Building a Comprehensive Reference Tandem MS Library of

Building a Comprehensive Reference Tandem MS Library of Peptides, Glycans and Glycopeptides for Therapeutic Antibodies Material Measurement Laboratory Q. Dong; M. Lorna A. De Leoz; L. E. Kilpatrick; Y. Liang; X. Yang; S. E. Stein Material Measurement Lab

The NIST/EPA/NIH Mass Spectral Library (SRD 1 A) Reference Standard Mass Spectra for Chemical Identification Fragments of charged molecules provide reproducible, discriminating fingerprints for molecules in complex mixtures in: • Environmental Analysis • Forensics • Medical Research • Chemical Manufacturing • Homeland Security • Food Analysis Content • NIST-evaluated spectra for 212, 961 compounds • Accompanying chemical name(s) and identifiers • Chemical structures and retention information • Validated software for spectrum matching Usage • World’s most widely used MS library • >5, 000 NEW installations per year • Integrated into instrument vendor software systems • Common element in GC-MS ‘gold standard’ identifications Biomolecular Measurement Division Material Measurement Lab
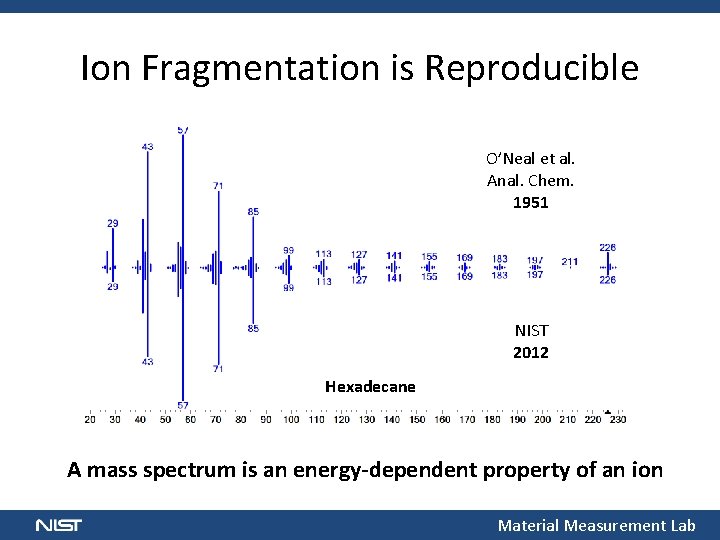
Ion Fragmentation is Reproducible O’Neal et al. Anal. Chem. 1951 NIST 2012 Hexadecane A mass spectrum is an energy dependent property of an ion Material Measurement Lab
Material Measurement Lab
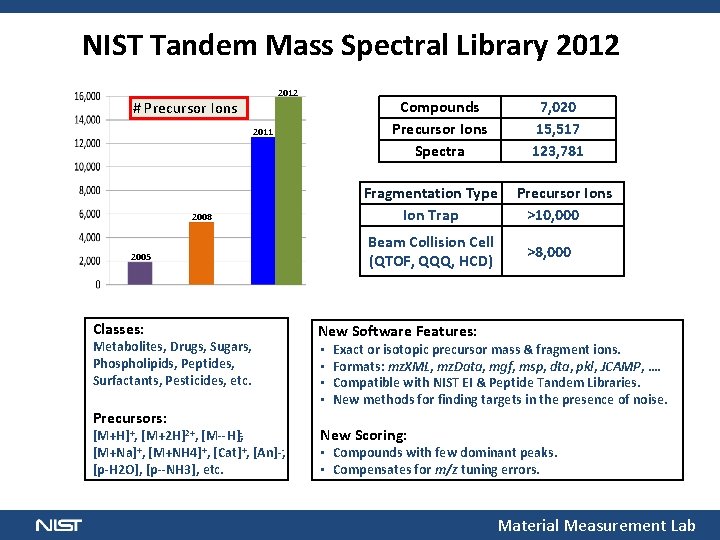
NIST Tandem Mass Spectral Library 2012 # Precursor Ions Compounds Precursor Ions Spectra 2011 Fragmentation Type Precursor Ions Ion Trap >10, 000 2008 Beam Collision Cell >8, 000 (QTOF, QQQ, HCD) 2005 Classes: Metabolites, Drugs, Sugars, Phospholipids, Peptides, Surfactants, Pesticides, etc. Precursors: [M+H]+, [M+2 H]2+, [M H] , [M+Na]+, [M+NH 4]+, [Cat]+, [An] , [p H 2 O], [p NH 3], etc. 7, 020 15, 517 123, 781 New Software Features: • • Exact or isotopic precursor mass & fragment ions. Formats: mz. XML, mz. Data, mgf, msp, dta, pkl, JCAMP, …. Compatible with NIST EI & Peptide Tandem Libraries. New methods for finding targets in the presence of noise. New Scoring: • Compounds with few dominant peaks. • Compensates for m/z tuning errors. Material Measurement Lab
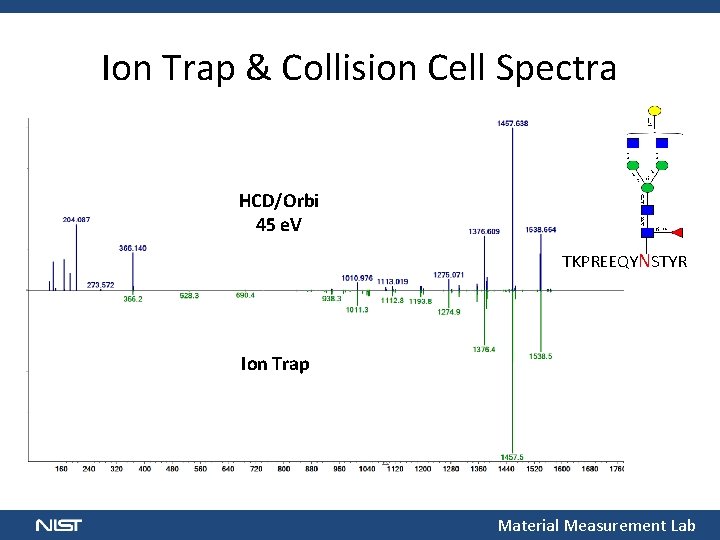
Ion Trap & Collision Cell Spectra HCD/Orbi 45 e. V TKPREEQYNSTYR Ion Trap Material Measurement Lab
![Energy Dependence Collision Energy [M+2 Na]2+ 58 e. V 65 e. V 72 e. Energy Dependence Collision Energy [M+2 Na]2+ 58 e. V 65 e. V 72 e.](http://slidetodoc.com/presentation_image_h/20f29af143cbc1138dcfc661134f7341/image-7.jpg)
Energy Dependence Collision Energy [M+2 Na]2+ 58 e. V 65 e. V 72 e. V Material Measurement Lab

