c DNA Microarrays an introduction Henrik Bengtsson hbmaths

c. DNA Microarrays - an introduction Henrik Bengtsson hb@maths. lth. se Bioinformatics Group Mathematical Statistics, Centre for Mathematical Sciences Lund University

Outline • • • The Genomic Code The Central Dogma of Biology The c. DNA Microarray Technique Data Analysis of c. DNA Microarray Data Statistical Problems Take-home message
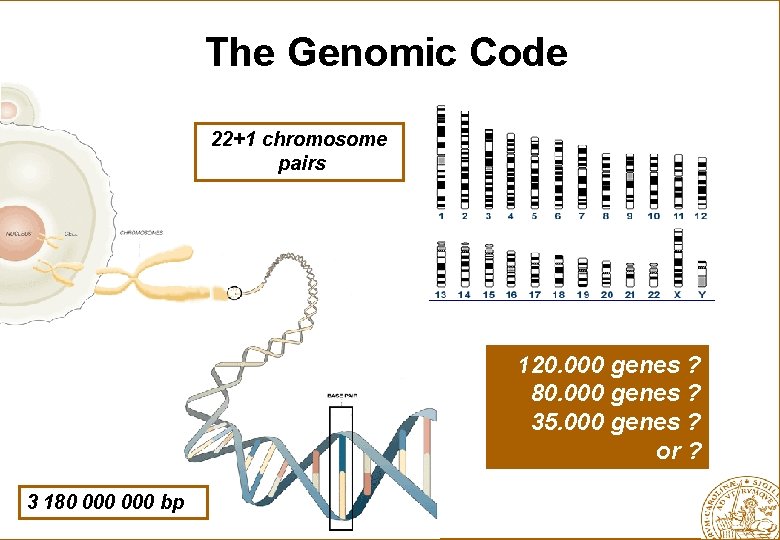
The Genomic Code 22+1 chromosome pairs 120. 000 genes ? 80. 000 genes ? 35. 000 genes ? or ? 3 180 000 bp
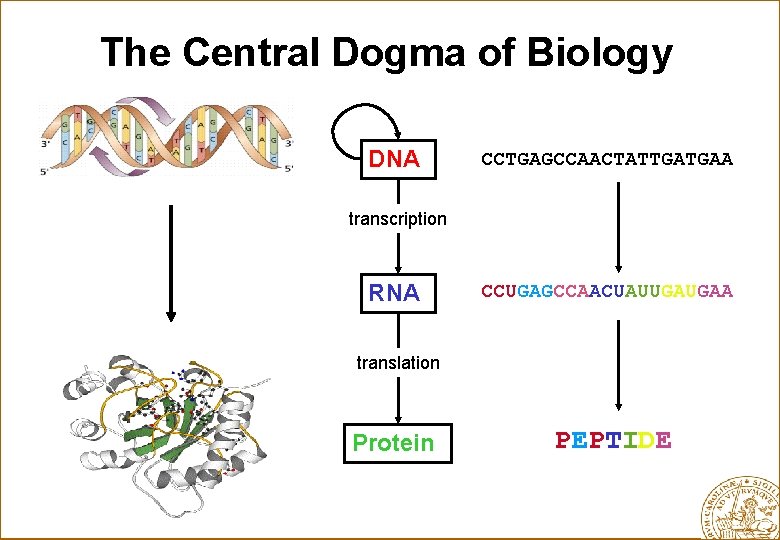
The Central Dogma of Biology DNA CCTGAGCCAACTATTGATGAA transcription RNA CCUGAGCCAACUAUUGAUGAA translation Protein PEPTIDE

The c. DNA Microarray Technique • High-throughput measuring - 5000 -20000 gene expressions at the same time • Identify genes that behaves different in different cell populations - tumor cells vs healthy cells - brain cells vs liver cells - same tissue different organisms • Time series experiments - gene expressions over time after treatment • . . .
