Canadian Bioinformatics Workshops www bioinformatics ca Day 2

Canadian Bioinformatics Workshops www. bioinformatics. ca Day 2 Section 8 1
Day 2 Section 8 2

Introduction to statistics for Genome. Wide Association Studies (GWAS) Day 2 Section 8 3

Outline • • • Background on GWAS Presentation of Gen. ABEL Data checking with Gen. ABEL Data analysis with Gen. ABEL Display of results Day 2 Section 8 4

R Packages for GWAS • Plink developped by the M. I. T. but only available for linux platform only. (http: //pngu. mgh. harvard. edu/~purcell/plink/). • SNPassoc (Juan R. González 1, et al. Bioinformatics, 2007 23(5): 654 -655) • Gen. ABEL (Aulchenko Y. S. , Ripke S. , Isaacs A. , van Duijn C. M. Bioinformatics. 2007, 23(10): 1294 -6. ) Day 2 Section 8 5

What is a GWAS? • A genome-wide association study is an approach that involves rapidly scanning markers across genome (≈0. 5 M or 1 M) of many people (≈2 K) to find genetic variations associated with a particular disease. • A large number of subjects are needed because (1)associations between SNPs and causal variants are expected to show low odds ratios, typically below 1. 5 (2)In order to obtain a reliable signal, given the very large number of tests that are required, associations must show a high level of significance to survive the multiple testing correction • Such studies are particularly useful in finding genetic variations that contribute to common, complex diseases Day 2 Section 8 6

What is a GWAS? Day 2 Section 8 7

Why are such studies possible now? The completion of the Human Genome Project in 2003 and the International Hap. Map Project in 2005, researchers now have a set of research tools that make it possible to find the genetic contributions to common diseases Day 2 Section 8 8

GWAS for complex diseases Day 2 Section 8 9
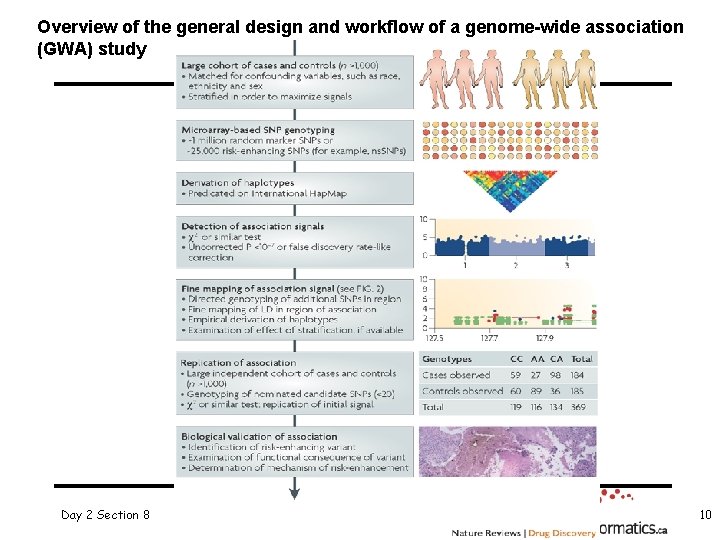
Overview of the general design and workflow of a genome-wide association (GWA) study Day 2 Section 8 10

What have GWAS found? • In 2005, it was learned through GWAS that age-related macular degeneration is associated with variation in the gene for complement factor H, which produces a protein that regulates inflammation (Klein et al. (2005) Science, 308, 385– 389) • In 2007, the Wellcome Trust Case-Control Consortium (WTCCC) carried out GWAS for the diseases coronary heart disease, type 1 diabetes, type 2 diabetes, rheumatoid arthritis, Crohn's disease, bipolar disorder and hypertension. This study was successful in uncovering many new disease genes underlying these diseases. • See next page for more publications in GWAS Day 2 Section 8 11