NIST Library of Peptide Ion Fragmentation spectra (SRD 1 C) Reference Standard tandem Mass Spectra for peptide and protein measurements • Biomedical Research • • Protein Identification Characterization of Protein Modifications • Bio-manufacturing • Homeland Security Content • Spectra for 8 species, including human • 324, 352 human peptide ions (24% coverage of human proteome) compiled from >30 M replicates • Validated software for spectrum matching Usage • Most widely used peptide tandem MS database • Freely distributed and accessible on the web • Integrated into Thermo Scientific. TM Proteome Discoverer software • Compatible with many open-source projects Biomolecular Measurement Division Material Measurement Lab
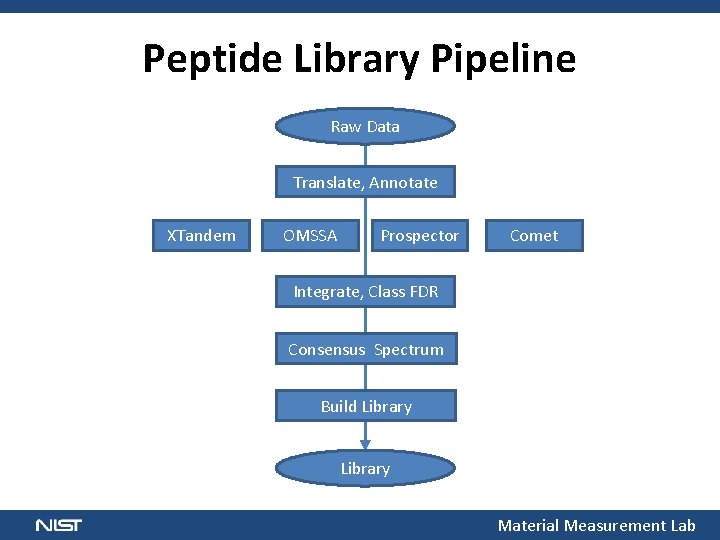
Peptide Library Pipeline Raw Data Translate, Annotate XTandem OMSSA Prospector Comet Integrate, Class FDR Consensus Spectrum Build Library Material Measurement Lab
peptide. nist. gov NISTMSQC: Full Analysis of LC-MS/MS data Library/quality metrics “Performance Metrics for Liquid Chromatography-Tandem Mass Spectrometry Systems in Proteomics Analyses”, Molecular & Cellular Proteomics, 9, 225, 2010 Material Measurement Lab
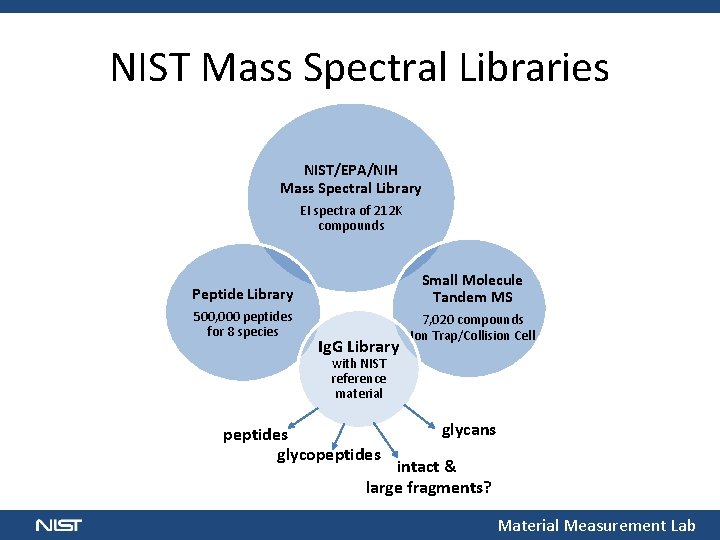
NIST Mass Spectral Libraries NIST/EPA/NIH Mass Spectral Library EI spectra of 212 K compounds Peptide Library Small Molecule Tandem MS 500, 000 peptides for 8 species 7, 020 compounds Ion Trap/Collision Cell Ig. G Library with NIST reference material peptides glycopeptides glycans intact & large fragments? Material Measurement Lab

