ca BIG the cancer Biomedical Informatics Grid Ken

ca. BIG: the cancer Biomedical Informatics Grid Ken Buetow NCICB/NCI/NIH/DHHS

NCI biomedical informatics § Goal: A virtual web of interconnected data, individuals, and organizations redefines how research is conducted, care is provided, and patients/participants interact with the biomedical research enterprise
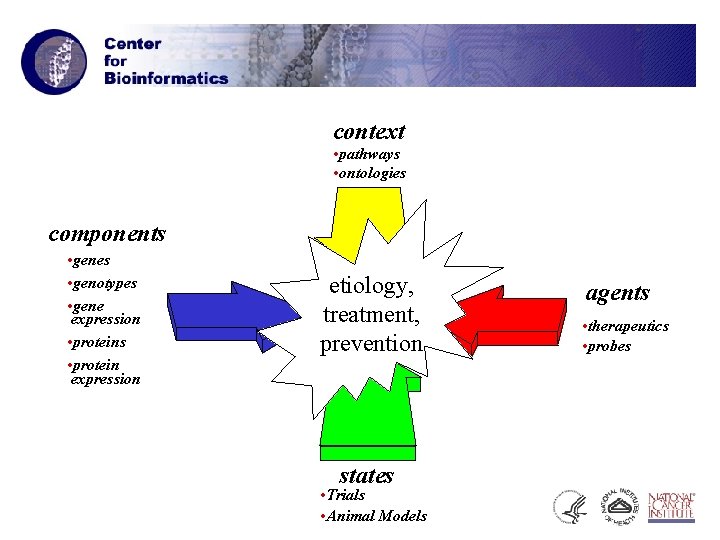
context • pathways • ontologies components • genes • genotypes • gene expression • proteins • protein expression etiology, treatment, prevention states • Trials • Animal Models agents • therapeutics • probes
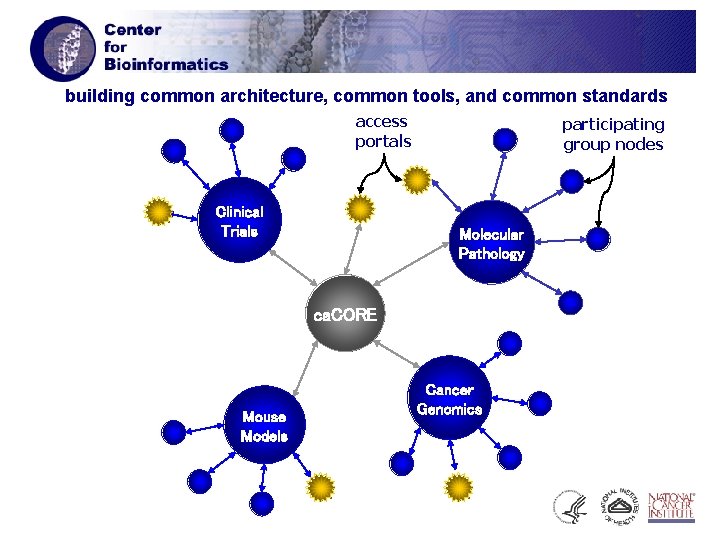
building common architecture, common tools, and common standards access portals Clinical Trials participating group nodes Molecular Pathology ca. CORE Mouse Models Cancer Genomics

