotides CACGTG NNNNN CACGTG E-Box Rat L-PK Rat ACC PI Rat FAS Rat S 14 Mouse S 14 Transcription CACGGGGCACTCCCGTG CATGTGAAAACGTCGTG CATGTGCCACAGGCGTG CACGTGGTGGCCCTGTG CACGCTGGAGTCAGCCC
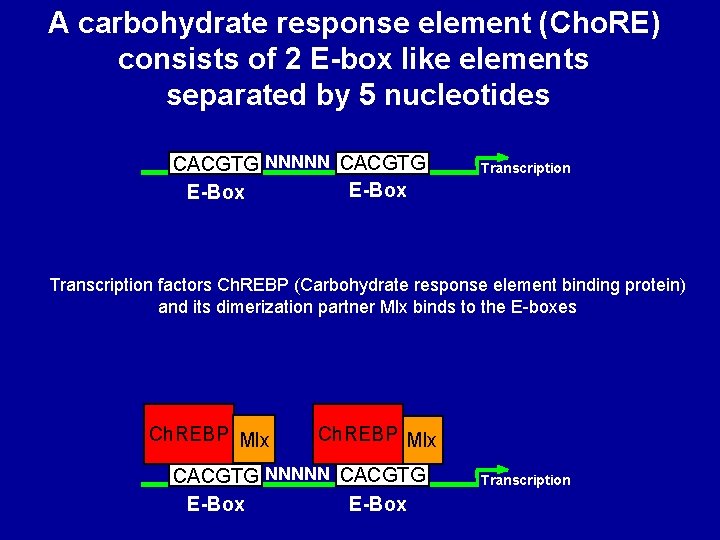
A carbohydrate response element (Cho. RE) consists of 2 E-box like elements separated by 5 nucleotides CACGTG NNNNN CACGTG E-Box Transcription factors Ch. REBP (Carbohydrate response element binding protein) and its dimerization partner Mlx binds to the E-boxes Ch. REBP Mlx CACGTG NNNNN CACGTG E-Box Transcription
![Glucose-dependent binding of Ch. REBP/Mlx to Cho. RE Ch. REBP Mlx Low [glucose] CACGTG Glucose-dependent binding of Ch. REBP/Mlx to Cho. RE Ch. REBP Mlx Low [glucose] CACGTG](http://slidetodoc.com/presentation_image_h2/3e16b75ba890fb61ba518dc351ddfaf2/image-11.jpg)
Glucose-dependent binding of Ch. REBP/Mlx to Cho. RE Ch. REBP Mlx Low [glucose] CACGTG NNNNN CACGTG E-Box High [glucose] Ch. REBP Mlx Transcription Ch. REBP Mlx CACGTG NNNNN CACGTG E-Box Transcription The direct cause of the effect is not glucose itself, but rather a compound generated in glucose metabolism, possibly xylulose 5 -phosphate

Enzymes that were previously found to be induced at the transcriptional level by glucose: Liver-type pyruvate kinase - Regulatory enzym in glycolysis Acetyl Co. A carboxylase Fatty acid synthase Regulatory enzymes in fatty acid synthesis Hypothesis: Ch. REBP mediates glucose-induced glycolysis and lipogenesis in the liver
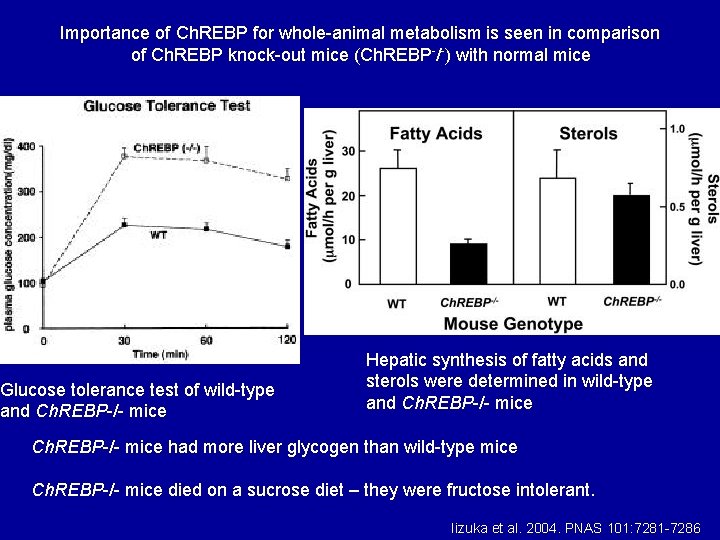
Importance of Ch. REBP for whole-animal metabolism is seen in comparison of Ch. REBP knock-out mice (Ch. REBP-/-) with normal mice Glucose tolerance test of wild-type and Ch. REBP-/- mice Hepatic synthesis of fatty acids and sterols were determined in wild-type and Ch. REBP-/- mice had more liver glycogen than wild-type mice Ch. REBP-/- mice died on a sucrose diet – they were fructose intolerant. Iizuka et al. 2004. PNAS 101: 7281 -7286

Metabolic control analysis S E 1 A 1 E 2 A 2 E 3 A 3 E 4 … En P Flux J Control of flux J through a pathway is distributed among all the enzymes A key regulatory enzyme is characterized by a large change in flux, when the enzymatic activity in the cell changes (by amount, allosteric effectors, or covalent modification). Example: phosphofructokinase in glycolysis Once one enzymatic bottleneck for flux is removed, another enzyme may become limiting for the flux.

