Building biological networks from diverse genomic data Chad

Building biological networks from diverse genomic data Chad Myers Department of Computer Science, Lewis-Sigler Institute for Integrative Genomics Princeton University PRIME Workshop on Pathway Databases and Modeling Tools June 16, 2006
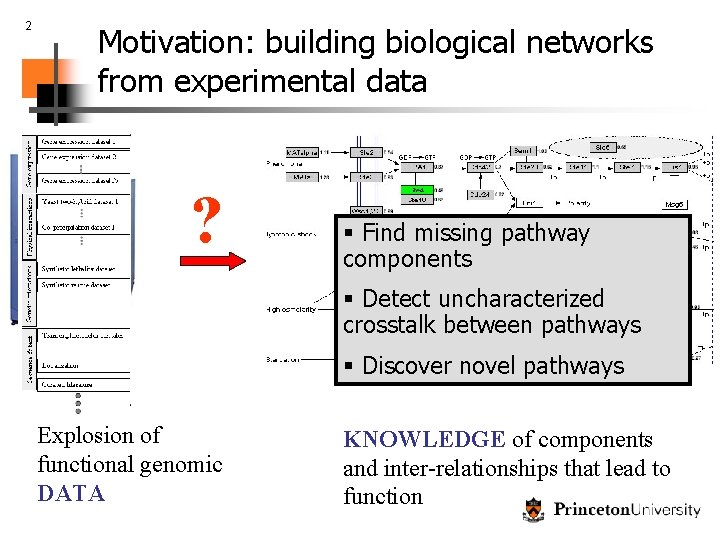
2 Motivation: building biological networks from experimental data ? § Find missing pathway components § Detect uncharacterized crosstalk between pathways § Discover novel pathways Explosion of functional genomic DATA KNOWLEDGE of components and inter-relationships that lead to function
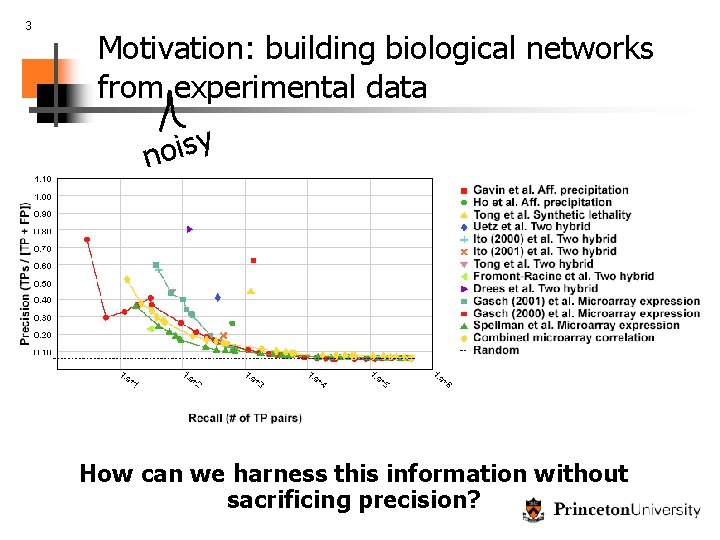
3 Motivation: building biological networks from experimental data y s i o n How can we harness this information without sacrificing precision?