Why Libraries? • Contain ‘fingerprints’ of previously identified components in a sample (digest) – Store, organize, annotate, extend and re-identify • Identify all known components – Independent of settings • Length, MCs, charge, protease, modification, … – More reliable (use peak intensities and priors) • Common resource among different labs – Data standard Material Measurement Lab
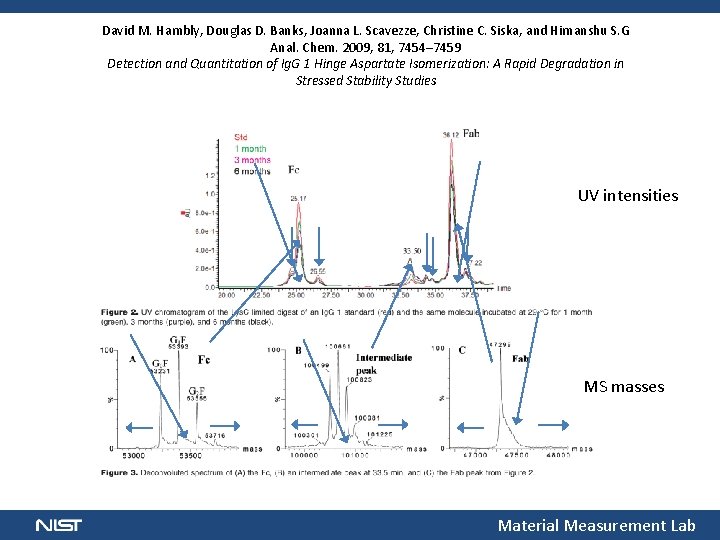
David M. Hambly, Douglas D. Banks, Joanna L. Scavezze, Christine C. Siska, and Himanshu S. G Anal. Chem. 2009, 81, 7454– 7459 Detection and Quantitation of Ig. G 1 Hinge Aspartate Isomerization: A Rapid Degradation in Stressed Stability Studies UV intensities MS masses Material Measurement Lab

