CAMPBELL BIOLOGY IN FOCUS URRY CAIN WASSERMAN MINORSKY

CAMPBELL BIOLOGY IN FOCUS URRY • CAIN • WASSERMAN • MINORSKY • REECE 13 The Molecular Basis of Inheritance Questions prepared by Douglas Darnowski, Indiana University Southeast James Langeland, Kalamazoo College Murty S. Kambhampati, Southern University at New Orleans Roberta Batorsky, Temple University © 2016 Pearson Education, Inc. SECOND EDITION

Who conducted the X-ray diffraction studies that were key to the discovery of the structure of DNA? A. B. C. D. E. Griffith Franklin Meselson and Stahl Chargaff Mc. Clintock © 2016 Pearson Education, Inc.

Who conducted the X-ray diffraction studies that were key to the discovery of the structure of DNA? A. B. C. D. E. Griffith Franklin Meselson and Stahl Chargaff Mc. Clintock © 2016 Pearson Education, Inc.

How do the leading, and the lagging strands differ? A. The leading strand is synthesized in the same direction as the movement of the replication fork, and the lagging strand is synthesized in the opposite direction. B. The leading strand is synthesized at twice the rate of the lagging strand. C. The leading strand is synthesized in short fragments that are ultimately stitched together, whereas the lagging strand is synthesized continuously. D. The leading strand is synthesized by adding nucleotides to the 3′ end of the growing strand, and the lagging strand is synthesized by adding nucleotides to the 5′ end. © 2016 Pearson Education, Inc.

How do the leading, and the lagging strands differ? A. The leading strand is synthesized in the same direction as the movement of the replication fork, and the lagging strand is synthesized in the opposite direction. B. The leading strand is synthesized at twice the rate of the lagging strand. C. The leading strand is synthesized in short fragments that are ultimately stitched together, whereas the lagging strand is synthesized continuously. D. The leading strand is synthesized by adding nucleotides to the 3′ end of the growing strand, and the lagging strand is synthesized by adding nucleotides to the 5′ end. © 2016 Pearson Education, Inc.

What enzyme does a gamete-producing cell include that compensates for replication-associated shortening? A. B. C. D. E. DNA polymerase II ligase telomerase DNA nuclease proofreading enzyme © 2016 Pearson Education, Inc.

What enzyme does a gamete-producing cell include that compensates for replication-associated shortening? A. B. C. D. E. DNA polymerase II ligase telomerase DNA nuclease proofreading enzyme © 2016 Pearson Education, Inc.

Suppose a double-stranded DNA molecule was shown to have 15% adenine bases. What would be the expected percentage of guanine bases in that molecule? A. B. C. D. 15% 35% 85% not enough information © 2016 Pearson Education, Inc.

Suppose a double-stranded DNA molecule was shown to have 15% adenine bases. What would be the expected percentage of guanine bases in that molecule? A. B. C. D. 15% 35% 85% not enough information © 2016 Pearson Education, Inc.