Interoperability Courtesy: Charlie Mead § in·ter·op·er·a·bil·i·ty - ability of a system. . . to use the parts or equipment of another system Source: Merriam-Webster web site § interoperability - ability of two or more systems or components to exchange information and to use the information that has been exchanged. Source: IEEE Standard Computer Dictionary: A Compilation of IEEE Standard Computer Glossaries, IEEE, 1990] Syntactic interoperability Semantic interoperability
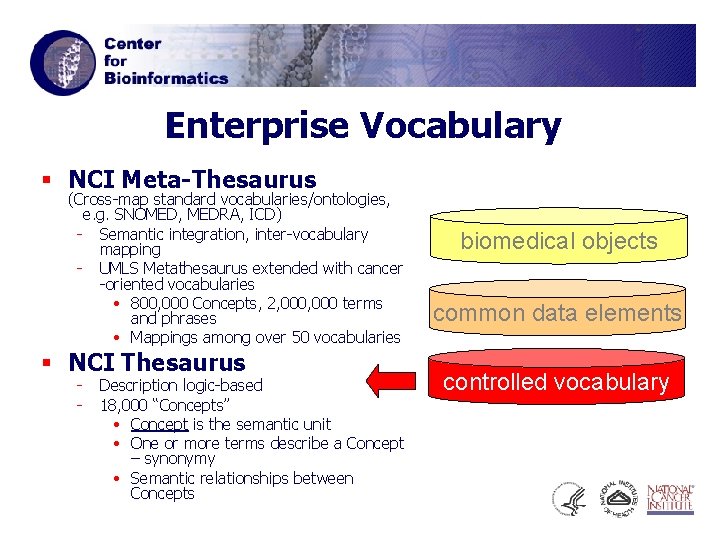
Enterprise Vocabulary § NCI Meta-Thesaurus (Cross-map standard vocabularies/ontologies, e. g. SNOMED, MEDRA, ICD) - Semantic integration, inter-vocabulary mapping - UMLS Metathesaurus extended with cancer -oriented vocabularies • 800, 000 Concepts, 2, 000 terms and phrases • Mappings among over 50 vocabularies § NCI Thesaurus - Description logic-based 18, 000 “Concepts” • Concept is the semantic unit • One or more terms describe a Concept – synonymy • Semantic relationships between Concepts biomedical objects common data elements controlled vocabulary
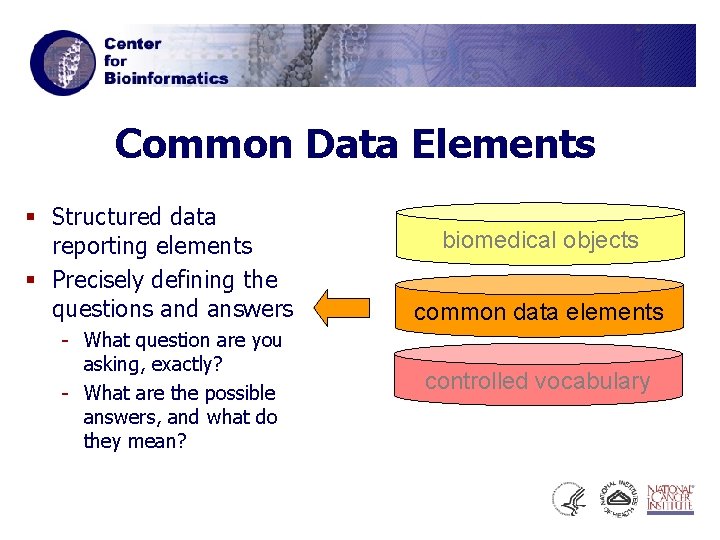
Common Data Elements § Structured data reporting elements § Precisely defining the questions and answers - What question are you asking, exactly? - What are the possible answers, and what do they mean? biomedical objects common data elements controlled vocabulary
Biomedical Information Objects § § § Data service infrastructure developed using OMG’s Model Driven Architecture approach Object models expressed in UML represent actual biomedical research entities such as genes, sequences, chromosomes, sequences, cellular pathways, ontologies, clinical protocols, etc. The object models form the basis for uniform APIs (Java, SOAP, HTTP-XML, Perl) that provide an abstraction layer and interfaces for developers to access information without worrying about the backend data stores biomedical objects common data elements controlled vocabulary

