Canadian Bioinformatics Workshops www bioinformatics ca In collaboration

Canadian Bioinformatics Workshops www. bioinformatics. ca

In collaboration with Cold Spring Harbor Laboratory & New York Genome Center
Module #: Title of Module 3

Module 1 bioinformatics. ca

You are free to: Copy, share, adapt, or re-mix; Photograph, film, or broadcast; Blog, live-blog, or post video of; This presentation. Provided that: You attribute the work to its author and respect the rights and licenses associated with its components. Slide Concept by Cameron Neylon, who has waived all copyright and related or neighbouring rights. This slide onlycc. Zero. Social Media Icons adapted with permission from originals by Christopher Ross. Original images are available under GPL at; http: //www. thisismyurl. com/free-downloads/15 -free-speech-bubble-icons-for-popular-websites Module 1 bioinformatics. ca

E-mail francis@oicr. on. ca @bffo #CBW 15 Module 1 bioinformatics. ca

Disclaimer • I do not (and will not) profit in any way, shape or form, from any of the brands, products or companies I may mention. Module 1 bioinformatics. ca

Module 1. 1 Overview of Workshop BF Francis Ouellette High Throughput Biology: From Sequence to Networks April 27 -May 3, 2015

Outline • • Bioinformatics History of bioinformatics. ca Cloud computing Getting on Amazon Web Services Module 1 bioinformatics. ca
What biologist do: • • • Make observations Make hypothesis Test them Challenge them Conclude things Write papers http: //goo. gl/7 s. CUI Module 1 bioinformatics. ca
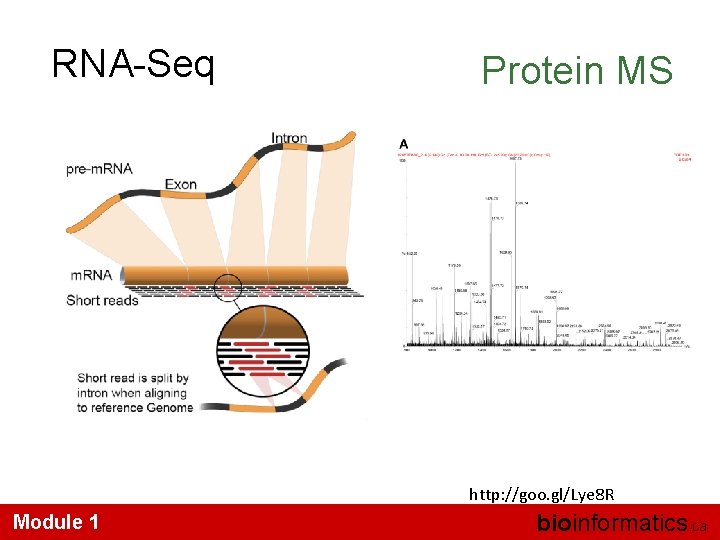
RNA-Seq Protein MS http: //goo. gl/Lye 8 R Module 1 bioinformatics. ca
Interaction and Pathway Space Module 1 bioinformatics. ca
Central Dogma DNA RNA protein Module 1 bioinformatics. ca
Central Dogma DNA RNA protein Module 1 Then you write a paper about it bioinformatics. ca