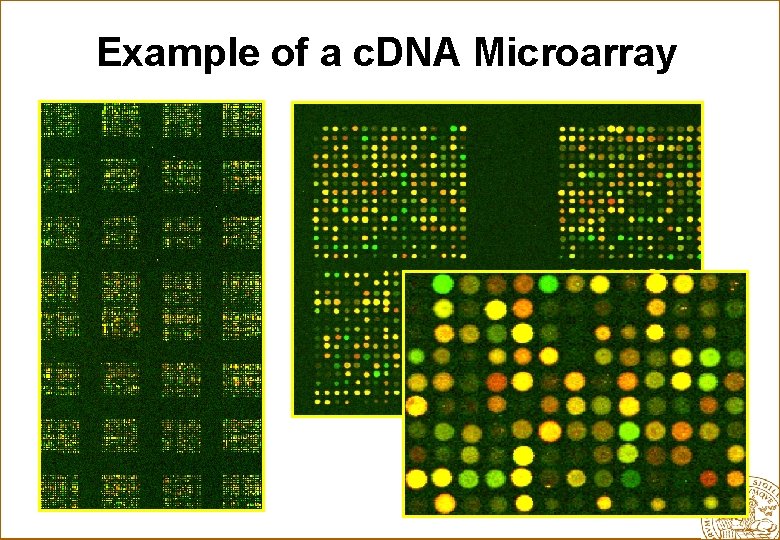
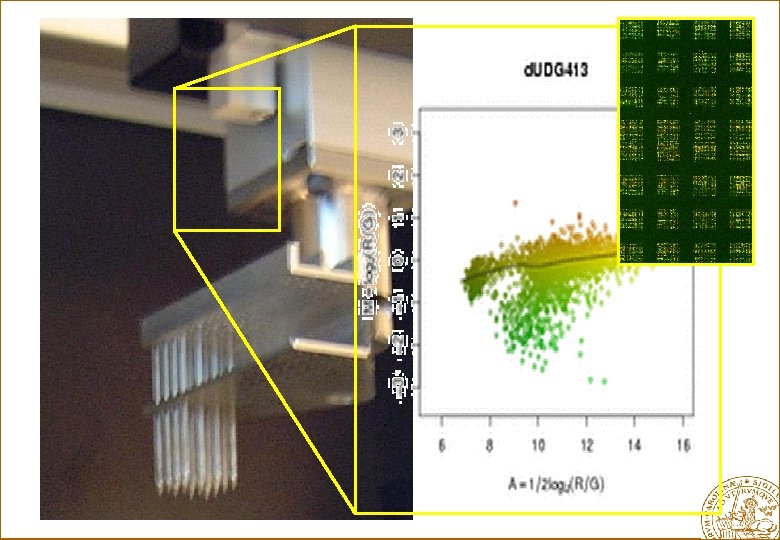
Example of a c. DNA Microarray
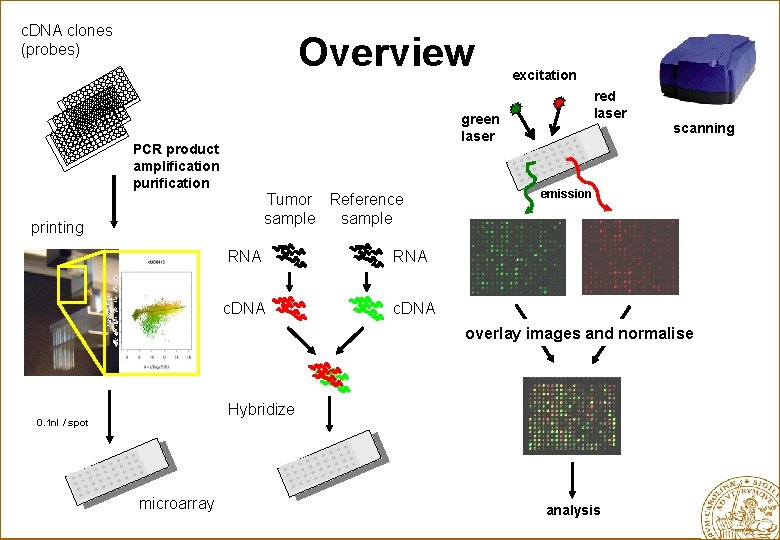
c. DNA clones (probes) Overview excitation red laser green laser PCR product amplification purification Tumor Reference sample printing RNA c. DNA scanning emission overlay images and normalise Hybridize 0. 1 nl / spot microarray analysis

Creating the slides
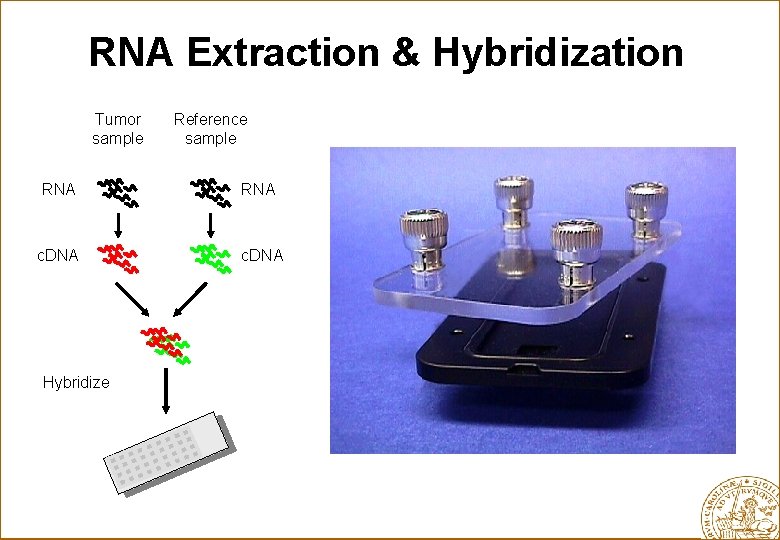
