Carbohydrates are the most abundant biomolecules on Earth

Carbohydrates are the most abundant biomolecules on Earth (each year, photosynthesis converts more than 100 billion metric tons of C 02 and H 20 into cellulose and other plant products). Certain carbohydrates (sugar and starch) are a dietary staple in most parts of the world, and the oxidation of carbohydrates is the central energy yielding pathway in most non photosynthetic cells. Monosaccharides, or simple sugars, consist of a single polyhydroxy aldehyde or ketone unit. The most abundant monosaccharide in nature is the six carbon sugar D glucose. Monosaccharides of four or more carbons tend to have cyclic structures. Oligosaccharides consist of short chains of mono saccharide units, or residues, joined by characteristic linkages called glycosidic bonds. The rnost abundant are the disaccharides, with two monosaccharide units. Typical is sucrose (cane sugar), which consists of the six carbon sugars D glucose and D fructose. The polysaccharides are sugar polymers containing more than 20 or more monosaccharide units; some have hundreds or thousands of units. Some polysaccharides, such as cellulose, are linear chains; others, such as glycogen, are branched. Both glycogen and cellulose consist of recurring units of D glucose, but they differ in the type of glycosidic linkage and consequently have strikingly different properties and biological roles.

The backbones of common mono saccharides are unbranched carbon chains in which all the carbon atoms are linked by single bonds. In the open chain form, one of the carbon atoms is double bonded to an oxygen atom to form a carbonyl group; each of the other carbon atoms has a hydroxyl group. If the carbonyl group is at an end of the carbon chain (that is, in an aldehyde group) the monosaccharide is an aldose if the carbonyl group is at any other position (in a ketone group) the monosaccharide is a ketose. The simplest monosaccharides are the two three carbon trioses: glyceraldehyde, an aldotriose, and dihydroxyacetone , a ketotriose. Monosaccharides with four, five, six, and seven carbon atoms in their backbones are called, respectively, tetroses, pentoses , hexoses, and heptoses. There aldoses and ketoses of each of these chain lengths: aldotetroses and ketotetroses, aldopentoses and ketopentoses, and so on.
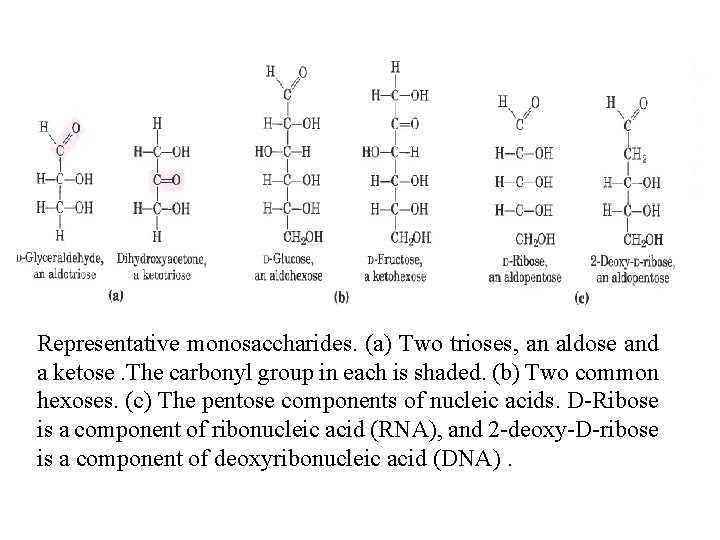
Representative monosaccharides. (a) Two trioses, an aldose and a ketose. The carbonyl group in each is shaded. (b) Two common hexoses. (c) The pentose components of nucleic acids. D Ribose is a component of ribonucleic acid (RNA), and 2 deoxy D ribose is a component of deoxyribonucleic acid (DNA).

Aldoses and ketoses having from three to six carbon atoms, shown as projection formulas. The carbon atoms in red are chiral centers. In these D isomers, the chiral carbon most distant from the carbonyl carbon has the same configuration as the chiral carbon in D glyceraldehyde.

Two sugars that differ only in the configuration around one carbon atom are called epimers; D glucose and D mannose, which differ only in the stereochemistry at C 2, are epimers, as are D glucose and D galactose (which differ at C 4)

In aqueousofsolution monosaccharides with An fivealdehyde or more carbon atoms the Formation hemiacetals and hemiketals. or ketone caninreact backbone occurinpredominantly as cyclic (ring) or structures in respectively, which the with an alcohol a 1: 1 ratio to yield a hemiacetal hemiketal, carbonyla group has formed a covalent bondcarbon with the oxygen of of a hydroxyl creating new chiral center at the carbonyl. Substitution a second group along the chain. The an formation these. When ring structures the resultis alcohol molecule produces acetal orofketal. the secondis alcohol of aofgeneral between the alcohols aldehydes or ketones bond to form part anotherreaction sugar molecule, b+ond and produced is a glycosidic derivatives called hemiacetals or hemiketals, which contain an additional asymmetric carbon atom and thus can exist in two stereoisomers forms.
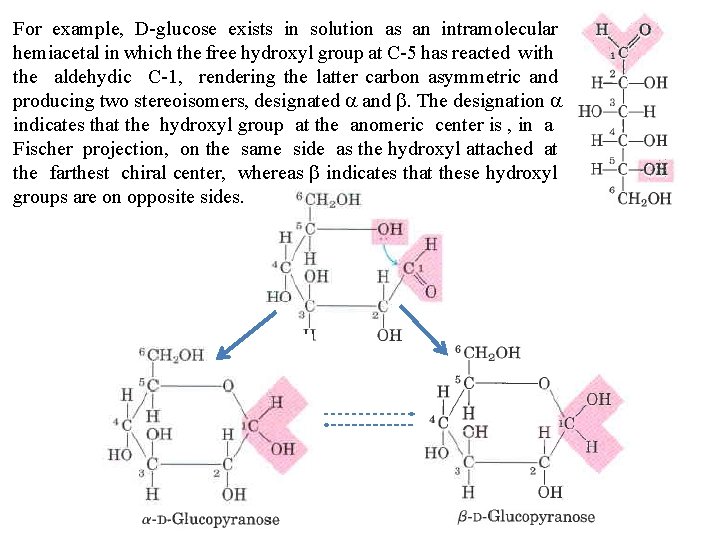
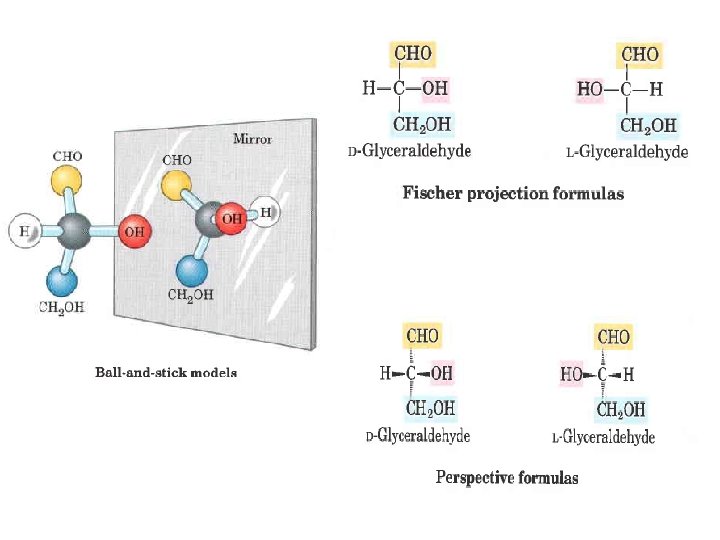
For example, D glucose exists in solution as an intramolecular hemiacetal in which the free hydroxyl group at C 5 has reacted with the aldehydic C 1, rendering the latter carbon asymmetric and producing two stereoisomers, designated a and b. The designation a indicates that the hydroxyl group at the anomeric center is , in a Fischer projection, on the same side as the hydroxyl attached at the farthest chiral center, whereas b indicates that these hydroxyl groups are on opposite sides.
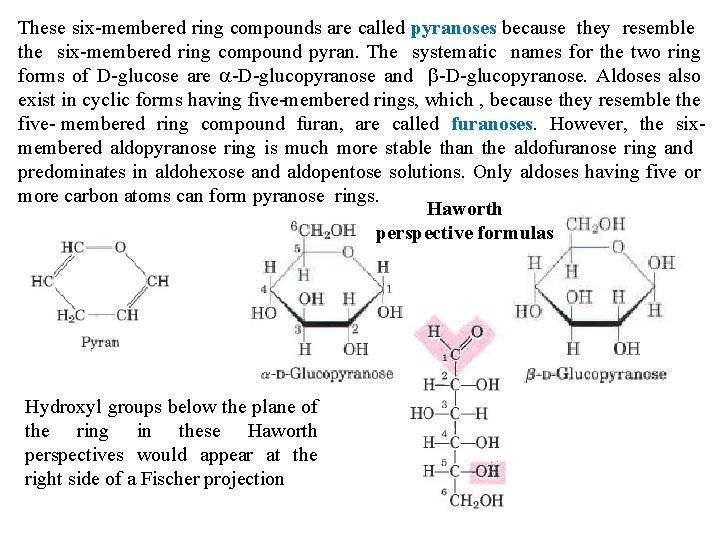
These six membered ring compounds are called pyranoses because they resemble Isomerie forms of monosaccharides that differ in their configuration about the six membered ring compound pyran. Theonly systematic names for the twothe ring hemiacetal or hemiketal carbon atom are called anomers. The hemiacetal forms of D glucose are a D glucopyranose and b D glucopyranose. Aldoses(or also carbonyl) carbon atom is called the anomeric carbon. a and bthey anomers of D exist in cyclic forms having five membered rings, which. The , because resemble the glucose interconvert aqueous solution processfuranoses. called mutarotation five membered ring incompound furan, by area called However, Thus, the asix solution ofaldopyranose a D glucose ring and ais solution of b D glucose form identical membered much more stable than eventually the aldofuranose ring and equilibrium mixtures havingand identical optical solutions. properties. Only aldoses having five or predominates in aldohexose aldopentose more carbon atoms can form pyranose rings. Haworth perspective formulas Hydroxyl groups below the plane of the ring in these Haworth perspectives would appear at the right side of a Fischer projection
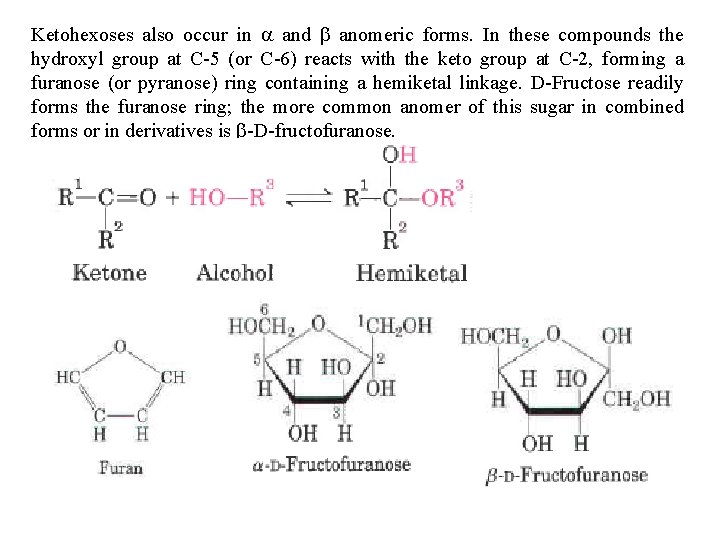
Ketohexoses also occur in a and b anomeric forms. In these compounds the hydroxyl group at C 5 (or C 6) reacts with the keto group at C 2, forming a furanose (or pyranose) ring containing a hemiketal linkage. D Fructose readily forms the furanose ring; the more common anomer of this sugar in combined forms or in derivatives is b D fructofuranose.
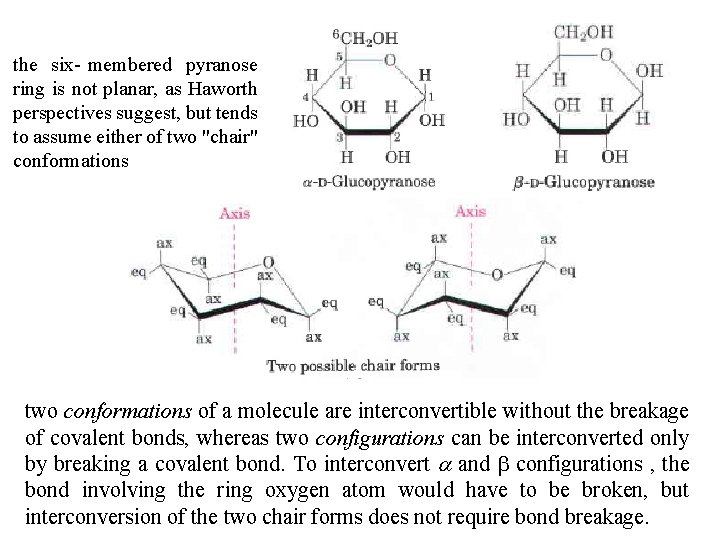
the six membered pyranose ring is not planar, as Haworth perspectives suggest, but tends to assume either of two "chair" conformations two conformations of a molecule are interconvertible without the breakage of covalent bonds, whereas two configurations can be interconverted only by breaking a covalent bond. To interconvert a and b configurations , the bond involving the ring oxygen atom would have to be broken, but interconversion of the two chair forms does not require bond breakage.

Conformational chair forms of the pyranose ring. Bonds to substituents and hydrogen atoms on the ring carbons may be either axial (ax), projecting parallel to the vertical axis through the ring, or equatorial (eq), projecting roughly perpendicular to this axis. Two conformers such are these are not readily interconvertible without breaking the ring. The preferred chair conformation of a D glucopyranose. Generally, substituents in the equatorial positions are less sterically hindered by neighboring substituents, and conformers with bulky substituents in equatorial positions are favored.

there a number of sugar derivatives in which a hydroxyl group in the parent compound is replaced with another substituent, or a carbon atom is oxdized to a carboxyl group. In glucosamine, galactosamine, and mannosamine, the hydroxyl at C 2 of the parent compound is replaced with an amino group. The amino group is nearly always condensed with acetic acid, as in N acetylglucosamine. This glucosamine derivative is part of many structural polymers, including those of the bacterial cell wall Bacterial cell walls also contain a derivative of glucosamine, N acetylmuramic acid, in which lactic acid (a three carbon carboxylic acid) is ether link ed to the oxygen at C 3 of N acetylglucosarnine.
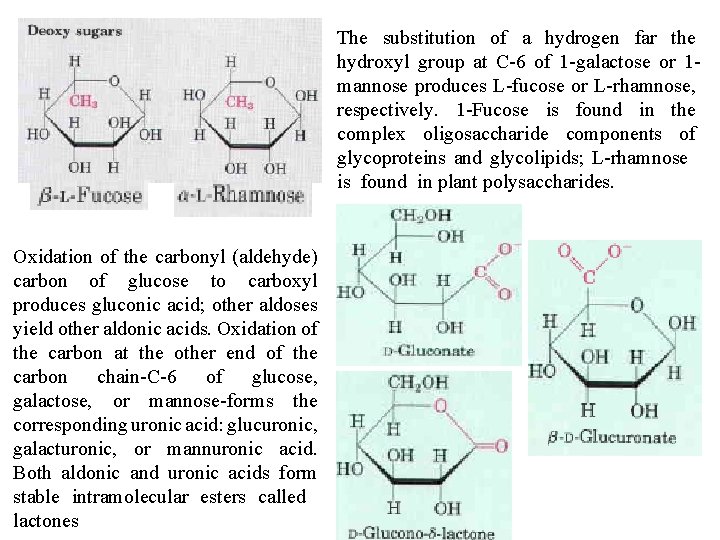
The substitution of a hydrogen far the hydroxyl group at C 6 of 1 galactose or 1 mannose produces L fucose or L rhamnose, respectively. 1 Fucose is found in the complex oligosaccharide components of glycoproteins and glycolipids; L rhamnose is found in plant polysaccharides. Oxidation of the carbonyl (aldehyde) carbon of glucose to carboxyl produces gluconic acid; other aldoses yield other aldonic acids. Oxidation of the carbon at the other end of the carbon chain C 6 of glucose, galactose, or mannose forms the corresponding uronic acid: glucuronic, galacturonic, or mannuronic acid. Both aldonic and uronic acids form stable intramolecular esters called lactones
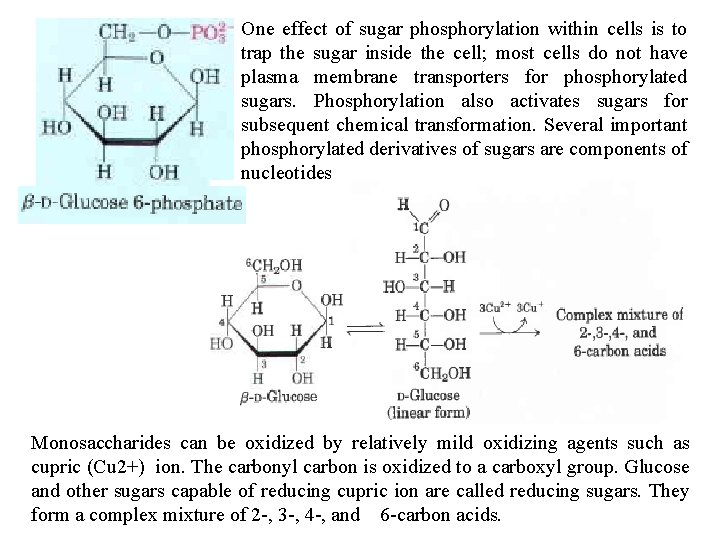
One In effect the synthesis of sugar and phosphorylation metabolism of within carbohydrates, cells is to trap the intermediates sugar inside the arecell; verymost oftencells notdothenotsugars have plasma them membrane selves but transporters their phosphorylated for phosphorylated derivatives. sugars. Condensation Phosphorylation of phosphoric also activates acid with sugars one of the for subsequent hydroxyl chemical groupstransformation. of a sugar forms Several a phosphate important phosphorylated ester, as in glucose derivatives 6 phosphate of sugars. are Sugar components phosphates of nucleotides are relatively stable at neutral p. H and bear a negative charge. Monosaccharides can be oxidized by relatively mild oxidizing agents such as cupric (Cu 2+) ion. The carbonyl carbon is oxidized to a carboxyl group. Glucose and other sugars capable of reducing cupric ion are called reducing sugars. They form a complex mixture of 2 , 3 , 4 , and 6 carbon acids.

Disaccharid consist of two monosaccharides joined covalently by an O glycosidic bond , which is formed when a hydroxyl group of one sugar reacts with the anomeric carbon of the other. This reaction represents the formation of an acetal from a hemiacetal (such as glucopyranose) and an alcohol (a hydroxyl group of the second sugar molecule), and the resulting compound is called a glycoside. Glycosidic bonds are readily hydrolyzed by acid but resist cleavage by base.
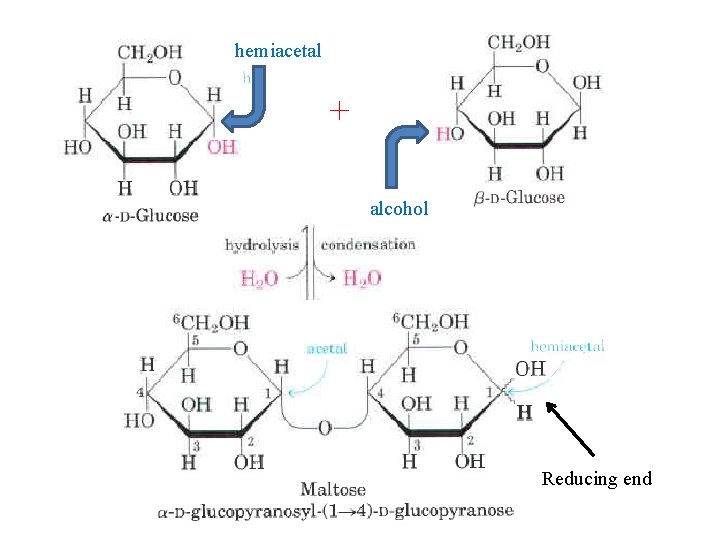
hemiacetal + alcohol Reducing end
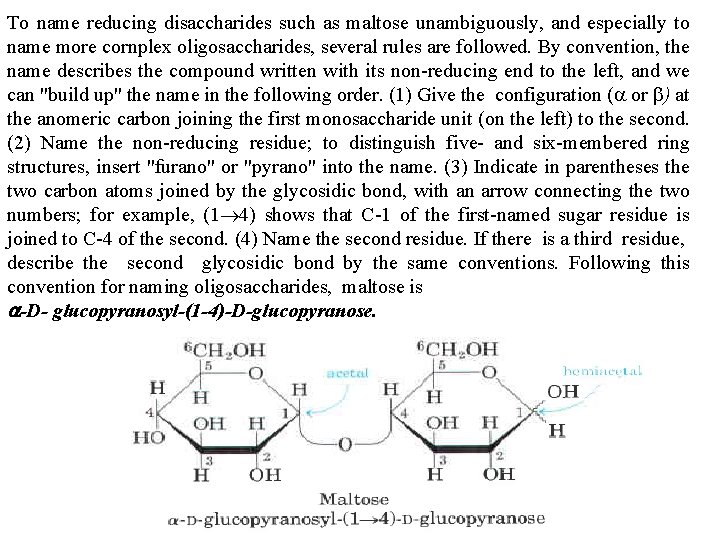
To name reducing disaccharides such as maltose unambiguously, and especially to name more cornplex oligosaccharides, several rules are followed. By convention, the name describes the compound written with its non reducing end to the left, and we can "build up" the name in the following order. (1) Give the configuration (a or b) at the anomeric carbon joining the first monosaccharide unit (on the left) to the second. (2) Name the non reducing residue; to distinguish five and six membered ring structures, insert "furano" or "pyrano" into the name. (3) Indicate in parentheses the two carbon atoms joined by the glycosidic bond, with an arrow connecting the two numbers; for example, (1 4) shows that C 1 of the first named sugar residue is joined to C 4 of the second. (4) Name the second residue. If there is a third residue, describe the second glycosidic bond by the same conventions. Following this convention for naming oligosaccharides, maltose is a D glucopyranosyl (1 4) D glucopyranose.
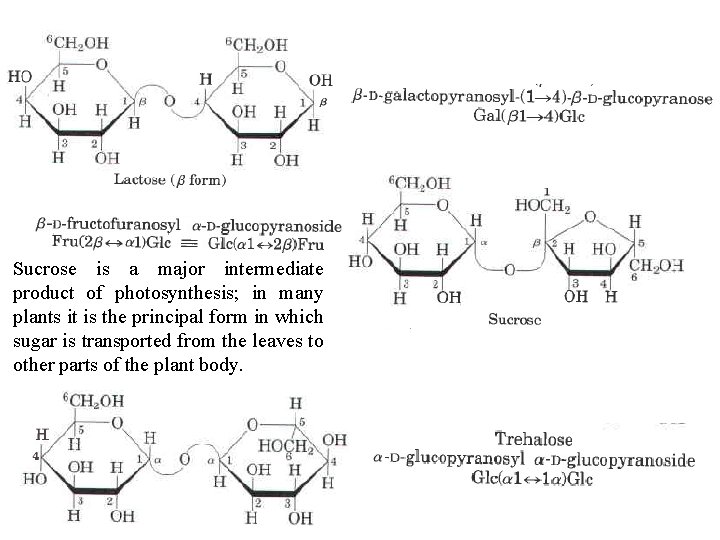
Sucrose is a major intermediate product of photosynthesis; in many plants it is the principal form in which sugar is transported from the leaves to other parts of the plant body.