4 Directed network discovery: involving the biologist in the search process n Previous approaches to network analysis from genomic data: n largely undirected global approaches that detect interesting network features n Incorporating expert direction can: n Improve sensitivity and precision by using context information n Focus on relevant information for biologist user (allows interactivity) Previous work: Bader et al. (2003), Asthana et al. (2004) Yamanashi et al. (2004, 2005), Kato et al. (2005) Two-hybrid interaction network, yeast (SH 3 domain) Boone lab
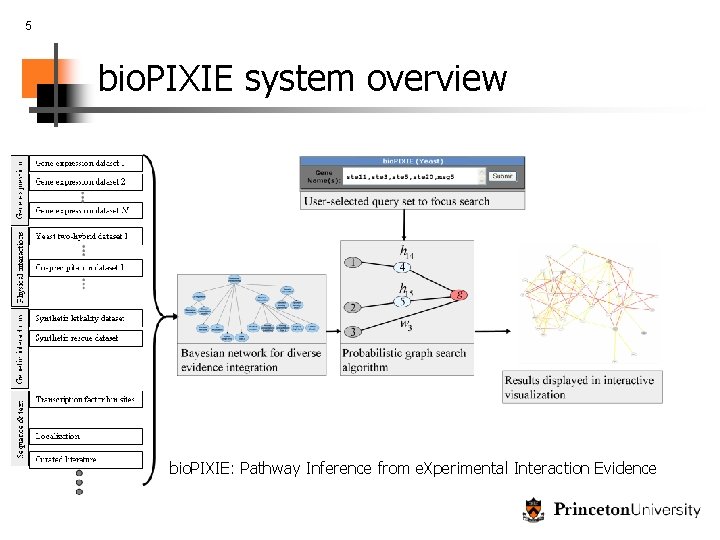
5 bio. PIXIE system overview bio. PIXIE: Pathway Inference from e. Xperimental Interaction Evidence

6 Overview n How do we integrate heterogeneous evidence? n Expert-driven network discovery n Making it usable: practical visualization and other interface considerations n Does it work? (evaluation experiments and biological validation) n Challenges/opportunities and future work

7 Heterogeneous data integration n Diverse forms of data: what’s a unifying framework? physical binding cellular localization genetic interaction sequence (TF motifs, coding, …) expression Map to associations of genes/proteins n Variable coverage, reliability, and relevance n Integration scheme should utilize information in data when available, but be robust when missing Bayes net