RNA Extraction & Hybridization Tumor sample Reference sample RNA c. DNA Hybridize
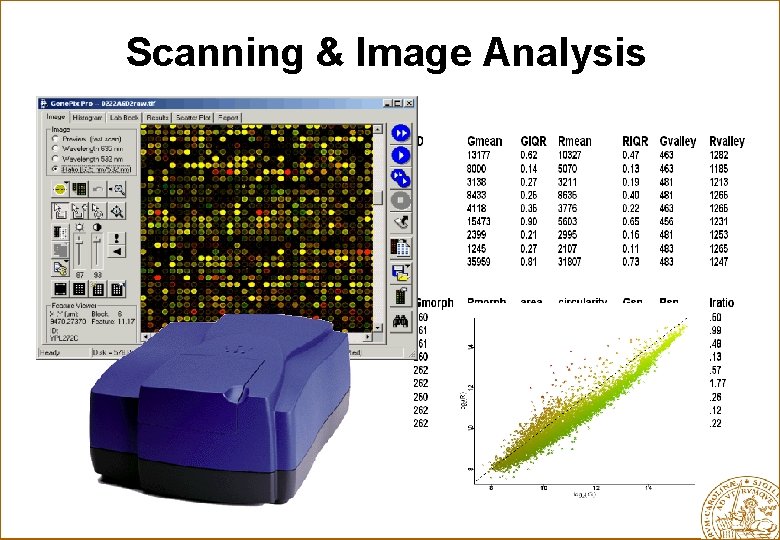
Scanning & Image Analysis
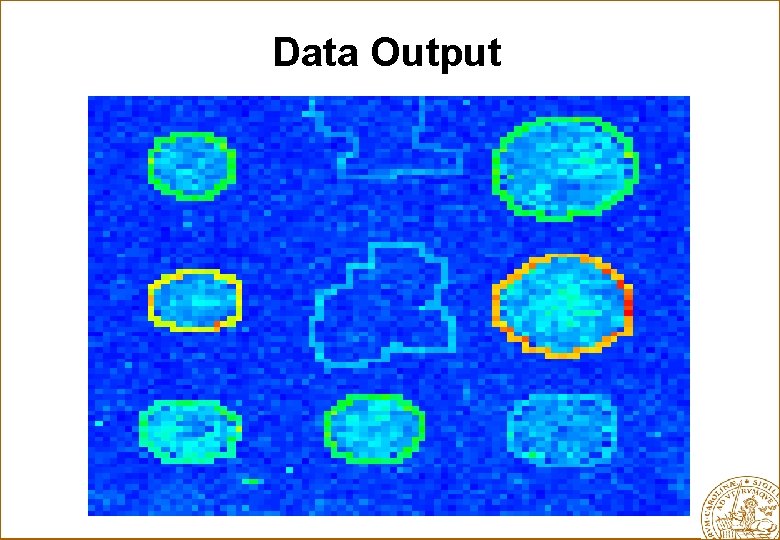
Data Output
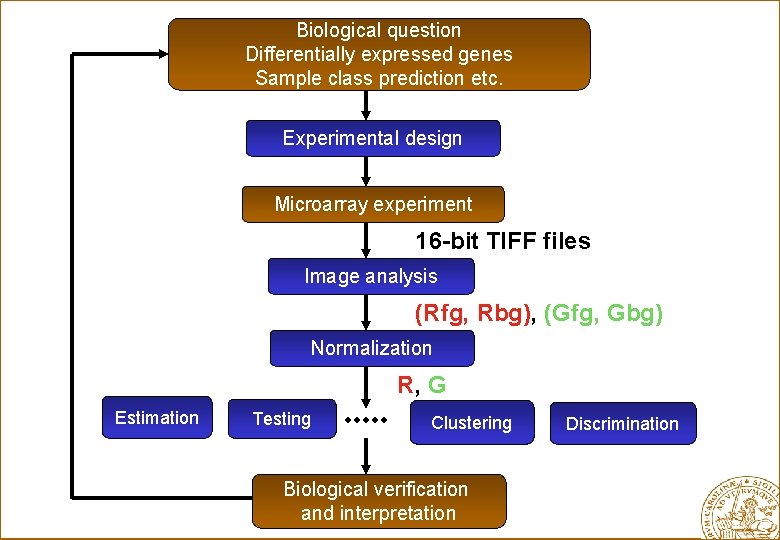
Biological question Differentially expressed genes Sample class prediction etc. Experimental design Microarray experiment 16 -bit TIFF files Image analysis (Rfg, Rbg), (Gfg, Gbg) Normalization