Polysaccharides Most carbohydrates found in nature occur as polysac charides, polymers of medium to high molecular weight. Polysaccharides, also called glycans, glycans differ from each other in the identity of their recurring monosaccharide units, in the length of their chains, in the types of bonds linking the units, and in the degree of branching. Homopolysaccharides contain only a single monomeric species; heteropolysaccharides contain two or more different kinds. Some homopolysaccharides serve as storage forms of monosaccharides that are used as fuels; starch and glycogen are homopolysaccharides of this type. Other homopolysaccharides (cellulose) serve as structural elements in plant cell walls.

The most important storage polysaccharides are starch in plant cells and glycogen in animal cells. Both polysaccharides occur intracellularly as large clusters or granules. Starch and glycogen molecules are heavily hydrated, because they have many exposed hydroxyl groups available to hydrogen bond with water. Most plant cells have the ability to form starch, and starch storage is especially abundant in tubers (underground stems), such as potatoes, and in seeds. Starch contains two types of glucose polymer, amylose and amylopectin amylose consists of long, linear (unbranched) chains of D glucose residues connected by (a l 4) linkages (as in maltose). Such chains vary in molecular weight from a few thousand to more than a million. Amylopectin also has a high molecular weight (up to 200 million) but unlike amylose is highly branched. The glycosidic linkages joining successive glucose residues in amylopectin chains are (al 4); the branch points (occurring every 24 to 30 residues) are (a 1 6) linkages.
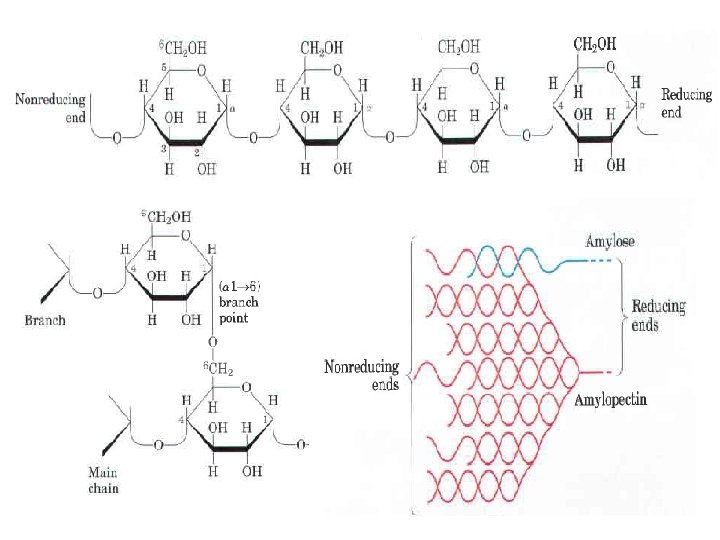

Why not store glucose in its monomeric form? It has been calculated that a glucose concentration of 0. 4 M correspond to an actual concentration of glycogen ( Starch), which is insoluble and contributes little to the osmolarity of the cytosol, of about 0. 01 m. M, is. If the cytosol contained 0. 4 M glucose, the osmolarity would be threateningly elevated, leading to osmotic entry of water that might rupture the cell. Furthermore, with an intracellular glucose concentration of 0. 4 M the free energy change for glucose uptake into cells against this very high concentration gradient would be prohibitively large. Dextrans are bacterial and yeast polysaccharides made up of (a l 6) linked poly D glucose; all have (a l 3) branches, and some also have (a l 2) or (a l 4) branches. Syn theticdextrans are used in several commerciai products (for example, Sephadex) that serve in the fractionation of proteins by size exclusion chromatography. The dextrans in these products are chemically cross linked to form insoluble materials of various porosities, admitting macromolecules of various sizes.
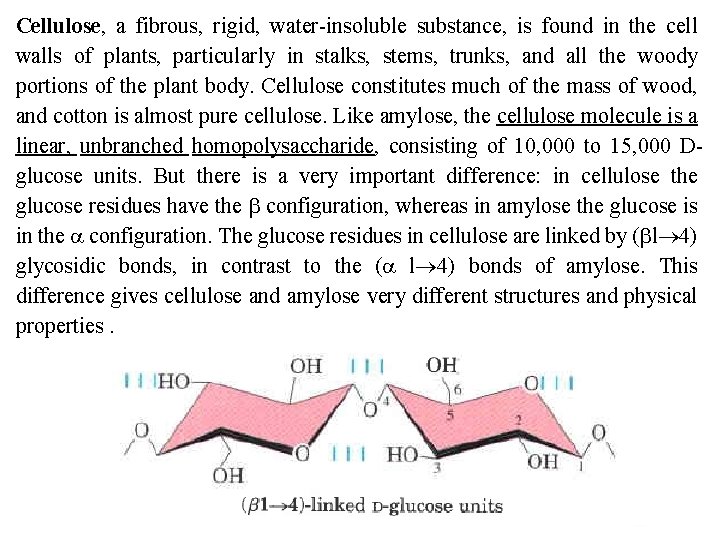
Cellulose, a fibrous, rigid, water insoluble substance, is found in the cell walls of plants, particularly in stalks, stems, trunks, and all the woody portions of the plant body. Cellulose constitutes much of the mass of wood, and cotton is almost pure cellulose. Like amylose, the cellulose molecule is a linear, unbranched homopolysaccharide, consisting of 10, 000 to 15, 000 D glucose units. But there is a very important difference: in cellulose the glucose residues have the b configuration, whereas in amylose the glucose is in the a configuration. The glucose residues in cellulose are linked by (bl 4) glycosidic bonds, in contrast to the (a l 4) bonds of amylose. This difference gives cellulose and amylose very different structures and physical properties.

The folding of polysaccharides in three dimensional structures follows the same principles as those governing polypeptide structure: subunits with a more or less rigid structure dictated by covalent bonds form three dimensional macromolecular structures that are stabilized by weak interactions within or between molecules: hydrogen bonds and hydrophobic and van der Waals interactions, and, for polymers with charged subunits, electrostatic interactions. Because polysaccharides have so many hydroxyl groups, hydrogen bonding has an especially important influence on their structure. Glycogen, starch, and cellulose are composed of pyranoside subunits (having six membered rings). Such molecules can be represented as a series of rigid pyranose rings connected by an oxygen atom bridging two carbon atoms (the glycosidic bond). There is, in principle, free rotation about both C O bonds linking the residues, but as in polypeptides, rotation about each bond is limited by steric hindrance by substituents. The three dimensional structures of these molecules can be described in terms of the dihedral angles, f and , about the glycosidic bond analogous to angles f and made by the peptide bond
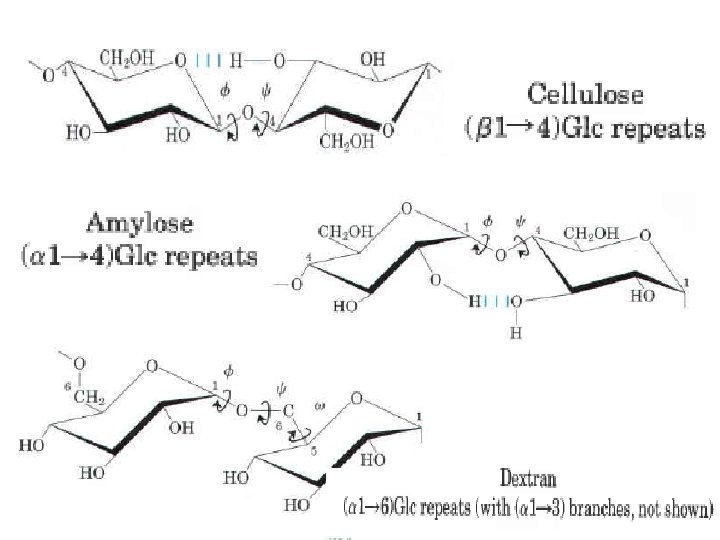
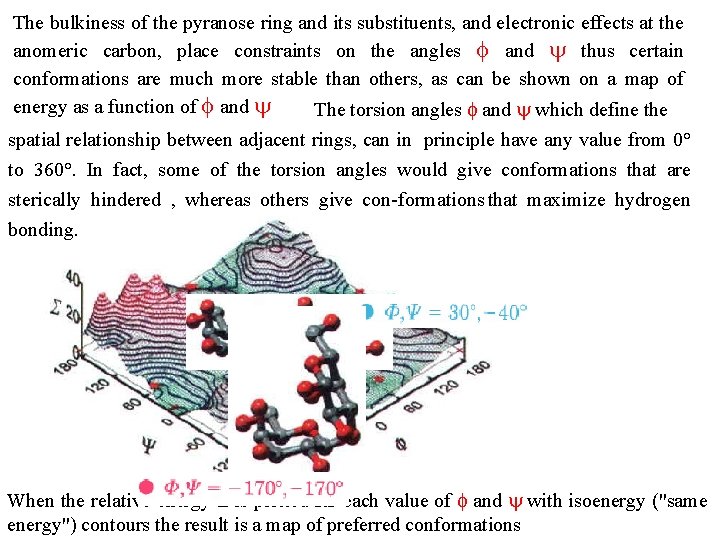
The bulkiness of the pyranose ring and its substituents, and electronic effects at the anomeric carbon, place constraints on the angles f and thus certain conformations are much more stable than others, as can be shown on a map of energy as a function of f and The torsion angles f and which define the spatial relationship between adjacent rings, can in principle have any value from 0° to 360°. In fact, some of the torsion angles would give conformations that are sterically hindered , whereas others give con formations that maximize hydrogen bonding. When the relative energy S is plotted far each value of f and with isoenergy ("same energy") contours the result is a map of preferred conformations
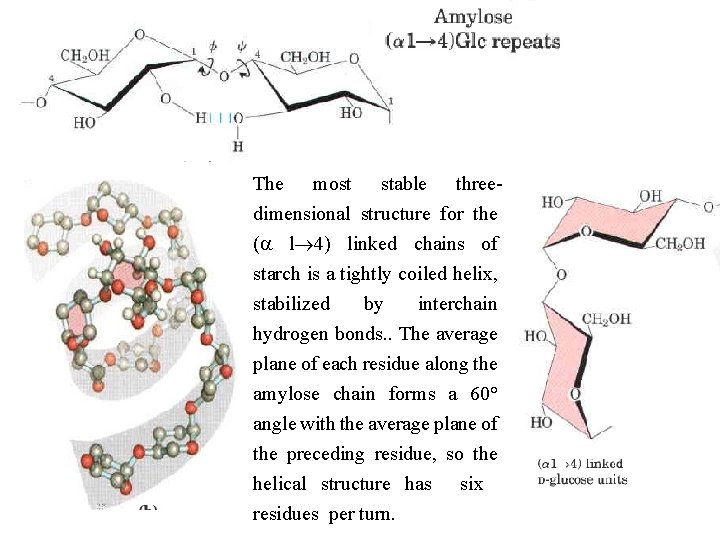
The most stable three dimensional structure for the (a l 4) linked chains of starch is a tightly coiled helix, stabilized by interchain hydrogen bonds. . The average plane of each residue along the amylose chain forms a 60° angle with the average plane of the preceding residue, so the helical structure has six residues per turn.
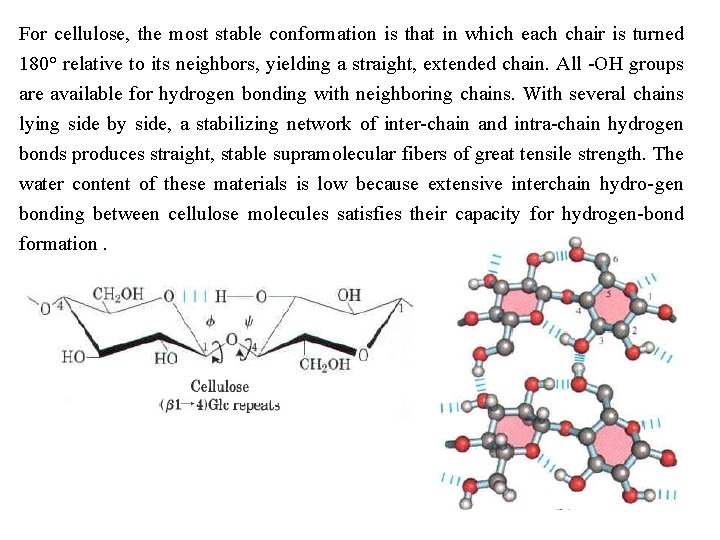
For cellulose, the most stable conformation is that in which each chair is turned 180° relative to its neighbors, yielding a straight, extended chain. All OH groups are available for hydrogen bonding with neighboring chains. With several chains lying side by side, a stabilizing network of inter chain and intra chain hydrogen bonds produces straight, stable supramolecular fibers of great tensile strength. The water content of these materials is low because extensive interchain hydro gen bonding between cellulose molecules satisfies their capacity for hydrogen bond formation.

Lipids Biological lipids are a chemically diverse group of compounds, the common and defining feature of which is their insolubility in water. The biological functions of the lipids are as diverse as their chemistry. Fats and oils are the principal stored forms of energy in many organisms. Phospholipids and sterols are major structural elements of biological membranes. Other lipids , although present in relatively small quantities, play crucial roles as enzyme cofactors, electron carriers, light absorbing pigments, etc Storage Lipids The fats and oils used almost universally as stored forms of energy in living organisms are derivatives of fatty acids The fatty acids are hydrocarbon derivatives, at about the same low oxidation state (that is, as highly reduced) as the hydrocarbons in fossil fuels. The complete cellular oxidation of fatty acids (to CO 2 and H 2 O), is highly exergonic.

Fatty acids are carboxylic acids with hydrocarbon chains ranging from 4 to 36 carbons long (C 4 to C 36). In some fatty acids, this chain is unbranched and fully saturated (contains no double bonds); in others the chain contains one or more double bonds. A few contain three carbon rings, hydroxyl groups, or methyl group branches. A simplified nomenclature for un branched fatty acids specifies the chain length and number of double bonds, separated by a colon for example, the 16 carbon saturated palmitic acid is abbreviated 16: 0, and the 18 carbon oleic acid, with one double bond, is 18: 1. The positions of any double bonds are specified relative to the carboxyl carbon, nurnbered 1, by superscript nurnbers following D (delta); a 20 carbon fatty acid with one double bond between C 9 and C 10 (C 1 being the carboxyl carbon) and another between C 12 and C 13 is designated 20: 2 (D 9, 12)
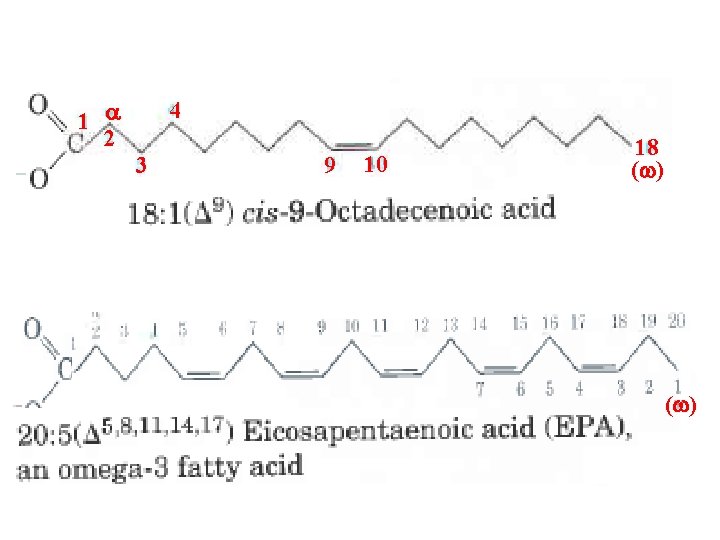
1 a 2 4 3 9 10 18 (w)