Some of the things we do when we try and understand the cell … • We do experiments • Some of these are bioinformatics experiments • We all want these to be reproducible • We want people to find our data • We want people to find our methods • … and we want them to be able to rerun our experiments, validate our work, move the science forward.
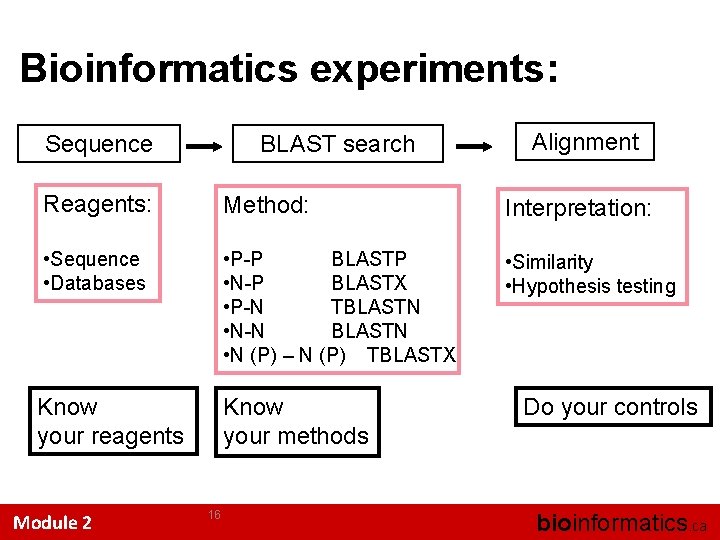
Bioinformatics experiments: Sequence BLAST search Alignment Reagents: Method: Interpretation: • Sequence • Databases • P-P BLASTP • N-P BLASTX • P-N TBLASTN • N-N BLASTN • N (P) – N (P) TBLASTX • Similarity • Hypothesis testing Know your reagents Know your methods Module 2 16 Do your controls bioinformatics. ca

What is Bioinformatics? Think – Pair – Share! Introduction 1. 0 Module 1 17 bioinformatics. ca

Bioinformatics is about integrating biological themes together with the help of computer tools and biological databases, and gaining new knowledge about the system in study. Module 1 bioinformatics. ca

1998 Module 1 bioinformatics. ca
1999 – 2007 Bioinformatics Proteomics Genomics Developing the Tools Module 1 bioinformatics. ca

2008– present Module 1 bioinformatics. ca

• • • Analysis of Metagenomic Data - 3 Bioinformatics of Cancer Genomics - 5 Exploratory Analysis of Biological Data using R - 2 High-throughput Biology: From Sequence to Networks - 7 Informatics and Statistics for Metabolomics - 2 Informatics for RNA-seq Analysis- 2 Informatics on High-Throughput Sequencing Data– 2 Introduction to R – 1 Microarray Expression Analysis - 2 Pathway and Network Analysis of -omic Data – 3 Module 1 bioinformatics. ca
bioinformatics. ca Module 1 bioinformatics. ca
http: //bioinformatics. ca/workshops/2014

E-mail: course_info@bioinformatics. ca Web: http: //bioinformatics. ca Workshop announcement mailing list: http: //bioinformatics. ca/mailman/listinfo/announce

Soap-Box time! • Open Access, Open Data and Open Source are essential for Science. • Openness is a responsibility, an obligation, and something that comes with the privilege of doing publicly funded work. Open Source Open Access Open Data Opencourseware

• If databases get it wrong, the onus is on on the user to let the databases know that it is wrong! http: //goo. gl/b. Gj. MH Module 1 bioinformatics. ca

• If databases get it wrong, the onus is on on the user to let the databases know that it is wrong! any db ………………………. . http: //goo. gl/b. Gj. MH Module 1 bioinformatics. ca

Q: Why do we have Bioinformatics? A: Open Data from Genomic and Proteomics Technologies Module 1 bioinformatics. ca

Module 1. 2 Overview of Cloud Computing BF Francis Ouellette High-Throughput Biology: From Sequence to Networks April 27 -May 3, 2015