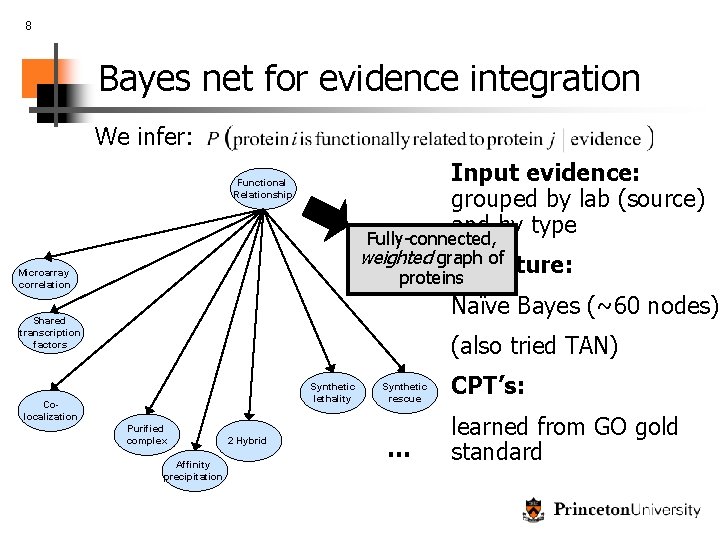
8 Bayes net for evidence integration We infer: Input evidence: grouped by lab (source) and by type Functional Relationship Fully-connected, weighted graph of Structure: proteins Microarray correlation Naïve Bayes (~60 nodes) Shared transcription factors (also tried TAN) Synthetic lethality Colocalization Purified complex Affinity precipitation 2 Hybrid Synthetic rescue … CPT’s: learned from GO gold standard

9 Overview n How do we integrate heterogeneous evidence? n Expert-driven network discovery n Making it usable: practical visualization and other interface considerations n Does it work? (evaluation experiments and biological validation) n Challenges/opportunities and future work
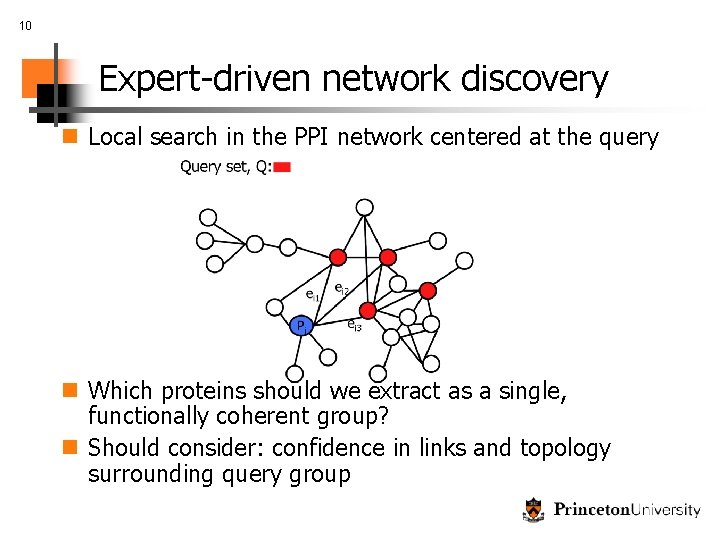
10 Expert-driven network discovery n Local search in the PPI network centered at the query n Which proteins should we extract as a single, functionally coherent group? n Should consider: confidence in links and topology surrounding query group
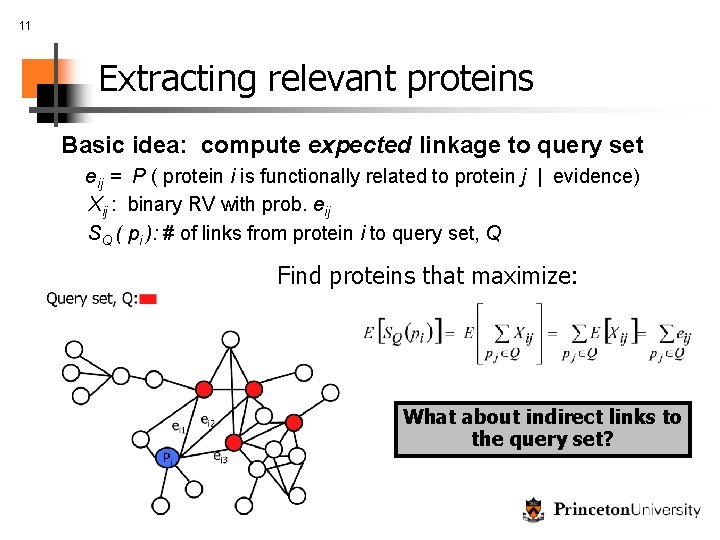