Standards supporting infrastructure § Enterprise Vocabulary Services (EVS) - Browsers - APIs § cancer Bioinformatics Infrastructure Objects (ca. BIO) - Applications - APIs § cancer Data Standards Repository (ca. DSR) - CDEs Case Report Forms Object models ISO 11179 model
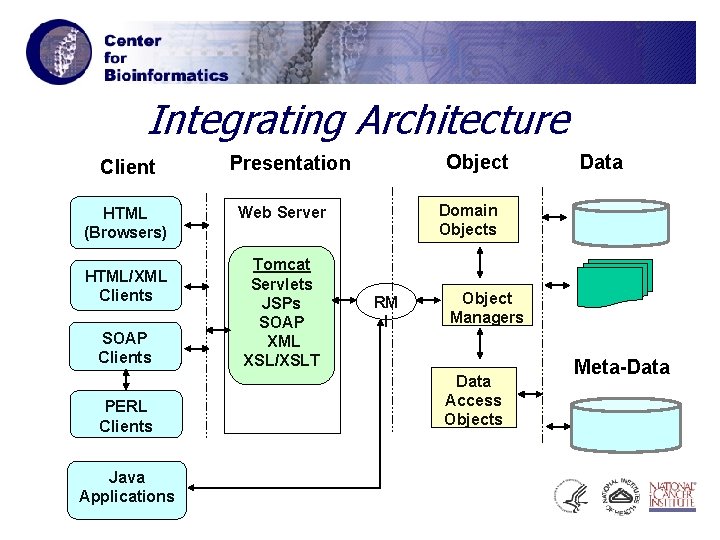
Integrating Architecture Client HTML (Browsers) HTML/XML Clients SOAP Clients PERL Clients Java Applications Object Presentation Domain Objects Web Server Tomcat Servlets JSPs SOAP XML XSL/XSLT Data RM I Object Managers Data Access Objects Meta-Data
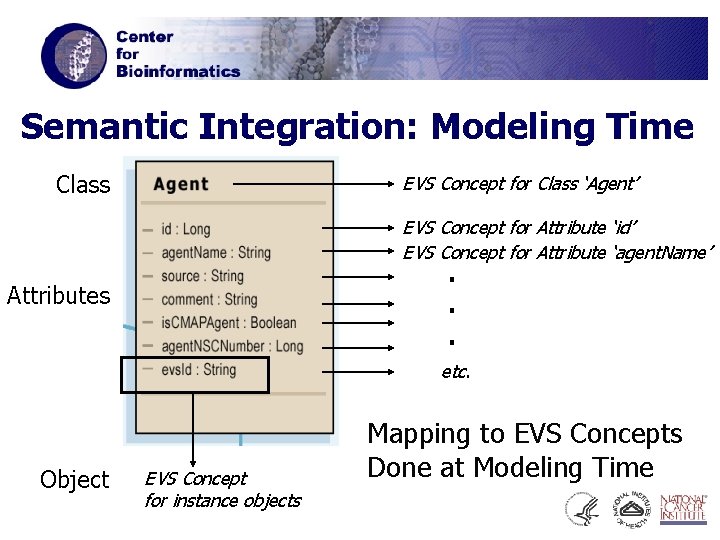
Semantic Integration: Modeling Time Class EVS Concept for Class ‘Agent’ EVS Concept for Attribute ‘id’ EVS Concept for Attribute ‘agent. Name’ . . . Attributes etc. Object EVS Concept for instance objects Mapping to EVS Concepts Done at Modeling Time
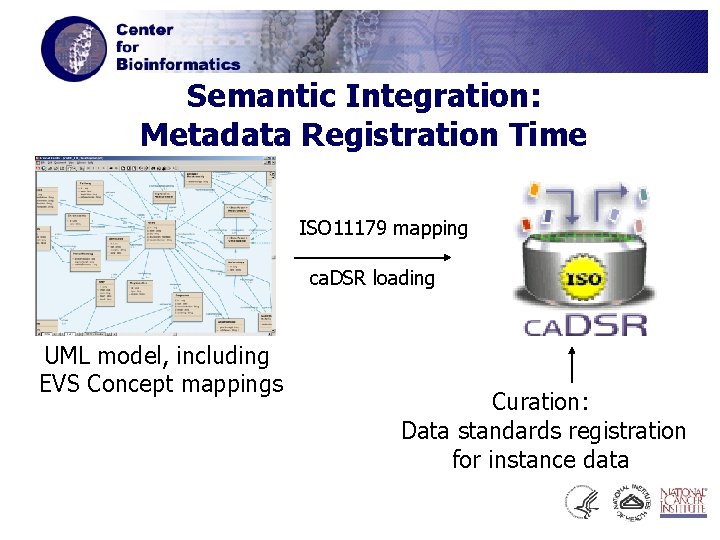