The physical properties of the fatty acids, are largely determined by the length and degree of unsaturation of the hydrocarbon chain. The nonpolar hydrocarbon chain accounts for the poor solubility of fatty acids in water. Lauric acid (12: 0, Mr 200), for example, has a solubility in water of 0. 063 mg/g much less than that of glucose (Mr 180), which is 1, 100 mg/g. The longer the fatty acyl chain and the fewer the double bonds, the lower is the solubility in water. The carboxylic acid group is polar (and ionized at neutral p. H) and accounts for the slight solubility of short chain fatty acids in water. Melting points are also strongly influenced by the length and degree of unsaturation of the hydrocarbon chain. At room temperature (25 °C) , the saturated fatty acids from 12: 0 to 24: 0 have a waxy consistency, whereas unsaturated fatty acids of these lengths are oily liquids. This difference in melting points is due to different degrees of packing of the fatty acid molecules In the fully saturated compounds, free rotation around each carbon bond gives the hydrocarbon chain great flexibility; the most stable conformation is the fully extended form, in which the steric hindrance of neighboring atoms is minimized. These molecules can pack together tightly in nearly crystalline arrays, with atoms all along their lengths in van der Waals contact with the atoms of neighboring molecules
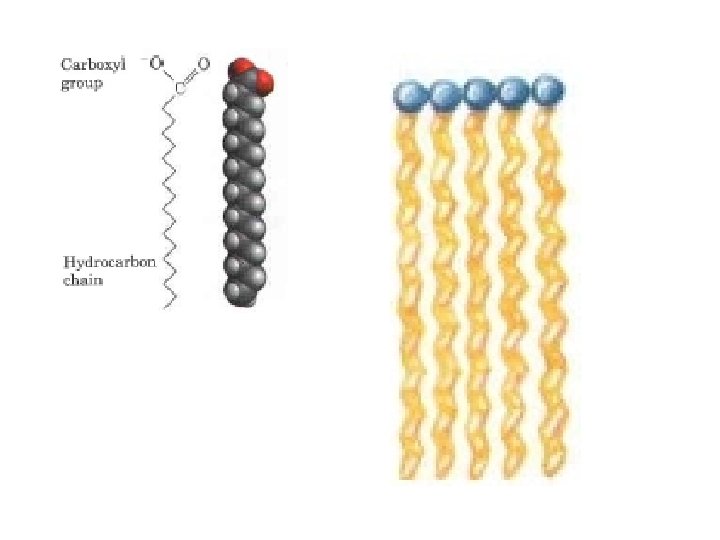
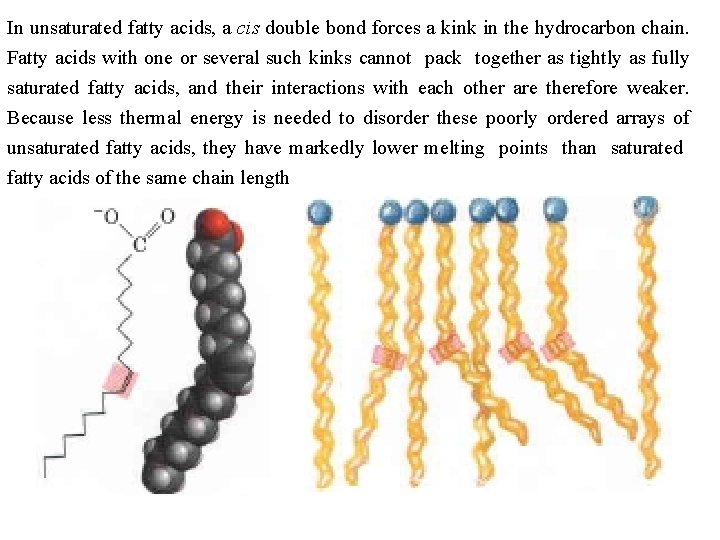
In unsaturated fatty acids, a cis double bond forces a kink in the hydrocarbon chain. Fatty acids with one or several such kinks cannot pack together as tightly as fully saturated fatty acids, and their interactions with each other are therefore weaker. Because less thermal energy is needed to disorder these poorly ordered arrays of unsaturated fatty acids, they have markedly lower melting points than saturated fatty acids of the same chain length
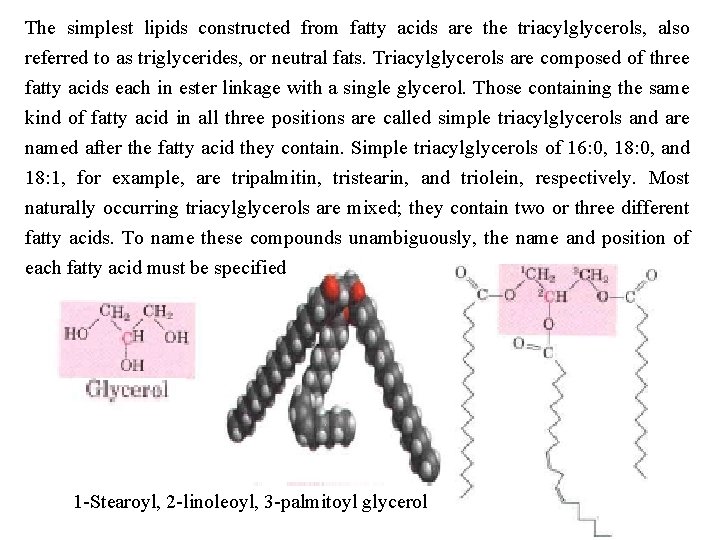
The simplest lipids constructed from fatty acids are the triacylglycerols, also referred to as triglycerides, or neutral fats. Triacylglycerols are composed of three fatty acids each in ester linkage with a single glycerol. Those containing the same kind of fatty acid in all three positions are called simple triacylglycerols and are named after the fatty acid they contain. Simple triacylglycerols of 16: 0, 18: 0, and 18: 1, for example, are tripalmitin, tristearin, and triolein, respectively. Most naturally occurring triacylglycerols are mixed; they contain two or three different fatty acids. To name these compounds unambiguously, the name and position of each fatty acid must be specified 1 Stearoyl, 2 linoleoyl, 3 palmitoyl glycerol
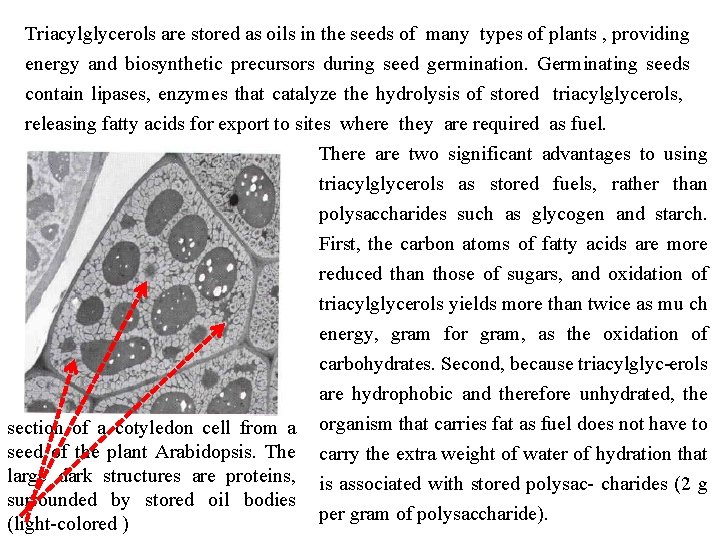
Triacylglycerols are stored as oils in the seeds of many types of plants , providing energy and biosynthetic precursors during seed germination. Germinating seeds contain lipases, enzymes that catalyze the hydrolysis of stored triacylglycerols, releasing fatty acids for export to sites where they are required as fuel. There are two significant advantages to using triacylglycerols as stored fuels, rather than polysaccharides such as glycogen and starch. First, the carbon atoms of fatty acids are more reduced than those of sugars, and oxidation of triacylglycerols yields more than twice as mu ch energy, gram for gram, as the oxidation of carbohydrates. Second, because triacylglyc erols are hydrophobic and therefore unhydrated, the section of a cotyledon cell from a organism that carries fat as fuel does not have to seed of the plant Arabidopsis. The carry the extra weight of water of hydration that large dark structures are proteins, is associated with stored polysac charides (2 g surrounded by stored oil bodies per gram of polysaccharide). (light colored )

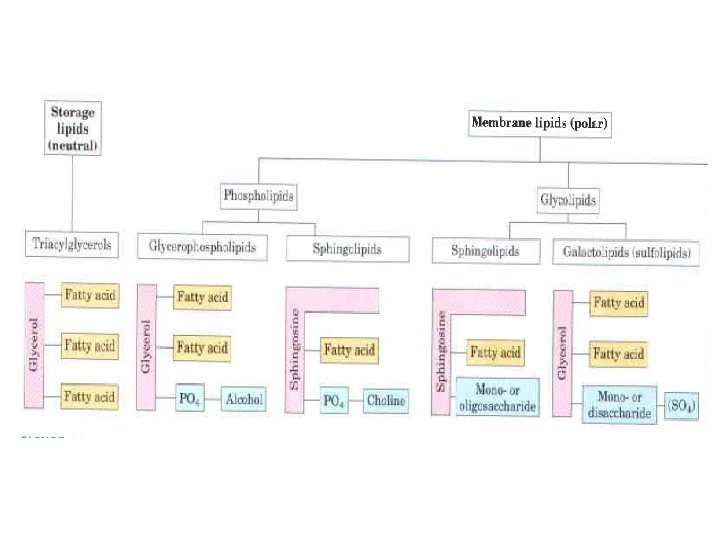
Structural Lipids in Membranes The central architectural feature of biological membranes is a double layer of lipids, which acts as a barrier to the passage of polar molecules and ions. Membrane lipids are amphipathic: one end of the molecule is hydrophobic, the other hydrophilic. Their hydrophobic interactions with each other and their hydrophilic interactions with water direct their packing into sheets called membrane bi layers. General types of membrane lipids: glycerophospholipids, in which the hydrophobic regions are composed of two fatty acids joined to glycerol; galactolipids and sulfolipids, which also contain two fatty acids esterified to glycerol, but lack the characteristic phosphate of phospholipids; sphingolipids, in which a single fatty acid is joined to a fatty amine, sphingosine; and sterols, compounds characterized by a rigid system of four fused hydrocarbon rings. The hydrophilic moieties in these amphipathic compounds may be as simple as a single OH group at one end of the sterol ring system, or they may be much more complex. In glycerophospholipids and some sphingolipids, a polar head group is joined to the hydrophobic moiety by a phosphodiester linkage; these are the phospholipids. Other sphingolipids lack phosphate but have a simple sugar or complex oligosaccharide at their polar ends; these are the glycolipid).

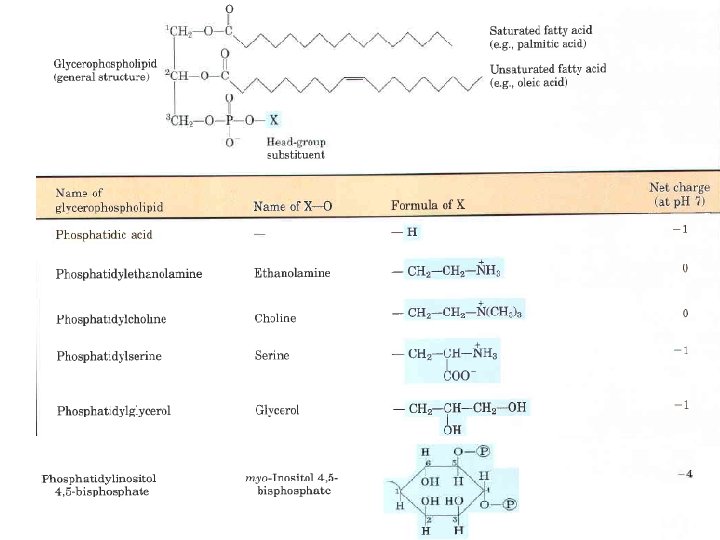
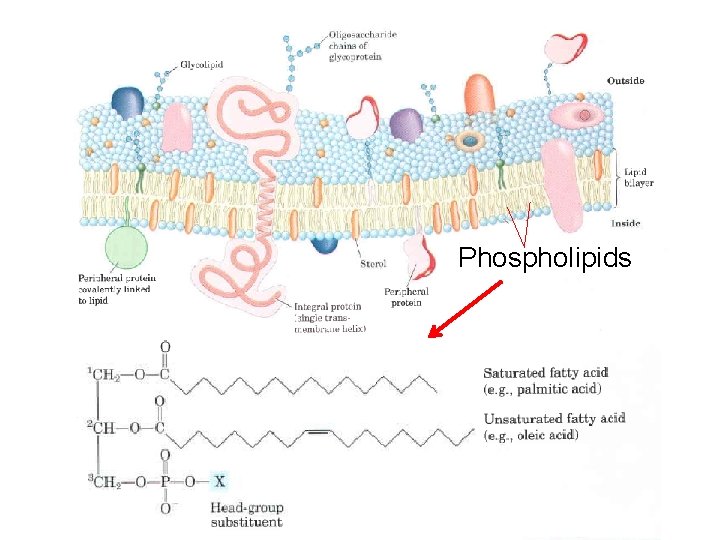
Glycerophospholipids, also called phosphoglycerides, are membrane lipids in which two fatty acids are attached in ester linkage to the first and second carbons of glycerol, and a highly polar or charged group is attached through a phosphodiester linkage to the third carbon. Glycerophospholipids are named as derivatives of the parent compound, phosphatidic acid, according to the polar alcohol in the head group. Saturated fatty acid (e. g. , palmitic acid) H Unsaturated fatty acid (e. g. , oleic acid)

Phosphatidylcholine and phosphatidylethanolamine have choline and ethanolamine as their polar head groups, for example. ln all these compounds , the head group is joined to glycerol through a phosphodiester bond, in which the phosphate group bears a negative charge at neutral p. H. The polar alcohol may be negatively charged (as in phosphatidylinositol 4, 5 bisphosphate), neutral (phosphatidylserine), or positively charged (phosphatidylcholine, phosphatidylethanolamine). The fatty acids in glycerophospholipids can be any of a wide variety, so a given phospholipid (phosphatidylcholine, for example) may consist of several molecular species, each with its unique complement of fatty acids. The distribution of molecular species is specific for different organisms, different tissues of the same organism, and different glycerophospholipids in the same cell or tissue. In general, glycerophospholipids contain a C 16 or C 18 saturated fatty acid at C 1 and a C 18 or C 20 unsaturated fatty acid at C 2.
Phospholipids
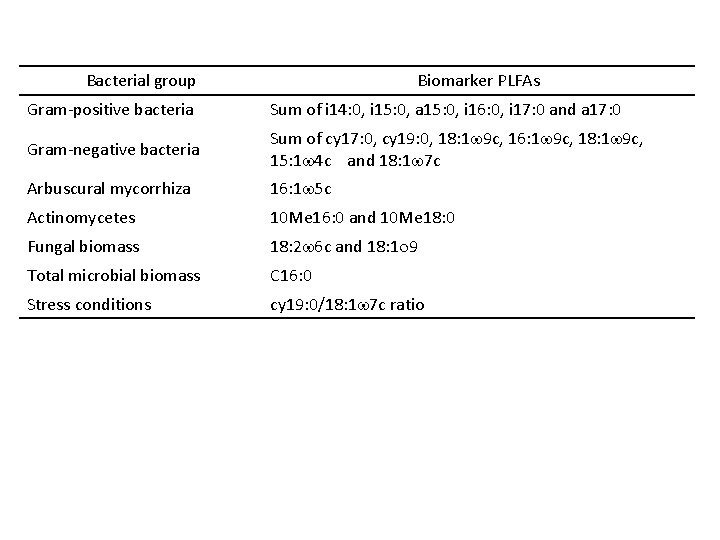
Bacterial group Biomarker PLFAs Gram-positive bacteria Sum of i 14: 0, i 15: 0, a 15: 0, i 16: 0, i 17: 0 and a 17: 0 Gram-negative bacteria Sum of cy 17: 0, cy 19: 0, 18: 1 w 9 c, 16: 1 w 9 c, 18: 1 w 9 c, 15: 1 w 4 c and 18: 1 w 7 c Arbuscural mycorrhiza 16: 1 w 5 c Actinomycetes 10 Me 16: 0 and 10 Me 18: 0 Fungal biomass 18: 2 w 6 c and 18: 1 o 9 Total microbial biomass C 16: 0 Stress conditions cy 19: 0/18: 1 w 7 c ratio
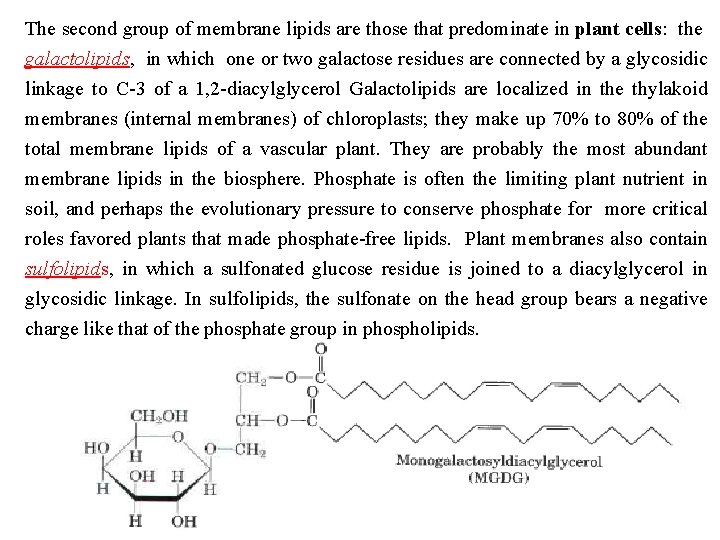
The second group of membrane lipids are those that predominate in plant cells: the galactolipids, in which one or two galactose residues are connected by a glycosidic linkage to C 3 of a 1, 2 diacylglycerol Galactolipids are localized in the thylakoid membranes (internal membranes) of chloroplasts; they make up 70% to 80% of the total membrane lipids of a vascular plant. They are probably the most abundant membrane lipids in the biosphere. Phosphate is often the limiting plant nutrient in soil, and perhaps the evolutionary pressure to conserve phosphate for more critical roles favored plants that made phosphate free lipids. Plant membranes also contain sulfolipids, in which a sulfonated glucose residue is joined to a diacylglycerol in glycosidic linkage. In sulfolipids, the sulfonate on the head group bears a negative charge like that of the phosphate group in phospholipids.
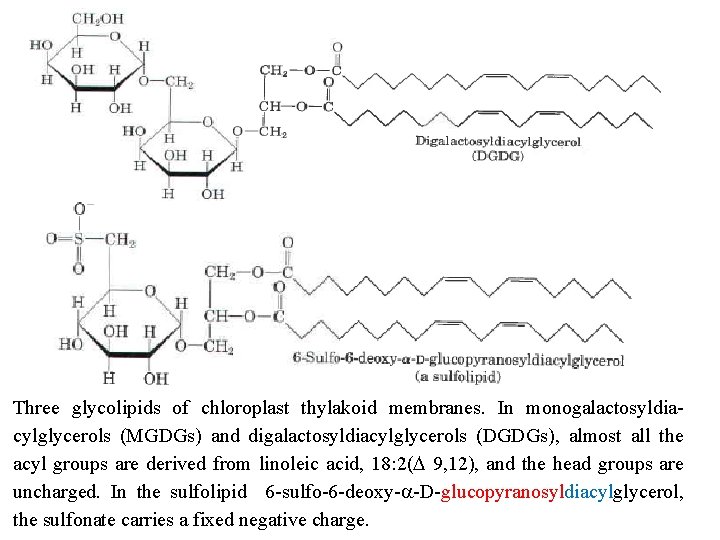
Three glycolipids of chloroplast thylakoid membranes. In monogalactosyldia cylglycerols (MGDGs) and digalactosyldiacylglycerols (DGDGs), almost all the acyl groups are derived from linoleic acid, 18: 2(D 9, 12), and the head groups are uncharged. In the sulfolipid 6 sulfo 6 deoxy a D glucopyranosyldiacylglycerol, the sulfonate carries a fixed negative charge.