Examples of GWAS • • Association scan of 14, 500 nonsynonymous SNPs in four diseases identifies autoimmunity variants. Nat Genet. 2007 Genome-wide association study of 14, 000 cases of seven common diseases and 3, 000 shared controls. Wellcome Trust Case Control Consortium Nature. 2007; 447; 661 -78 Genomewide association analysis of coronary artery disease. Samani et al. N Engl J Med. 2007; 357; 443 -53 Sequence variants in the autophagy gene IRGM and multiple other replicating loci contribute to Crohn's disease susceptibility. Parkes et al. Nat Genet. 2007; 39; 830 -2 Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Todd et al. Nat Genet. 2007; 39; 857 -64 A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Frayling et al. Science. 2007; 316; 889 -94 Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Zeggini et al. Science. 2007; 316; 1336 -41 Scott et al. (2007) A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science, 316, 1341– 1345. Day 2 Section 8 12
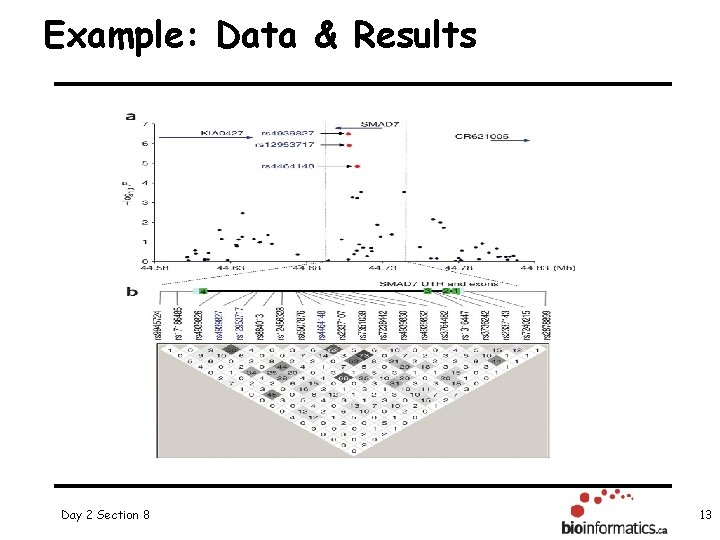
Example: Data & Results Day 2 Section 8 13

Problem(s) How to make inference about SNP-Disease associations ? Which computational tools to use? Day 2 Section 8 14

Features of Gen. ABEL • Specifically designed for GWAS • Provides specific facilities for storage and manipulation of large data • Very fast tests for GWAS • Specific functions to analyze and display the results • More efficient than the library “genetics” Day 2 Section 8 15
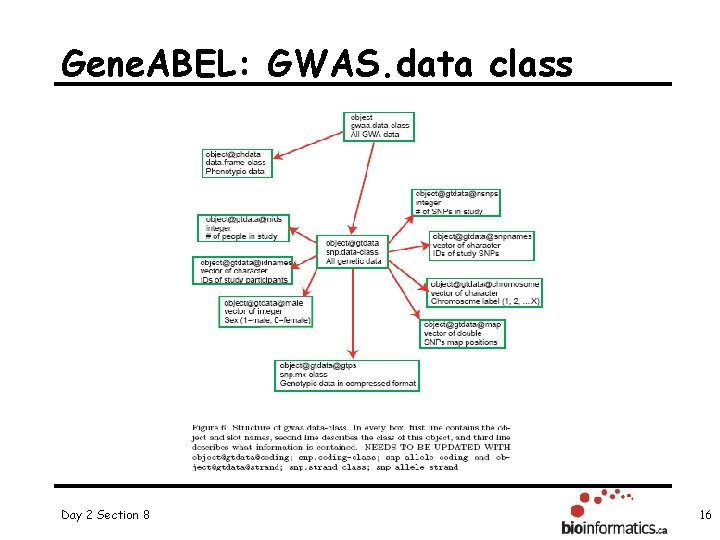
Gene. ABEL: GWAS. data class Day 2 Section 8 16

