C elegans RNASeq experimental design 31518 Lab Methods

C. elegans RNA-Seq experimental design 3/15/18 Lab Methods in Genomics
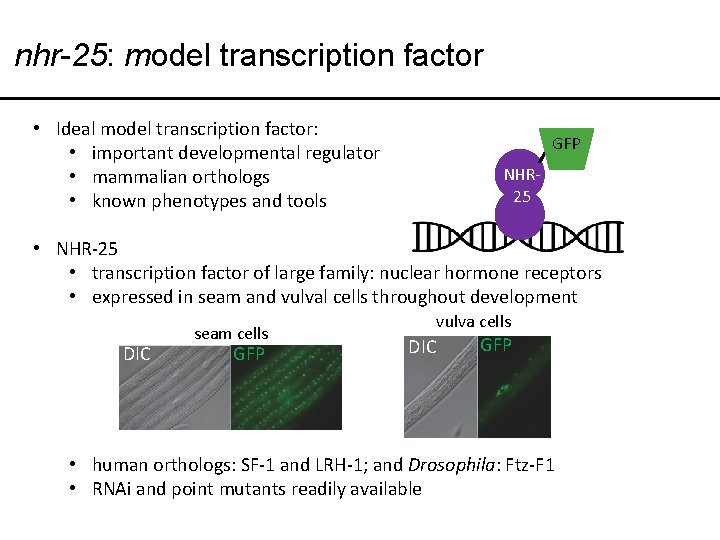
nhr-25: model transcription factor • Ideal model transcription factor: • important developmental regulator • mammalian orthologs • known phenotypes and tools GFP NHR 25 • NHR-25 • transcription factor of large family: nuclear hormone receptors • expressed in seam and vulval cells throughout development DIC seam cells GFP vulva cells DIC GFP • human orthologs: SF-1 and LRH-1; and Drosophila: Ftz-F 1 • RNAi and point mutants readily available
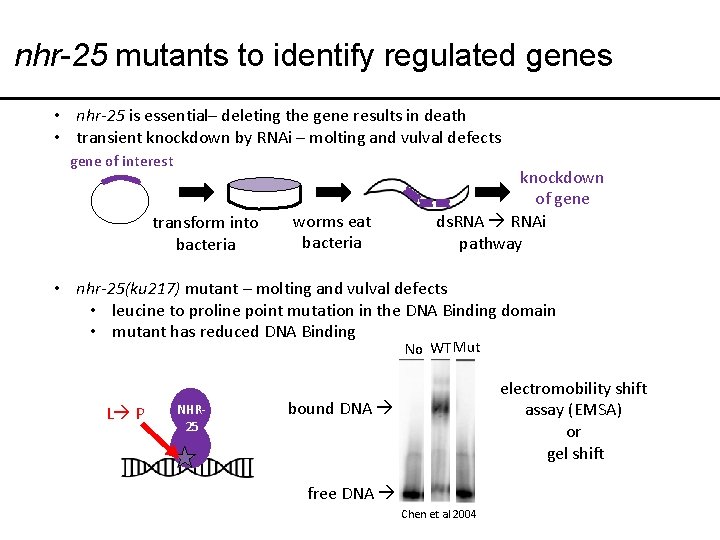
nhr-25 mutants to identify regulated genes • nhr-25 is essential– deleting the gene results in death • transient knockdown by RNAi – molting and vulval defects gene of interest transform into bacteria worms eat bacteria knockdown of gene ds. RNAi pathway • nhr-25(ku 217) mutant – molting and vulval defects • leucine to proline point mutation in the DNA Binding domain • mutant has reduced DNA Binding No WT Mut L P NHR 25 electromobility shift assay (EMSA) or gel shift bound DNA free DNA Chen et al 2004
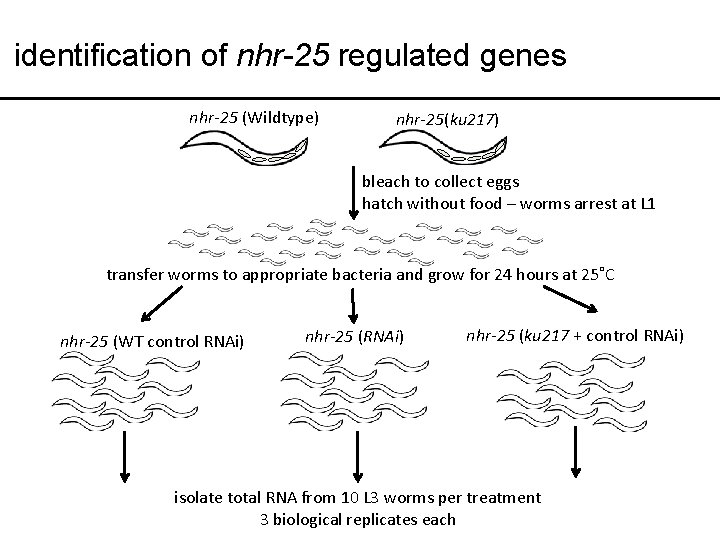
identification of nhr-25 regulated genes nhr-25 (Wildtype) nhr-25(ku 217) bleach to collect eggs hatch without food – worms arrest at L 1 transfer worms to appropriate bacteria and grow for 24 hours at 25˚C nhr-25 (WT control RNAi) nhr-25 (ku 217 + control RNAi) isolate total RNA from 10 L 3 worms per treatment 3 biological replicates each
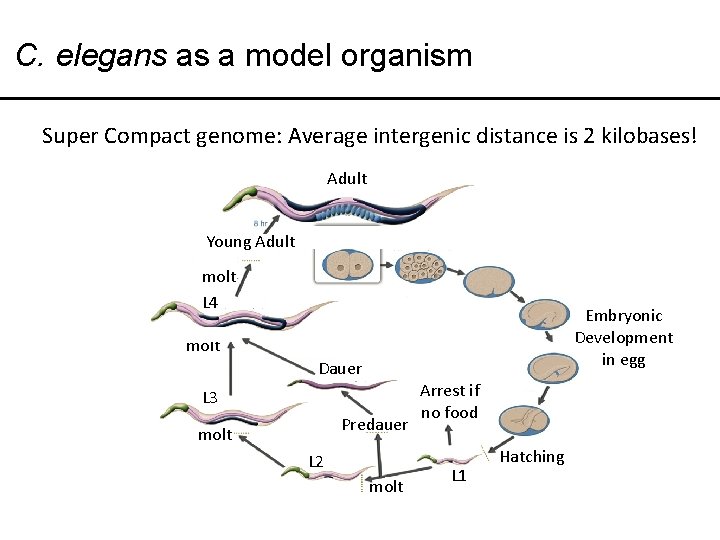
C. elegans as a model organism Super Compact genome: Average intergenic distance is 2 kilobases! Adult Young Adult molt L 4 Embryonic Development in egg molt Dauer L 3 Predauer molt L 2 molt Arrest if no food L 1 Hatching
- Slides: 5