BSB 512 Nuclear Magnetic Resonance Spectroscopy TTh 2

BSB 512 Nuclear Magnetic Resonance Spectroscopy TTh 2: 20 - 3: 25 pm Lecture notes available at http: //sos. bio. sunysb. edu/bsb 512 Lecture 1: Lecture 2 -4: Lecture 5: Lecture 6: Basics Protein structure determination Relaxation and dynamics Lab Session: Sample preparation. Pulse Programs Probe selection. Tuning. Shimming Pulse calibration. Data collection.

References http: //www. cis. rit. edu/htbooks/nmr Books Wuthrich, K. Levitt, MH Cavanagh J. et al. Ernst, R. et al. Bax, A NMR of Proteins and Nucleic Acids Spin Dynamics Protein NMR Spectroscopy Principles of NMR in One and Two Dimensions Two dimensional NMR in Liquids

NMR Spectroscopy: Some history 1915 Einstein and de Hass - Correlation between magnetic moment and spin angular momentum 1922 Stern and Gerlach - Spins are quantized 1946 Bloch and Purcell - First NMR experiment 1967 Richard Ernst - Fourier transformations 1971 Jean Jeener - Two dimensional NMR - COSY 19721976 Richard Ernst - First two dimensional NMR experiment 19731986 Kurt Wuthrich - First independent NMR - X-ray comparison

High resolution NMR of proteins • Observe protons (1 H) • This differs from x-ray diffraction where one determines structure based on the electron density from the electron rich atoms (C, N, O). • Protein is solubilized in water.