Sphingolipids, the other large class of membrane lipids, also have a polar head group and two nonpolar tails, but unlike glycerophospholipids and galactolipids they contain no glycerol. Sphingolipids are composed of one molecule of the long chain amino alcohol sphingosine (also called 4 sphingenine) or one of its derivatives, one molecule of a long chain fatty acid, and a polar head group that is joined by a glycosidic linkage or with a phosphodiester Sphingosine H Ceramide Carbons C 1, C 2, and C 3 of the sphingosine molecule are structurally analogous to the three carbons of glycerol in glycerophospholipids. When a fatty acid is attached in amide linkage to the NH 2 on C 2, the resulting compound is a ceramide, which is structurally similar to a diacylglycerol. Ceramide is the structural parent of all sphingolipids
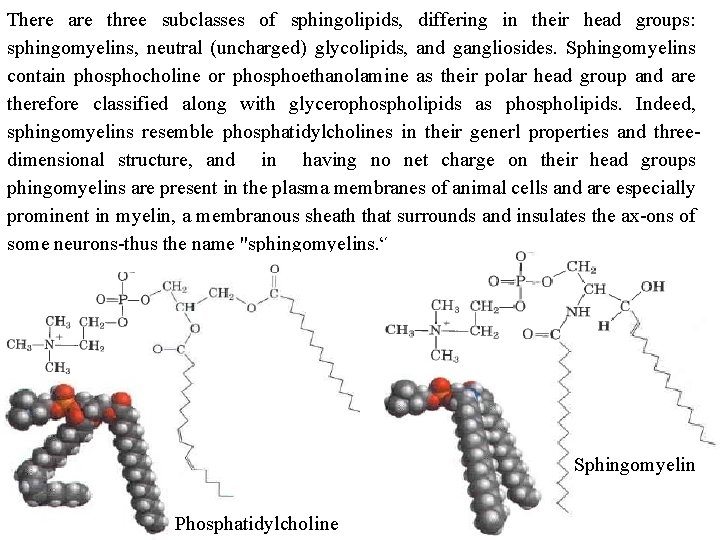
There are three subclasses of sphingolipids, differing in their head groups: sphingomyelins, neutral (uncharged) glycolipids, and gangliosides. Sphingomyelins contain phosphocholine or phosphoethanolamine as their polar head group and are therefore classified along with glycerophospholipids as phospholipids. Indeed, sphingomyelins resemble phosphatidylcholines in their generl properties and three dimensional structure, and in having no net charge on their head groups phingomyelins are present in the plasma membranes of animal cells and are especially prominent in myelin, a membranous sheath that surrounds and insulates the ax ons of some neurons thus the name "sphingomyelins. “ Sphingomyelin Phosphatidylcholine
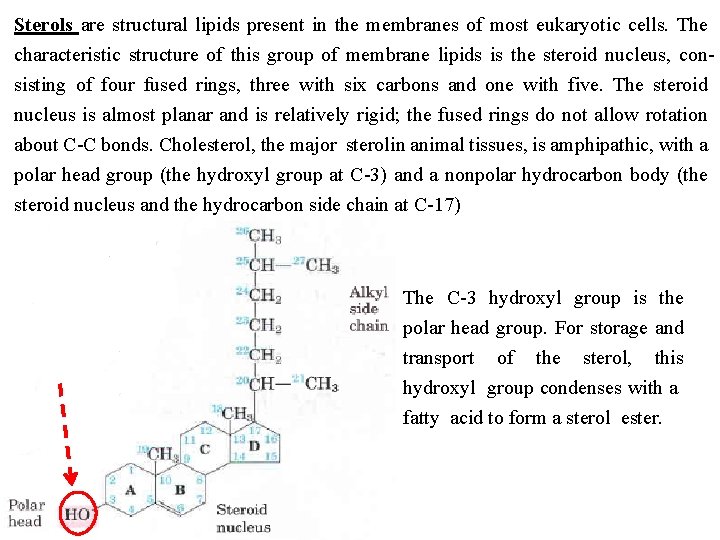
Sterols are structural lipids present in the membranes of most eukaryotic cells. The characteristic structure of this group of membrane lipids is the steroid nucleus, con sisting of four fused rings, three with six carbons and one with five. The steroid nucleus is almost planar and is relatively rigid; the fused rings do not allow rotation about C C bonds. Cholesterol, the major sterolin animal tissues, is amphipathic, with a polar head group (the hydroxyl group at C 3) and a nonpolar hydrocarbon body (the steroid nucleus and the hydrocarbon side chain at C 17) The C 3 hydroxyl group is the polar head group. For storage and transport of the sterol, this hydroxyl group condenses with a fatty acid to form a sterol ester.
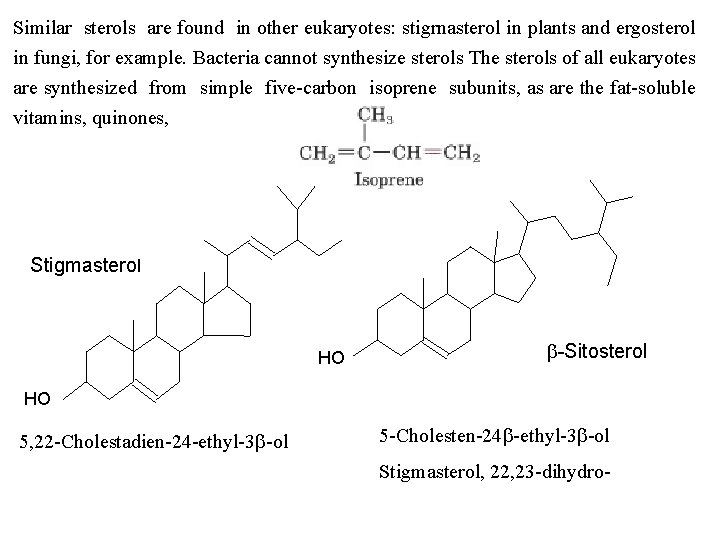
Similar sterols are found in other eukaryotes: stigrnasterol in plants and ergosterol in fungi, for example. Bacteria cannot synthesize sterols The sterols of all eukaryotes are synthesized from simple five carbon isoprene subunits, as are the fat soluble vitamins, quinones, Stigmasterol HO b-Sitosterol HO 5, 22 Cholestadien 24 ethyl 3 b ol 5 Cholesten 24 b ethyl 3 b ol Stigmasterol, 22, 23 dihydro
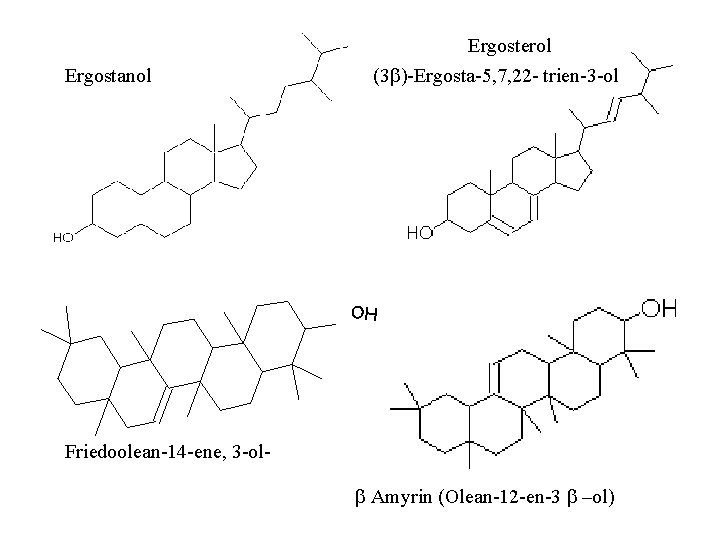
Ergostanol Ergosterol (3 b) Ergosta 5, 7, 22 trien 3 ol OH Friedoolean 14 ene, 3 ol b Amyrin (Olean 12 en 3 b –ol)

Lipids as Signals, Cofactors, and Pigments: storage lipids and structural lipids are major cellular components; membrane lipids make up 5% to 10% of the dry mass of most cells, and storage lipids more than 80% of the mass of an germinating seed. With some important exceptions, these lipids play a passive role in the cell; lipid fuels are stored until oxidized by enzymes, and membrane lipids form impermeable barriers around cells and cellular compartments. Another group of lipids, present in much smaller amounts, have active roles in the metabolic traffic as metabolites and messengers. Some serve as potent signals as hormones, or as intracellular messengers generated in response to an extracellular signal. Others function as enzyme cofactors in electron transfer reactions in chloroplasts and mitochondria, or in the transfer of sugar moieties in a variety of glycosylation reactions. A third group consists of lipids with a system of conjugated double bonds: pigment molecules that absorb visible light. Some of these act as light capturing pigments in vision and photosynthesis; others produce natural colorations, such as the orange of pumpkins and carrots and the yellow of canary feathers. Finally, a very large group of volatile lipids produced in plants serve as signals that pass through the air, allowing plants to communicate with each other, and to invite or repel insects and animal
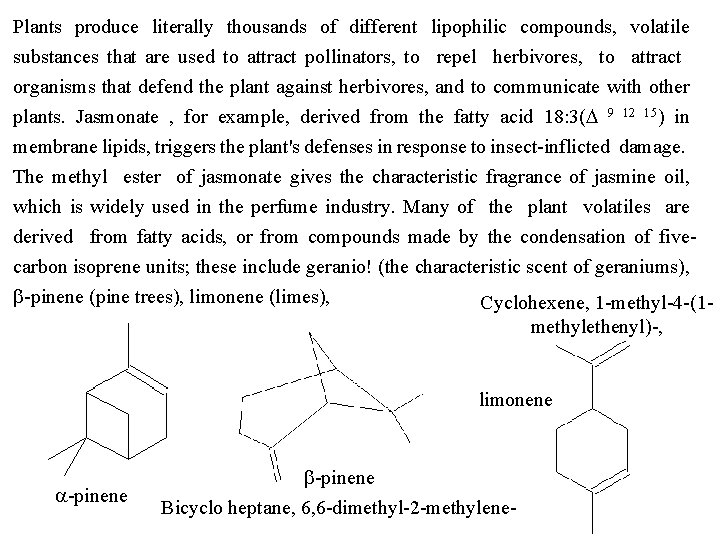
Plants produce literally thousands of different lipophilic compounds, volatile substances that are used to attract pollinators, to repel herbivores, to attract organisms that defend the plant against herbivores, and to communicate with other plants. Jasmonate , for example, derived from the fatty acid 18: 3(D 9 12 15) in membrane lipids, triggers the plant's defenses in response to insect inflicted damage. The methyl ester of jasmonate gives the characteristic fragrance of jasmine oil, which is widely used in the perfume industry. Many of the plant volatiles are derived from fatty acids, or from compounds made by the condensation of five carbon isoprene units; these include geranio! (the characteristic scent of geraniums), b pinene (pine trees), limonene (limes), Cyclohexene, 1 methyl 4 (1 methylethenyl) , limonene a pinene b pinene Bicyclo heptane, 6, 6 dimethyl 2 methylene
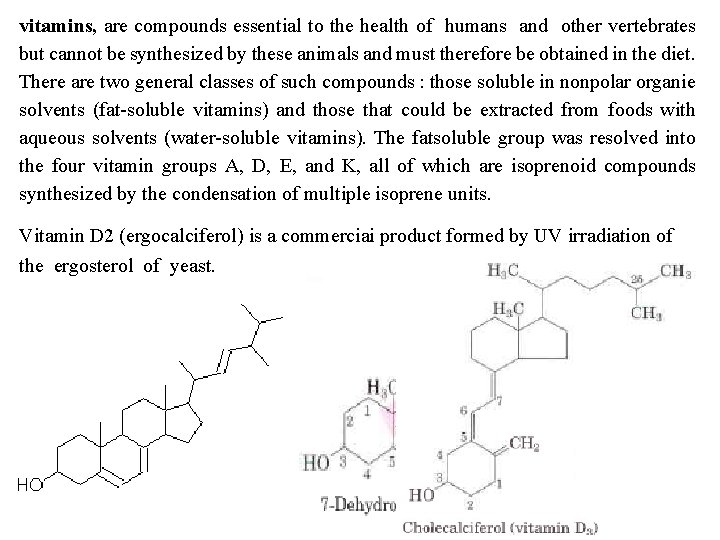
vitamins, are compounds essential to the health of humans and other vertebrates but cannot be synthesized by these animals and must therefore be obtained in the diet. There are two general classes of such compounds : those soluble in nonpolar organie solvents (fat soluble vitamins) and those that could be extracted from foods with aqueous solvents (water soluble vitamins). The fatsoluble group was resolved into the four vitamin groups A, D, E, and K, all of which are isoprenoid compounds synthesized by the condensation of multiple isoprene units. Vitamin D 2 (ergocalciferol) is a commerciai product formed by UV irradiation of the ergosterol of yeast.
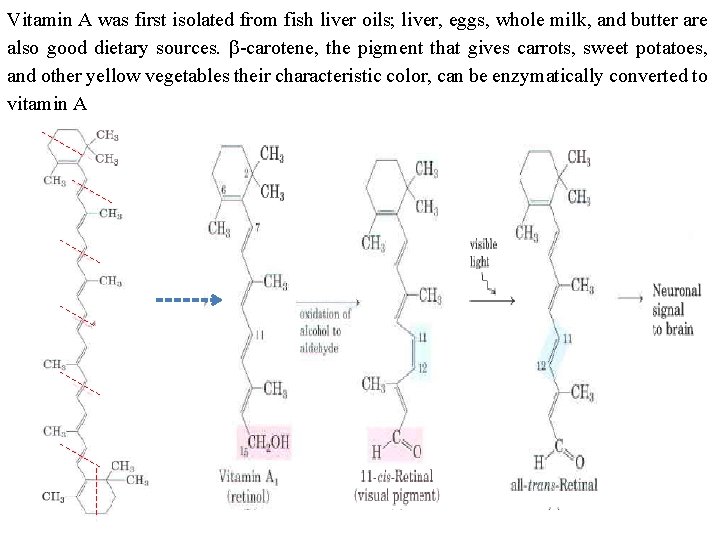
Vitamin A was first isolated from fish liver oils; liver, eggs, whole milk, and butter are also good dietary sources. b carotene, the pigment that gives carrots, sweet potatoes, and other yellow vegetables their characteristic color, can be enzymatically converted to vitamin A

Vitamins E and K and the Lipid Quinones Are Oxidation Reduction Cofactors: Vitamin E is the collective name for a group of closely related lipids called tocopherols, all of which contain a substituted aromatic ring and a long isoprenoid side chain. Because they are hydrophobic, tocopherols associate with cell membranes, Tocopherols are biological antioxidants. The aromatic ring reacts with and destroys the most reactive forms of oxygen radicals and other free radicals, protecting unsaturated fatty acids from oxidation and preventing oxidative damage to membrane lipids, which can cause cell fragility. Tocopherols are found in eggs and vegetable oils and are especially abundant in wheat germ.
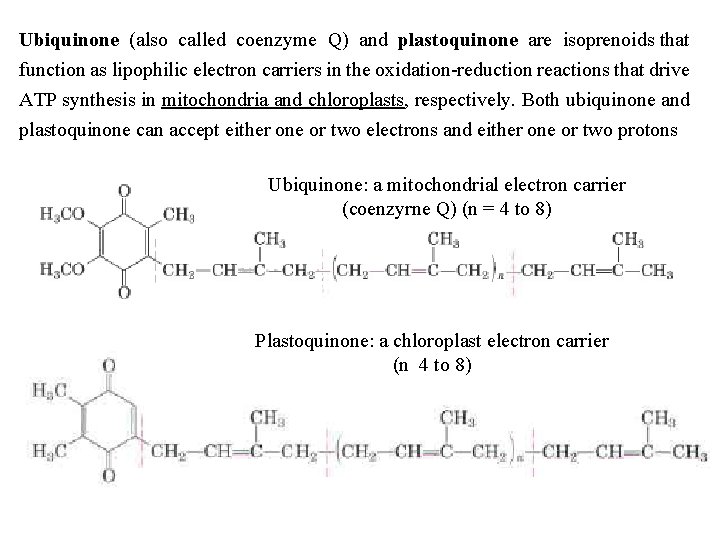
Ubiquinone (also called coenzyme Q) and plastoquinone are isoprenoids that function as lipophilic electron carriers in the oxidation reduction reactions that drive ATP synthesis in mitochondria and chloroplasts, respectively. Both ubiquinone and plastoquinone can accept either one or two electrons and either one or two protons Ubiquinone: a mitochondrial electron carrier (coenzyrne Q) (n = 4 to 8) Plastoquinone: a chloroplast electron carrier (n 4 to 8)
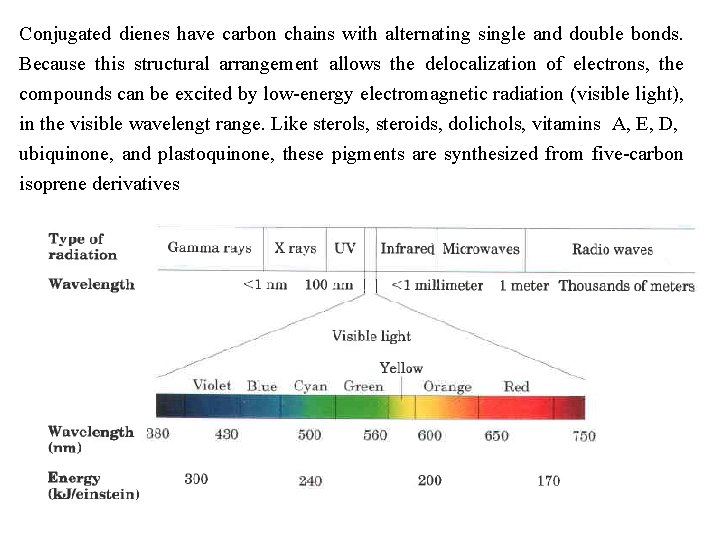
Conjugated dienes have carbon chains with alternating single and double bonds. Because this structural arrangement allows the delocalization of electrons, the compounds can be excited by low energy electromagnetic radiation (visible light), in the visible wavelengt range. Like sterols, steroids, dolichols, vitamins A, E, D, ubiquinone, and plastoquinone, these pigments are synthesized from five carbon isoprene derivatives
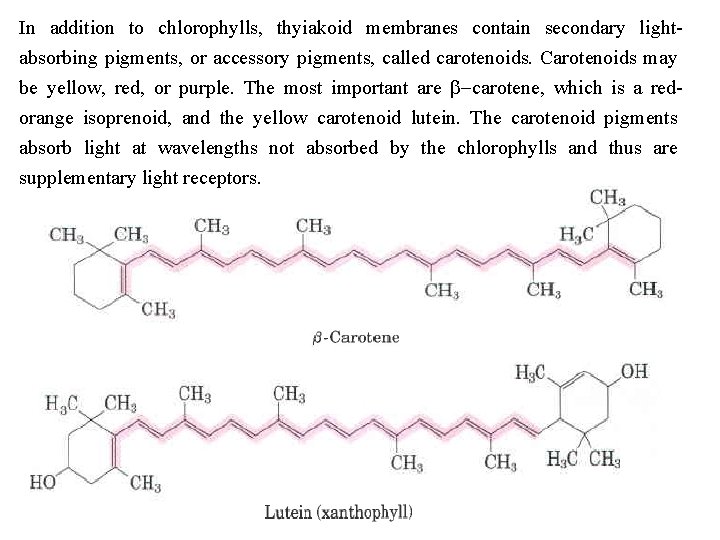
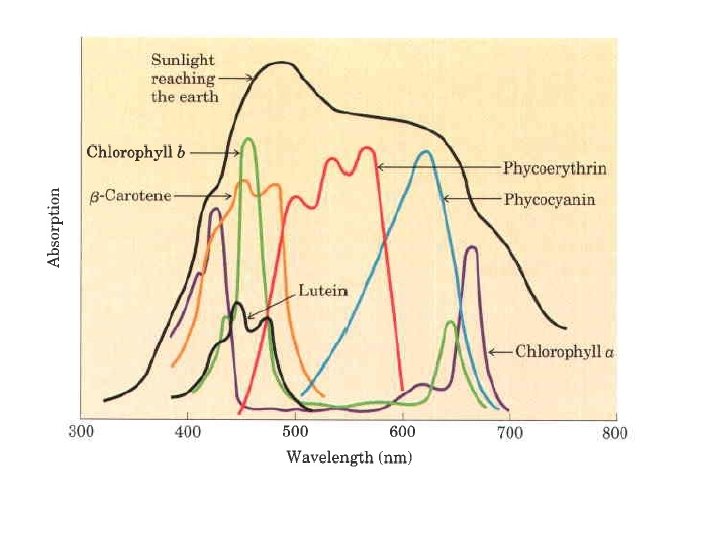
In addition to chlorophylls, thyiakoid membranes contain secondary light absorbing pigments, or accessory pigments, called carotenoids. Carotenoids may be yellow, red, or purple. The most important are b-carotene, which is a red orange isoprenoid, and the yellow carotenoid lutein. The carotenoid pigments absorb light at wavelengths not absorbed by the chlorophylls and thus are supplementary light receptors.

Membranes define the external boundaries of cells and regulate the molecular traffic across that boundary; in eukaryotic cells, they divide the internal space into discrete compartments to segregate processes and components. They organize complex reaction sequences and are central to both biological energy conservation and cell to cell communication. The biological activities of membranes flow from their remarkable physical properties. Membranes are flexible, self sealing allowing the shape changes that accompany cell growth; are selectively permeable to polar solutes, they retain certain compounds and ions within cells and within specific cellular compartments, while excluding others. Membranes are not merely passive barriers. They include an array of proteins specialized for promoting or catalyzing various cellular processes. At the cell surface, transporters move specific organic solutes and inorganic ions across the membrane. Within the cell membranes organize cellular processes such as the synthesis of lipids and certain proteins, and the energy transductions in mitochondria and chloroplasts
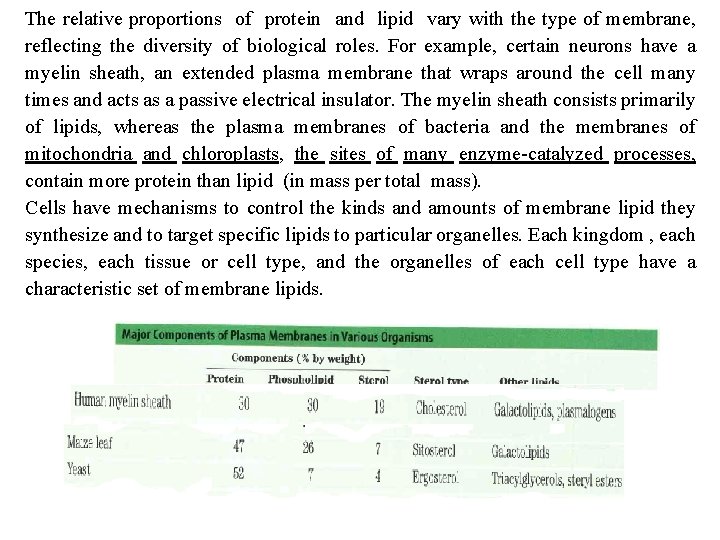
The relative proportions of protein and lipid vary with the type of membrane, reflecting the diversity of biological roles. For example, certain neurons have a myelin sheath, an extended plasma membrane that wraps around the cell many times and acts as a passive electrical insulator. The myelin sheath consists primarily of lipids, whereas the plasma membranes of bacteria and the membranes of mitochondria and chloroplasts, the sites of many enzyme catalyzed processes, contain more protein than lipid (in mass per total mass). Cells have mechanisms to control the kinds and amounts of membrane lipid they synthesize and to target specific lipids to particular organelles. Each kingdom , each species, each tissue or cell type, and the organelles of each cell type have a characteristic set of membrane lipids.