Challenges with Improvements in Instrument Performance • With increase in sensitivity and mass accuracy/resolution: – More components with increasing detail revealed • Document in library – view and re-use • All components may be ‘annotated’ – What are they? Are they significant? • “Truth in Labeling” • Libraries can hold this information in reusable format Material Measurement Lab

Single Protein Digest Library All identifiable, recurring products in the digest of 1 protein • Peptides – – Use available MS ‘sequencing’ methods for tentative IDs Employ wide range of digestion conditions Apply filters to reject ‘homologies’ For each peptide store: • Spectrum (ion trap and collision cell fragmentation) • Occurrence frequencies, intensity, retention • Glycopeptides – Identify by Oxonium and Loss-Patterns - then extract • Glycans – From known samples: See Poster (Lorna De-Leoz) Material Measurement Lab
Start with Human Serum Albumin (HSA) Material Measurement Lab
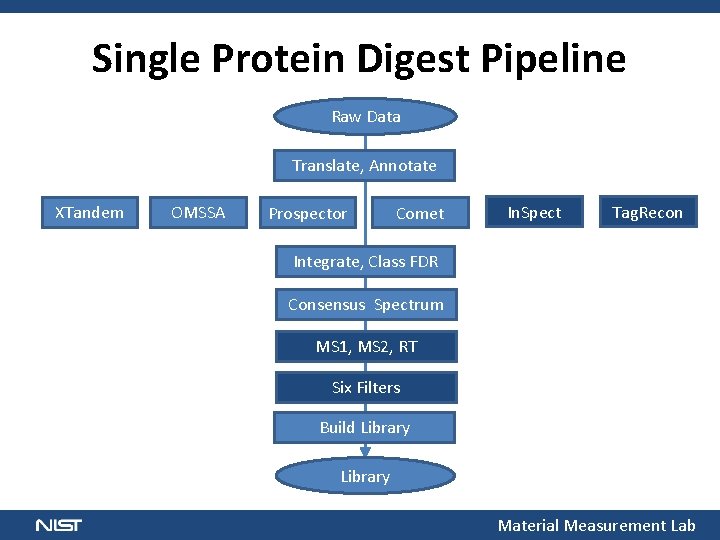
Single Protein Digest Pipeline Raw Data Translate, Annotate XTandem OMSSA Prospector Comet In. Spect Tag. Recon Integrate, Class FDR Consensus Spectrum MS 1, MS 2, RT Six Filters Build Library Material Measurement Lab
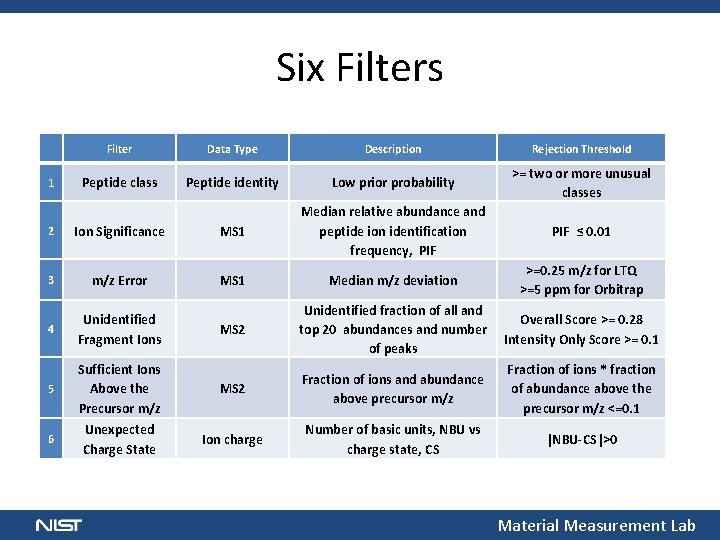
Six Filters Filter Data Type Description Rejection Threshold 1 Peptide class Peptide identity Low prior probability >= two or more unusual classes PIF ≤ 0. 01 2 Ion Significance MS 1 Median relative abundance and peptide ion identification frequency, PIF 3 m/z Error MS 1 Median m/z deviation 4 Unidentified Fragment Ions MS 2 Unidentified fraction of all and top 20 abundances and number of peaks Overall Score >= 0. 28 Intensity Only Score >= 0. 1 5 Sufficient Ions Above the Precursor m/z MS 2 Fraction of ions and abundance above precursor m/z Fraction of ions * fraction of abundance above the precursor m/z <=0. 1 6 Unexpected Charge State Ion charge Number of basic units, NBU vs charge state, CS |NBU CS|>0 >=0. 25 m/z for LTQ >=5 ppm for Orbitrap Material Measurement Lab