Exploring GWAS. data class objects R code: library("Gen. ABEL") data(ge 03 d 2 ex) # phenotype data summary(ge 03 d 2 ex@phdata) R output Day 2 Section 8 17

Exploring GWAS. data class objects R code: library("Gen. ABEL") data(ge 03 d 2 ex) # phenotype data summary(ge 03 d 2 ex@phdata) # number of people in study ge 03 d 2 ex@gtdata@nids # number of SNPs ge 03 d 2 ex@gtdata@nsnps # SNP names ge 03 d 2 ex@gtdata@snpnames[1: 10] # Chromosome labels ge 03 d 2 ex@gtdata@chromosome[1: 10] # SNPs map positions ge 03 d 2 ex@gtdata@map[1: 10] Day 2 Section 8 18

Descriptive statistics: phenotypes R code: descriptive. trait(ge 03 d 2 ex) R output type 2 diabetes status R code: descriptives. trait(ge 03 d 2 ex, by=ge 03 d 2 ex@phdata$dm 2)) Day 2 Section 8 = by case-control status 19

Descriptive statistics: markers R code: descriptives. marker(ge 03 d 2 ex) R output Day 2 Section 8 20
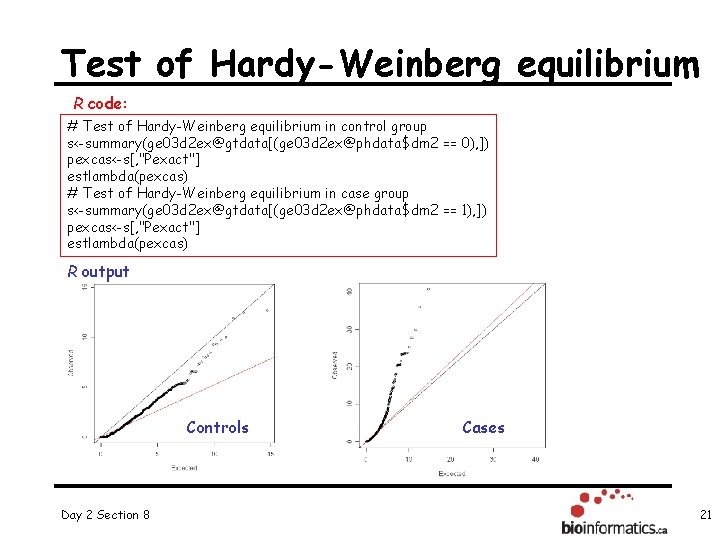
Test of Hardy-Weinberg equilibrium R code: # Test of Hardy-Weinberg equilibrium in control group s<-summary(ge 03 d 2 ex@gtdata[(ge 03 d 2 ex@phdata$dm 2 == 0), ]) pexcas<-s[, "Pexact"] estlambda(pexcas) # Test of Hardy-Weinberg equilibrium in case group s<-summary(ge 03 d 2 ex@gtdata[(ge 03 d 2 ex@phdata$dm 2 == 1), ]) pexcas<-s[, "Pexact"] estlambda(pexcas) R output Controls Day 2 Section 8 Cases 21