Membranes are impermeable to most polar or charged solutes, but permeable to nonpolar compounds; they are 3 to 5 nm (30 to 50 Å) thick and appear trilaminar when viewed in cross section with the electron microscope. The combined evidence from electron microscopy and studies of' chemical composition, as well as physical studies of permeability and the motion of individual protein and lipid molecules within membranes, led to the development of the fluid mosaic model for the structure of biological membranes. Phospholipids form a bilayer in which the nonpolar regions of the lipid molecules in each layer face the core of the bilayer and their polar head groups face outward, interacting with the aqueous phase on either side. Proteins are embedded in this bilayer sheet, held by hydrophobic interactions between the membrane lipids and hydrophobic domains in the proteins. Some proteins protrude from only one side of the membrane; others have domains exposed on both sides: the protein domains exposed on one side of the bilayer are different from those exposed on the other side, reflecting functional asymmetry. The individual lipid and protein units in a membrane form a fluid mosaic with a pattern is free to change constantly. The membrane mosaic is fluid because most of the interactions among its components are non-covalent, leaving individual lipid and protein molecules free to move laterally in the plane of the membrane.
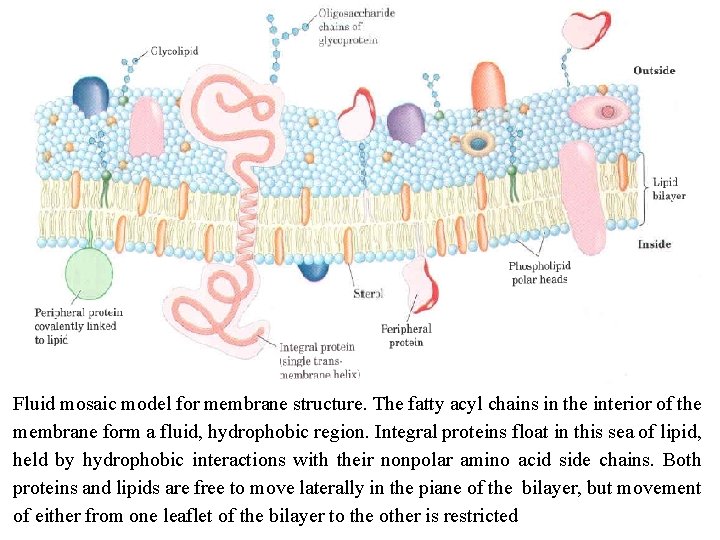
Fluid mosaic model for membrane structure. The fatty acyl chains in the interior of the membrane form a fluid, hydrophobic region. Integral proteins float in this sea of lipid, held by hydrophobic interactions with their nonpolar amino acid side chains. Both proteins and lipids are free to move laterally in the piane of the bilayer, but movement of either from one leaflet of the bilayer to the other is restricted

Glycerophospholipids, sphingolipids, and sterols are virtually insoluble in water. When mixed with water, they spontaneously form microscopic lipid aggregates, clustering together, with their hydrophobic moieties in contact with each other and their hydrophilic groups interacting with the surrounding water. This clustering reduces the amount of hydrophobic surface exposed to water and thus minimizes the number of molecules in the shell of ordered water at the lipid water interface, resulting in an increase in entropy. Hydrophobic interactions among lipid molecules provide thermodynamic driving force for the formation and maintenance of these clusters.
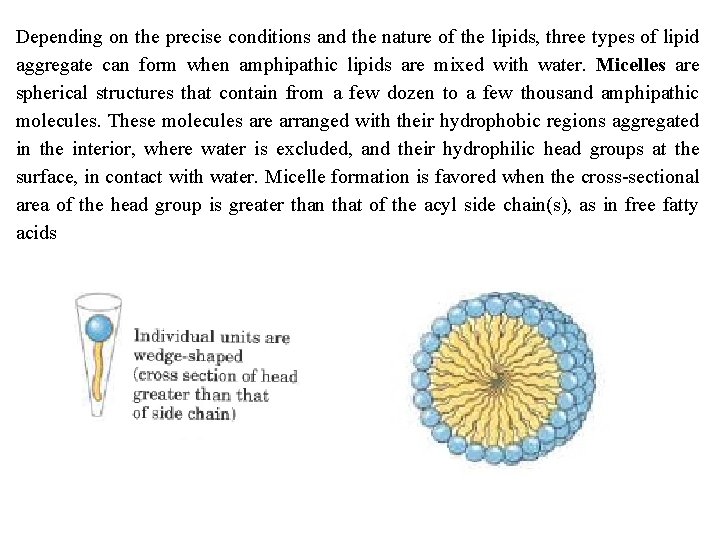
Depending on the precise conditions and the nature of the lipids, three types of lipid aggregate can form when amphipathic lipids are mixed with water. Micelles are spherical structures that contain from a few dozen to a few thousand amphipathic molecules. These molecules are arranged with their hydrophobic regions aggregated in the interior, where water is excluded, and their hydrophilic head groups at the surface, in contact with water. Micelle formation is favored when the cross sectional area of the head group is greater than that of the acyl side chain(s), as in free fatty acids
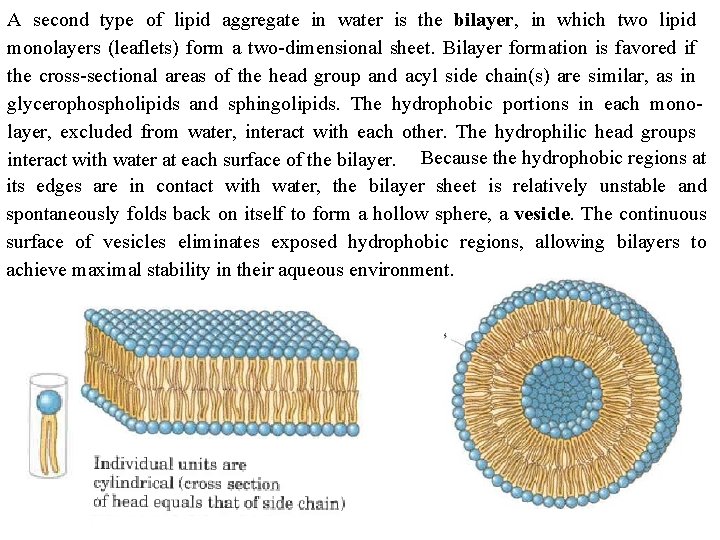
A second type of lipid aggregate in water is the bilayer, in which two lipid monolayers (leaflets) form a two dimensional sheet. Bilayer formation is favored if the cross sectional areas of the head group and acyl side chain(s) are similar, as in glycerophospholipids and sphingolipids. The hydrophobic portions in each mono layer, excluded from water, interact with each other. The hydrophilic head groups interact with water at each surface of the bilayer. Because the hydrophobic regions at its edges are in contact with water, the bilayer sheet is relatively unstable and spontaneously folds back on itself to form a hollow sphere, a vesicle. The continuous surface of vesicles eliminates exposed hydrophobic regions, allowing bilayers to achieve maximal stability in their aqueous environment.
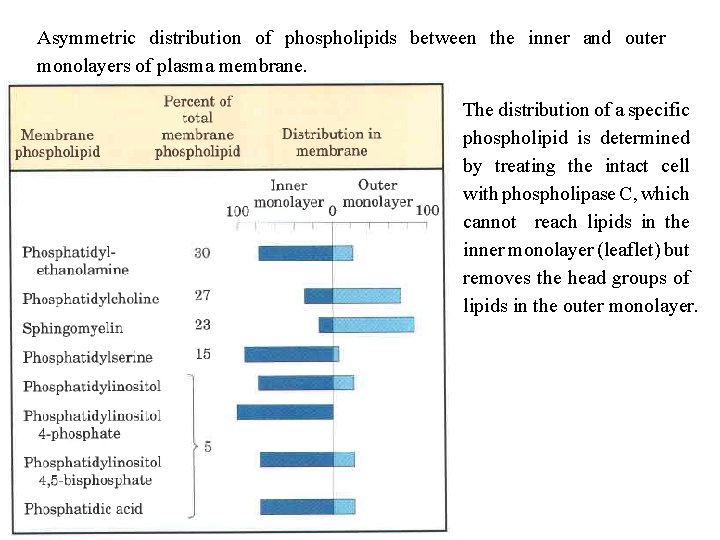
Asymmetric distribution of phospholipids between the inner and outer monolayers of plasma membrane. The distribution of a specific phospholipid is determined by treating the intact cell with phospholipase C, which cannot reach lipids in the inner monolayer (leaflet) but removes the head groups of lipids in the outer monolayer.

Three Types of Membrane Proteins Differ in Their Association with the Membrane Integral membrane proteins are very firmly associated with the lipid bilayer, and are removable only by agents that interfere with hydrophobic interactions, such as detergents, organic solvents, or denaturants. Peripheral membrane proteins associate with the membrane through electrostatic interactions and hydrogen bonding with the hydrophilic domains of integral proteins and with the polar head groups of membrane lipids. They can be released by relatively mild treatments that interfere with electrostatic interactions or break hydrogen bonds. Amphitropic proteins are found both in the cytosol and in association with membranes. Their affinity for membranes results in some cases from the protein's noncovalent interaction with a membrane protein or lipid, and in other cases from the presence of one or more lipids covalently attached to the amphitropic protein. Generally, the reversible association of amphitropic proteins with the membrane is regulated; for example, phosphory 1 ation or ligand binding can farce a conformational change in the protein, exposing a membrane binding site that was previously inaccessible.

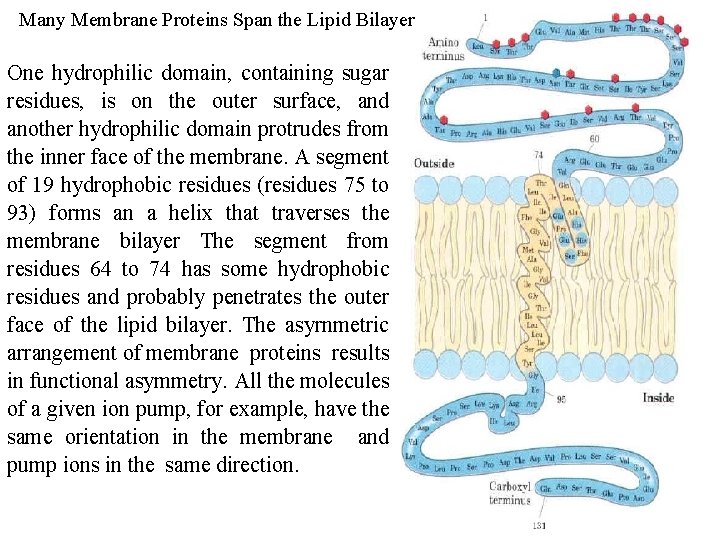
Many Membrane Proteins Span the Lipid Bilayer One hydrophilic domain, containing sugar residues, is on the outer surface, and another hydrophilic domain protrudes from the inner face of the membrane. A segment of 19 hydrophobic residues (residues 75 to 93) forms an a helix that traverses the membrane bilayer The segment from residues 64 to 74 has some hydrophobic residues and probably penetrates the outer face of the lipid bilayer. The asyrnmetric arrangement of membrane proteins results in functional asymmetry. All the molecules of a given ion pump, for example, have the same orientation in the membrane and pump ions in the same direction.
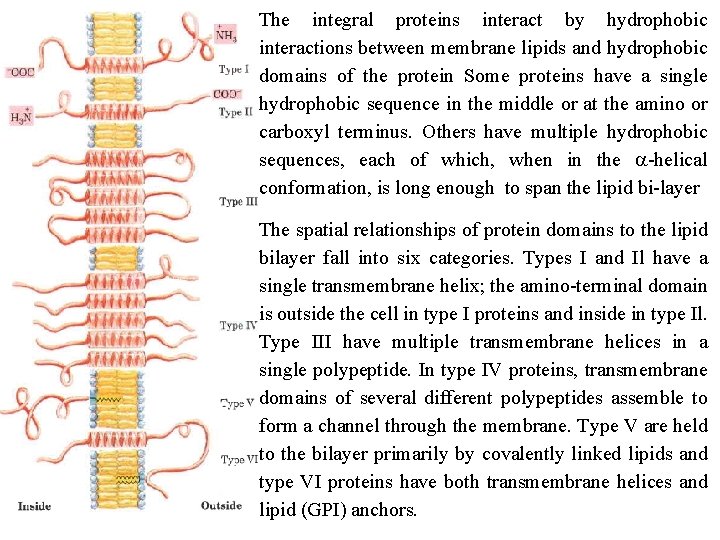
The integral proteins interact by hydrophobic interactions between membrane lipids and hydrophobic domains of the protein Some proteins have a single hydrophobic sequence in the middle or at the amino or carboxyl terminus. Others have multiple hydrophobic sequences, each of which, when in the a helical conformation, is long enough to span the lipid bi layer The spatial relationships of protein domains to the lipid bilayer fall into six categories. Types I and Il have a single transmembrane helix; the amino terminal domain is outside the cell in type I proteins and inside in type Il. Type III have multiple transmembrane helices in a single polypeptide. In type IV proteins, transmembrane domains of several different polypeptides assemble to form a channel through the membrane. Type V are held to the bilayer primarily by covalently linked lipids and type VI proteins have both transmembrane helices and lipid (GPI) anchors.
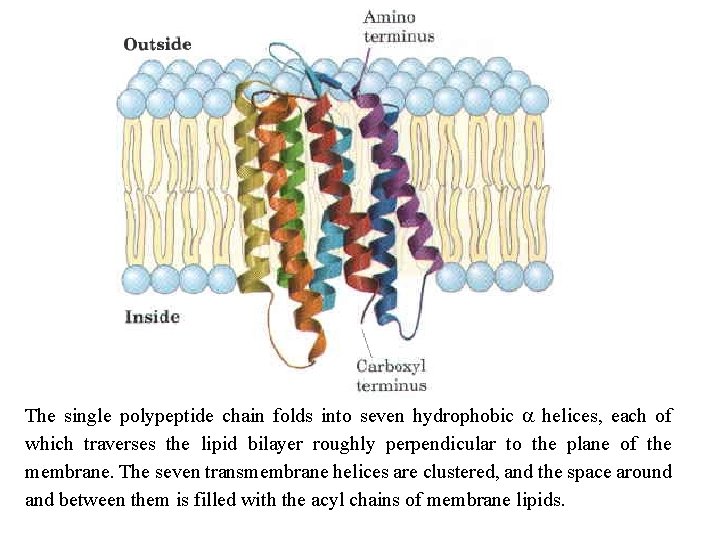
The single polypeptide chain folds into seven hydrophobic a helices, each of which traverses the lipid bilayer roughly perpendicular to the plane of the membrane. The seven transmembrane helices are clustered, and the space around and between them is filled with the acyl chains of membrane lipids.

Crystallized membrane proteins solved by crystallography often include molecules of phospholipids. Many of these phospholipid molecules lie on the protein surface, their head groups interacting with polar amino acid residues at the inner and outer membrane water interfaces and their side chains associated with non polar residues. These annular lipids form a bilayer shell (annulus) around the protein, oriented roughly as expected for phospholipids in a bi layer. Lipid annul associated with integral membrane proteins. The crystal structure of aquaporin, a transmembrane water channel, includes a shell of phospholipids positioned with their head groups (blue) at the expected positions on the inner and outer membrane surfaces and their hydrophobic acyl chains (gold) intimately associateci with the surface of the protein exposed to the bilayer. The lipid forms a "grease seal" around the protein, which is depicted as a green surface representation
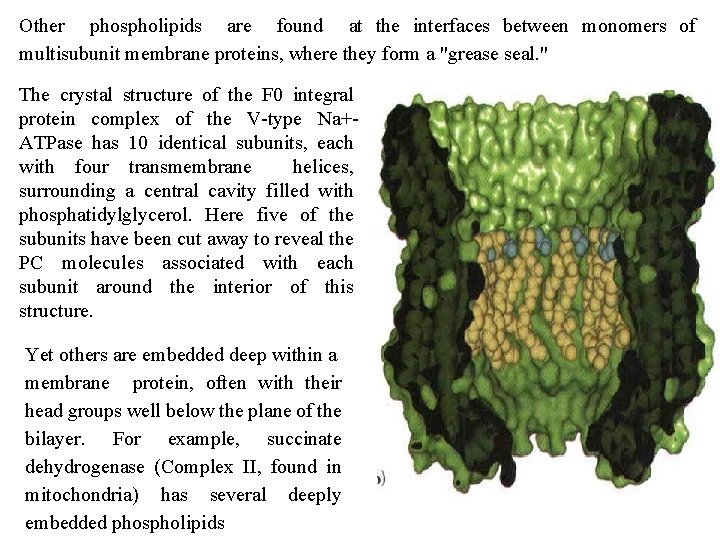
Other phospholipids are found at the interfaces between monomers of multisubunit membrane proteins, where they form a "grease seal. " The crystal structure of the F 0 integral protein complex of the V type Na+ ATPase has 10 identical subunits, each with four transmembrane helices, surrounding a central cavity filled with phosphatidylglycerol. Here five of the subunits have been cut away to reveal the PC molecules associated with each subunit around the interior of this structure. Yet others are embedded deep within a membrane protein, often with their head groups well below the plane of the bilayer. For example, succinate dehydrogenase (Complex II, found in mitochondria) has several deeply embedded phospholipids
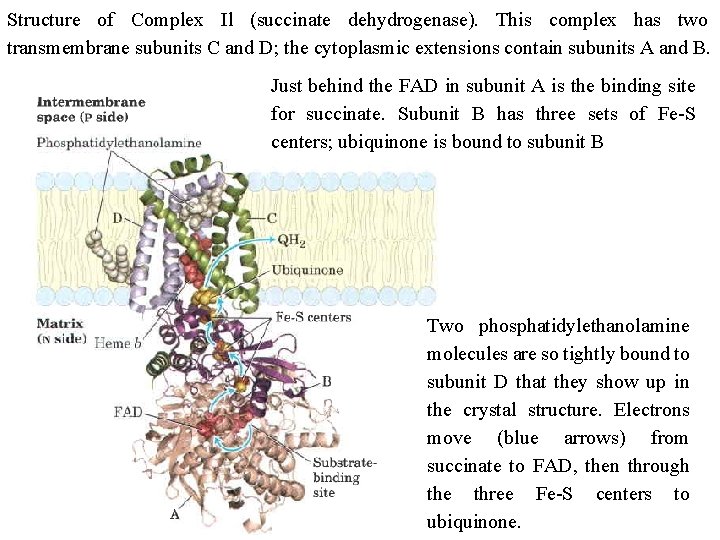
Structure of Complex Il (succinate dehydrogenase). This complex has two transmembrane subunits C and D; the cytoplasmic extensions contain subunits A and B. Just behind the FAD in subunit A is the binding site for succinate. Subunit B has three sets of Fe S centers; ubiquinone is bound to subunit B Two phosphatidylethanolamine molecules are so tightly bound to subunit D that they show up in the crystal structure. Electrons move (blue arrows) from succinate to FAD, then through the three Fe S centers to ubiquinone.

Determination of the three dimensional structure of a membrane protein that is, its topology, is generally much more difficult than determining its amino acid sequence. The amino acid sequences are known for thousands of membrane proteins, but relatively few three dimensional structures have been established by crystallography or NMR spectroscopy. The presence of unbroken sequences of more than 20 hydrophobic residues in a membrane protein is commonly taken as evidence that these sequences traverse the lipid bilayer, acting as hydrophobic anchors or forming transmembrane channels. Virtually all integral proteins have at least one such sequence. An a helical sequence of 20 to 25 residues is just long enough to span the thickness (30 Å) of the lipid bilayer (recall that the length of an a helix is 1. 5 Å 0. 15 nm per amino acid residue). A polypeptide chain surrounded by lipids, having no water molecules with which to hydrogen bond, will tend to form a helices or b sheets, in which intra chain hydrogen bonding is maximized. If the side chains of all amino acids in a helix are non polar, hydrophobic interactions with the surrounding lipids further stabilize the helix.