High resolution NMR of proteins • Observe protons • Assign proton resonances to indivdual amino acids. Proton resonances are often resolved by differences in chemical shifts. • Measure intra-residue and inter-residue proton to proton distances through dipolar couplings. • Measure torsion angles through J-couplings. • Use distance and torsion angle constraints to determine secondary and tertiary structure.
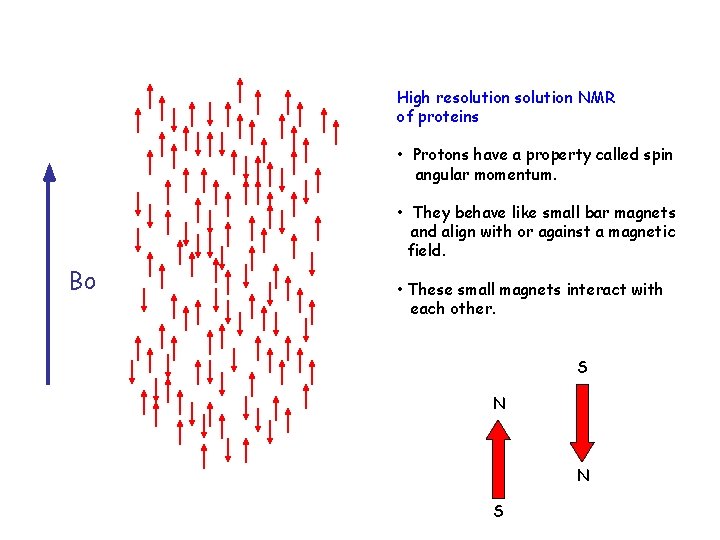
High resolution NMR of proteins • Protons have a property called spin angular momentum. • They behave like small bar magnets and align with or against a magnetic field. Bo • These small magnets interact with each other. S N N S
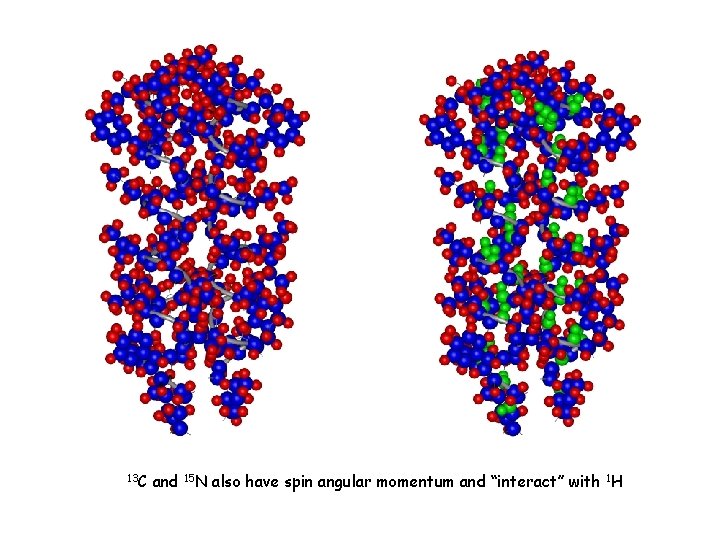
13 C and 15 N also have spin angular momentum and “interact” with 1 H
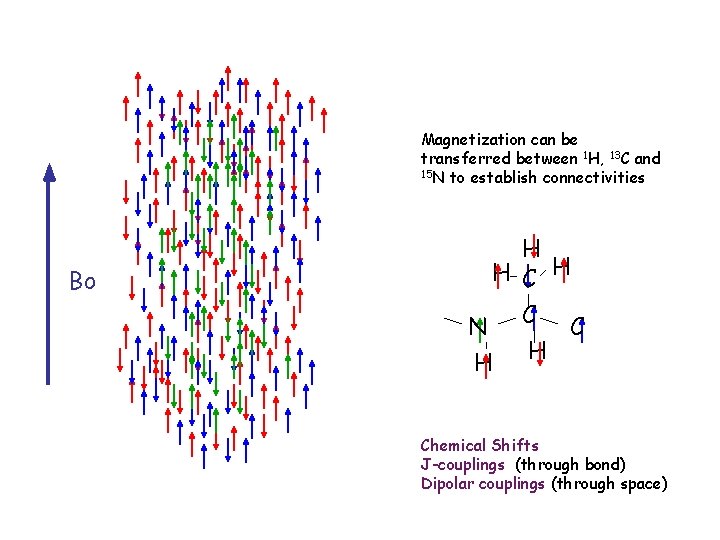
Magnetization can be transferred between 1 H, 13 C and 15 N to establish connectivities Bo H H C H N C C H H Chemical Shifts J-couplings (through bond) Dipolar couplings (through space)
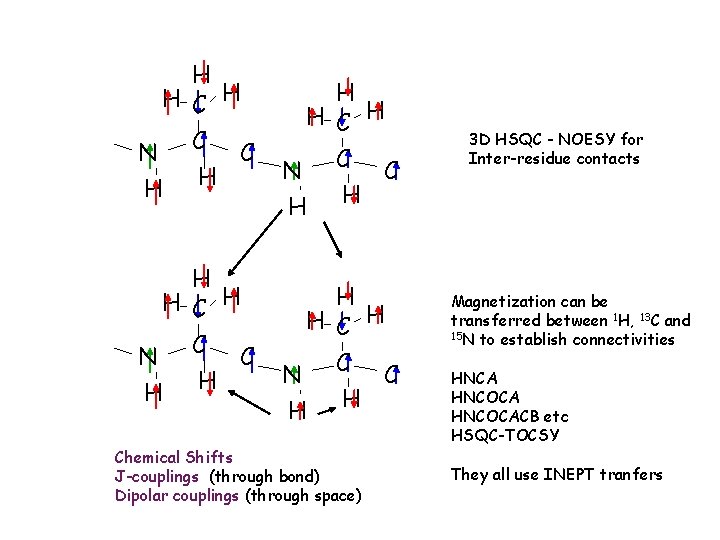
H H C H N C C H H Chemical Shifts J-couplings (through bond) Dipolar couplings (through space) 3 D HSQC - NOESY for Inter-residue contacts Magnetization can be transferred between 1 H, 13 C and 15 N to establish connectivities HNCA HNCOCACB etc HSQC-TOCSY They all use INEPT tranfers