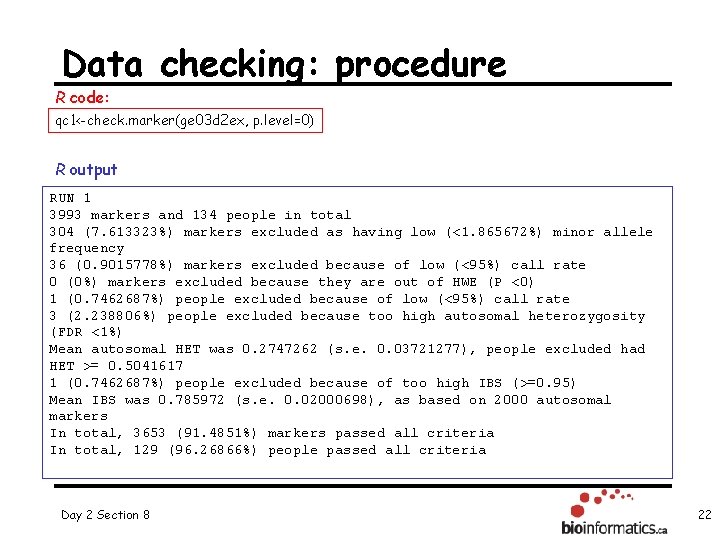
Data checking: procedure R code: qc 1<-check. marker(ge 03 d 2 ex, p. level=0) R output RUN 1 3993 markers and 134 people in total 304 (7. 613323%) markers excluded as having low (<1. 865672%) minor allele frequency 36 (0. 9015778%) markers excluded because of low (<95%) call rate 0 (0%) markers excluded because they are out of HWE (P <0) 1 (0. 7462687%) people excluded because of low (<95%) call rate 3 (2. 238806%) people excluded because too high autosomal heterozygosity (FDR <1%) Mean autosomal HET was 0. 2747262 (s. e. 0. 03721277), people excluded had HET >= 0. 5041617 1 (0. 7462687%) people excluded because of too high IBS (>=0. 95) Mean IBS was 0. 785972 (s. e. 0. 02000698), as based on 2000 autosomal markers In total, 3653 (91. 4851%) markers passed all criteria In total, 129 (96. 26866%) people passed all criteria Day 2 Section 8 22
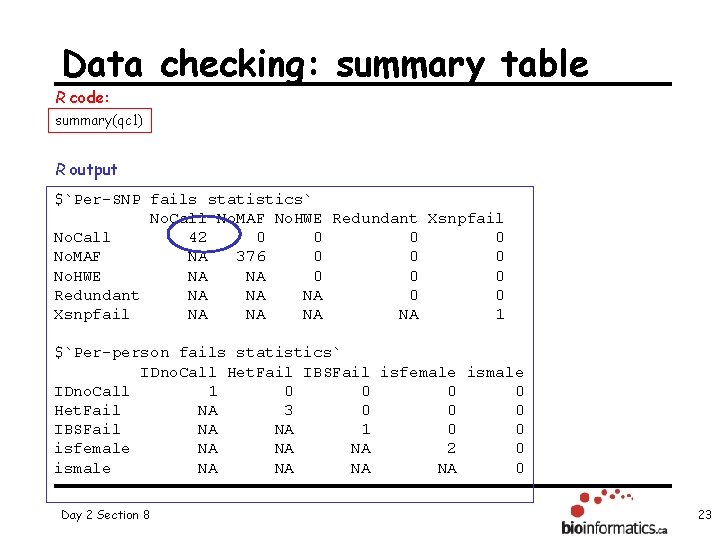
Data checking: summary table R code: summary(qc 1) R output $`Per-SNP fails statistics` No. Call No. MAF No. HWE Redundant Xsnpfail No. Call 42 0 0 0 No. MAF NA 376 0 0 No. HWE NA 0 0 Redundant NA NA 0 0 Xsnpfail NA NA 1 $`Per-person fails statistics` IDno. Call Het. Fail IBSFail isfemale ismale IDno. Call 1 0 0 0 0 Het. Fail NA 3 0 0 0 IBSFail NA NA 1 0 0 isfemale NA NA NA 2 0 ismale NA NA NA 0 Day 2 Section 8 23