Consider the replication bubble diagrammed at the right. Which letters represent leading strands? A. B. C. D. W and X Y and Z W and Z X and Y © 2016 Pearson Education, Inc. 3′ 5′ W Y X Z 5′ 3′
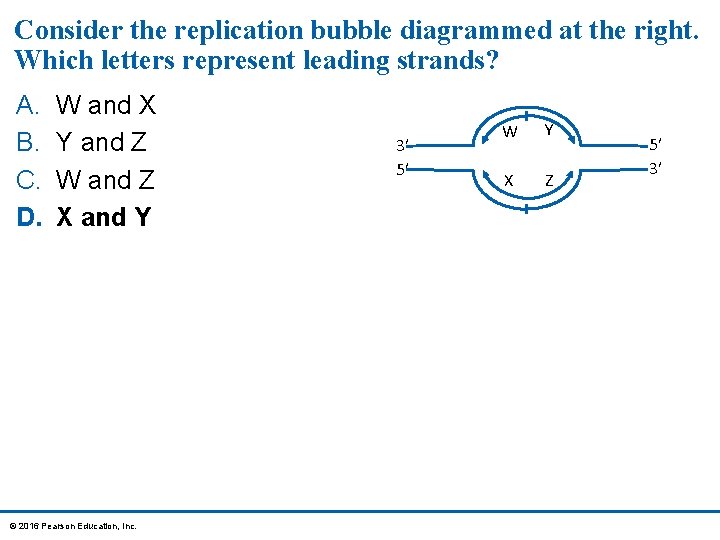
Consider the replication bubble diagrammed at the right. Which letters represent leading strands? A. B. C. D. W and X Y and Z W and Z X and Y © 2016 Pearson Education, Inc. 3′ 5′ W Y X Z 5′ 3′
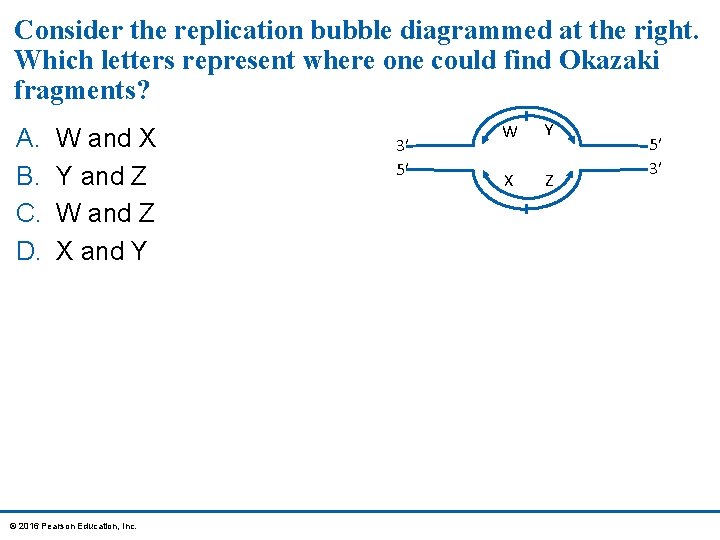
Consider the replication bubble diagrammed at the right. Which letters represent where one could find Okazaki fragments? A. B. C. D. W and X Y and Z W and Z X and Y © 2016 Pearson Education, Inc. 3′ 5′ W Y X Z 5′ 3′
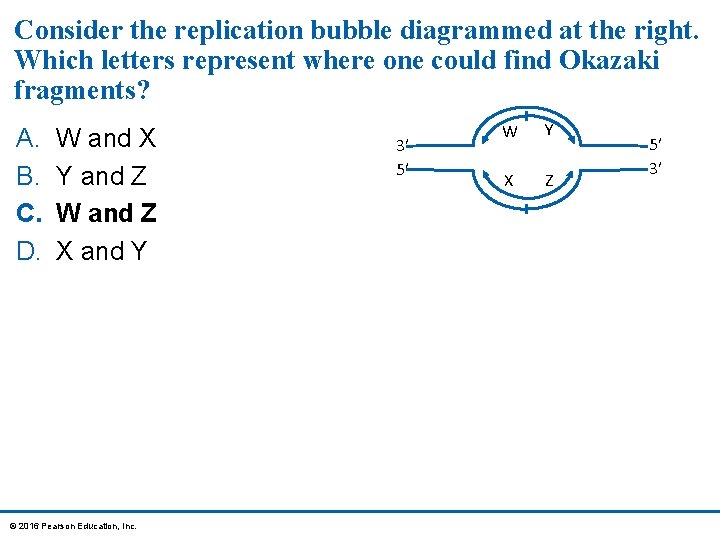
Consider the replication bubble diagrammed at the right. Which letters represent where one could find Okazaki fragments? A. B. C. D. W and X Y and Z W and Z X and Y © 2016 Pearson Education, Inc. 3′ 5′ W Y X Z 5′ 3′