Concept 1: Some nuclei have non-zero spin quantum numbers. e- Nuclei with odd mass numbers have half-integer spin quantum numbers. i. e. 13 C, 1 H, 31 P are spin I = 1/2 17 O is spin I = 5/2 Nuclei with an even mass number and an even charge number have spin quantum numbers of zero. ie. 12 C Nuclei with an even mass number and an odd charge number have integer spin quantum numbers. i. e. 2 H is spin I = 1 Electrons also have a spin quantum number of 1/2
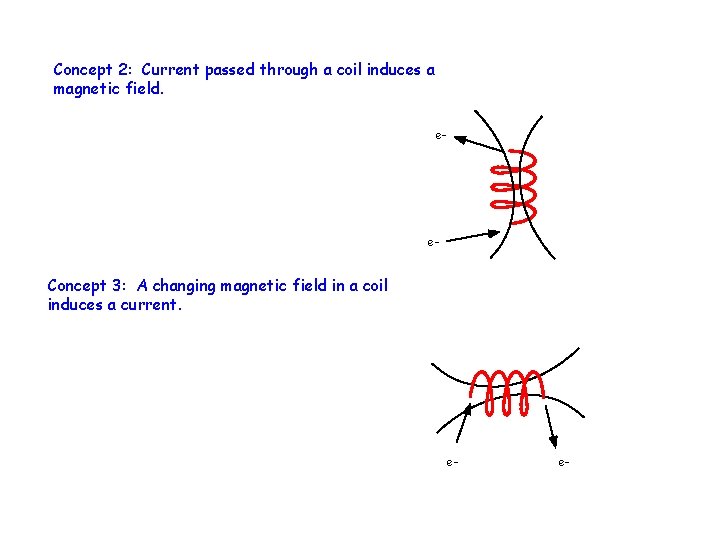
Concept 2: Current passed through a coil induces a magnetic field. e- e- Concept 3: A changing magnetic field in a coil induces a current. e- e-
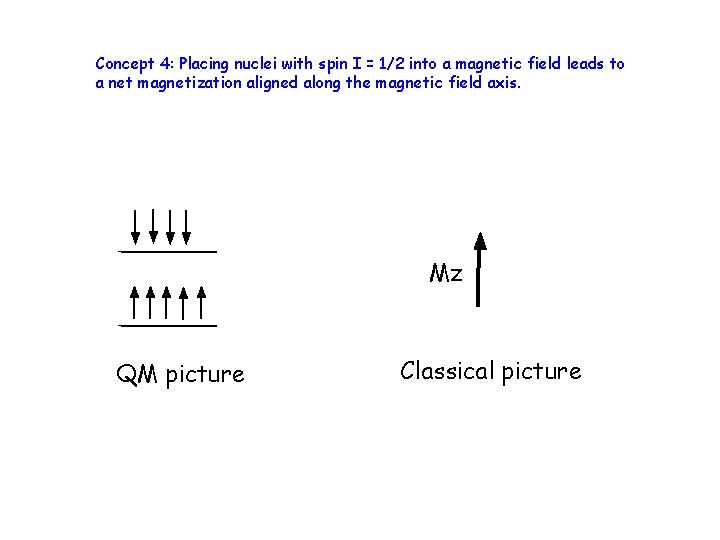
Concept 4: Placing nuclei with spin I = 1/2 into a magnetic field leads to a net magnetization aligned along the magnetic field axis. Mz QM picture Classical picture
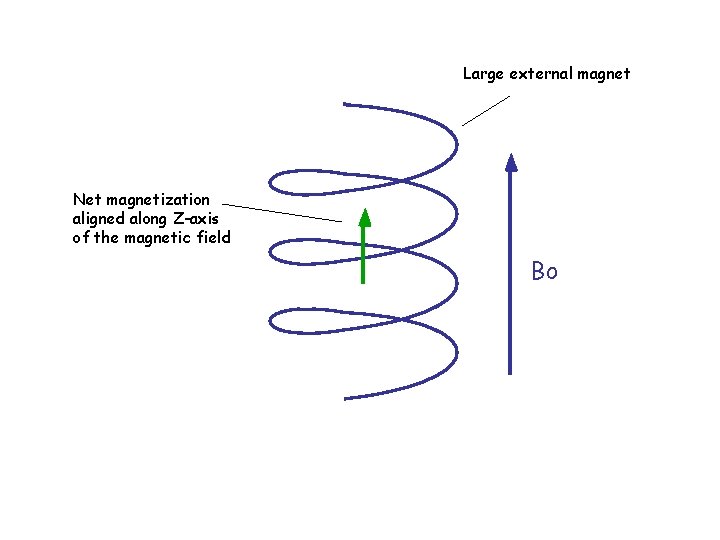
Large external magnet Net magnetization aligned along Z-axis of the magnetic field Bo
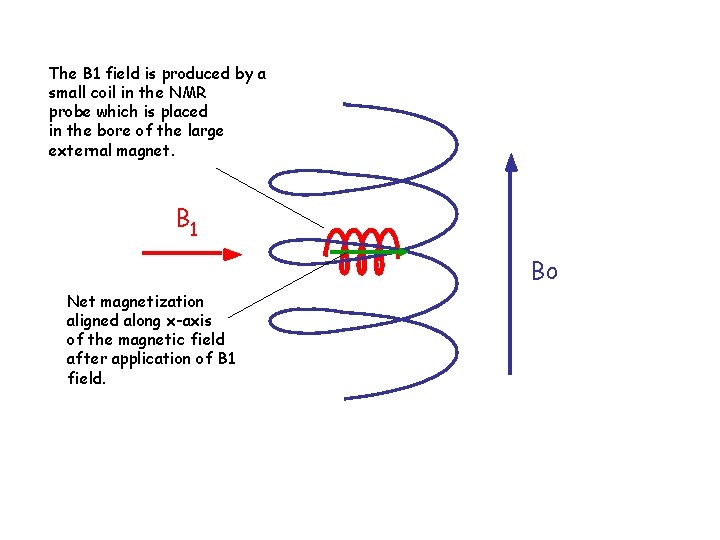
The B 1 field is produced by a small coil in the NMR probe which is placed in the bore of the large external magnet. B 1 Bo Net magnetization aligned along x-axis of the magnetic field after application of B 1 field.