Semantic Integration: Metadata Registration Time ISO 11179 mapping ca. DSR loading UML model, including EVS Concept mappings Curation: Data standards registration for instance data
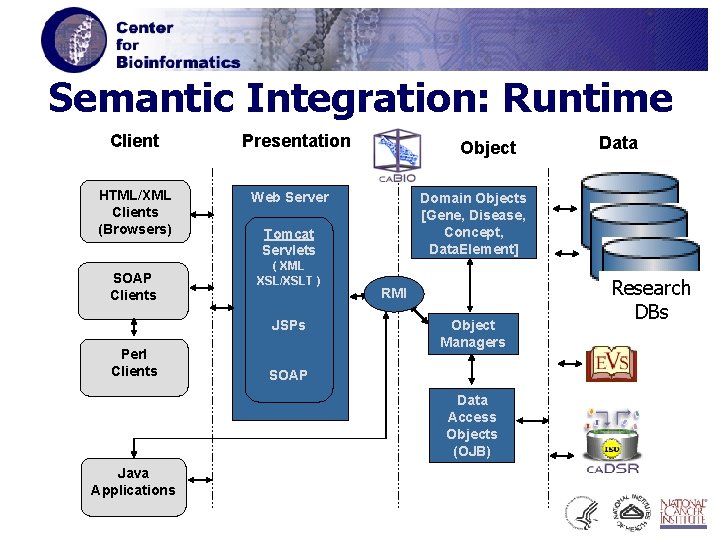
Semantic Integration: Runtime Client HTML/XML Clients (Browsers) SOAP Clients Presentation Web Server Domain Objects [Gene, Disease, Concept, Data. Element] Tomcat Servlets ( XML XSL/XSLT ) JSPs Perl Clients Object RMI Object Managers SOAP Data Access Objects (OJB) Java Applications Data Research DBs
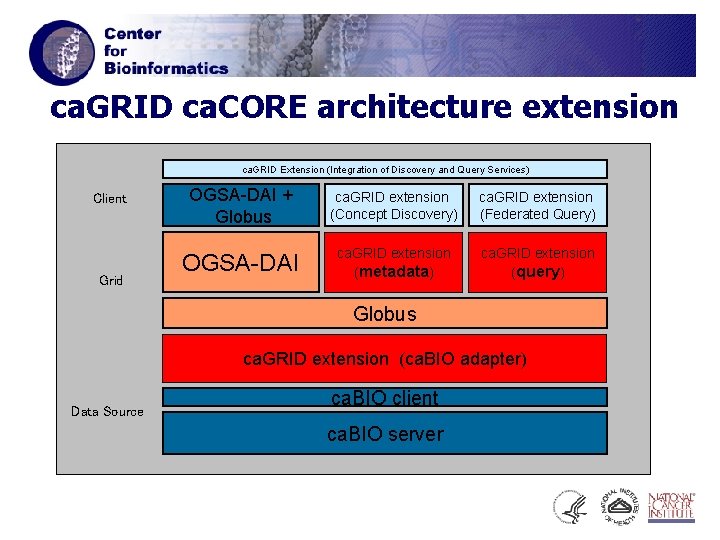
ca. GRID ca. CORE architecture extension ca. GRID Extension (Integration of Discovery and Query Services) Client Grid OGSA-DAI + Globus ca. GRID extension (Concept Discovery) ca. GRID extension (Federated Query) OGSA-DAI ca. GRID extension (metadata) ca. GRID extension (query) Globus ca. GRID extension (ca. BIO adapter) Data Source ca. BIO client ca. BIO server