Biological Modifications • Methionine Oxidation – ETYGEMADCCAK • Cysteinylation – ALVLIAFAQYLQQCPFEDHVK • Phosphorylation – TCVADESAENCDK • Acetylation – LKCASLQK Material Measurement Lab

Some Analytical Modifications • Methionine Oxidation – ETYGEMADCCAK • Transpeptidation • Glutamine Deamination • Iron – QTALVELVK – TPVSDRR – AAFTECCQAADK • Pyro-carbamidomethyl – CCTESLVNR • Sodiation • Underalkylation – CCTESLVNR • Carbamylation • Overalkylation • Tris-derived N-term (=70) – LVNEVTEFAK – SLHTLFGDK – Method/Sample Dependent Material Measurement Lab

Indigestible Peptides • Hyper-missed cleavage peptides – DVFLGMFLYEYARRHPDYSVVLLLRLAKTYETTLEK/4+ – NYAEAKDVFLGMFLYEYARRHPDYSVVLLLR/4+ – QNCELFEQLGEYKFQNALLVRYTKKVPQVSTPTLVEVSR/5+ • Formed quickly, not affected by digestion time Material Measurement Lab

Ig. G Modifications • Large-Scale Identification and Quantification of Covalent Modifications in Therapeutic Proteins – “In an LC/MS/MS analysis of a tryptic digestion of an Ig. G 2 mono-clonal antibody, 1712 peptide ions … and 227 modifications were identified …” • Zhongqi Zhang, Anal. Chem. 2009, 81, 8354– 8364 • From protein and processing – Most from processing Material Measurement Lab

Ig. G Tryptic Peptide Libraries • Six Commercial Drugs – 1 Constant Region, Individual Variable Region Libraries • Cetuximab Example – Spectra for 975 Different Ions – – – – 156 Simple Tryptics 175 Modifications 200 ‘In-sample’ Semitryptics 169 ‘In-source’ Semitryptics 101 Unexpected Missed Cleavage 41 Under-/Over-alkylation 133 Unidentified Modifications Material Measurement Lab