11 Extracting relevant proteins Basic idea: compute expected linkage to query set eij = P ( protein i is functionally related to protein j | evidence) Xij : binary RV with prob. eij SQ ( pi ): # of links from protein i to query set, Q Find proteins that maximize: What about indirect links to the query set?
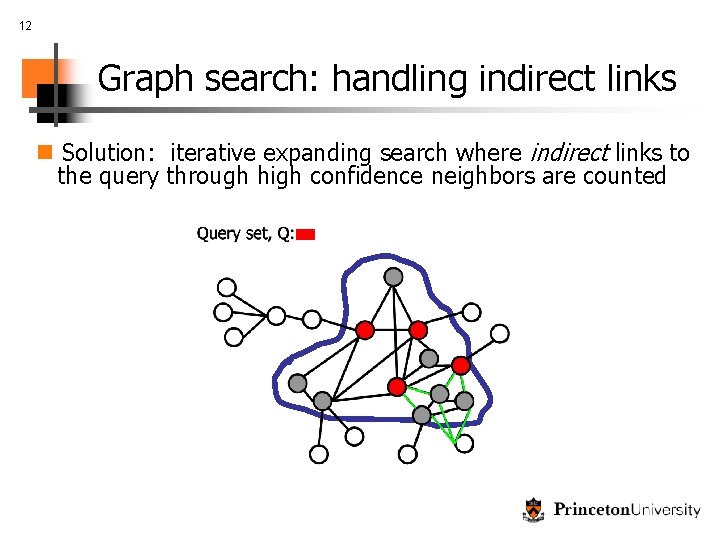
12 Graph search: handling indirect links n Solution: iterative expanding search where indirect links to the query through high confidence neighbors are counted

13 Overview n How do we integrate heterogeneous evidence? n Expert-driven network discovery n Making it usable: practical visualization and other interface considerations n Does it work? (evaluation experiments and biological validation) n Challenges/opportunities and future work

14 Making bio. PIXIE usable Guiding principles: § Accessibility (users can access most recent data with little effort) § Simplicity vs. flexibility § Drill-down (details, e. g. supporting exp. data, hidden until requested) § Browseable
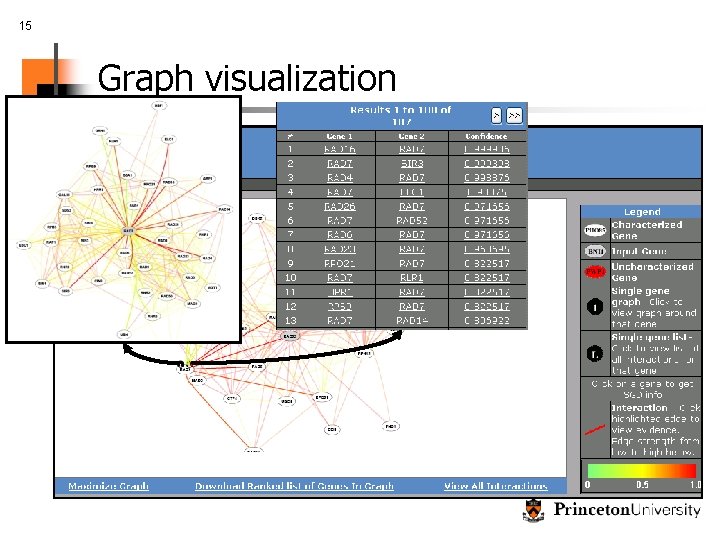
15 Graph visualization

16 Overview n How do we integrate heterogeneous evidence? n Expert-driven network discovery n Making it usable: practical visualization and other interface considerations n Does it work? (evaluation experiments and biological validation) n Challenges/opportunities and future work
