Bsub Cyc A ModelOrganism Database for Bacillus subtilis

Bsub. Cyc – A Model-Organism Database for Bacillus subtilis Ingrid M. Keseler SRI International

Why Bacillus subtilis? • Gram-positive bacterium • Long history of basic research • Interesting life cycle (sporulation, competence, biofilm) • Model organism for a variety of important pathogens (e. g. Staphylococcus aureus) and bioterrorism agents (e. g. Bacillus anthracis) • Important industrial microorganism: production of enzymes (e. g. Novozymes, Genencor)

Bacillus subtilis Vital Stats • One of the earliest published genome sequences 1997 Nov 20; 390(6657): 249 -56. Nature. • One of the earliest published genome sequences – recently re-sequenced and reannotated Microbiology. 2009 Jun; 155(Pt 6): 1758 -75. • 4, 215, 606 bp, 4, 400+ genes • Pub. Med keyword search “subtilis” – 25, 753 publications on 10/22/2010
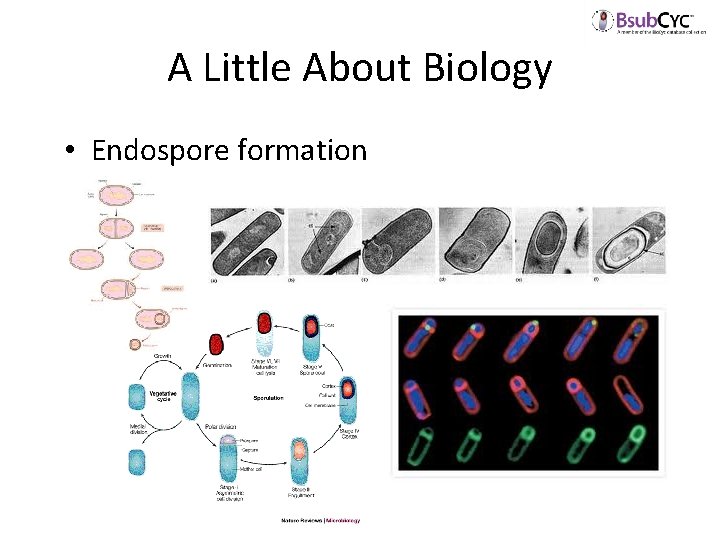
A Little About Biology • Endospore formation

A Little More About Biology • Biofilm formation

Building Bsub. Cyc • Based on Gen. Bank AL 009126. 3 GI: 225184640, deposited by the Genoscope group October 1, 2009 • Resulting in • 1222 metabolic reactions • 214 metabolic pathways Bsub. Cyc. org

Metabolic Overview

Further Improvements • Gene name synonyms from Geno. List • Links to other databases: • • Subti. Wiki (U. Goettingen) Geno. List (Pasteur Institute) Subtilis. Wiki (Texas A&M) STRING (EMBL)