Several simple methods of analyzing amino acid sequences yield reasonably accurate predictions of secondary structure for transmembrane proteins. The relative polarity of each amino acid has been determined experimentally by measuring the free energy change accompanying the movement of that amino acid side chain from a hydrophobic solvent into water This DG of transfer, which can be expressed as a hydropathy index, ranges from very exergonic for charged or polar residues to very endergonic for aromatic or aliphatic hydrocarbon side chains
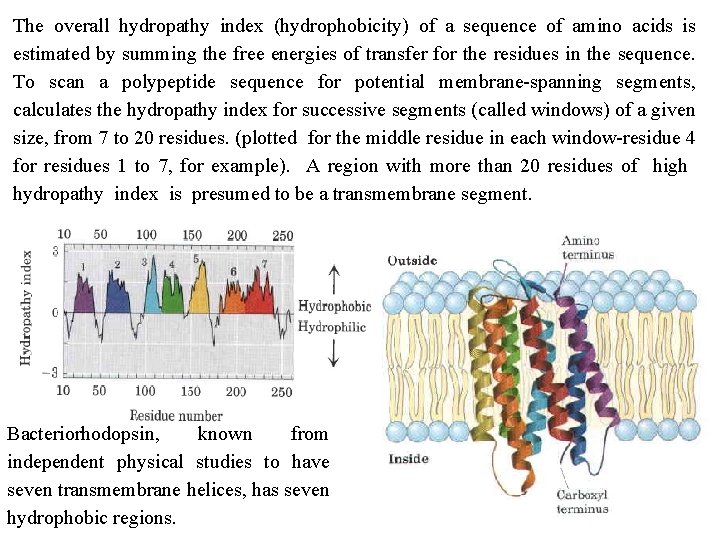
The overall hydropathy index (hydrophobicity) of a sequence of amino acids is estimated by summing the free energies of transfer for the residues in the sequence. To scan a polypeptide sequence for potential membrane spanning segments, calculates the hydropathy index for successive segments (called windows) of a given size, from 7 to 20 residues. (plotted for the middle residue in each window residue 4 for residues 1 to 7, for example). A region with more than 20 residues of high hydropathy index is presumed to be a transmembrane segment. Bacteriorhodopsin, known from independent physical studies to have seven transmembrane helices, has seven hydrophobic regions.
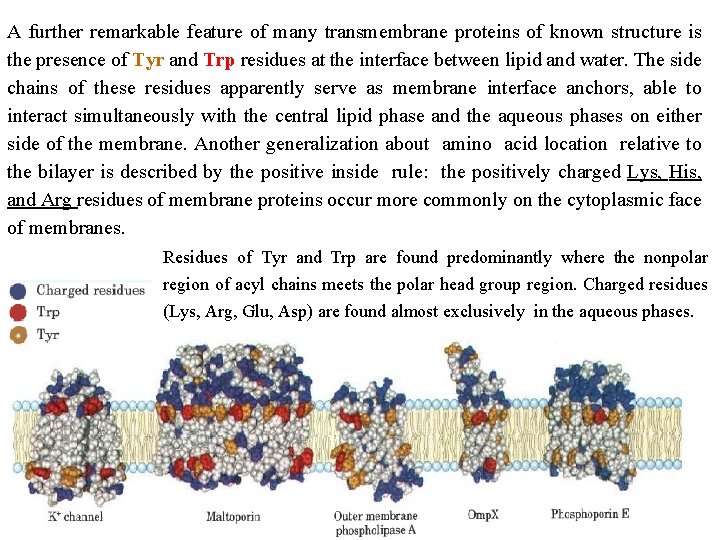
A further remarkable feature of many transmembrane proteins of known structure is the presence of Tyr and Trp residues at the interface between lipid and water. The side chains of these residues apparently serve as membrane interface anchors, able to interact simultaneously with the central lipid phase and the aqueous phases on either side of the membrane. Another generalization about amino acid location relative to the bilayer is described by the positive inside rule: the positively charged Lys, His, and Arg residues of membrane proteins occur more commonly on the cytoplasmic face of membranes. Residues of Tyr and Trp are found predominantly where the nonpolar region of acyl chains meets the polar head group region. Charged residues (Lys, Arg, Glu, Asp) are found almost exclusively in the aqueous phases.
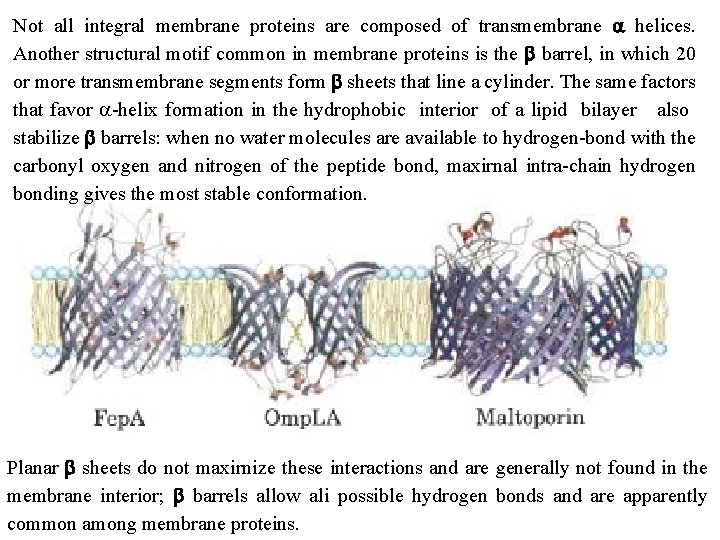
Not all integral membrane proteins are composed of transmembrane a helices. Another structural motif common in membrane proteins is the b barrel, in which 20 or more transmembrane segments form b sheets that line a cylinder. The same factors that favor a helix formation in the hydrophobic interior of a lipid bilayer also stabilize b barrels: when no water molecules are available to hydrogen bond with the carbonyl oxygen and nitrogen of the peptide bond, maxirnal intra chain hydrogen bonding gives the most stable conformation. Planar b sheets do not maxirnize these interactions and are generally not found in the membrane interior; b barrels allow ali possible hydrogen bonds and are apparently common among membrane proteins.

Some membrane proteins contain one or more covalently linked lipids, which may be of several types: long chain fatty acids, isoprenoids, sterols, or glycosylated derivatives of phosphatidylinositol (GPIs). The attached lipid provides a hydrophobic anchor that inserts into the lipid bilayer and holds the protein at the membrane surface. The strength of the hydrophobic interaction between a bilayer and a single hydrocarbon chain linked to a protein is barely enough to anchor the protein securely, but many proteins have more than one attached lipid moiety. Other interactions, such as ionic attractions between positively charged Lys residues in the protein and negatively charged lipid head groups, probably contribute to the stability of the attachment. The association of these lipid linked proteins with the membrane is certainly weaker than that for integral membrane proteins and is, in at least some cases, reversible. But treatment with alkaline carbonate does not release GPI linked proteins, which are therefore, by the working definition, integral proteins.
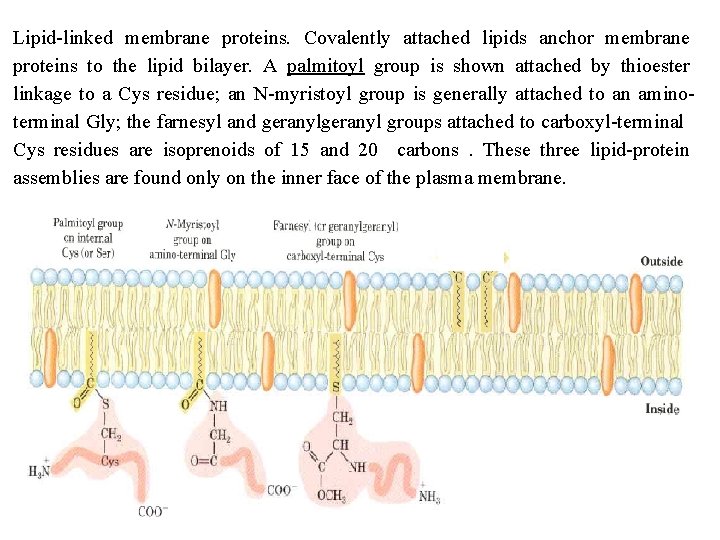
Lipid linked membrane proteins. Covalently attached lipids anchor membrane proteins to the lipid bilayer. A palmitoyl group is shown attached by thioester linkage to a Cys residue; an N myristoyl group is generally attached to an amino terminal Gly; the farnesyl and geranyl groups attached to carboxyl terminal Cys residues are isoprenoids of 15 and 20 carbons. These three lipid protein assemblies are found only on the inner face of the plasma membrane.
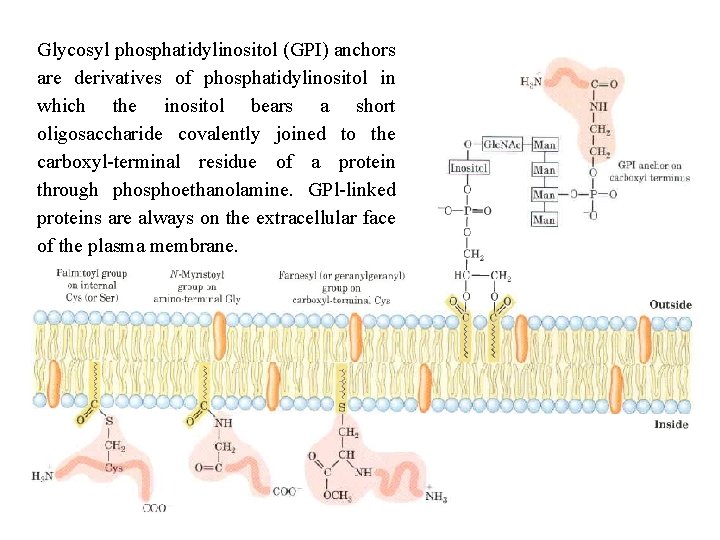
Glycosyl phosphatidylinositol (GPI) anchors are derivatives of phosphatidylinositol in which the inositol bears a short oligosaccharide covalently joined to the carboxyl terminal residue of a protein through phosphoethanolamine. GPl linked proteins are always on the extracellular face of the plasma membrane.

One remarkable feature of all biological membranes is their flexibility, their ability to change shape without losing their integrity and becoming leaky. The basis for this property is the non covalent interactions among lipids in the bilayer and the mobìlity allowed to ìndìvidual lìpìds because they are not covalently anchored to one another. Although the lipid bilayer structure is quite stable, its individual phospholipid and sterol molecules have much freedom of motion. The structure and flexibility of the lipid bilayer depend on the kinds of lipids present, and change with temperature. Below normal physiological temperatures, the lipids in a bilayer form a semisolid gel phase, in which all types of motion of individual lipid molecu 1 es are strong 1 y constrained; the bilayer is paracrystalline polar head groups are uniformly arrayed at the surface, and the acyl chains are nearly motionless and packed with regular geometry.

Above physiological temperatures, individual hydrocarbon chains of fatty acids are in constant motion produced by rotation about the carbon bonds of the long acyl side chains. In this liquid disordered state, or fluid state the interior of the bilayer is more fluid than solid and the bilayer is like a sea of constantly moving lipid. At intermediate (physio 1 ogica 1) temperatures, the lipids exist in a liquid ordered state; there is less thermal motion in the acyl chains of the lipid bilayer, but lateral movement in the p 1 ane of the bilayer still takes place.

At temperatures in the physiological range, long chain saturated fatty acids (such as 16: 0 and 18: 0) pack into a liquid ordered array, but the kinks in unsaturated fatty acids interfere with packing, favoring the liquid disordered state. Shorter chain fatty acy 1 groups have the same effect. The sterol content of a membrane is another important determinant of lipid state. The rigid planar structure of the steroid nucleus, inserted between fatty acyl side chains, reduces the freedom of neighboring acy 1 chains to move by rotation about their carbon bonds, forcing the chains into their fully extended conformation. The presence of sterols therefore reduces the fluidity in the core of the bilayer, thus favoring the liquid ordered phase, and increases the thickness of the lipid leaflet
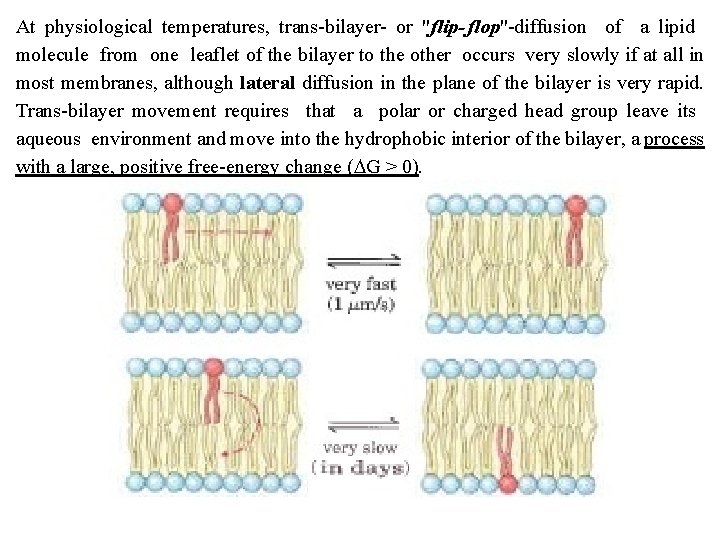
At physiological temperatures, trans bilayer or "flip flop" diffusion of a lipid molecule from one leaflet of the bilayer to the other occurs very slowly if at all in most membranes, although lateral diffusion in the plane of the bilayer is very rapid. Trans bilayer movement requires that a polar or charged head group leave its aqueous environment and move into the hydrophobic interior of the bilayer, a process with a large, positive free energy change (DG > 0).

Several families of proteins, including the flippases, floppases, and scramblases, facilitate the trans bilayer movement of lipids, providing a path that is energetically more favorable and much faster than the uncatalyzed movement. The combination of asymmetric biosynthesis of membrane lipids, very slow uncatalyzed flip flop diffusion, and the presence of selective, energy dependent lipid translocators is responsible for the trans bilayer asymmetry in lipid composition. Flippases catalyze translocation of the aminophospholipids phosphatidylethanolamine and phosphatidylserine from the extracellular to the cytosolic leaflet of the plasma membrane, contributing to the asymmetric distribution of phospholipids: phosphatidylethanolamine and phosphatidylserine primarily in the cytosolic leaflet, and the sphingolipids and phosphatidylcholine in the outer leaflet. Floppases move plasma membrane phospholipids from the cytosolic to the extracellular leaflet, and like flippases are ATP depend ent. Floppases are members of the ABC transporter family, all of which actively transport hydrophobic substrates out ward across the plasma membrane. Scramblases are proteins that move any membrane phospholipid across the bilayer down its concentration gradient (from the leaflet where it has a higher concentration to the leaflet where it has a lower concentration); their activity is not dependent on ATP.
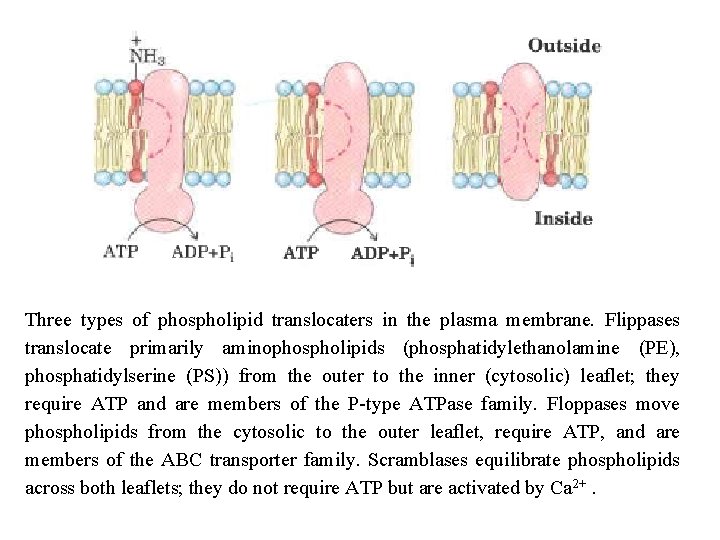
Three types of phospholipid translocaters in the plasma membrane. Flippases translocate primarily aminophospholipids (phosphatidylethanolamine (PE), phosphatidylserine (PS)) from the outer to the inner (cytosolic) leaflet; they require ATP and are members of the P type ATPase family. Floppases move phospholipids from the cytosolic to the outer leaflet, require ATP, and are members of the ABC transporter family. Scramblases equilibrate phospholipids across both leaflets; they do not require ATP but are activated by Ca 2+.
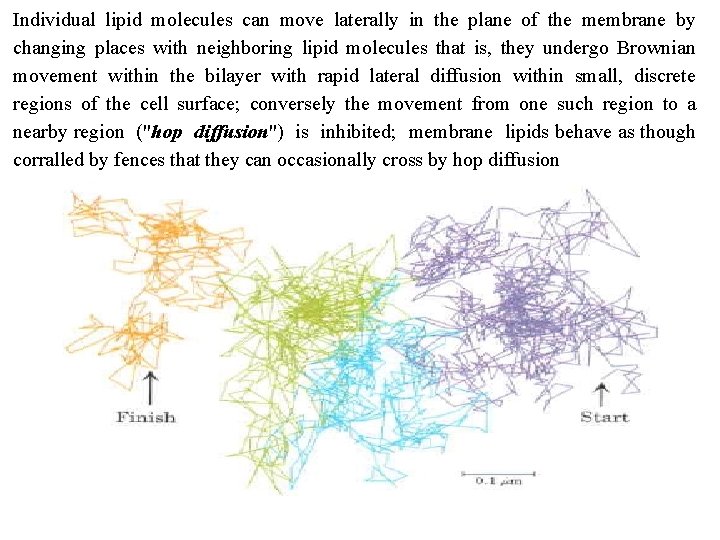
Individual lipid molecules can move laterally in the plane of the membrane by changing places with neighboring lipid molecules that is, they undergo Brownian movement within the bilayer with rapid lateral diffusion within small, discrete regions of the cell surface; conversely the movement from one such region to a nearby region ("hop diffusion") is inhibited; membrane lipids behave as though corralled by fences that they can occasionally cross by hop diffusion


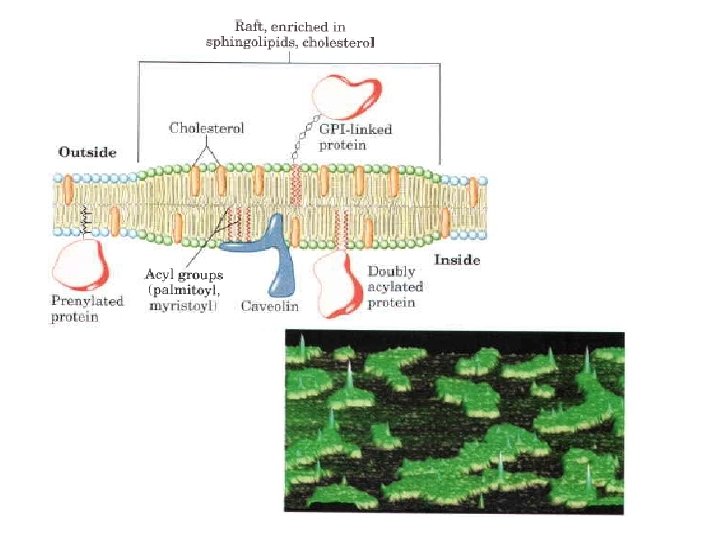
Sphingolipids and Cholesterol Cluster Together in Membrane Rafts Glycosphingolipids, which typically contain longchain saturated fatty acids, form transient clusters in the outer leaflet that largely exclude glycerophospholipids, which typically contain one unsaturated fatty acyl group and a shorter saturated acyl group. The long, saturated acyl groups of sphingolipids can form more compact, more stable associations with the long ring system of cholesterol than can the shorter, often unsaturated, chains of phospholipids. The cholesterol sphingolipid microdomains in the outer monolayer of the plasma membrane, are slightly thicker and more ordered (less fluid) than neighboring microdomains rich in phospholipids and are more difficult to dissolve with nonionic detergents; they behave like liquid ordered sphingolipid rafts adrift on an ocean of liquid disordered phospholipids. These lipid rafts are remarkably enriched in two classes of integral membrane proteins: those anchored to the membrane by two covalently attached long chain saturated fatty acids (two palmitoyl groups or a palmitoyl and a myristoyl group) and GPI anchored proteins. Presumably these lipid anchors, like the acyl chains of sphingolipids, form more stable associations with the cholesterol and long acyl groups in rafts than with the surrounding phospholipids.
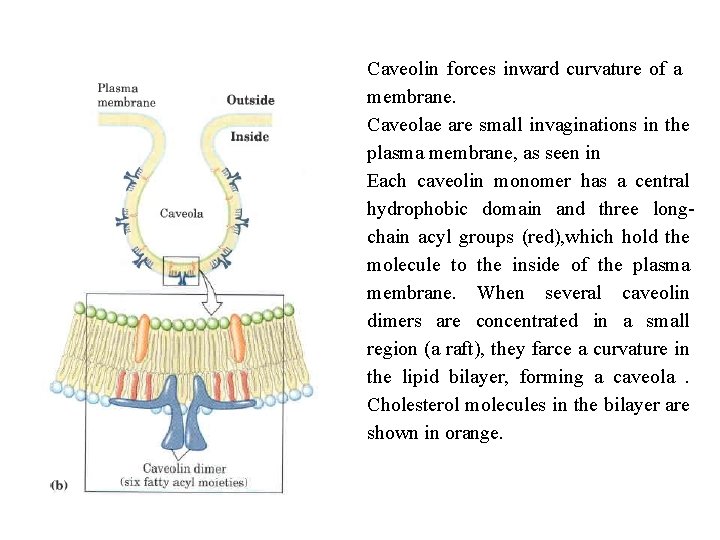
Caveolin forces inward curvature of a membrane. Caveolae are small invaginations in the plasma membrane, as seen in Each caveolin monomer has a central hydrophobic domain and three long chain acyl groups (red), which hold the molecule to the inside of the plasma membrane. When several caveolin dimers are concentrated in a small region (a raft), they farce a curvature in the lipid bilayer, forming a caveola. Cholesterol molecules in the bilayer are shown in orange.