Metabolic control analysis S E 1 A 1 E 2 A 2 E 3 A 3 E 4 … En Flux J Theoretically, if the enzyme activities of all the enzymes in the pathway were changed by the same degree (and if S were not limiting), then the flux would change by the same degree. Efficient increase of flux through pathway is brought about by increasing the enzyme activities or enzyme concentrations of all regulatory enzymes in a pathway. P
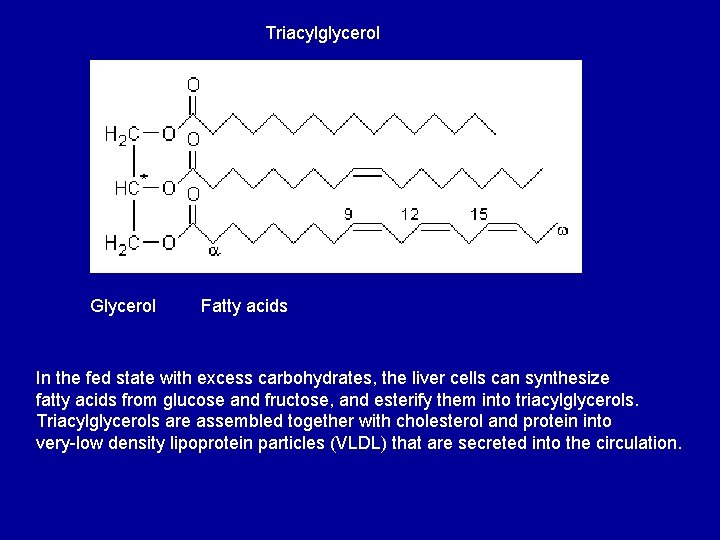
Triacylglycerol Glycerol Fatty acids In the fed state with excess carbohydrates, the liver cells can synthesize fatty acids from glucose and fructose, and esterify them into triacylglycerols. Triacylglycerols are assembled together with cholesterol and protein into very-low density lipoprotein particles (VLDL) that are secreted into the circulation.