Cloud computing … and new software paradigm • Data sets are reaching the Petabyte scale. • Data (and the security rules that come with it) will be somewhere, and you will move your software to it. • Software development paradigm will change: no more reading of files into RAM, processing, and then writing output: you need to think about processing streaming data coming from a sequencing machine somewhere on the net.
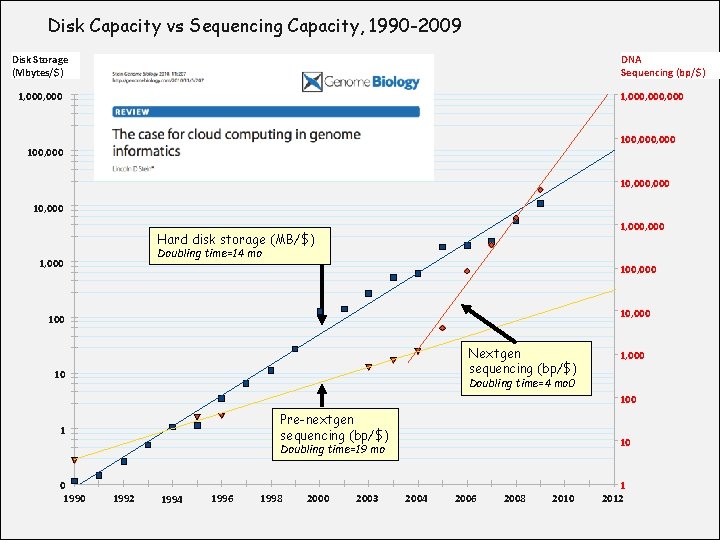
Disk Capacity vs Sequencing Capacity, 1990 -2009 Disk Storage (Mbytes/$) DNA Sequencing (bp/$) 1, 000, 000 100, 000 10, 000 1, 000 Hard disk storage (MB/$) Doubling time=14 mo 1, 000 100, 000 100 Nextgen sequencing (bp/$) 10 1, 000 Doubling time=4 mo 0 100 Pre-nextgen sequencing (bp/$) 1 10 Doubling time=19 mo 0 1990 Module 1 1992 1994 1996 1998 2000 2003 2004 2006 2008 2010 1 2012 bioinformatics. ca

About DNA and computers • We now have ~ $1000 genome, but now need to think more about the cost of the analysis. • The doubling time of the reduction of sequencing in cost is in the “many months” range. • The doubling time of storage and network bandwidth is “very small number of years” range. • The doubling time of CPU speed is 18 months. • The cost of sequencing a base pair will equal the cost of storing a base pair by in the next “very small number” of years. Module 1 bioinformatics. ca

What is the general biomedical scientists to do? • Too much data and not enough computer infrastructure in most labs – – Where do they go? Write more grants? Get more hardware? Look to the sky? Module 1 bioinformatics. ca
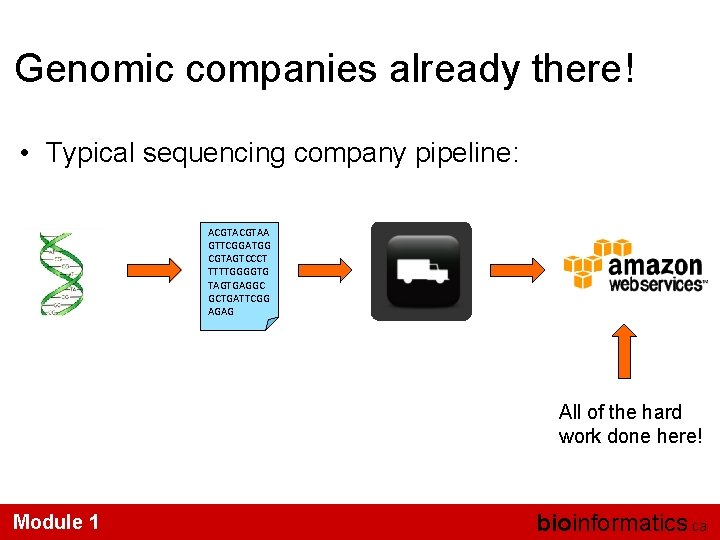
Genomic companies already there! • Typical sequencing company pipeline: ACGTAA GTTCGGATGG CGTAGTCCCT TTTTGGGGTG TAGTGAGGC GCTGATTCGG AGAG All of the hard work done here! Module 1 bioinformatics. ca