Data checking: output • The procedure provides the list of individuals (idok) and SNPs (snpok) who passed all QC criteria. • It is then possible to obtain a “clean” dataset: R code: data 1<-ge 03 d 2 ex[qc 1$idok, qc 1$snpok] Day 2 Section 8 24
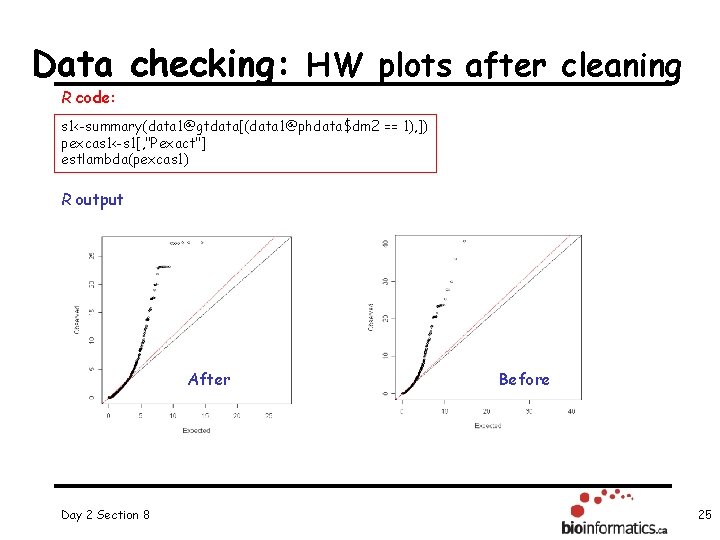
Data checking: HW plots after cleaning R code: s 1<-summary(data 1@gtdata[(data 1@phdata$dm 2 == 1), ]) pexcas 1<-s 1[, "Pexact"] estlambda(pexcas 1) R output After Day 2 Section 8 Before 25
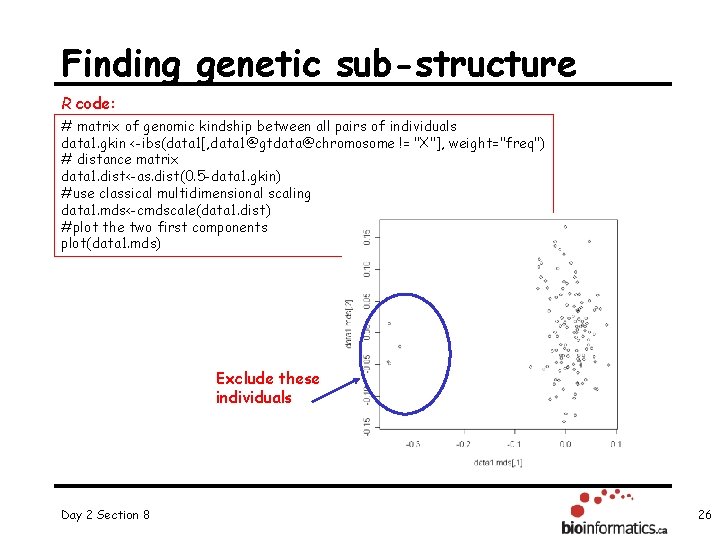
Finding genetic sub-structure R code: # matrix of genomic kindship between all pairs of individuals data 1. gkin <-ibs(data 1[, data 1@gtdata@chromosome != "X"], weight="freq") # distance matrix data 1. dist<-as. dist(0. 5 -data 1. gkin) #use classical multidimensional scaling data 1. mds<-cmdscale(data 1. dist) #plot the two first components plot(data 1. mds) Exclude these individuals Day 2 Section 8 26