Ch. REBP • Mlx Is the Principal Mediator of Glucose-induced Gene Expression in the Liver* Lin Ma 1, Luke N. Robinson 1, and Howard C. Towle 2
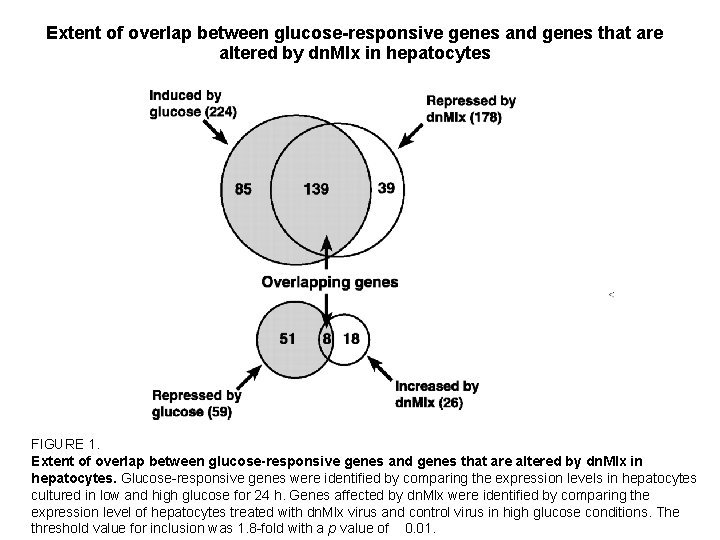
Extent of overlap between glucose-responsive genes and genes that are altered by dn. Mlx in hepatocytes FIGURE 1. Extent of overlap between glucose-responsive genes and genes that are altered by dn. Mlx in hepatocytes. Glucose-responsive genes were identified by comparing the expression levels in hepatocytes cultured in low and high glucose for 24 h. Genes affected by dn. Mlx were identified by comparing the expression level of hepatocytes treated with dn. Mlx virus and control virus in high glucose conditions. The threshold value for inclusion was 1. 8 -fold with a p value of 0. 01.
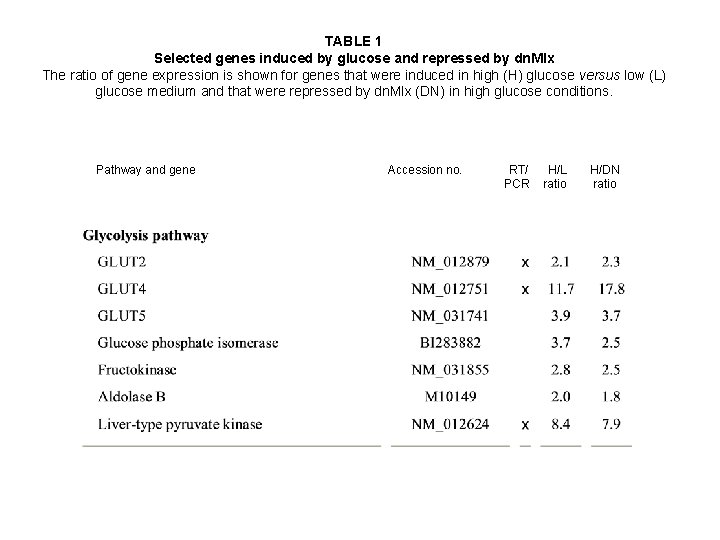
TABLE 1 Selected genes induced by glucose and repressed by dn. Mlx The ratio of gene expression is shown for genes that were induced in high (H) glucose versus low (L) glucose medium and that were repressed by dn. Mlx (DN) in high glucose conditions. Pathway and gene Accession no. RT/ PCR H/L ratio H/DN ratio
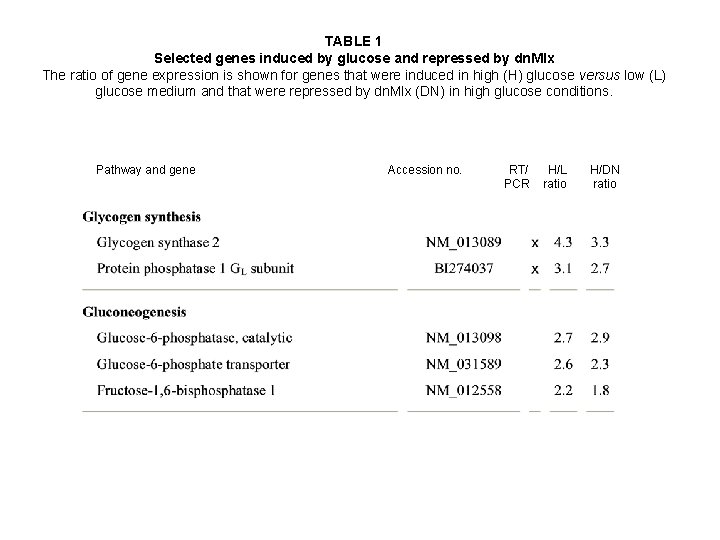