Most people already there! • • Google docs Dropbox Netflix Twitter Module 1 bioinformatics. ca

Amazon Web Services (AWS) • • • Infinite storage (scalable): S 3 (simple storage service) Compute per hour: EC 2 (elastic cloud computing) Ready when you are High Performance Computing Multiple football fields of HPC throughout the world HPC are expanded at one contained at a time: http: //goo. gl/7 PVAl Module 1 bioinformatics. ca

Some of the challenges with cloud computing: • • • Not cheap! Getting files to and from there Not the best solution for everybody Standardization PHI: personal health information & security concerns In the USA: Patriot act Module 1 bioinformatics. ca

Some of the advantages with cloud computing: • At the CBW: we received a grant from Amazon, so supported by ‘AWS in Education grant award. • There are better ways of transferring large files, and now AWS makes it free to upload files. • A number of datasets exist on AWS (e. g. 1000 genome data). • Many useful bioinformatics AMI’s (Amazon Machine Images) exist on AWS: e. g. cloudbiolinux & Cloud. Man (Galaxy) • Many flavors of cloud available, not just AWS Module 1 bioinformatics. ca

In this workshop: • Some tools (data) are • on your computer • on the web • on the cloud. • You will become efficient at traversing these various spaces, and finding resources you need, and using what is best for you. • There are different ways of using the cloud: 1. Command line (like your own very powerful Unix box) 2. With a web-browser (e. g. Galaxy): not in this workshop Module 1 bioinformatics. ca

“Big Data” is a relative term! • This is what a 5 MB hard drive looked like in 1956! • What will it be in 2056? http: //goo. gl/f 1 Pk. V Module 1 bioinformatics. ca

Min. ION from Oxford Nanopore http: //www. nanoporetech. com/technology/minion-a-miniaturised-sensing-instrument

Things we have set up: • Loaded data files to an AWS • We brought up an Ubuntu (Linux) instance, and loaded a whole bunch of software for NGS analysis. • We then cloned this, and made separate instances for everybody in the class. • We’ve simplified the security: you basically all have the same login and file access, and opened ports. In your own world you would be more secure. Module 1 bioinformatics. ca
For this workshop: all on Wiki! http: //bioinformatics. ca/workshop_wiki/ Login: Firstname. Lastname Password: guest Module 1 bioinformatics. ca
Module 1 bioinformatics. ca
Module 1 bioinformatics. ca
Module 1 bioinformatics. ca
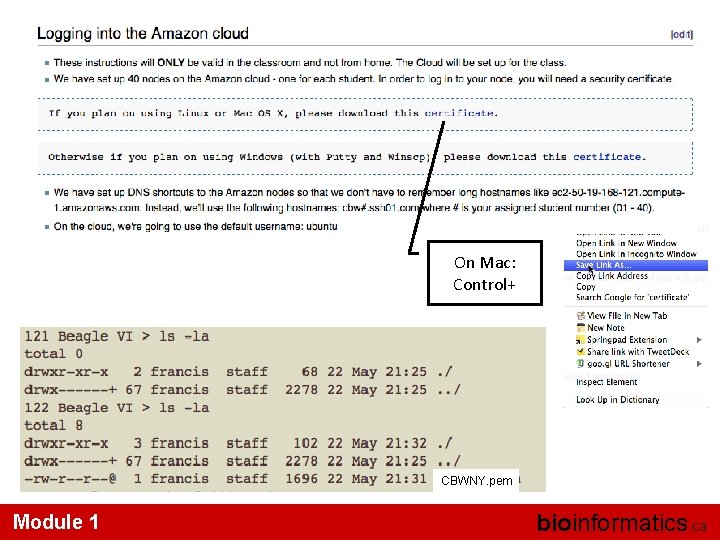
On Mac: Control+ CBWNY. pem Module 1 bioinformatics. ca
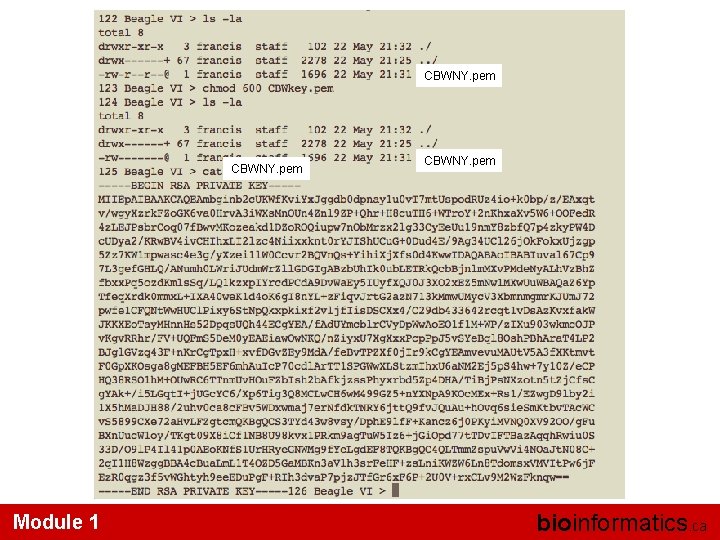
CBWNY. pem Module 1 CBWNY. pem bioinformatics. ca