Remove outliers R code: km<-kmeans(data 1. mds, centers=2, nstart=1000) cl 1<-names(which(km$cluster==1)) cl 2<-names(which(km$cluster==2)) data 2<-data 1[cl 1, ] Then, repeat the QC analysis allowing for HWE checks (using controls and exclude markers with FDR 0. 2) R code: qc 2<-check. marker(data 2, hweids=(data 2@phdata$dm 2 ==0), fdr=0. 2) summary(qc 2) R output Day 2 Section 8 27
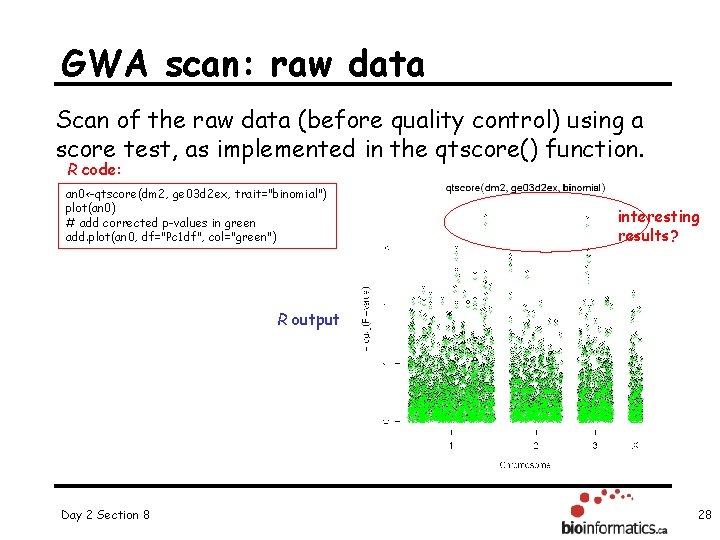
GWA scan: raw data Scan of the raw data (before quality control) using a score test, as implemented in the qtscore() function. R code: an 0<-qtscore(dm 2, ge 03 d 2 ex, trait="binomial") plot(an 0) # add corrected p-values in green add. plot(an 0, df="Pc 1 df", col="green") interesting results? R output Day 2 Section 8 28

GWA scan: raw data Scan of the raw data (before quality control) using a score test, as implemented in the qtscore() function. R code: #descriptive table descriptives. scan(an 0) R output: Top 10 results Day 2 Section 8 29
![GWA scan: cleaned data R code: data 2<-data 2[qc 2$idok, qc 2$snpok] # plot GWA scan: cleaned data R code: data 2<-data 2[qc 2$idok, qc 2$snpok] # plot](http://slidetodoc.com/presentation_image_h/2d8f7fe082bbdae00170bc72d77f9e79/image-30.jpg)
GWA scan: cleaned data R code: data 2<-data 2[qc 2$idok, qc 2$snpok] # plot an 1<-qtscore(dm 2, data 2, trait="binomial") plot(an 1) # add corrected p-values add. plot(an 1, df="Pc 1 df", col="green") interesting results R output Day 2 Section 8 30
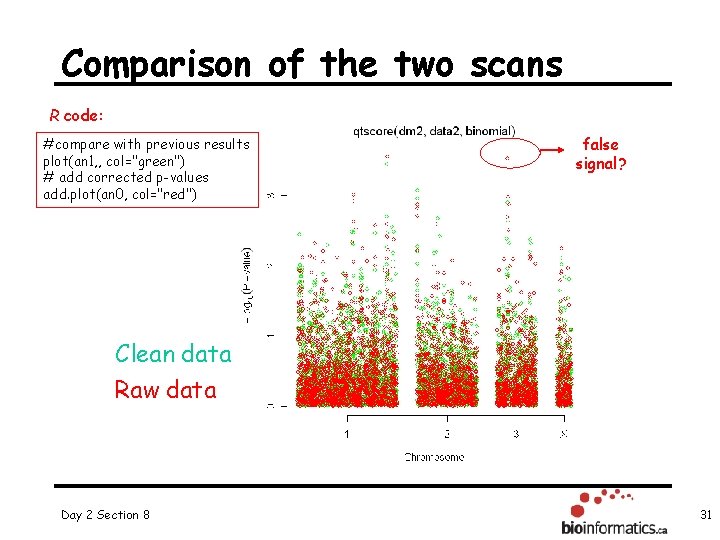
Comparison of the two scans R code: #compare with previous results plot(an 1, , col="green") # add corrected p-values add. plot(an 0, col="red") false signal? Clean data Raw data Day 2 Section 8 31