TABLE 1 Selected genes induced by glucose and repressed by dn. Mlx The ratio of gene expression is shown for genes that were induced in high (H) glucose versus low (L) glucose medium and that were repressed by dn. Mlx (DN) in high glucose conditions. Pathway and gene Accession no. RT/ PCR H/L ratio H/DN ratio
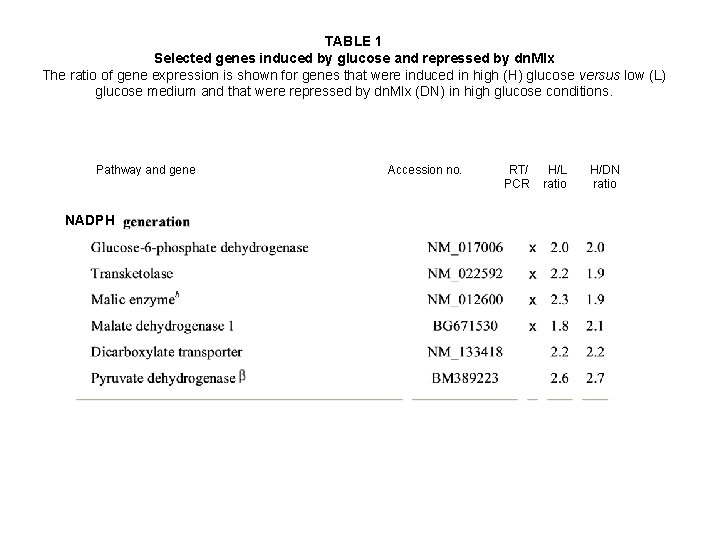
TABLE 1 Selected genes induced by glucose and repressed by dn. Mlx The ratio of gene expression is shown for genes that were induced in high (H) glucose versus low (L) glucose medium and that were repressed by dn. Mlx (DN) in high glucose conditions. Pathway and gene NADPH Accession no. RT/ PCR H/L ratio H/DN ratio
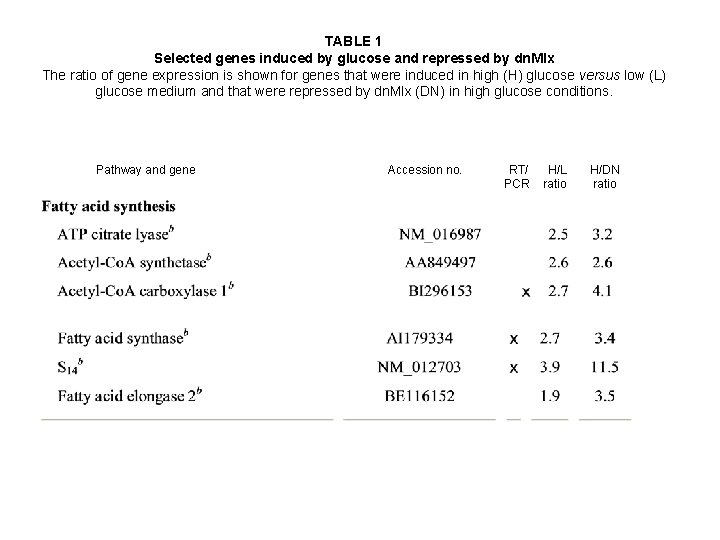
TABLE 1 Selected genes induced by glucose and repressed by dn. Mlx The ratio of gene expression is shown for genes that were induced in high (H) glucose versus low (L) glucose medium and that were repressed by dn. Mlx (DN) in high glucose conditions. Pathway and gene Accession no. RT/ PCR H/L ratio H/DN ratio
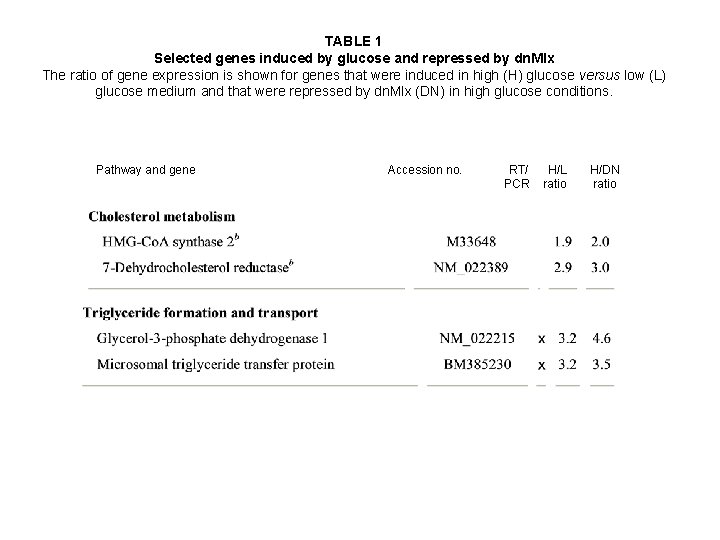
TABLE 1 Selected genes induced by glucose and repressed by dn. Mlx The ratio of gene expression is shown for genes that were induced in high (H) glucose versus low (L) glucose medium and that were repressed by dn. Mlx (DN) in high glucose conditions. Pathway and gene Accession no. RT/ PCR H/L ratio H/DN ratio