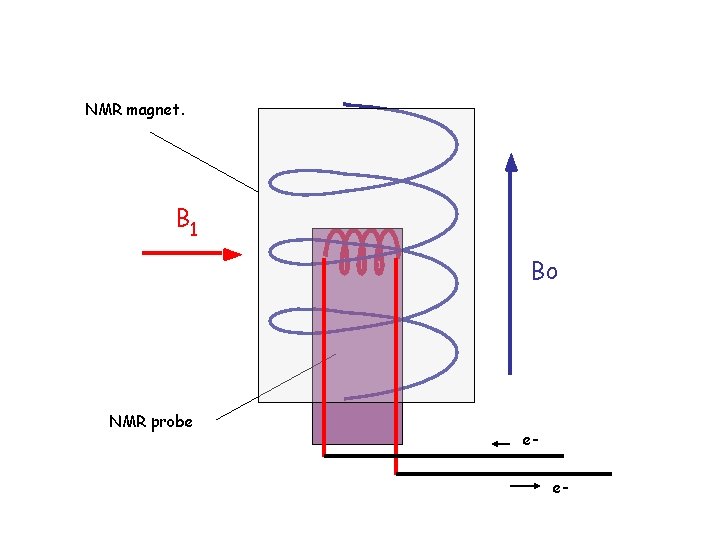
NMR magnet. B 1 Bo NMR probe ee-
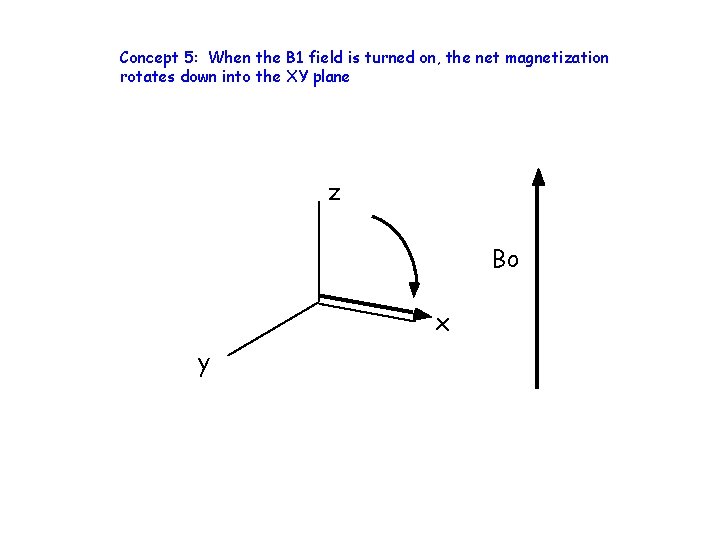
Concept 5: When the B 1 field is turned on, the net magnetization rotates down into the XY plane z Bo x y

Concept 6: When the B 1 field is turned off, the net magnetization relaxes back to the Z axis with the time constant T 1 z T 1 Bo x y T 1 is the “longitudinal” relaxation time constant which results from “spin-lattice” relaxation
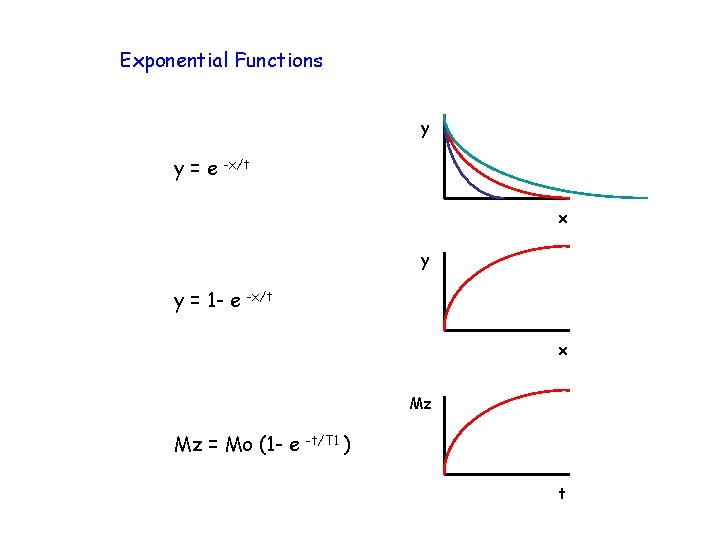
Exponential Functions y y=e -x/t x y y = 1 - e -x/t x Mz Mz = Mo (1 - e -t/T 1 ) t