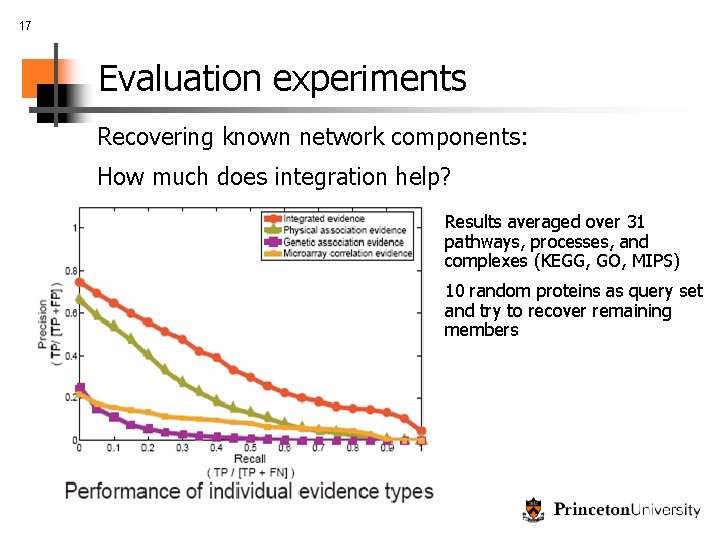
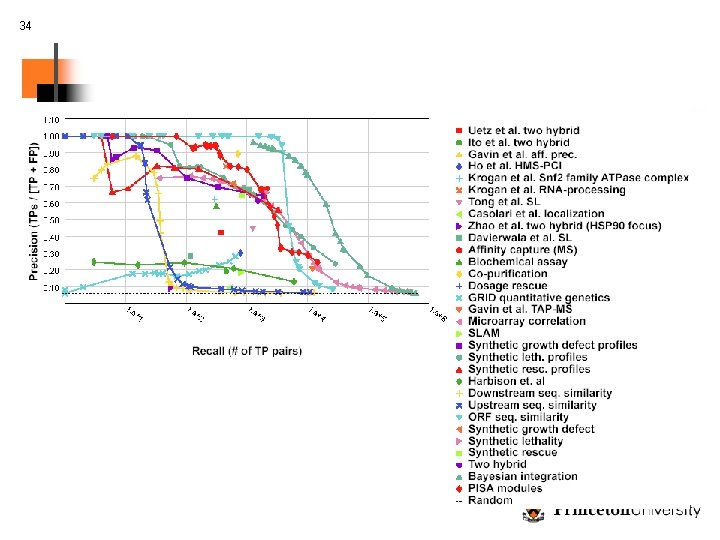
17 Evaluation experiments Recovering known network components: How much does integration help? Results averaged over 31 pathways, processes, and complexes (KEGG, GO, MIPS) 10 random proteins as query set and try to recover remaining members
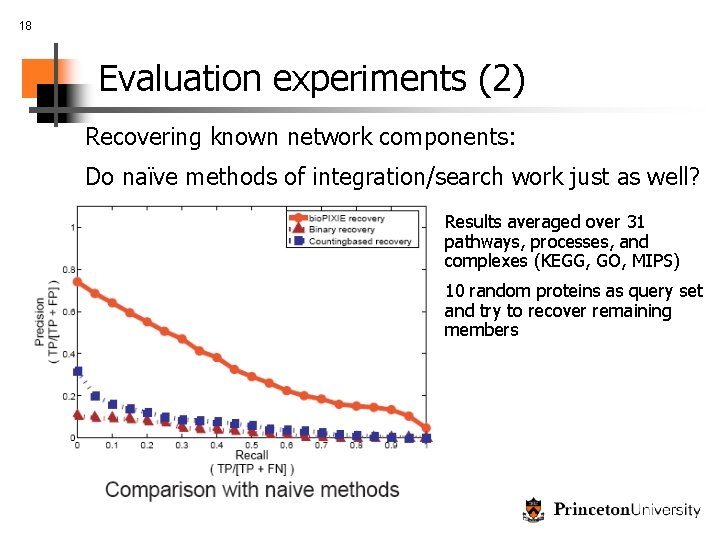
18 Evaluation experiments (2) Recovering known network components: Do naïve methods of integration/search work just as well? Results averaged over 31 pathways, processes, and complexes (KEGG, GO, MIPS) 10 random proteins as query set and try to recover remaining members

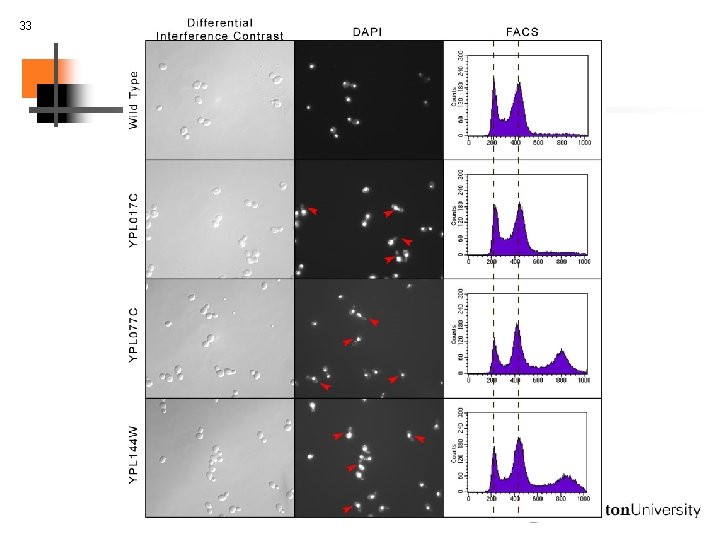
19 Biological validation: finding new components Using bio. PIXIE to characterize unknown genes S. cerevisiae uncharacterized gene, YPL 077 C Predicted involvement in chromosome segregation
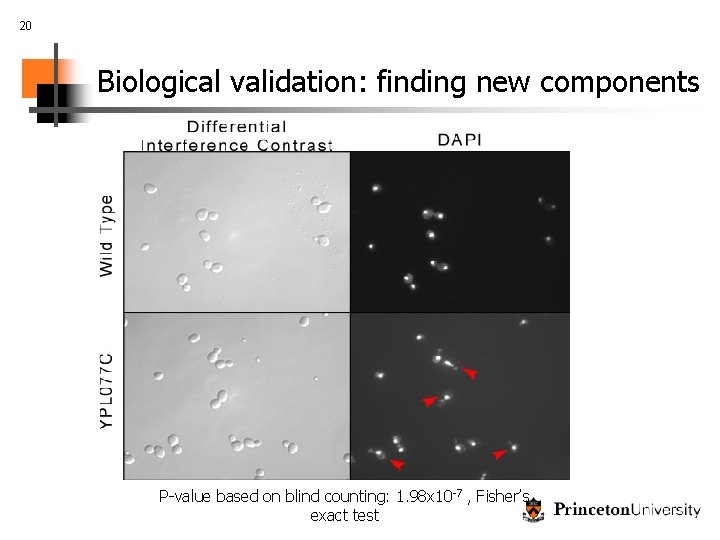
20 Biological validation: finding new components P-value based on blind counting: 1. 98 x 10 -7 , Fisher’s exact test
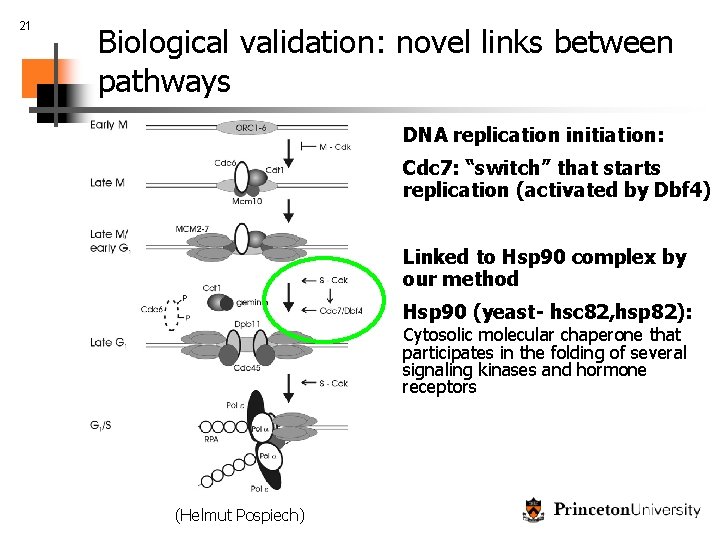