NCICB applications: • clincial trials support - C 3 DS • molecular pathology - ca. Array • cancer images - ca. Image • pre-clinical models - ca. Models. Db • laboratory support - ca. LIMS

Standards-based Data System for the conduct of clinical trials: • • • C 3 D (Cancer Central Clinical Database) – WWW-based e. CRF-based primary data capture by protocol C 3 PR (Cancer Central Clinical Participant Registry) – WWW-based Central registration of participants across protocols C 3 PA (Cancer Central Clinical Protocol Administration) – Scientific management system for clinical protocols C 3 TR (Cancer Central Clinical Tissue Repository) – Tissue repository C 3 DW (Cancer Central Clinical Data Warehouse) – De-identified patient information accessed via ca. BIO



Image Portal • The NCICB has developed an image portal to allow researchers to search for mouse and human images and annotations – Human and mouse images and annotations were provided by the MMHCC
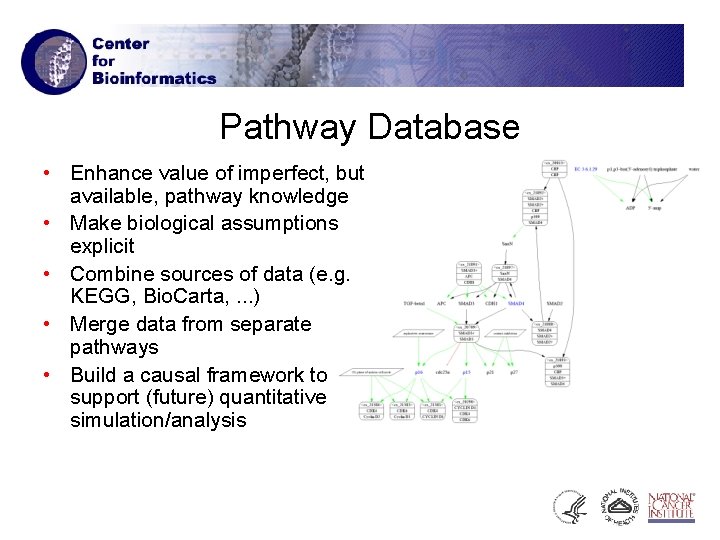
Pathway Database • Enhance value of imperfect, but available, pathway knowledge • Make biological assumptions explicit • Combine sources of data (e. g. KEGG, Bio. Carta, . . . ) • Merge data from separate pathways • Build a causal framework to support (future) quantitative simulation/analysis

Cancer Biomedical Informatics Grid (ca. BIG) § Common, widely distributed infrastructure permits cancer research community to focus on innovation § Shared vocabulary, data elements, data models facilitate information exchange § Collection of interoperable applications developed to common standard § Raw published cancer research data is available for mining and integration
ca. BIG will facilitate sharing of infrastructure, applications, and data

ca. BIG action plan § Establish pilot network of Cancer Centers - Groups agreeing to ca. BIG principles - Mixture of capabilities - Mixture of contributions § Expanding collection of participants § Establish consortium development process - Collecting and sharing expertise - Identifying and prioritizing community needs - Expanding development efforts § Moving at the speed of the internet…

Three Domain Workspaces and two Cross Cutting Workspaces have been launched during the Pilot phase DOMAIN WORKSPACE 1 Clinical Trial Management Systems addresses the need for consistent, open and comprehensive tools for clinical trials management. DOMAIN WORKSPACE 2 Integrative Cancer Research provides tools and systems to enable integration and sharing of information. DOMAIN WORKSPACE 3 Tissue Banks & Pathology Tools provides for the integration, development, and implementation of tissue and pathology tools. responsible for evaluating, developing, and integrating systems for vocabulary and ontology content, standards, and software systems for content delivery CROSS CUTTING WORKSPACE 1 Vocabularies & Common Data Elements developing architectural standards and architecture necessary for other workspaces. CROSS CUTTING WORKSPACE 2 Architecture