Salute Transport across Membranes Every living cell must acquire from its surroundings the raw materials for biosynthesis and for energy production, and must release to its environment the byproducts of metabolism. A few nonpolar compounds can dissolve in the lipid bilayer and cross the membrane unassisted, but for transmembrane movement of any polar compound or ion, a membrane protein is essential. In some cases a membrane protein simply facilitates the diffusion of a salute down its concentration gradient, but transport can also occur against a gradient of concentration, electrical charge, or both, in which case the process requires energy. The energy may come directly from ATP hydrolysis or may be supplied in the form of one salute moving down its electrochemical gradient, with the release of enough energy to drive another salute up its gradient. Ions may also move across membranes via ion channels formed by proteins, or they may be carried across by ionophores, small molecules that mask the charge of ions and allow them to diffuse through the lipid bilayer. With very few exceptions, the traffic of small molecules across the plasma membrane is mediated by proteins such as transmembrane channels, carriers, or pumps. Within the eukaryotic cell, different compartments have different concentrations of ions and of metabolic intermediates and products, and these, too, must move across intracellular membranes in tightly regulated, protein mediated processes.
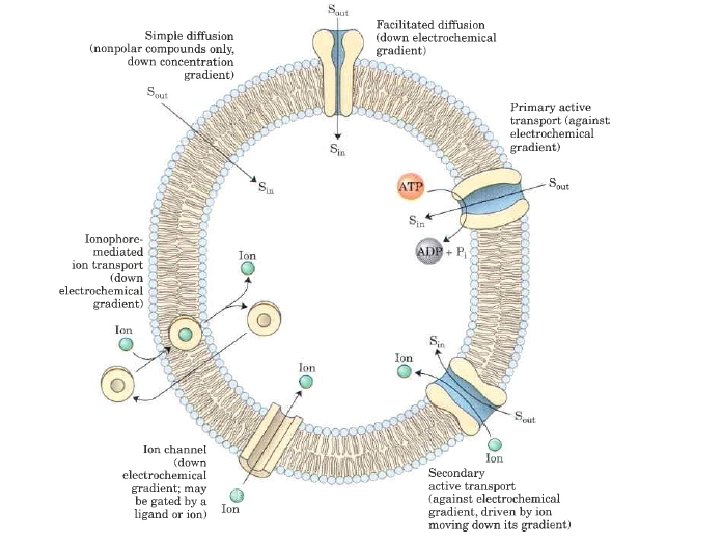
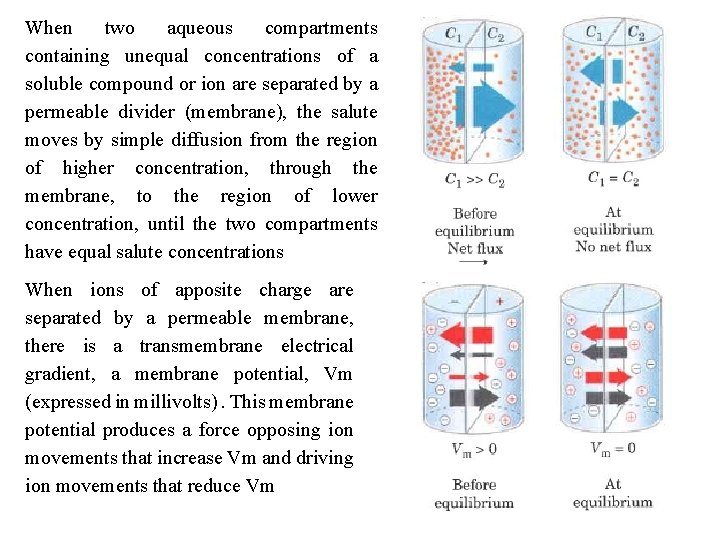
When two aqueous compartments containing unequal concentrations of a soluble compound or ion are separated by a permeable divider (membrane), the salute moves by simple diffusion from the region of higher concentration, through the membrane, to the region of lower concentration, until the two compartments have equal salute concentrations When ions of apposite charge are separated by a permeable membrane, there is a transmembrane electrical gradient, a membrane potential, Vm (expressed in millivolts). This membrane potential produces a force opposing ion movements that increase Vm and driving ion movements that reduce Vm
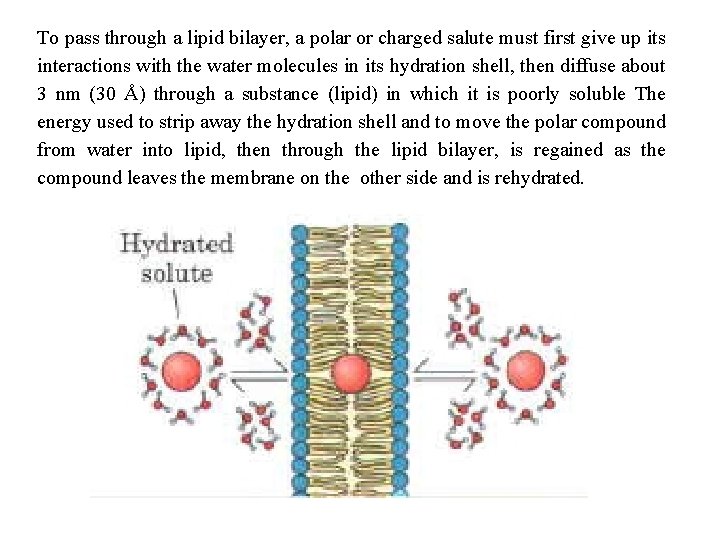
To pass through a lipid bilayer, a polar or charged salute must first give up its interactions with the water molecules in its hydration shell, then diffuse about 3 nm (30 Å) through a substance (lipid) in which it is poorly soluble The energy used to strip away the hydration shell and to move the polar compound from water into lipid, then through the lipid bilayer, is regained as the compound leaves the membrane on the other side and is rehydrated.
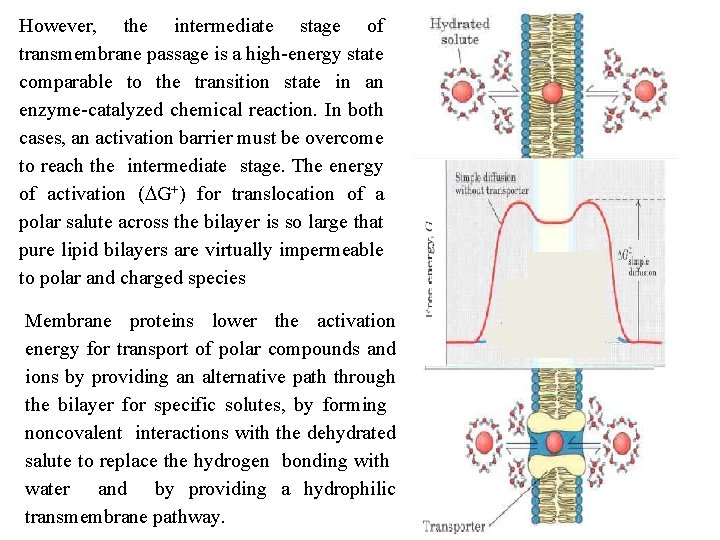
However, the intermediate stage of transmembrane passage is a high energy state comparable to the transition state in an enzyme catalyzed chemical reaction. In both cases, an activation barrier must be overcome to reach the intermediate stage. The energy of activation (DG+) for translocation of a polar salute across the bilayer is so large that pure lipid bilayers are virtually impermeable to polar and charged species Membrane proteins lower the activation energy for transport of polar compounds and ions by providing an alternative path through the bilayer for specific solutes, by forming noncovalent interactions with the dehydrated salute to replace the hydrogen bonding with water and by providing a hydrophilic transmembrane pathway.

Proteins that bring about this facilitated diffusion, or passive transport, are not enzymes in the usual sense; their "substrates" are moved from one compartment to another but are not chemically altered. Membrane proteins that speed the movement of a solute across a membrane by facilitating diffusion are called transporters or permeases. Like enzymes, transporters bind their substrates with stereochemical specificity through multiple weak, non covalent interactions. The negative free energy change associated with these weak interactions, DGbinding, counterbalances the positive free energy change that accompanies loss of the water of hydration from the substrate, DGdehydration thereby lowering activation energy for trans membrane passage. Transporters span the lipid bilayer several times, forming a transmembrane channel lined with hydrophilic amino acid side chains. The channel provides an alternative path for a specific substrate to move across the lipid bilayer without its having to dissolve in the bilayer, further lowering DG+ for transmembrane diffusion. The result is an increase of several to many orders of magnitude in the rate of transmembrane passage of the substrate.

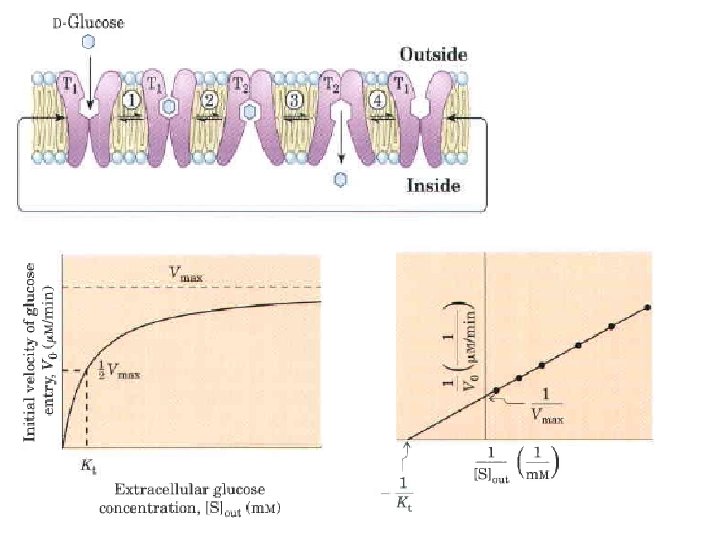
Carriers bind their substrates with high stereospecificity, catalyze transport at rates well below the limits of free diffusion, and are saturable in the same sense as are enzymes: there is some substrate concentration above which further increases will not produce a greater rate of transport. Channels generally allow transmembrane movement at rates orders of magnitude greater than those typical of carriers, rates approaching the limit of unhindered diffusion. Channels typically show less stereospecificity than carriers and are usually not saturable. Most channels are oligomeric complexes of several, often identical, subunits, whereas many carriers function as monomeric proteins. The classification as carrier or channel is the broadest distinction among transporters. Some channels are made primarily of helical transmembrane segments, others have b-barrel structures. Among the carriers, some simply facilitate diffusion down a concentration gradient: passive transporter superfamily. Active transporters can drive substrates across the membrane against a concentration gradient, some using energy provided directly by a chemical reaction (primary active transporters) and some coupling uphill transport of one substrate with downhill transport of another (secondary active transporters).
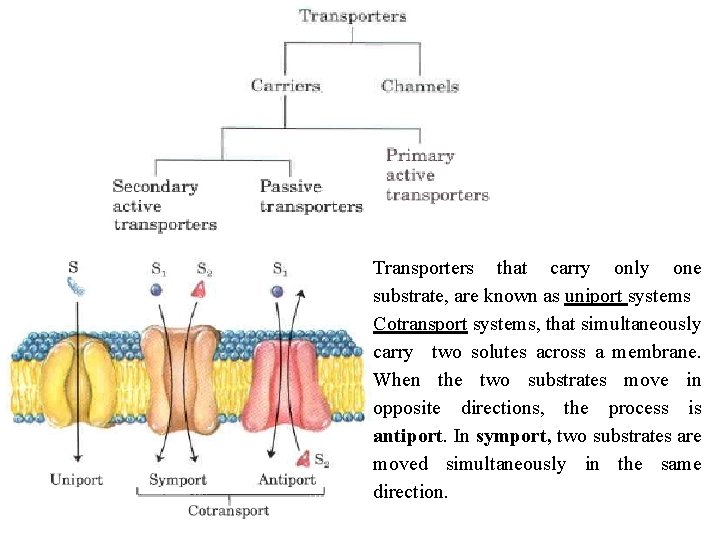
Transporters that carry only one substrate, are known as uniport systems Cotransport systems, that simultaneously carry two solutes across a membrane. When the two substrates move in opposite directions, the process is antiport. In symport, two substrates are moved simultaneously in the same direction.
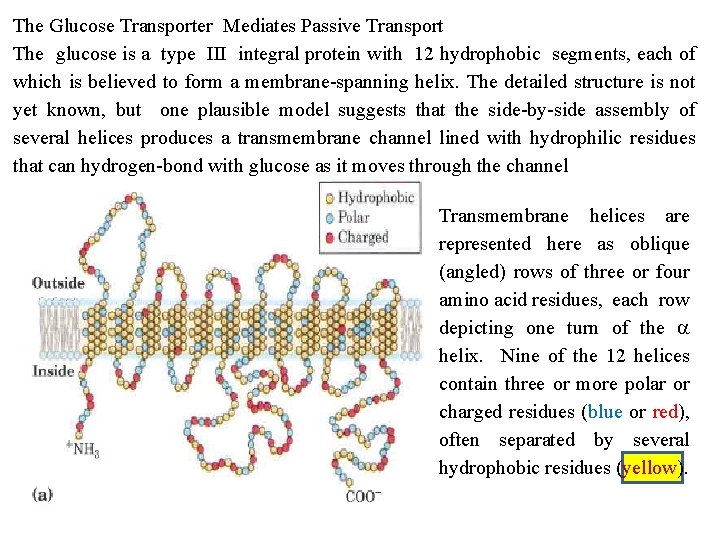
The Glucose Transporter Mediates Passive Transport The glucose is a type III integral protein with 12 hydrophobic segments, each of which is believed to form a membrane spanning helix. The detailed structure is not yet known, but one plausible model suggests that the side by side assembly of several helices produces a transmembrane channel lined with hydrophilic residues that can hydrogen bond with glucose as it moves through the channel Transmembrane helices are represented here as oblique (angled) rows of three or four amino acid residues, each row depicting one turn of the a helix. Nine of the 12 helices contain three or more polar or charged residues (blue or red), often separated by several hydrophobic residues (yellow).
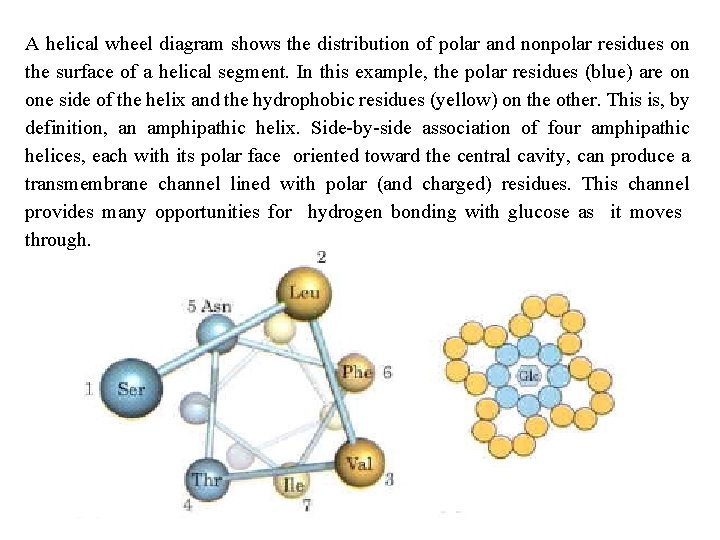
A helical wheel diagram shows the distribution of polar and nonpolar residues on the surface of a helical segment. In this example, the polar residues (blue) are on one side of the helix and the hydrophobic residues (yellow) on the other. This is, by definition, an amphipathic helix. Side by side association of four amphipathic helices, each with its polar face oriented toward the central cavity, can produce a transmembrane channel lined with polar (and charged) residues. This channel provides many opportunities for hydrogen bonding with glucose as it moves through.
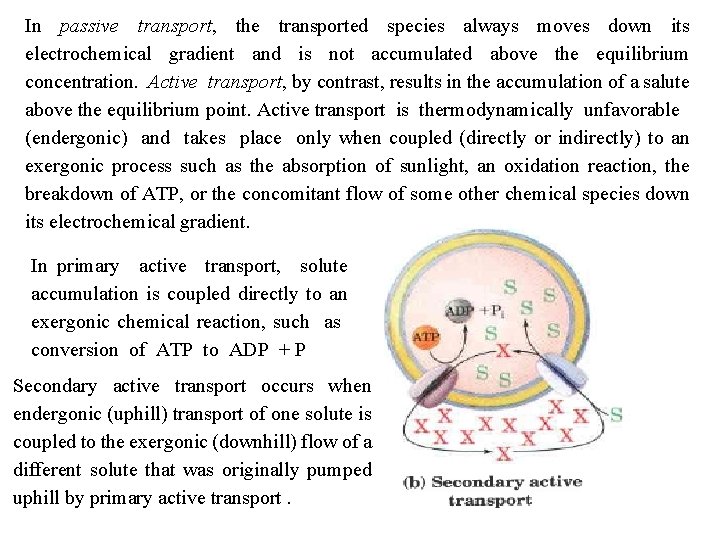
In passive transport, the transported species always moves down its electrochemical gradient and is not accumulated above the equilibrium concentration. Active transport, by contrast, results in the accumulation of a salute above the equilibrium point. Active transport is thermodynamically unfavorable (endergonic) and takes place only when coupled (directly or indirectly) to an exergonic process such as the absorption of sunlight, an oxidation reaction, the breakdown of ATP, or the concomitant flow of some other chemical species down its electrochemical gradient. In primary active transport, solute accumulation is coupled directly to an exergonic chemical reaction, such as conversion of ATP to ADP + P Secondary active transport occurs when endergonic (uphill) transport of one solute is coupled to the exergonic (downhill) flow of a different solute that was originally pumped uphill by primary active transport.

The amount of energy needed for the transport of a solute against a gradient can be calculated from the initial concentration gradient. The general equation for the free energy change in the chemical process that converts S to P is where DG’o is the standard free energy change, R is the gas constant, 8. 315 J/mol • K, and T is the absolute temperature. When the "reaction" is simply transport of a solute from a region where its concentration is C 1 to a region where its concentration is C 2, no bonds are made or broken and DG’o is zero. The free energy change for transport , DGt is then (for all uncharged solutes) If there is a 10 fold difference in concentration between two compartments, the cost of moving 1 mol of an uncharged solute at 25 °C uphill across a membrane sepa rating the compartments is

When the solute is an ion, its movement without an accompanying counterion results in the endergonic separation of positive and negative charges, producing an electrical potential; such a transport process is said to be electrogenic. The energetic cost of moving an ion depends on the electrochemical potential, the sum of the chemical and electrical gradients: where Z is the charge on the ion, F is the Faraday constant (96, 480 J/V • mol), and is the transmembrane electrical potential (in volts). Eukaryotic cells typically have plasma membrane potentials of about 0. 05 V (with the inside negative relative to the outside). Most cells maintain more than a 10 fold difference in ion concentrations across their plasma or intra cellular membranes, and for many cells and tissues active transport is therefore a major energy consuming process.

The family of active transporters called P type ATP asesare cation transporters that are reversibly phosphorylated by ATP (thus the name P type) as part of the transport cycle. Phosphorylation forces a conformational change that is central to movement of the cation across the membrane. The P type ATPases are widespread in eukaryotes and bacteria. The Ca 2+ ATPase (a uniporter for Ca 2+ ions) and the Na+K+ ATPase (an antiporter for Na+ and K+ ions) are P type. ATPases that maintain differences in ionic composition between the cytosol and the extracellular medium Lipid flippases, are structurally and functionally related to P type transporters. In vascular plants, a P type ATPase pumps protons out of cells, establishing an electrochemical difference of as much as 2 p. H units and 250 m. V across the plasma membrane. The best understood P type pumps are the Ca 2+ pumps that maintain a low concentration of Ca 2+ in the cytosol of virtually all cells. The plasma membrane Ca 2+ pump moves calcium ions out of the cell, and another P type pump in the endoplasmic reticulum moves Ca 2+ into the ER lumen.

The sarcoplasmic and endoplasmic reticulum calcium (SERCA) pumps are P type ATPases closely related in structure and mechanism. The SERCA pump of the sarcoplasmic reticulum, which comprises 80% of the protein in that membrane, is a single polypeptide (Mr 110, 000) that spans the membrane 10 times. Three cytosolic domains formed by long loops connect the transmembrane helices : the N domain, where the nucleotide ATP and Mg 2+ bind; the P domain, which contains the phosphorylated Asp residue characteristic of all P type ATPases; and the A (actuator) domain, which communicates movements of the N and P domains to the two Ca 2+ binding sites. The M domain contains the transmembrane helices and the Ca 2+ binding sites, which are located near the middle of the membrane bilayer, 40 to 50 Å from the phosphorylated Asp residue thus Asp phosphorylation dephosphorylation does not directly affect Ca 2 + binding.
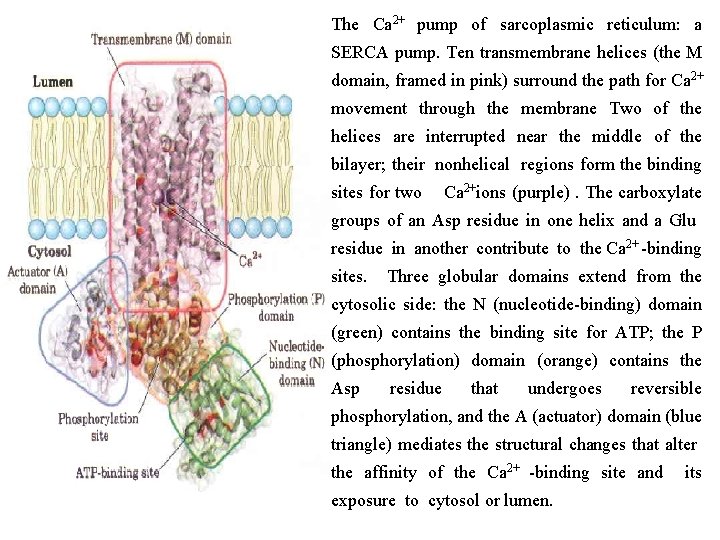
The Ca 2+ pump of sarcoplasmic reticulum: a SERCA pump. Ten transmembrane helices (the M domain, framed in pink) surround the path for Ca 2+ movement through the membrane Two of the helices are interrupted near the middle of the bilayer; their nonhelical regions form the binding sites for two Ca 2+ions (purple). The carboxylate groups of an Asp residue in one helix and a Glu residue in another contribute to the Ca 2+ binding sites. Three globular domains extend from the cytosolic side: the N (nucleotide binding) domain (green) contains the binding site for ATP; the P (phosphorylation) domain (orange) contains the Asp residue that undergoes reversible phosphorylation, and the A (actuator) domain (blue triangle) mediates the structural changes that alter the affinity of the Ca 2+ binding site and exposure to cytosol or lumen. its

The mechanism postulated for SERCA pumps takes into account the large conformational changes and the phosphorylation dephosphorylation of the critical Asp residue in the P domain that is known to occur during a catalytic cycle. Each catalytic cycle moves two Ca 2+ ions across the membrane and converts an ATP to ADP and Pi. The role of ATP binding and hydrolysis is to bring about the interconversion of two conformations (E 1 and E 2) of the transporter. In the E 1 conformation, the two Ca 2+ binding sites are exposed on the cytosolic side of the ER or sarcoplasmic reticulum and bind Ca 2+ with high affinity. ATP binding and Asp phosphorylation drive a conformational change from E l to E 2 in which the Ca 2+ binding sites are now exposed on the lumenal side of the membrane and their affinity for Ca 2+ is greatly reduced, causing Ca 2+ release into the lumen. By this mechanism, the energy released by hydrolysis of ATP during one phosphorylation dephosphorylation cycle drives Ca 2+ across the membrane against a large electrochemical gradient.