• ls -l (long listing) drwx------+ 67 francis -rw-r--r--@ 1 francis rwx : owner rwx : group rwx: world r read (4) w write (2) x execute (1) staff 2278 22 May 21: 25. . / 1696 22 May 21: 31 CBWNY. pem Which ever way you add these 3 numbers, you know which integers were used (6 is always 4+2, 5 is 4+1, 4 is by itself, 0 is none of them etc …) So, when you have: chmod 600 <file name> It is “rw” for the file owner only Module 1 bioinformatics. ca

Logging in to AWS Module 1 bioinformatics. ca

Windows Module 1 bioinformatics. ca
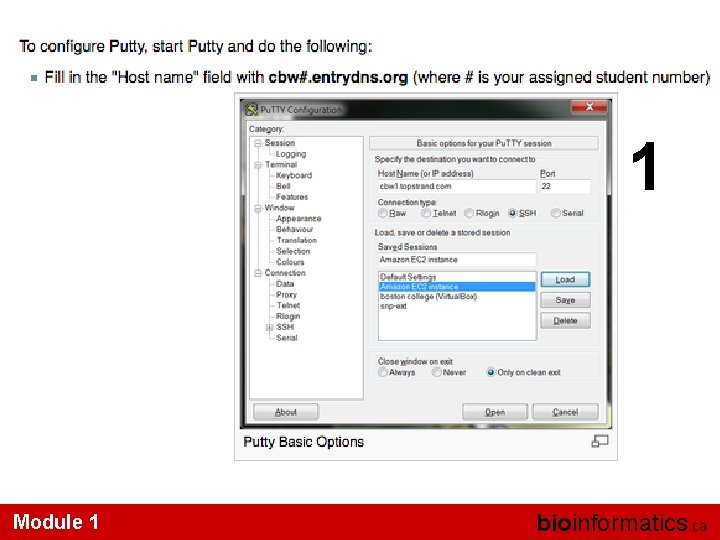
1 Module 1 bioinformatics. ca
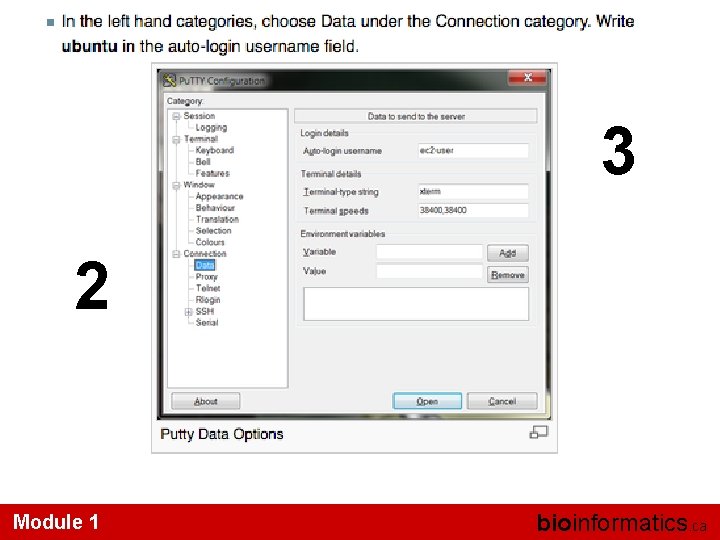