Key deliverables of ca. BIG pilot § Componentized, standards-based Clinical Trials Management System - e-IND filing/regulatory reporting with FDA - Electronic management of trials - Integration of diverse trials § Tissue Management System - Systematic description and characterization of tissue resources - Ability to link tissue resources to clinical and molecular correlative descriptions § “Plug and Play” analytic tool set - microarray proteomics pathways data analysis and statistical methods gene annotation § Diverse library of raw, structured data
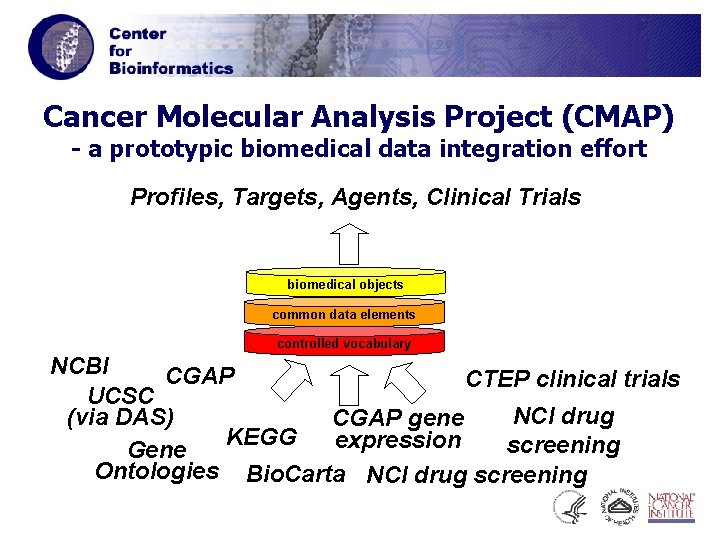
Cancer Molecular Analysis Project (CMAP) - a prototypic biomedical data integration effort Profiles, Targets, Agents, Clinical Trials biomedical objects common data elements controlled vocabulary NCBI CGAP CTEP clinical trials UCSC NCI drug (via DAS) CGAP gene KEGG expression screening Gene Ontologies Bio. Carta NCI drug screening