21 Biological validation: novel links between pathways DNA replication initiation: Cdc 7: “switch” that starts replication (activated by Dbf 4) Linked to Hsp 90 complex by our method Hsp 90 (yeast- hsc 82, hsp 82): Cytosolic molecular chaperone that participates in the folding of several signaling kinases and hormone receptors (Helmut Pospiech)
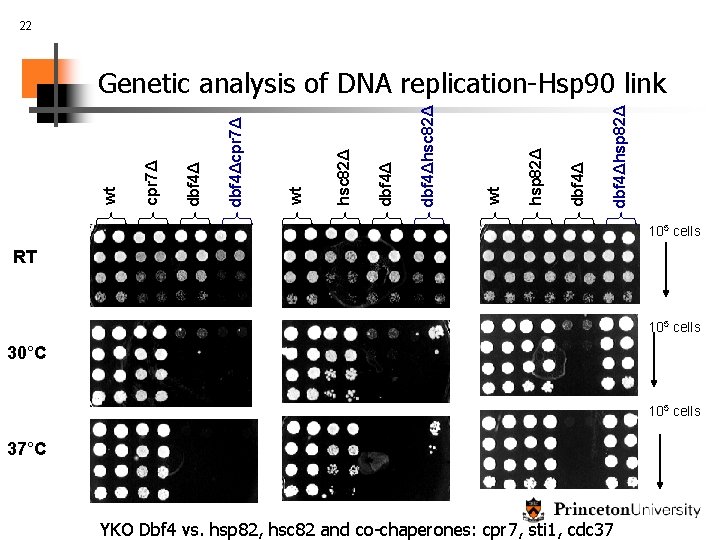
22 dbf 4Δhsp 82Δ dbf 4Δ hsp 82Δ wt dbf 4Δhsc 82Δ dbf 4Δ hsc 82Δ wt dbf 4Δcpr 7Δ dbf 4Δ cpr 7Δ wt Genetic analysis of DNA replication-Hsp 90 link 105 cells RT 105 cells 30°C 105 cells 37°C YKO Dbf 4 vs. hsp 82, hsc 82 and co-chaperones: cpr 7, sti 1, cdc 37

23 Overview n How do we integrate heterogeneous evidence? n Expert-driven network discovery n Making it usable: practical visualization and other interface considerations n Does it work? (evaluation experiments and biological validation) n Challenges/opportunities and future work

24 Practical challenges/opportunities § Visualizing complex networks of interactions in a meaningful way § how does it scale with added data? § easy user navigation around the network § Data-centric vs. established knowledge views How do we overlay current knowledge of pathways with predictions derived from experimental data?