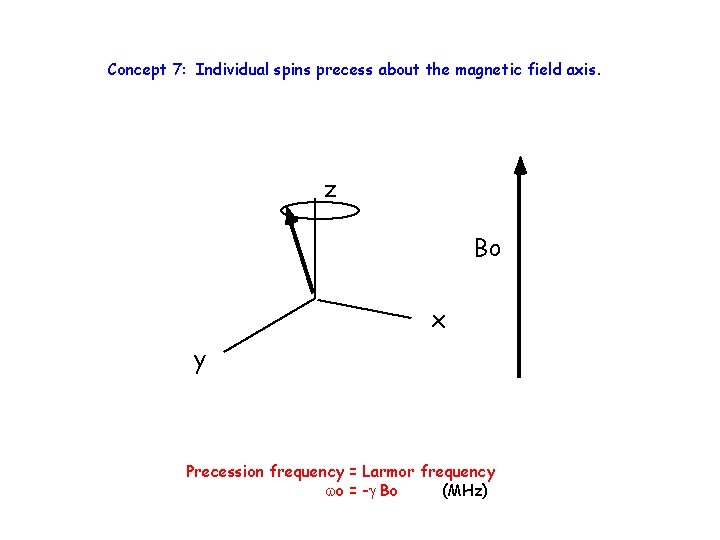
Concept 7: Individual spins precess about the magnetic field axis. z Bo x y Precession frequency = Larmor frequency wo = -g Bo (MHz)
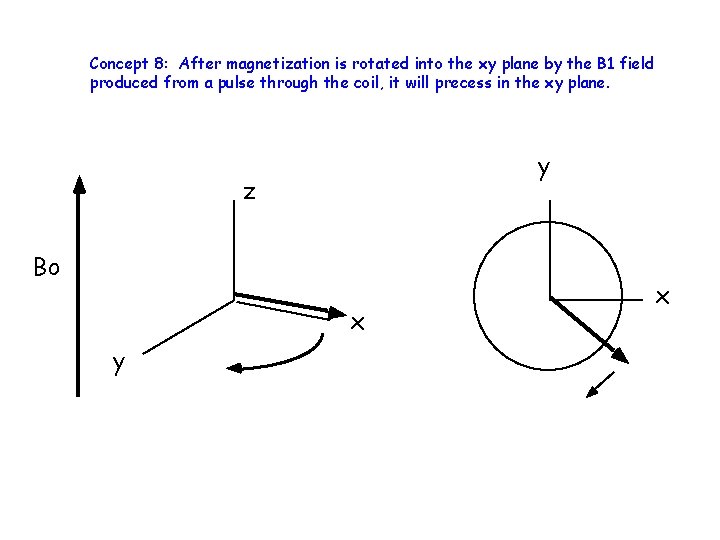
Concept 8: After magnetization is rotated into the xy plane by the B 1 field produced from a pulse through the coil, it will precess in the xy plane. y z Bo x y x
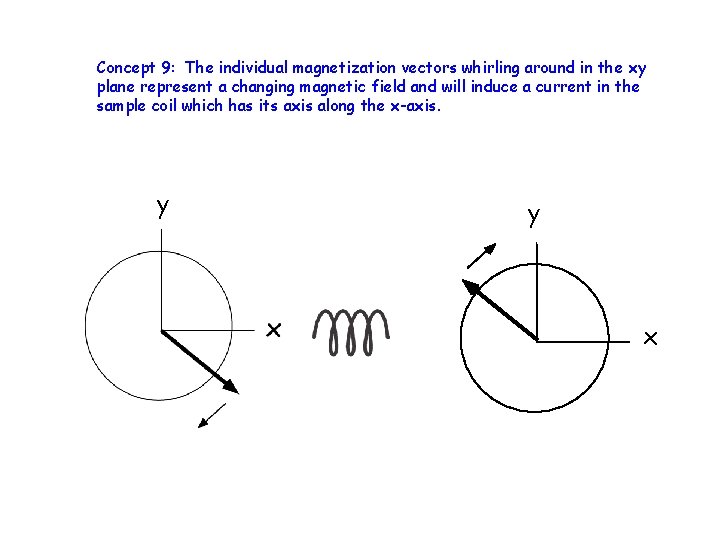
Concept 9: The individual magnetization vectors whirling around in the xy plane represent a changing magnetic field and will induce a current in the sample coil which has its axis along the x-axis. y y x