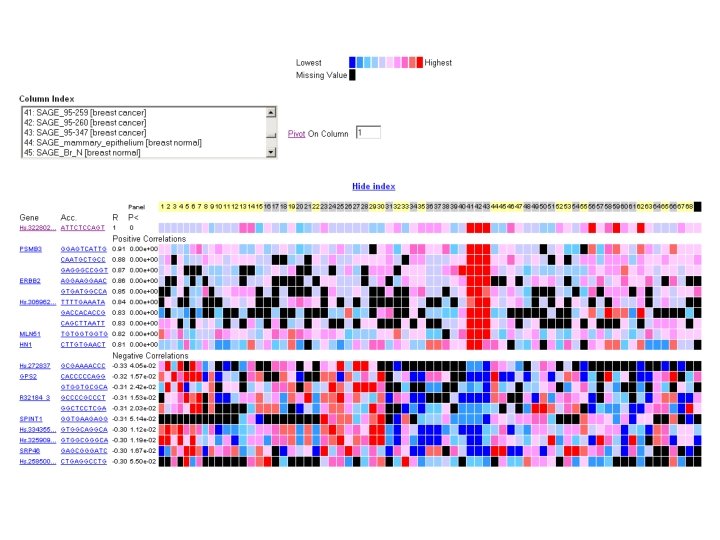


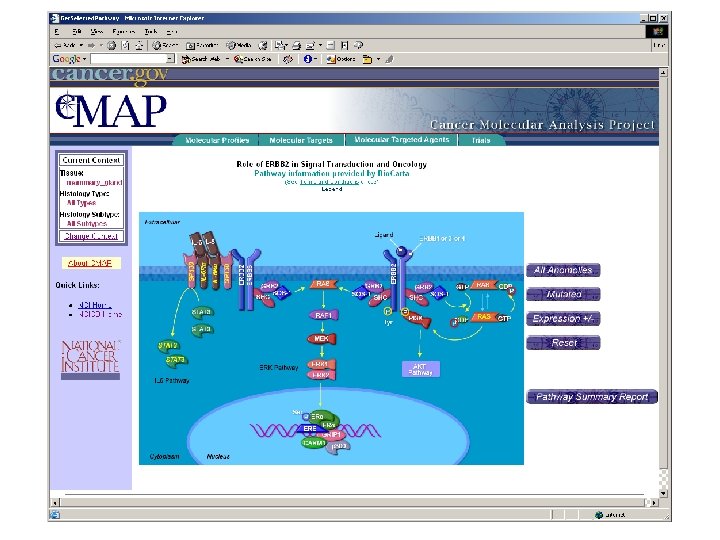
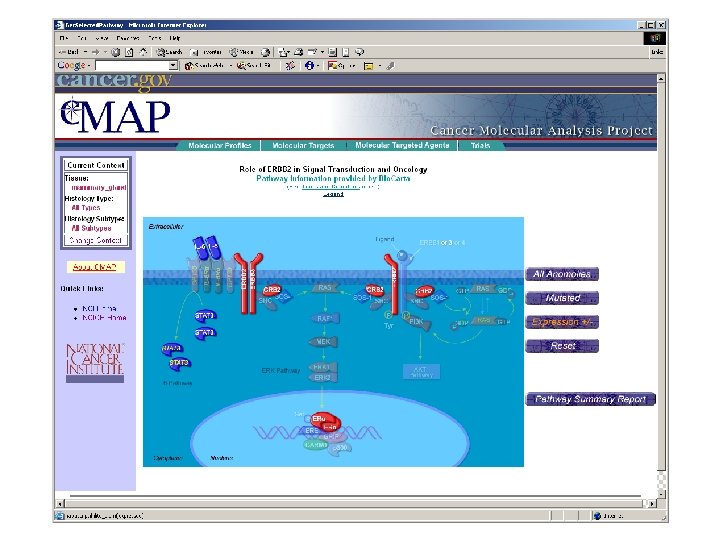


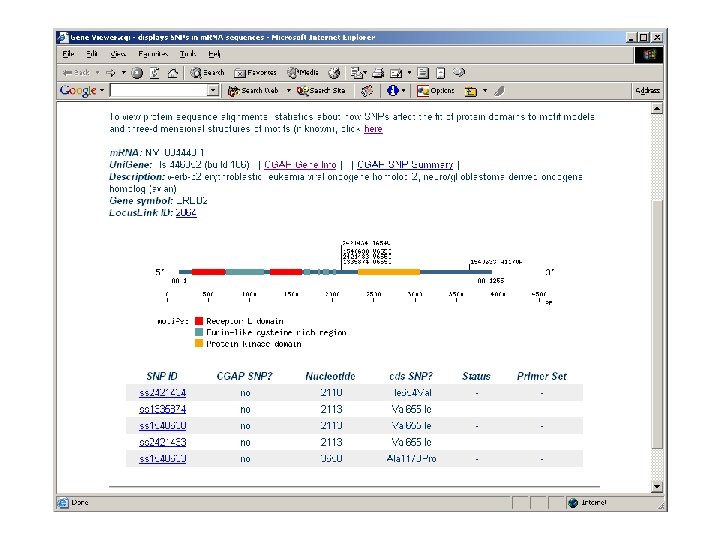



ca. BIG community contributions § Infrastructure - Ontologies - Databases § Applications - Clinical trials support - Analytic tools - Data mining § Data - Trials - Experimental outcomes • Genomic • Microarray • Proteomic


acknowledgements § § NCICB - Peter Covitz - Sue Dubman - Mary Jo Deering - Leslie Derr - Carl Schaefer - Christos Andonyadis - Mervi Heiskanen - Denise Hise - Kotien Wu - Fei Xu - Frank Hartel LPG/CCR - Michael Edmundson - Bob Clifford - Cu Nguyen http: //ncicb. nci. nih. gov http: //cmap. nci. nih. gov http: //ca. BIG. nci. nih. gov
- Slides: 41