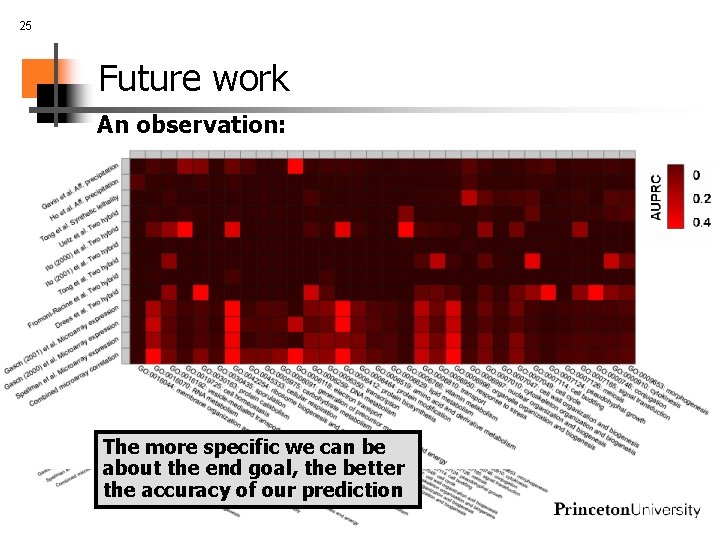
25 Future work An observation: The more specific we can be about the end goal, the better the accuracy of our prediction
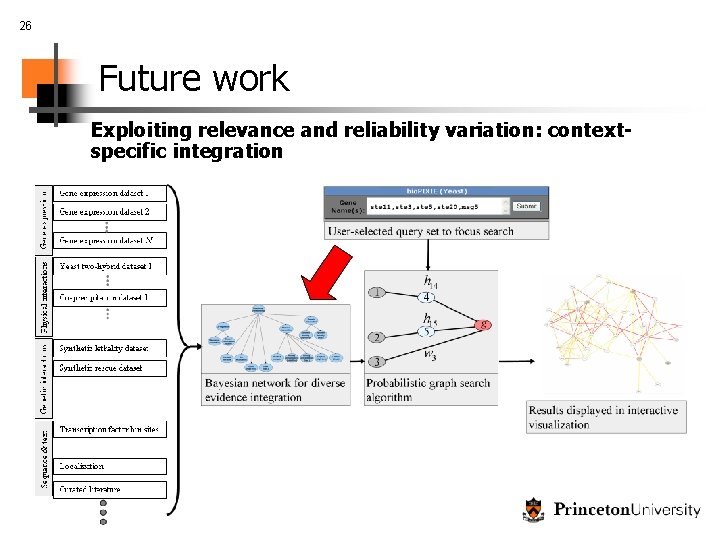
26 Future work Exploiting relevance and reliability variation: contextspecific integration

27 Summary bio. PIXIE can facilitate precise network discovery from experimental data using: n Bayesian data integration n Expert-directed search n Web-based dynamic interface bio. PIXIE is an effective tool for browsing genomic evidence and generating specific, testable hypotheses http: //pixie. princeton. edu

28 Acknowledgements Olga Troyanskaya Drew Robson Adam Wible Kara Dolinski Camelia Chiriac Matt Hibbs Curtis Huttenhower David Botstein Lab Thank you! Leonid Kruglyak Lab http: //pixie. princeton. edu
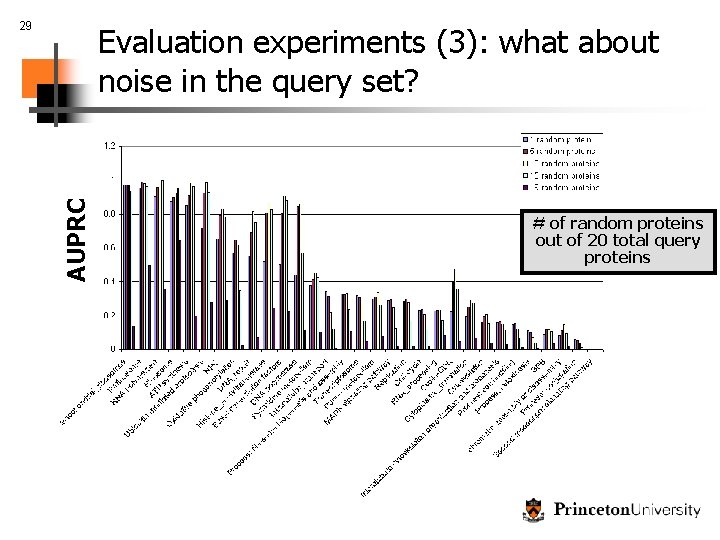
29 AUPRC Evaluation experiments (3): what about noise in the query set? # of random proteins out of 20 total query proteins