R, G Estimation Testing Clustering Biological verification and interpretation Discrimination
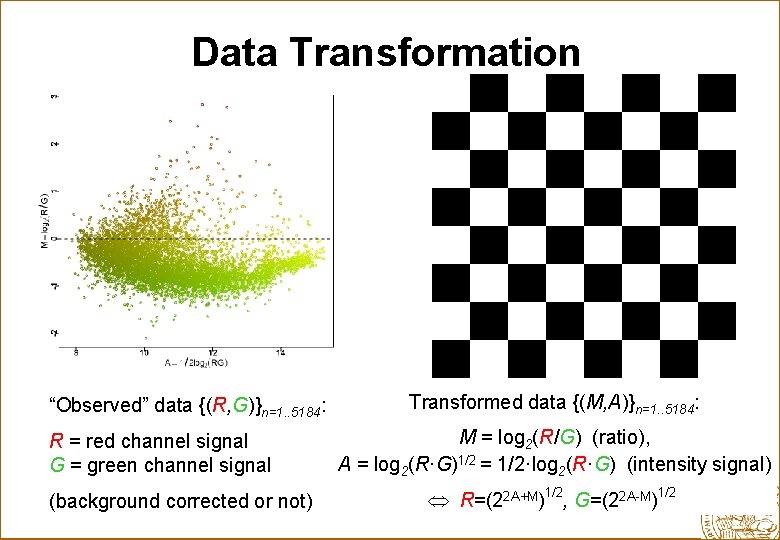
Data Transformation “Observed” data {(R, G)}n=1. . 5184: R = red channel signal G = green channel signal (background corrected or not) Transformed data {(M, A)}n=1. . 5184: M = log 2(R/G) (ratio), A = log 2(R·G)1/2 = 1/2·log 2(R·G) (intensity signal) R=(22 A+M)1/2, G=(22 A-M)1/2