Consider the replication bubble diagrammed at the right. Which diagram below depicts what this structure would look like when replication is complete? A. 3′ 5′ 5′ 3′ C. 3′ 5′ B. 5′ 3′ © 2016 Pearson Education, Inc. and 5′ 3′ D. 5′ 3′ 3′ 5′ and 5′ 3′
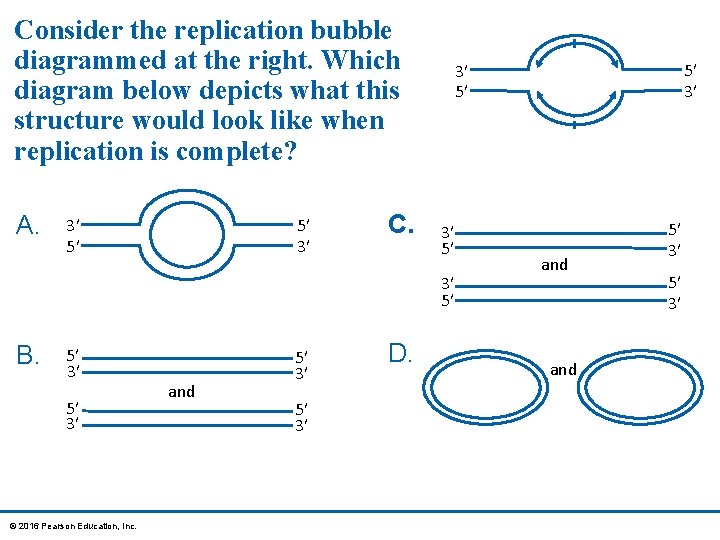
Consider the replication bubble diagrammed at the right. Which diagram below depicts what this structure would look like when replication is complete? A. 3′ 5′ 5′ 3′ C. 3′ 5′ B. 5′ 3′ © 2016 Pearson Education, Inc. and 5′ 3′ D. 5′ 3′ 3′ 5′ and 5′ 3′

Imagine a bacterial cell with a mutation that renders DNA Pol I completely nonfunctional (note that this would be a lethal mutation). What, precisely, would go wrong with replication in this cell? A. inability to unwind double helix B. inability to prime replication C. inability to extend the length of leading and lagging strands D. inability to replace primers © 2016 Pearson Education, Inc.

Imagine a bacterial cell with a mutation that renders DNA Pol I completely nonfunctional (note that this would be a lethal mutation). What, precisely, would go wrong with replication in this cell? A. inability to unwind double helix B. inability to prime replication C. inability to extend the length of leading and lagging strands D. inability to replace primers © 2016 Pearson Education, Inc.

DNA replication overall has very high fidelity. Which of the following phenomena or processes contribute to this high fidelity? More than one may apply. A. base pairing B. proofreading C. mismatch repair © 2016 Pearson Education, Inc.

DNA replication overall has very high fidelity. Which of the following phenomena or processes contribute to this high fidelity? More than one may apply. A. base pairing B. proofreading C. mismatch repair © 2016 Pearson Education, Inc.

Which of the following would typically not be used to clone DNA? A. B. C. D. plasmid vector telomerase restriction enzyme PCR © 2016 Pearson Education, Inc.

Which of the following would typically not be used to clone DNA? A. B. C. D. plasmid vector telomerase restriction enzyme PCR © 2016 Pearson Education, Inc.

Arrange the following terms in order, from smallest to largest size: nucleosome, metaphase chromosome, histone, 30 -nm fiber. A. histone, nucleosome, 30 -nm fiber, metaphase chromosome B. nucleosome, histone, 30 -nm fiber, metaphase chromosome C. histone, nucleosome, metaphase chromosome, 30 -nm fiber D. 30 -nm fiber, histone, nucleosome, metaphase chromosome © 2016 Pearson Education, Inc.

Arrange the following terms in order, from smallest to largest size: nucleosome, metaphase chromosome, histone, 30 -nm fiber. A. histone, nucleosome, 30 -nm fiber, metaphase chromosome B. nucleosome, histone, 30 -nm fiber, metaphase chromosome C. histone, nucleosome, metaphase chromosome, 30 -nm fiber D. 30 -nm fiber, histone, nucleosome, metaphase chromosome © 2016 Pearson Education, Inc.
- Slides: 23