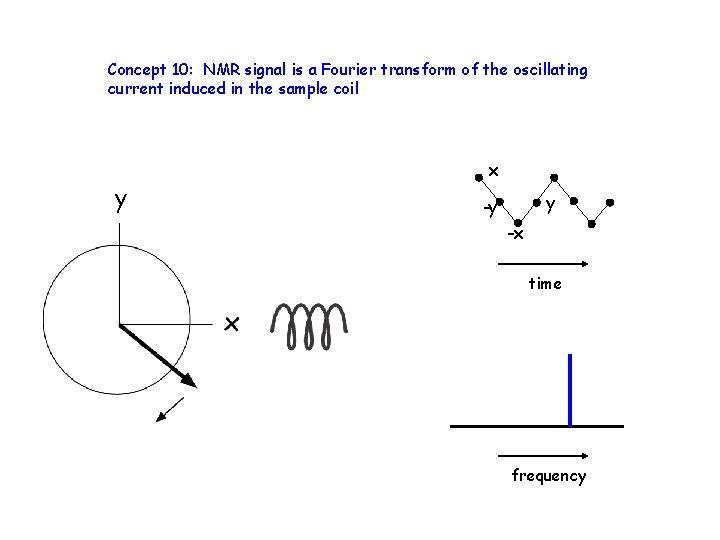
Concept 10: NMR signal is a Fourier transform of the oscillating current induced in the sample coil y x y -y -x time frequency

Chemical Shifts
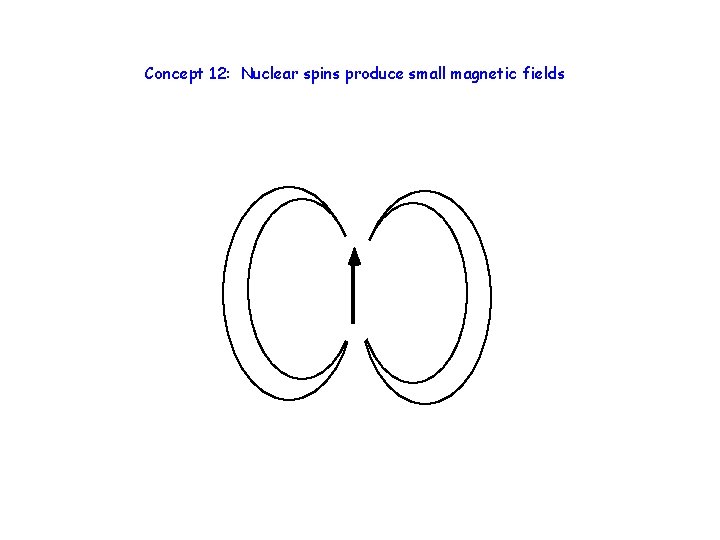
Concept 12: Nuclear spins produce small magnetic fields
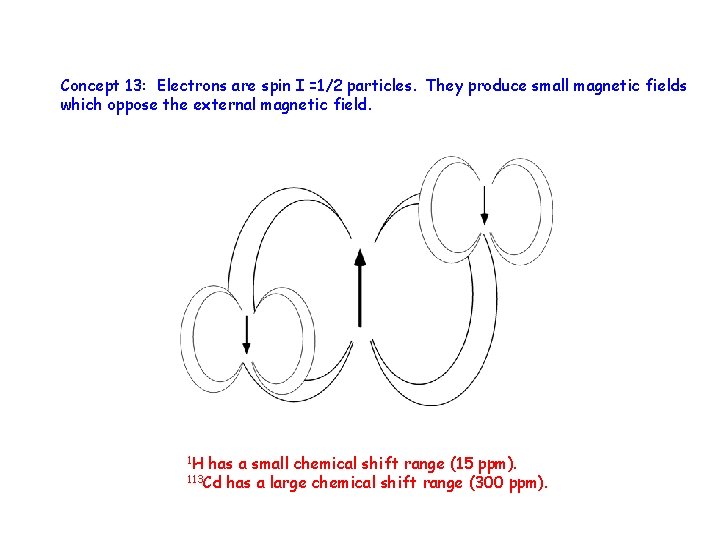
Concept 13: Electrons are spin I =1/2 particles. They produce small magnetic fields which oppose the external magnetic field. 1 H has a small chemical shift range (15 ppm). has a large chemical shift range (300 ppm). 113 Cd