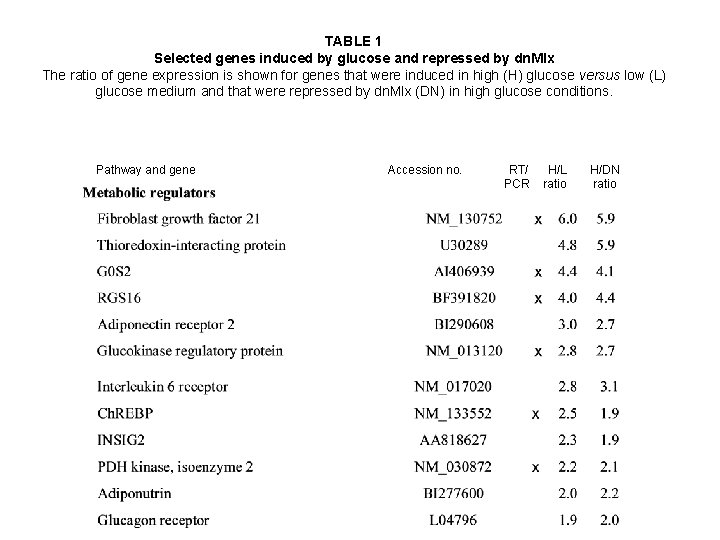
TABLE 1 Selected genes induced by glucose and repressed by dn. Mlx The ratio of gene expression is shown for genes that were induced in high (H) glucose versus low (L) glucose medium and that were repressed by dn. Mlx (DN) in high glucose conditions. Pathway and gene Accession no. RT/ PCR H/L ratio H/DN ratio
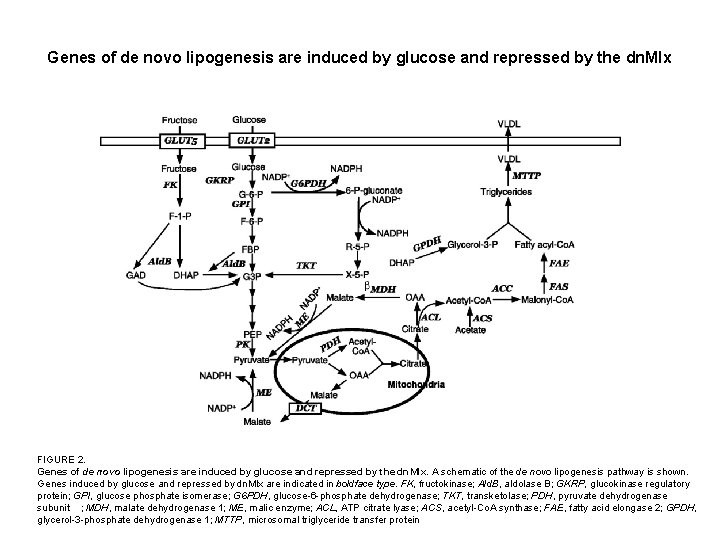
Genes of de novo lipogenesis are induced by glucose and repressed by the dn. Mlx FIGURE 2. Genes of de novo lipogenesis are induced by glucose and repressed by the dn. Mlx. A schematic of the de novo lipogenesis pathway is shown. Genes induced by glucose and repressed by dn. Mlx are indicated in boldface type. FK, fructokinase; Ald. B, aldolase B; GKRP, glucokinase regulatory protein; GPI, glucose phosphate isomerase; G 6 PDH, glucose-6 -phosphate dehydrogenase; TKT, transketolase; PDH, pyruvate dehydrogenase subunit ; MDH, malate dehydrogenase 1; ME, malic enzyme; ACL, ATP citrate lyase; ACS, acetyl-Co. A synthase; FAE, fatty acid elongase 2; GPDH, glycerol-3 -phosphate dehydrogenase 1; MTTP, microsomal triglyceride transfer protein