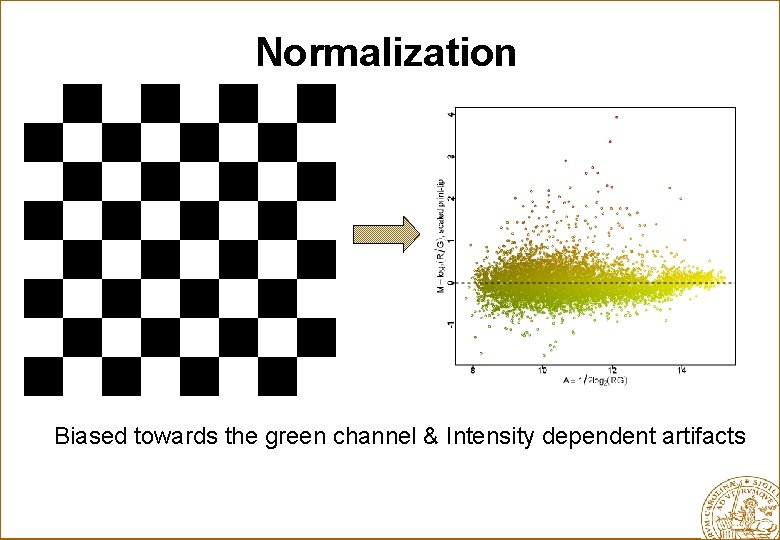
Normalization Biased towards the green channel & Intensity dependent artifacts
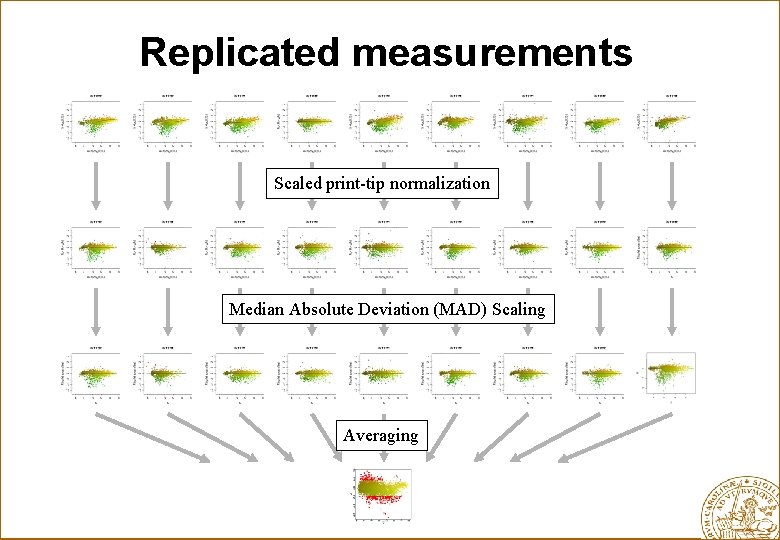
Replicated measurements Scaled print-tip normalization Median Absolute Deviation (MAD) Scaling Averaging
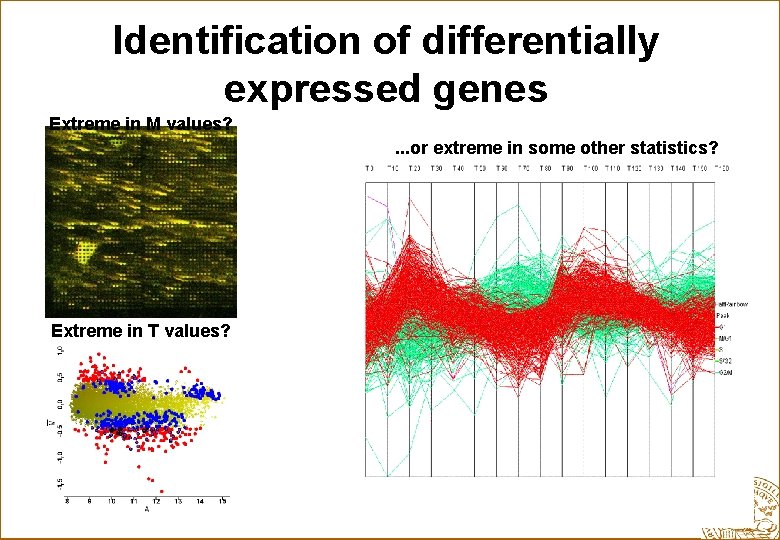
Identification of differentially expressed genes Extreme in M values? . . . or extreme in some other statistics? Extreme in T values?