Cetuximab Modifications Modification Site Different Ions Carbamyl N-term, K 32 Oxidation M, H, W 27 Cation: Na D, E 25 Gln->pyro-Glu Q at N-term 12 Cation: Ca D, E 12 Formyl N-term, K, S, T 10 Dehydration D 8 Pyro-carbamidomethyl C at N-term 7 Transpeptidation Arg and Lys at N or C-term 6 Glu->pyro-Glu E at N-term 2 Deamidation N, Q 2 Material Measurement Lab

Ig. G Digest Peptides Library Consensus Spectrum FNW(O)YVDGVEVHNAK/3+ Annotation Material Measurement Lab

Glycopeptide Features NEUTRAL MONOSACCHARIDE LOSSES OXONIUM IONS Theory m/z 163. 0601 m/z 204. 0867 m/z 274. 0921 m/z 292. 1026 m/z 366. 1387 m/z 454. 1555 m/z 657. 2349 Our addition m/z 138. 0550 m/z 168. 0655 m/z 186. 0761 -(2 H 2 O+CH 2 O) -2 H 2 O -H 2 O 1+ 2+ 3+ 4+ 203. 0794 101. 5397 67. 6931 50. 7698 162. 0528 81. 0264 54. 0176 40. 5132 146. 0579 73. 0289 48. 6859 36. 5144 291. 0954 145. 5477 97. 0318 72. 7738 Hexose Deoxyhexose (e. g. fucose) N-acetylhexosamine Sialic Acid (e. g. Neu. Ac) 35 e. V HCD MS 2 Neutral Losses Threshold: ~20 ppm Oxonium ion peaks All z=2 Threshold: ~60 ppm 81 81 81 101 Material Measurement Lab
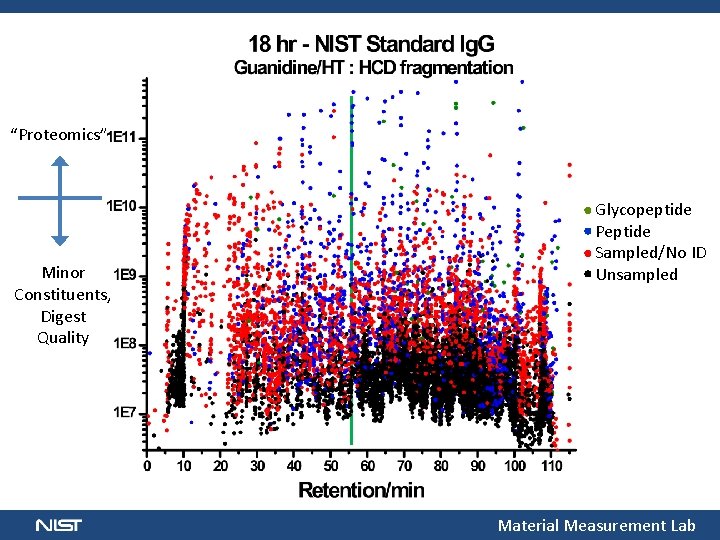
“Proteomics” Minor Constituents, Digest Quality Glycopeptide Peptide Sampled/No ID Unsampled Material Measurement Lab
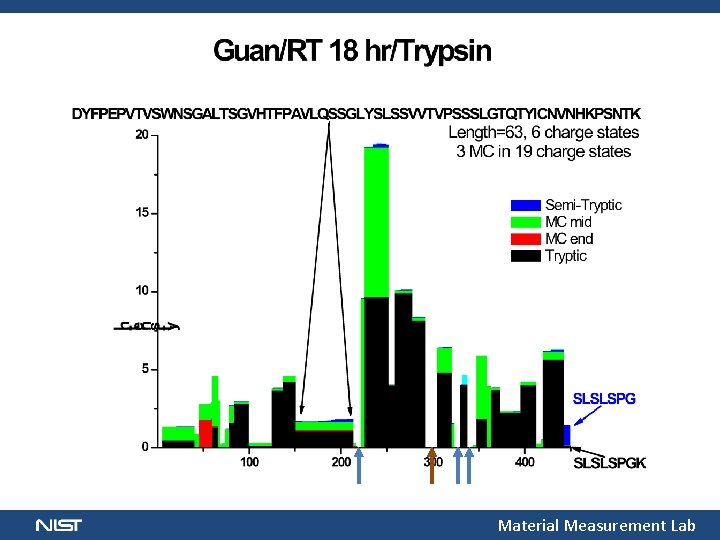
Material Measurement Lab
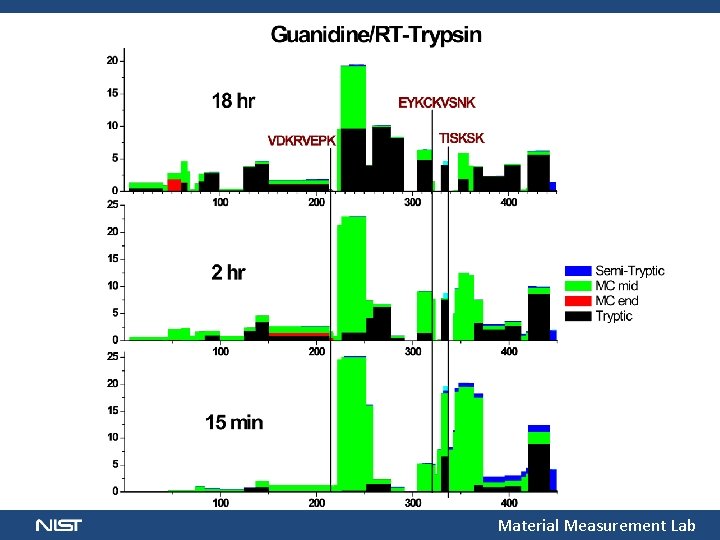
Material Measurement Lab
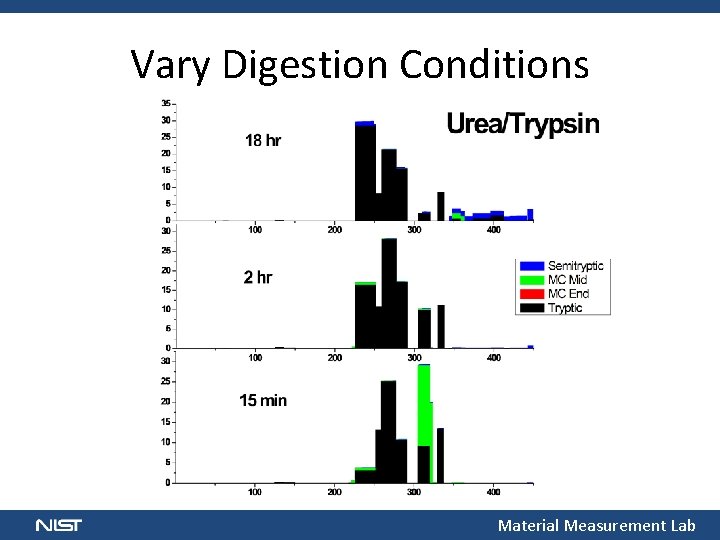
Vary Digestion Conditions Material Measurement Lab
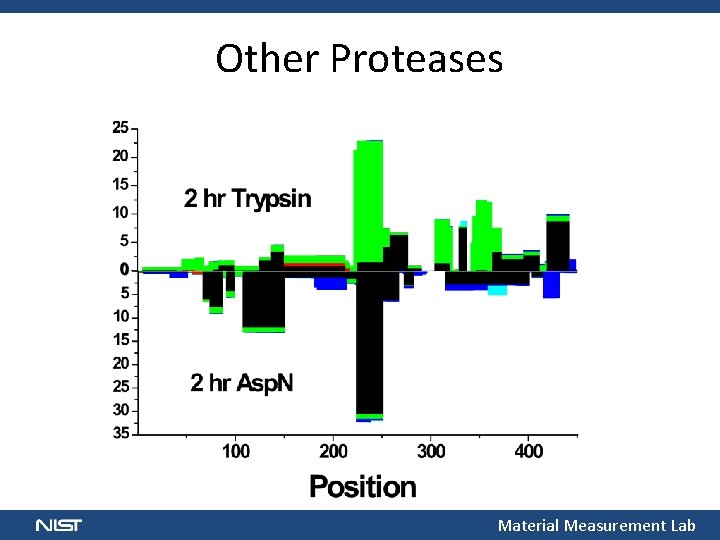
Other Proteases Material Measurement Lab