Importing Regulatory Information • DBTBS

Transcriptional Regulation • Imported from DBTBS (Database of Transcriptional regulation in Bacillus subtilis): • 1076 transcription start sites • 1226 transcription units • 703 transcription factor binding sites • Import is incomplete due to a variety of complications
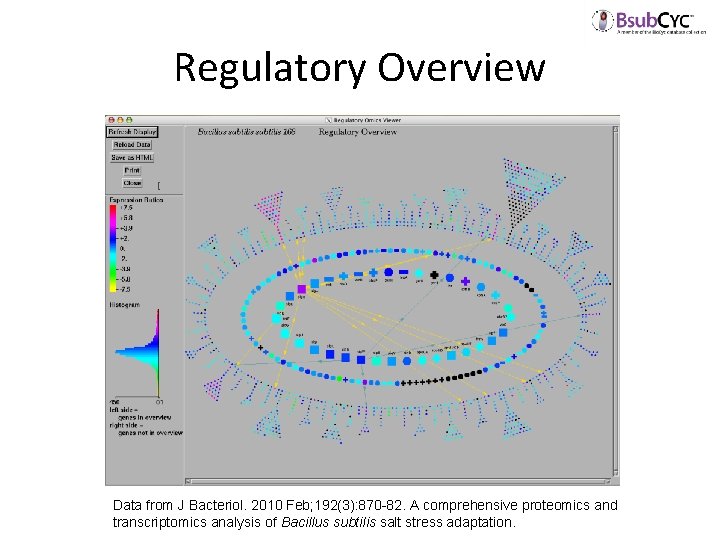
Regulatory Overview Data from J Bacteriol. 2010 Feb; 192(3): 870 -82. A comprehensive proteomics and transcriptomics analysis of Bacillus subtilis salt stress adaptation.
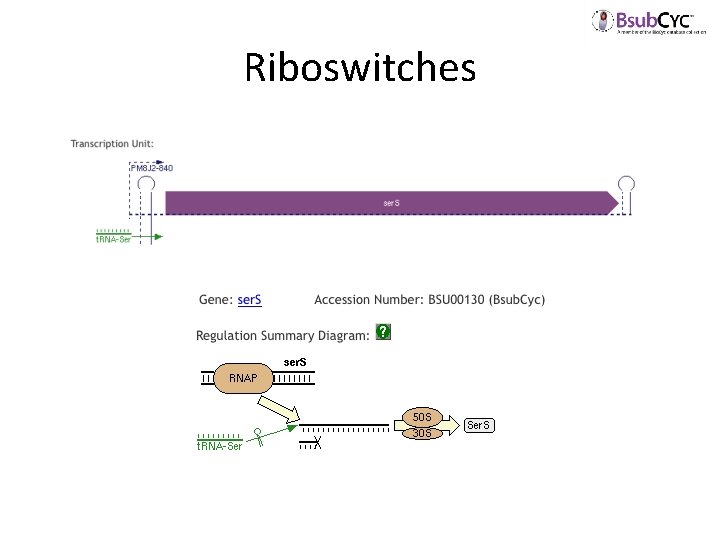
Riboswitches
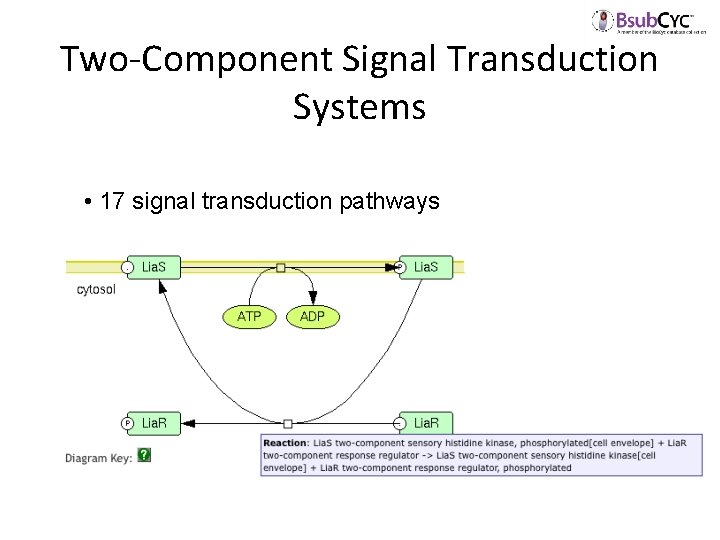
Two-Component Signal Transduction Systems • 17 signal transduction pathways

New Literature • New publications from 2009 and 2010 added to genes/proteins: • Publication year 2009: 87 • Publication year 2010: 169 • Approximate number of new gene functions since the beginning of this project: ~70? (gene name changed) • 2 new pathways
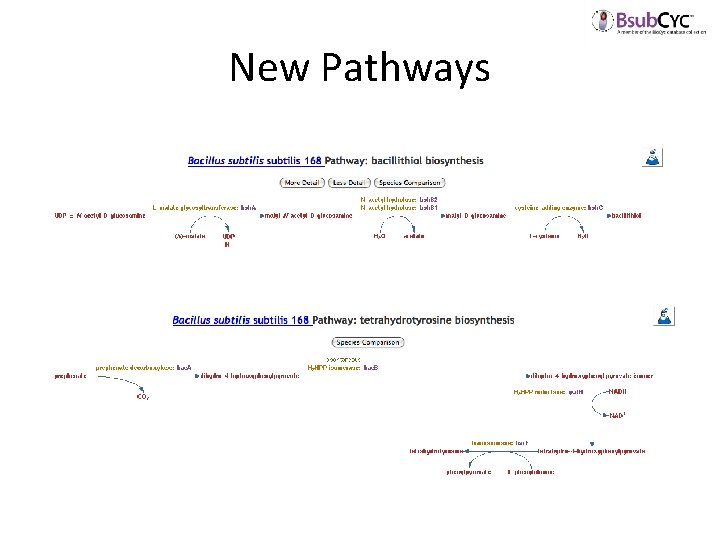
New Pathways

Future Directions • Work on metabolic pathways – recruit scientists with expertise • Improve transcriptional regulatory network • Post-transcriptional regulation • Import data from the co-PIs of the project • Funding!

Acknowledgements • Bioinformatics Research Group at SRI • • Tomer Altman Pallavi Kaipa Markus Krummenacker Ron Caspi • David Rudner, Anna-Barbara Hachmann (Harvard) • Jim Hu (Texas A&M) • Funding: NIH Grand Opportunity grant RC 2 GM 092616
- Slides: 17