List of genes that the biologist can understand verify with other experiments Gene: Mavg Aavg T SE 2341 -0. 86 10. 9 6412 -0. 75 11. 1 6123 -0. 70 9. 8 102 0. 65 10. 3 2020 0. 64 9. 3 3132 0. 62 9. 9 4439 -0. 62 9. 7 2031 -0. 61 10. 7 657 -0. 60 9. 2 502 0. 58 10. 0 1239 -0. 58 9. 8 5392 -0. 57 9. 9 3921 0. 52 11. 3. . . -18. 0 -14. 7 -12. 2 -14. 5 -11. 9 -14. 4 -14. 6 -13. 7 -13. 6 -12. 7 -11. 4 -20. 7 13. 5 0. 125 0. 102 0. 121 0. 136 0. 118 0. 090 0. 088 0. 087 0. 094 0. 101 0. 103 0. 057 0. 083
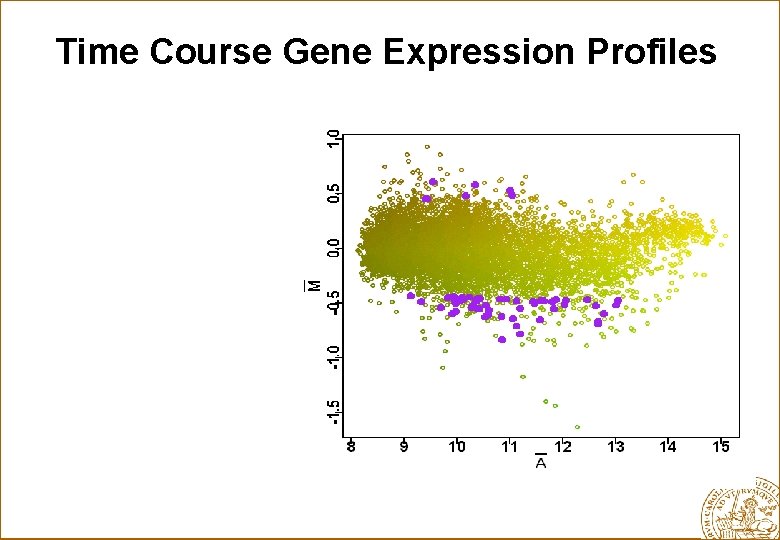
Time Course Gene Expression Profiles

Statistical Problems 1. Image analysis - what is foreground? - what is background? 2. Quality - which spots can we trust? - which slides can we trust? 3. Artifacts from preparing the RNA, the printing, the scanning etc. 4. Data cleanup 5. Normalization within an experiment: - when few genes change. - when many genes change. - dye-swap to minimize dye effects. 6. Normalization between experiments: - location and scale effects. 7. What is noise and what is variability? 10. Which genes are actually up- and down regulated? 11. P-values. 12. Planning of experiments: - what is best design? - what is an optimal sample sizes? 13. Classification: - of samples. - of genes. 14. Clustering: - of samples. - of genes. 15. Time course experiments. 16. Gene networks. - identification of pathways 17. .