NGA 42 G 0 NGA 2 F 1 G 0 F NGA 3 B 1, 2 G 0 B NA 2 G 1 F 1 G 1 F A 1 F 1 G 2 FS NA 2 F 1 G 2 F A 2 F 1 NGA 2 B 2 G 2 FS 2 G 0 B NGA 21 G 0 NGA 4 B 2 G 0 B 1 Purchased; 2 From Starter Glycan MS/MS Library Typical Precursor ions § § [M+2 H]2+ [M+H+K]2+ [M+H+Na]2+ [M+2 Na]2+ Additional precursor ions for sialylated glycans § § [M+2 Na-4 H]2[M+K-3 H]2[M-2 H]2[M+Na+K-4 H]2 - Fragmentation Methods § Ion Trap (Resonant) § ‘HCD’ at 12 energies (5 -70 e. V) the Consortium for Functional Glycomics through Dr. Jim Paulson Material Measurement Lab
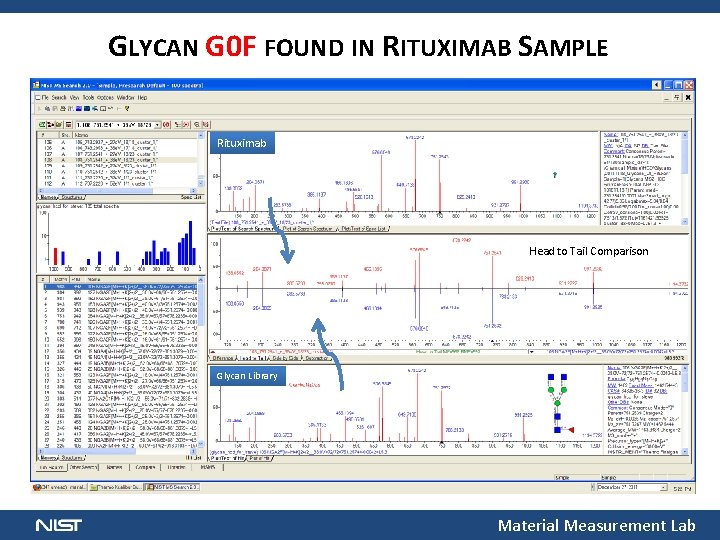
GLYCAN G 0 F FOUND IN RITUXIMAB SAMPLE Rituximab Head to Tail Comparison Glycan Library Material Measurement Lab

SRM/D SRD + SRM SRD • Standard Reference Materials + Data – Whole material + Data + Libraries MS Library, Thermodynamic Data, Chemistry Webbook, … • Standard Reference Materials – Target Analytes (Cholesterol, Vitamin D) • Often in Matrix (urine, plasma, …) – Whole Materials (Plasma Metabolites, …) • Identify and value-assign many components Material Measurement Lab

SRM/DFuture: - Metabolites SRM/D in. Ig. G Plasma http: //srm 1950. nist. gov http: //Ig. G. nist. gov Material Measurement Lab

NIST MS Data Center Lab Computer Material Measurement Lab
- Slides: 36