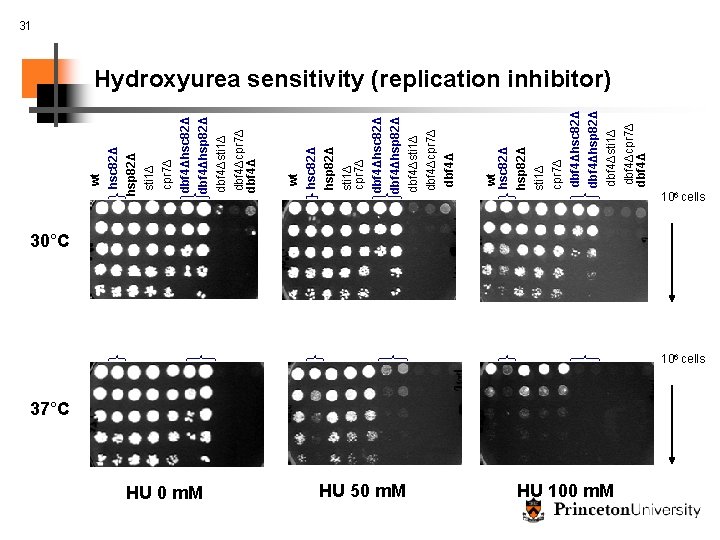
HU 0 m. M HU 50 m. M wt hsc 82Δ hsp 82Δ sti 1Δ cpr 7Δ dbf 4Δhsc 82Δ dbf 4Δhsp 82Δ dbf 4Δsti 1Δ dbf 4Δcpr 7Δ dbf 4Δ 31 Hydroxyurea sensitivity (replication inhibitor) 37°C HU 100 m. M 106 cells 30°C 106 cells

32 Is this interaction specific to DNA replication? wt hsc 82Δ hsp 82Δ sti 1Δ cpr 7Δ dbf 4Δhsc 82Δ dbf 4Δhsp 82Δ dbf 4Δsti 1Δ dbf 4Δcpr 7Δ dbf 4Δ Conclusions: wt hsc 82Δ hsp 82Δ sti 1Δ cpr 7Δ dbf 4Δhsc 82Δ dbf 4Δhsp 82Δ dbf 4Δsti 1Δ dbf 4Δcpr 7Δ dbf 4Δ MMS sensitivity (induces DNA damage) § Hsp 90 complex plays specific role in DNA replication 37°C § Hsc 82 and hsp 82 do not have identical function § Possible new link between signaling cascades, stress, and DNA replication MMS treatment has no apparent effect at RT, 30°C or 37°C (shown) § Our system generates specific, testable hypotheses 106 cells
33
34
- Slides: 33