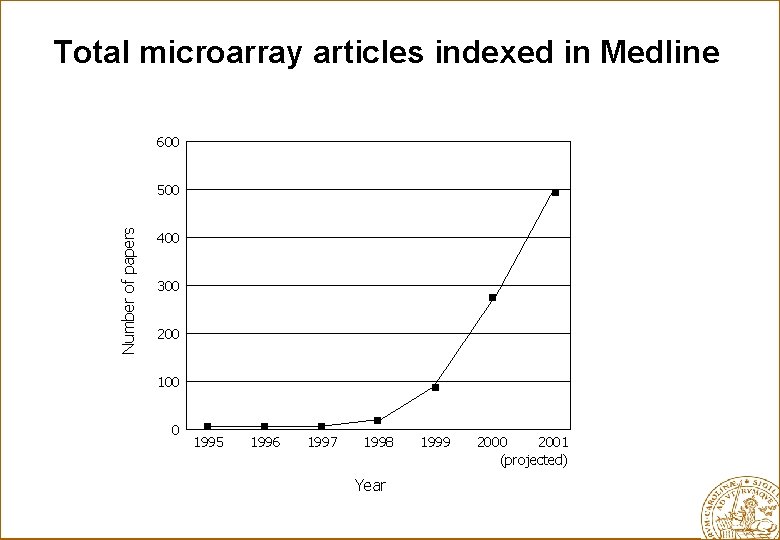
Total microarray articles indexed in Medline 600 Number of papers 500 400 300 200 100 0 1995 1996 1997 1998 Year 1999 2000 2001 (projected)

Acknowledgments/Collaborators Statistics Dept, UC Berkeley: Sandrine Dudoit Terry Speed Yee Hwa Yang Oncology Dept, Lund University: Pär-Ola Bendahl Åke Borg Johan Vallon-Christersson Enerst Gallo Research Inst. , California: Monica Moore Karen Berger Endocrinology, Lund University, Malmö: Leif Groop Peter Almgren Lawrence Berkeley National Laboratory: Saira Mian Matt Callow Mathematical Statistics, Chalmers University: Olle Nerman Staffan Nilsson Dragi Anevski CSIRO Image Analysis Group, Melbourne: Michael Buckley

Take-home message • • Bioinformatics is the future! More educated people are needed! Statistics is fun when it is applied! Master’s thesis project? Talk to us! http: //www. maths. lth. se/matstat/bioinformatics/

Finding genes in DNA sequence “This is one of the most challenging and interesting problems in computational biology at the moment. With so many genomes being sequenced so rapidly, it remains important to begin by identifying genes computationally. ” – Terry Speed.
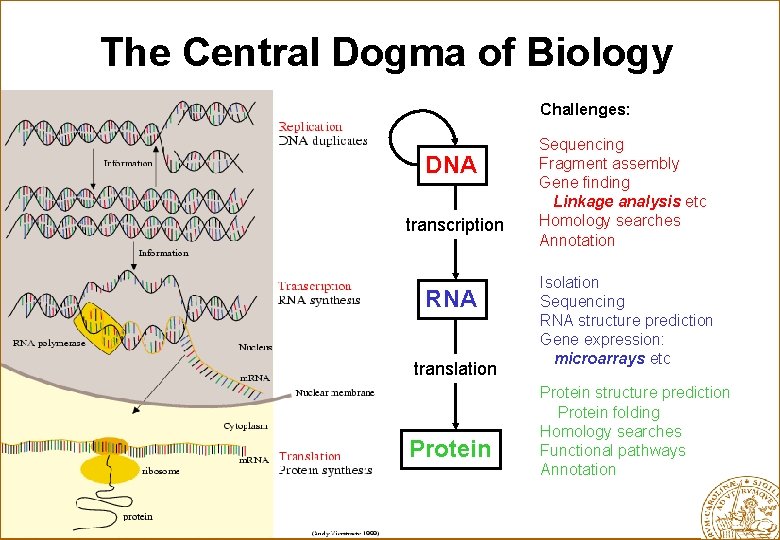
The Central Dogma of Biology Challenges: DNA transcription RNA translation Protein Sequencing Fragment assembly Gene finding Linkage analysis etc Homology searches Annotation Isolation Sequencing RNA structure prediction Gene expression: microarrays etc Protein structure prediction Protein folding Homology searches Functional pathways Annotation
- Slides: 25