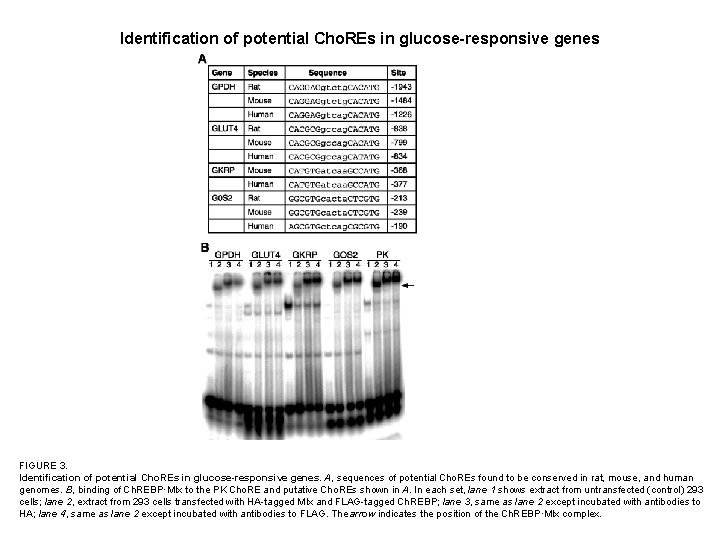
Identification of potential Cho. REs in glucose-responsive genes FIGURE 3. Identification of potential Cho. REs in glucose-responsive genes. A, sequences of potential Cho. REs found to be conserved in rat, mouse, and human genomes. B, binding of Ch. REBP·Mlx to the PK Cho. RE and putative Cho. REs shown in A. In each set, lane 1 shows extract from untransfected (control) 293 cells; lane 2, extract from 293 cells transfected with HA-tagged Mlx and FLAG-tagged Ch. REBP; lane 3, same as lane 2 except incubated with antibodies to HA; lane 4, same as lane 2 except incubated with antibodies to FLAG. The arrow indicates the position of the Ch. REBP·Mlx complex.
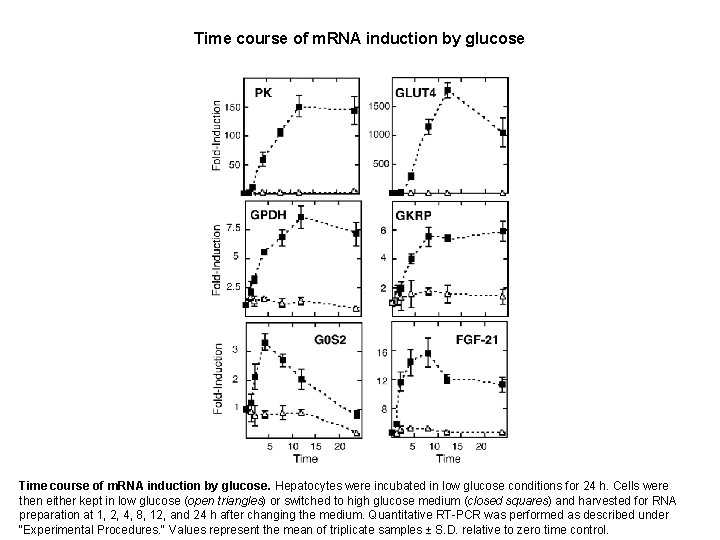
Time course of m. RNA induction by glucose. Hepatocytes were incubated in low glucose conditions for 24 h. Cells were then either kept in low glucose (open triangles) or switched to high glucose medium (closed squares) and harvested for RNA preparation at 1, 2, 4, 8, 12, and 24 h after changing the medium. Quantitative RT-PCR was performed as described under "Experimental Procedures. " Values represent the mean of triplicate samples ± S. D. relative to zero time control.