3 2 Module 1 bioinformatics. ca
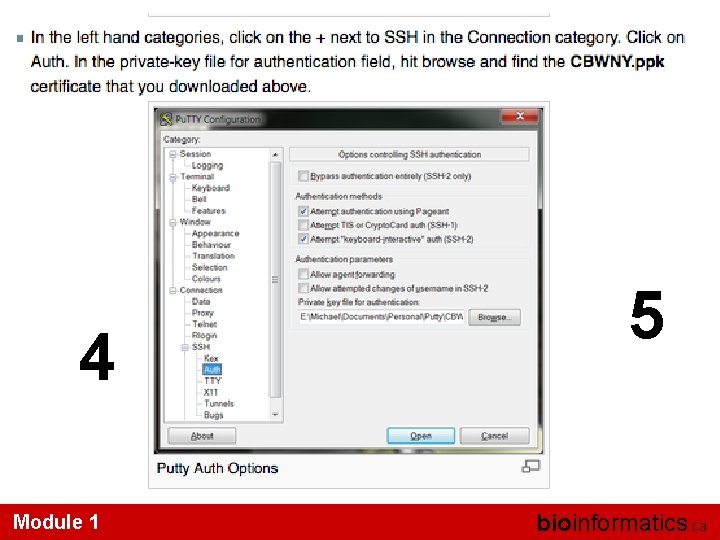
4 Module 1 5 bioinformatics. ca
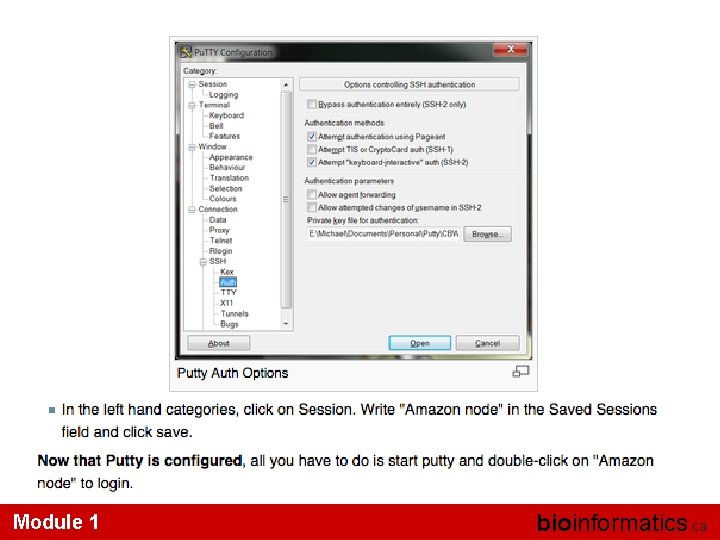
Module 1 bioinformatics. ca

So, at this point: • Your laptop is ready for the workshop • If it is not, you know where to get the information you need • You know how to use the wiki for this workshop • You know where all of the lectures are • You have read all of the pre-lecture material • If not, you know where the papers are, and you are a speed reader • You know how to login to AWS Module 1 bioinformatics. ca

We are on a Coffee Break & Networking Session Module 1 bioinformatics. ca
- Slides: 58