GWA scan: cleaned data R code: #descriptive table descriptives. scan(an 1) Clean data Raw data Day 2 Section 8 32
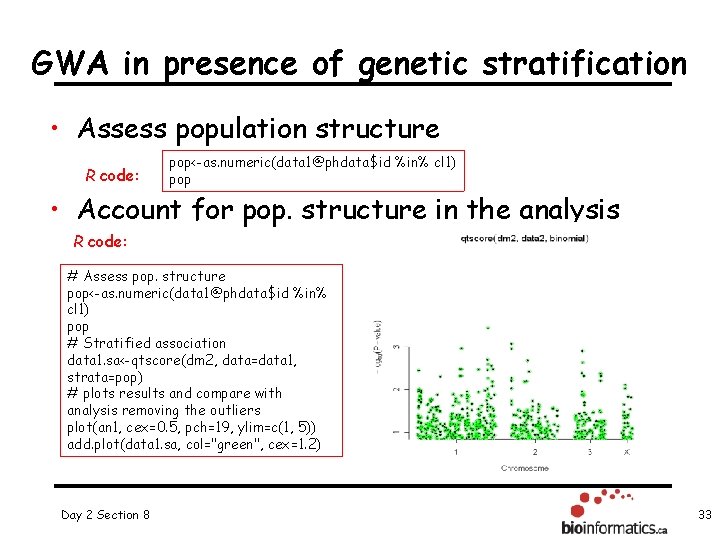
GWA in presence of genetic stratification • Assess population structure R code: pop<-as. numeric(data 1@phdata$id %in% cl 1) pop • Account for pop. structure in the analysis R code: # Assess pop. structure pop<-as. numeric(data 1@phdata$id %in% cl 1) pop # Stratified association data 1. sa<-qtscore(dm 2, data=data 1, strata=pop) # plots results and compare with analysis removing the outliers plot(an 1, cex=0. 5, pch=19, ylim=c(1, 5)) add. plot(data 1. sa, col="green", cex=1. 2) Day 2 Section 8 33
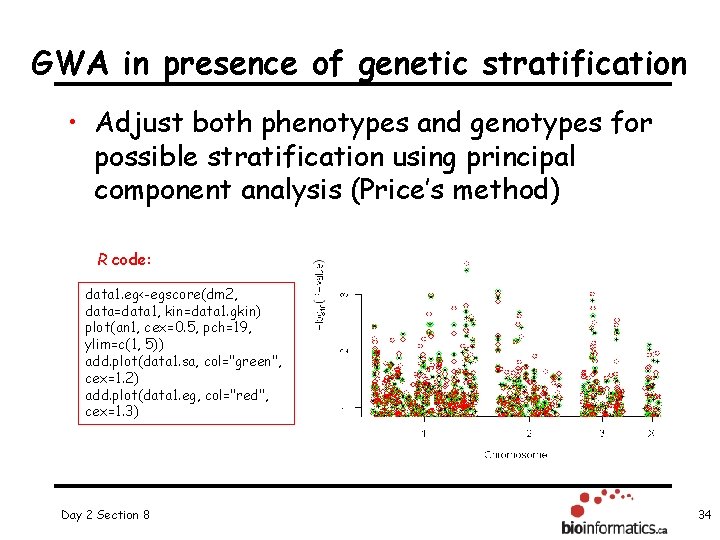
GWA in presence of genetic stratification • Adjust both phenotypes and genotypes for possible stratification using principal component analysis (Price’s method) R code: data 1. eg<-egscore(dm 2, data=data 1, kin=data 1. gkin) plot(an 1, cex=0. 5, pch=19, ylim=c(1, 5)) add. plot(data 1. sa, col="green", cex=1. 2) add. plot(data 1. eg, col="red", cex=1. 3) Day 2 Section 8 34

Other interesting features • • Genetic data imputations Meta-analysis of GWA scans Analysis of selected regions Conversion of plink files Day 2 Section 8 35

Conclusion • GWAS is becoming of major area of research • New computational tools and stat methods are needed • Gen. ABEL is an interesting program, especially for easy data cleaning and display of results • Plink has more features for stat analysis but not yet available in R for Windows! Day 2 Section 8 36

Thank you! Day 2 Section 8 37
- Slides: 37