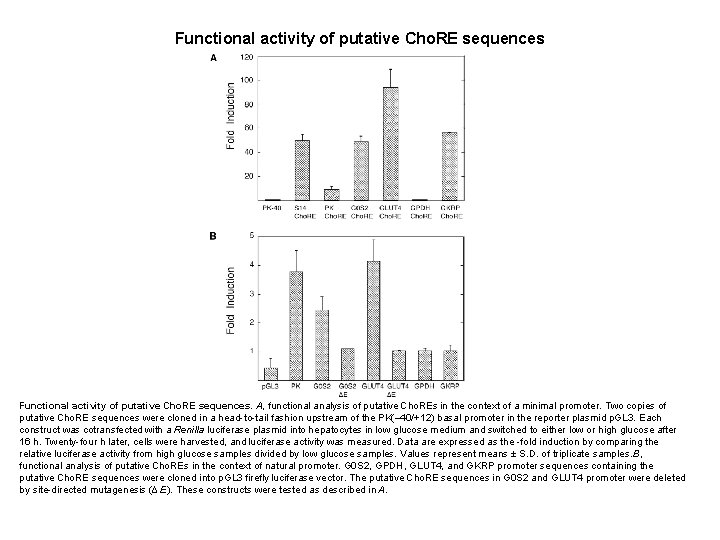
Functional activity of putative Cho. RE sequences. A, functional analysis of putative Cho. REs in the context of a minimal promoter. Two copies of putative Cho. RE sequences were cloned in a head-to-tail fashion upstream of the PK(– 40/+12) basal promoter in the reporter plasmid p. GL 3. Each construct was cotransfected with a Renilla luciferase plasmid into hepatocytes in low glucose medium and switched to either low or high glucose after 16 h. Twenty-four h later, cells were harvested, and luciferase activity was measured. Data are expressed as the -fold induction by comparing the relative luciferase activity from high glucose samples divided by low glucose samples. Values represent means ± S. D. of triplicate samples. B, functional analysis of putative Cho. REs in the context of natural promoter. G 0 S 2, GPDH, GLUT 4, and GKRP promoter sequences containing the putative Cho. RE sequences were cloned into p. GL 3 firefly luciferase vector. The putative Cho. RE sequences in G 0 S 2 and GLUT 4 promoter were deleted by site-directed mutagenesis (Δ E). These constructs were tested as described in A.
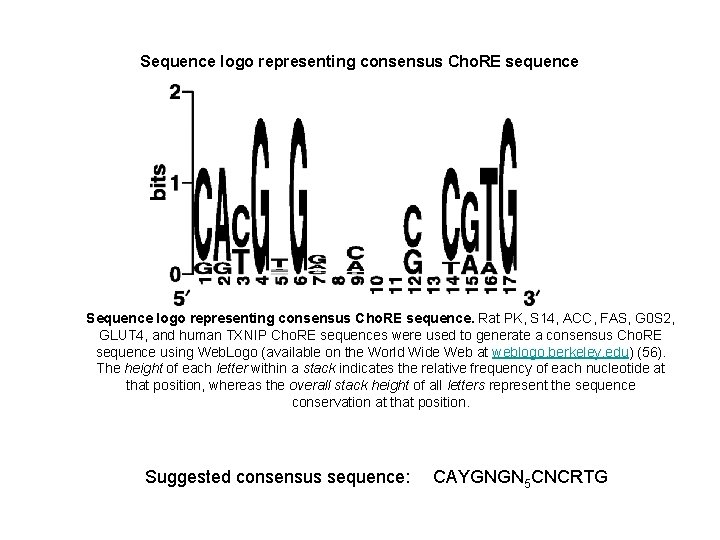
Sequence logo representing consensus Cho. RE sequence. Rat PK, S 14, ACC, FAS, G 0 S 2, GLUT 4, and human TXNIP Cho. RE sequences were used to generate a consensus Cho. RE sequence using Web. Logo (available on the World Wide Web at weblogo. berkeley. edu) (56). The height of each letter within a stack indicates the relative frequency of each nucleotide at that position, whereas the overall stack height of all letters represent the sequence conservation at that position. Suggested consensus sequence: CAYGNGN 5 CNCRTG

Conclusion: Glucose up-regulates genes encoding enzymes and proteins involved in the entire pathway of lipogenesis. Ch. REBP-Mlx is the principal mediator that transcriptionally regulates glucose-responsive genes.
- Slides: 30