The transport cycle begins with the protein in the El conformation, with the Ca 2+ binding sites facing the cytosol. Two Ca 2+ ions bind, then ATP binds to the transporter and phosphorylates Asp 351 , forming El P. Phosphorylation favors the second conformation, E 2 P, in which the Ca 2+ binding sites, now with a reduced affinity for Ca 2+, are accessible on the other side of the membrane (the lumen or extracellular space), and the released Ca 2+ diffuses away. Finally, E 2 P is dephosphorylated, returning the protein to the El conformation for another round of transport
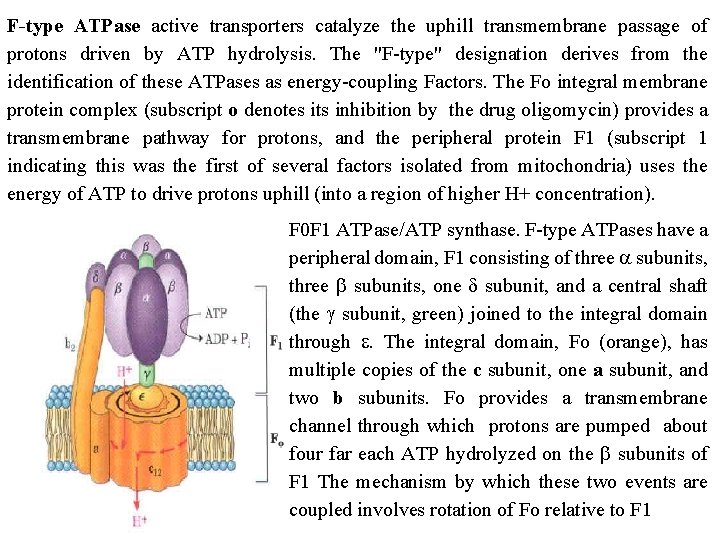
F type ATPase active transporters catalyze the uphill transmembrane passage of protons driven by ATP hydrolysis. The "F type" designation derives from the identification of these ATPases as energy coupling Factors. The Fo integral membrane protein complex (subscript o denotes its inhibition by the drug oligomycin) provides a transmembrane pathway for protons, and the peripheral protein F 1 (subscript 1 indicating this was the first of several factors isolated from mitochondria) uses the energy of ATP to drive protons uphill (into a region of higher H+ concentration). F 0 F 1 ATPase/ATP synthase. F type ATPases have a peripheral domain, F 1 consisting of three a subunits, three b subunits, one d subunit, and a central shaft (the g subunit, green) joined to the integral domain through e. The integral domain, Fo (orange), has multiple copies of the c subunit, one a subunit, and two b subunits. Fo provides a transmembrane channel through which protons are pumped about four far each ATP hydrolyzed on the b subunits of F 1 The mechanism by which these two events are coupled involves rotation of Fo relative to F 1
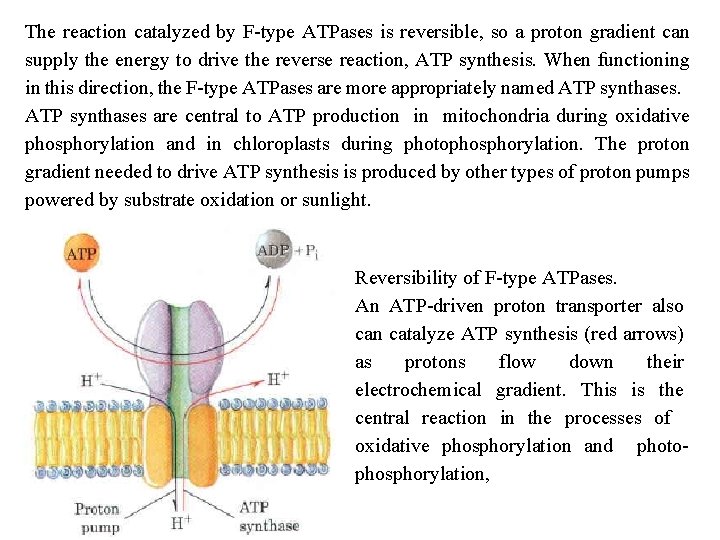
The reaction catalyzed by F type ATPases is reversible, so a proton gradient can supply the energy to drive the reverse reaction, ATP synthesis. When functioning in this direction, the F type ATPases are more appropriately named ATP synthases are central to ATP production in mitochondria during oxidative phosphorylation and in chloroplasts during photophosphorylation. The proton gradient needed to drive ATP synthesis is produced by other types of proton pumps powered by substrate oxidation or sunlight. Reversibility of F type ATPases. An ATP driven proton transporter also can catalyze ATP synthesis (red arrows) as protons flow down their electrochemical gradient. This is the central reaction in the processes of oxidative phosphorylation and photo phosphorylation,
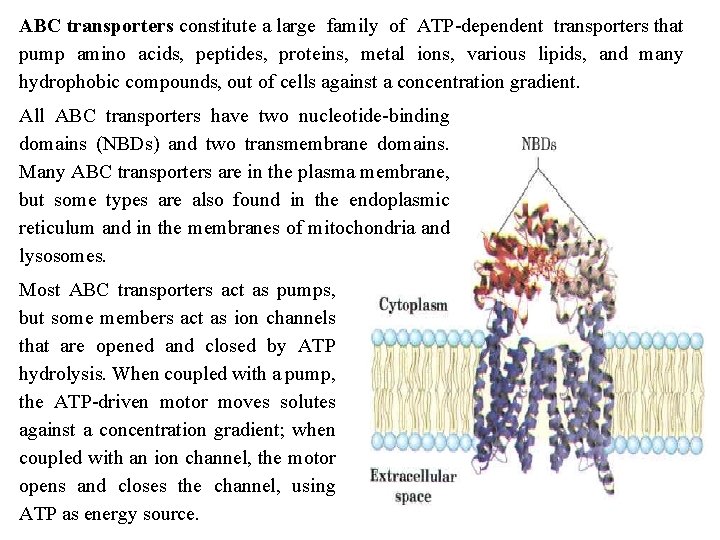
ABC transporters constitute a large family of ATP dependent transporters that pump amino acids, peptides, proteins, metal ions, various lipids, and many hydrophobic compounds, out of cells against a concentration gradient. All ABC transporters have two nucleotide binding domains (NBDs) and two transmembrane domains. Many ABC transporters are in the plasma membrane, but some types are also found in the endoplasmic reticulum and in the membranes of mitochondria and lysosomes. Most ABC transporters act as pumps, but some members act as ion channels that are opened and closed by ATP hydrolysis. When coupled with a pump, the ATP driven motor moves solutes against a concentration gradient; when coupled with an ion channel, the motor opens and closes the channel, using ATP as energy source.

A family of integral membrane proteins the aquaporins (AQPs), provide channels for rapid movement of water molecules across all plasma membranes. Aquaporins are found in all organisms. The plant Arabidopsis thaliana has 38 AQP genes, reflecting the critical roles of water movement in plant physiology. Changes in turgor pressure, require rapid movement of water across a membrane Water molecules flow through an AQP 1 channel at a rate of about 109 s 1 The low activation energy for passage of water through aquaporin channels (AG+< 15 k. J/mol) suggests that water moves through the channels in a continuous stream, in the direction dictated by the osmotic gradient. Aquaporins are higly selective do not allow passage of protons (hydronium ions, H 30+), which would collapse membrane electrochemical gradients. AQP 1 consists of four identica monomers each of which forms a transmembrane pore with a diameter sufficient to allow passage of water molecules in single file. Each monomer has six transmembrane helical segments and two shorter helices, both of which contain the sequence Asn Pro Ala (NPA). The six transmembrane helices form the pore through the monomer, and the two short loops containing the NPA sequences extend toward the middle of the bilayer from opposite sides. Their NPA regions overlap in the middle of the membrane to form part of the specificity filter the structure that al lows only water to pass
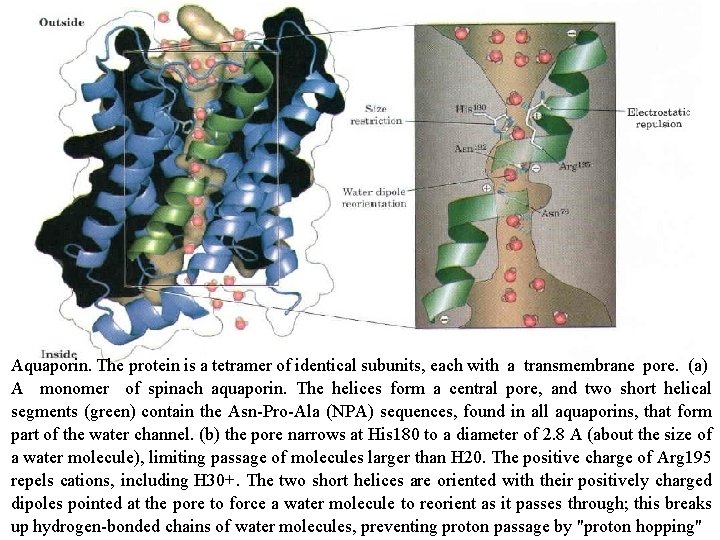
Aquaporin. The protein is a tetramer of identical subunits, each with a transmembrane pore. (a) A monomer of spinach aquaporin. The helices form a central pore, and two short helical segments (green) contain the Asn Pro Ala (NPA) sequences, found in all aquaporins, that form part of the water channel. (b) the pore narrows at His 180 to a diameter of 2. 8 A (about the size of a water molecule), limiting passage of molecules larger than H 20. The positive charge of Arg 195 repels cations, including H 30+. The two short helices are oriented with their positively charged dipoles pointed at the pore to force a water molecule to reorient as it passes through; this breaks up hydrogen bonded chains of water molecules, preventing proton passage by "proton hopping"

The water channel narrows to a diameter of 2. 8 Å near the center of the membrane, severely restricting the size of molecules that can travel through. The positive charge of a highly conserved Arg residue at this bottleneck discourages the passage of cations such as H 30+ The residues that line the channel of each AQP 1 monomer are generally nonpolar, but carbonyl oxygens in the peptide backbone, projecting into the narrow part of the channel at intervals, can hydrogen bond with individual water molecules as they pass through; the two Asn residues (Asn 76 and Asn 192) in the NPA loops also form hydrogen bonds with the water. The structure of the channel does not permit formation of a chain of water molecules close enough to allow proton hopping, which would effectively move protons across the membrane. Critical Arg and His residues and electric dipoles formed by the short helices of the NPA loops provide positive charges in positions that repel any protons that might leak through the pore, and prevent hydrogen bonding between adjacent water molecules. An aquaporin isolated from spinach is known to be "gated" open when two critical Ser residues near the intracellular end of the channel are phosphorylated, and closed when they are dephosphorylated. Phosphorylation favors a conformation that presses two nearby Leu residues and a His residue into the channel, blocking the movement of water past that point and effectively closing the channel.

Ion selective channels first are present in the plasma membranes of all cells, as well as in the intracellular membranes of eukaryotes provide another mechanism for moving inorganic ions across membranes. Ion channels, together with ion pumps such as the Na+K+ ATPase, determine a plasma membrane's permeability to specific ions and regulate the cytosolic concentration of ions and the membrane potential. Ion channels are distinct from ion transporters. First, the rate of flux through channels can be several orders of magnitude greater than the turnover number for a transporter 107 to 108 ions/s for an ion channel, approaching theoretical maximum for unrestricted diffusion. By contrast, the turnover rate of the Na+K+ ATPase is about 100 s 1 Second, ion channels are not saturable: rates do not approach a maximum at high substrate concentration. Third, they are gated in response to some cellular event. In ligand gated channels, binding of an extracellular or intracellular small molecule forces an allosteric transition in the protein, which opens or closes the channel. In voltage gated ion channels, a change in transmembrane electrical potential (Vm) causes a charged protein domain to move relative to the membrane, opening or closing the channel.

The Structure of a K+ Channel Among the members of this protein family, the similarities in sequence are greatest in the "pore region, " which contains the ion selectivity filter that allows K+ (radius 1. 33 Å) to pass 10, 000 times more readily than Na+ (radius 0. 95 Å ) at a rate (about 108 ions/s) approaching theoretical limit for unrestricted diffusion. The K+ channel consists of four identical subunits that span the membrane and form a cone within a cone surrounding the ion channel, with the wide end of the double cone facing the extracellular space. Each subunit has two transmembrane a helices as well as a third, shorter helix that contributes to the pore region. The outer cone is formed by one of the transmembrane helices of each subunit. The inner cone formed by the other four transmembrane helices surrounds the ion channel and cradles the ion selectivity filter.
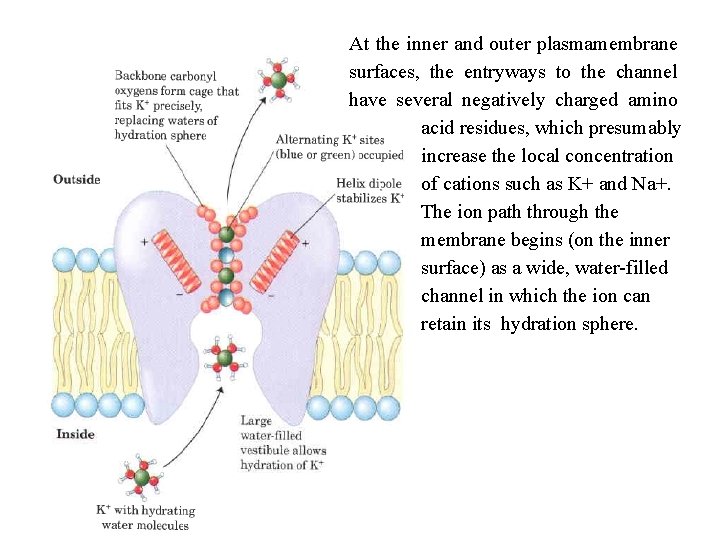
At the inner and outer plasmamembrane surfaces, the entryways to the channel have several negatively charged amino acid residues, which presumably increase the local concentration of cations such as K+ and Na+. The ion path through the membrane begins (on the inner surface) as a wide, water filled channel in which the ion can retain its hydration sphere.
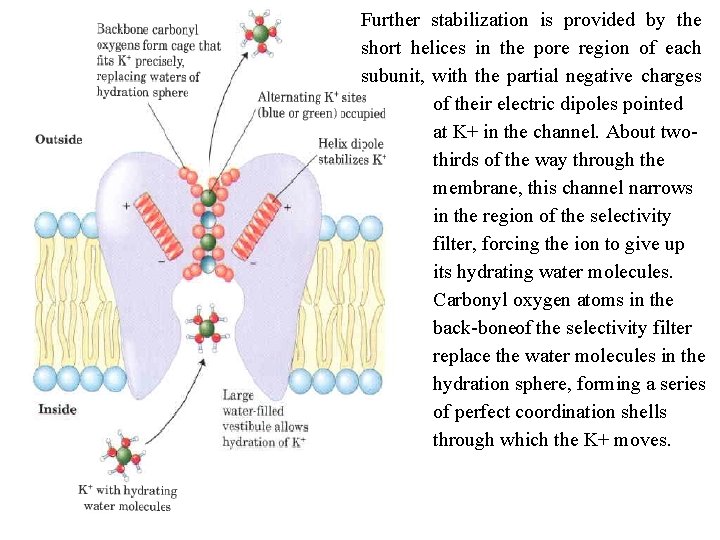
Further stabilization is provided by the short helices in the pore region of each subunit, with the partial negative charges of their electric dipoles pointed at K+ in the channel. About two thirds of the way through the membrane, this channel narrows in the region of the selectivity filter, forcing the ion to give up its hydrating water molecules. Carbonyl oxygen atoms in the back boneof the selectivity filter replace the water molecules in the hydration sphere, forming a series of perfect coordination shells through which the K+ moves.
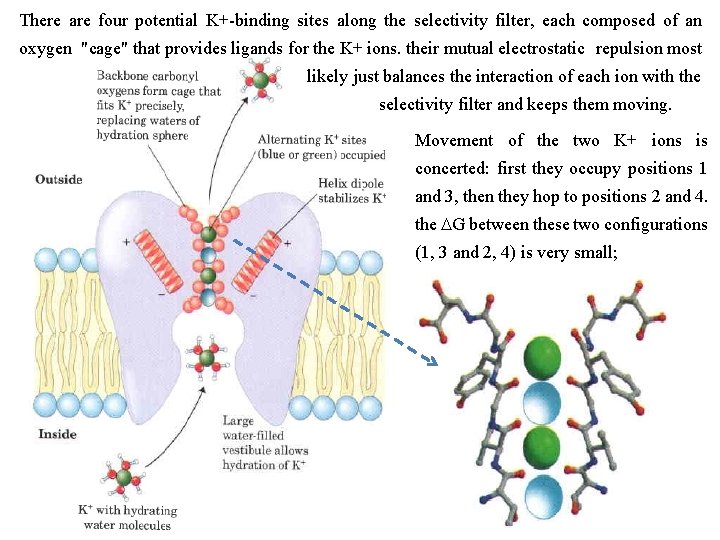
There are four potential K+ binding sites along the selectivity filter, each composed of an oxygen "cage" that provides ligands for the K+ ions. their mutual electrostatic repulsion most likely just balances the interaction of each ion with the selectivity filter and keeps them moving. Movement of the two K+ ions is concerted: first they occupy positions 1 and 3, then they hop to positions 2 and 4. the DG between these two configurations (1, 3 and 2, 4) is very small;
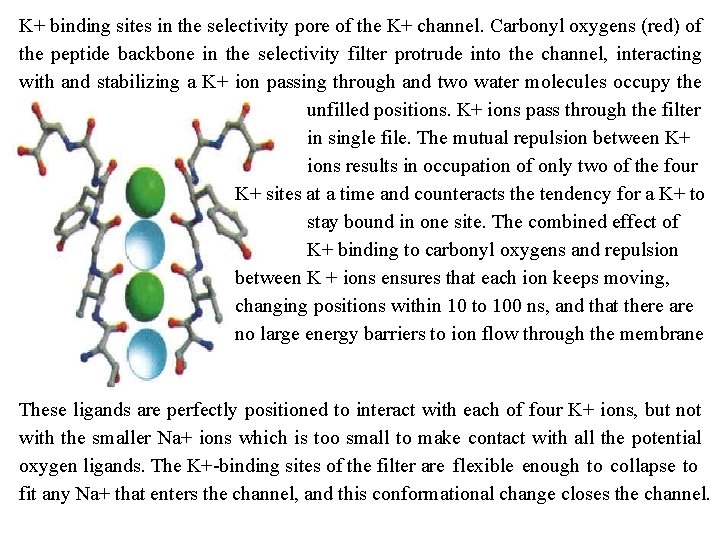
K+ binding sites in the selectivity pore of the K+ channel. Carbonyl oxygens (red) of the peptide backbone in the selectivity filter protrude into the channel, interacting with and stabilizing a K+ ion passing through and two water molecules occupy the unfilled positions. K+ ions pass through the filter in single file. The mutual repulsion between K+ ions results in occupation of only two of the four K+ sites at a time and counteracts the tendency for a K+ to stay bound in one site. The combined effect of K+ binding to carbonyl oxygens and repulsion between K + ions ensures that each ion keeps moving, changing positions within 10 to 100 ns, and that there are no large energy barriers to ion flow through the membrane These ligands are perfectly positioned to interact with each of four K+ ions, but not with the smaller Na+ ions which is too small to make contact with all the potential oxygen ligands. The K+ binding sites of the filter are flexible enough to collapse to fit any Na+ that enters the channel, and this conformational change closes the channel.

- Slides: 127