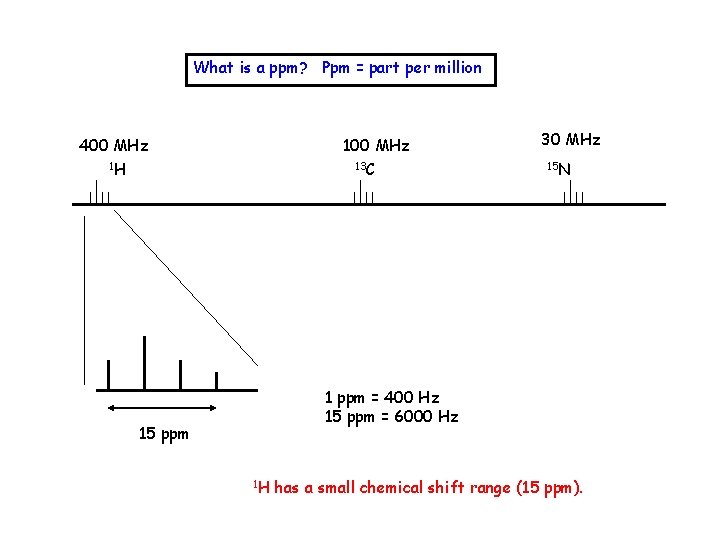
What is a ppm? Ppm = part per million 400 MHz 1 H 100 MHz 13 C 30 MHz 15 N 1 ppm = 400 Hz 15 ppm = 6000 Hz 15 ppm 1 H has a small chemical shift range (15 ppm).
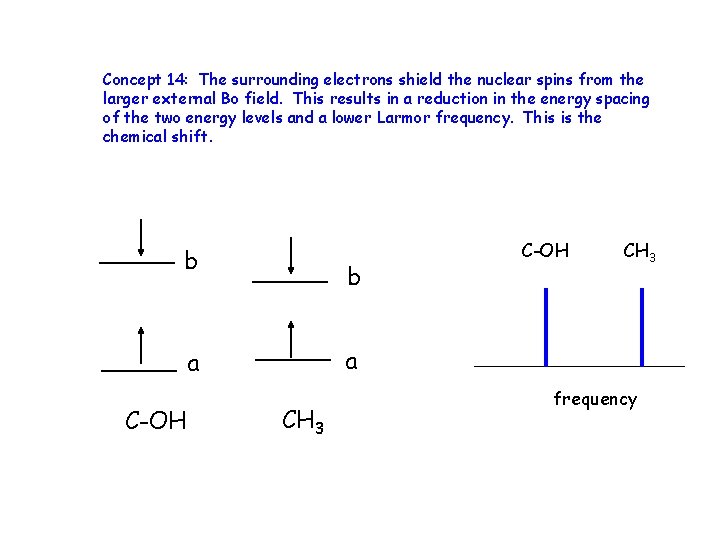
Concept 14: The surrounding electrons shield the nuclear spins from the larger external Bo field. This results in a reduction in the energy spacing of the two energy levels and a lower Larmor frequency. This is the chemical shift. b b CH 3 a a C-OH CH 3 frequency
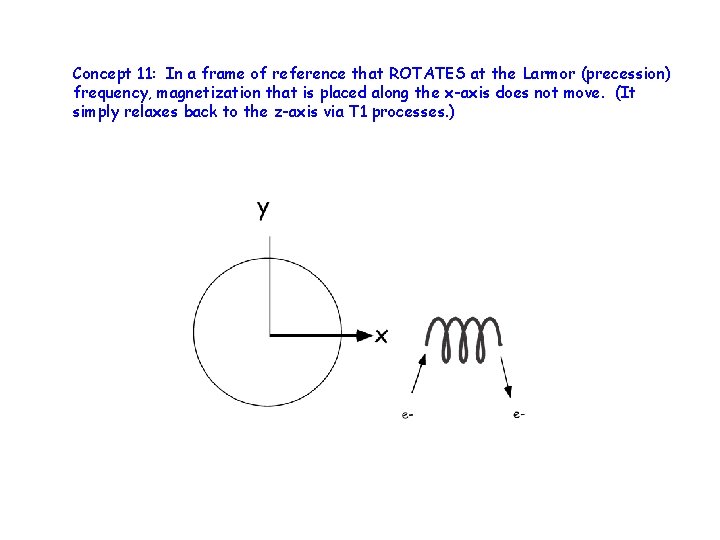
Concept 11: In a frame of reference that ROTATES at the Larmor (precession) frequency, magnetization that is placed along the x-axis does not move. (It simply relaxes back to the z-axis via T 1 processes. )
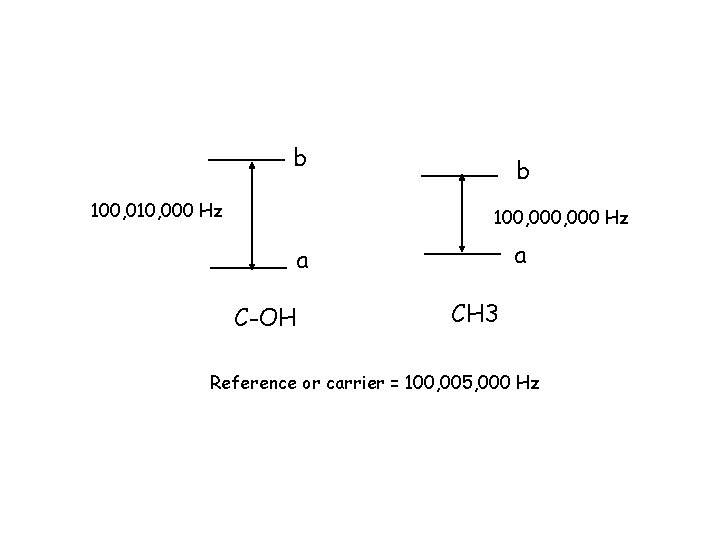
b 100, 010, 000 Hz b 100, 000 Hz a a C-OH CH 3 Reference or carrier = 100, 005, 000 Hz
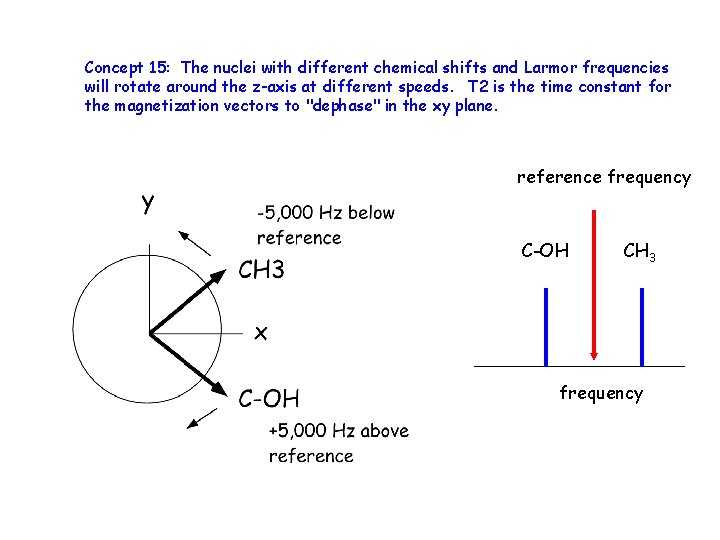
Concept 15: The nuclei with different chemical shifts and Larmor frequencies will rotate around the z-axis at different speeds. T 2 is the time constant for the magnetization vectors to "dephase" in the xy plane. reference frequency C-OH CH 3 frequency
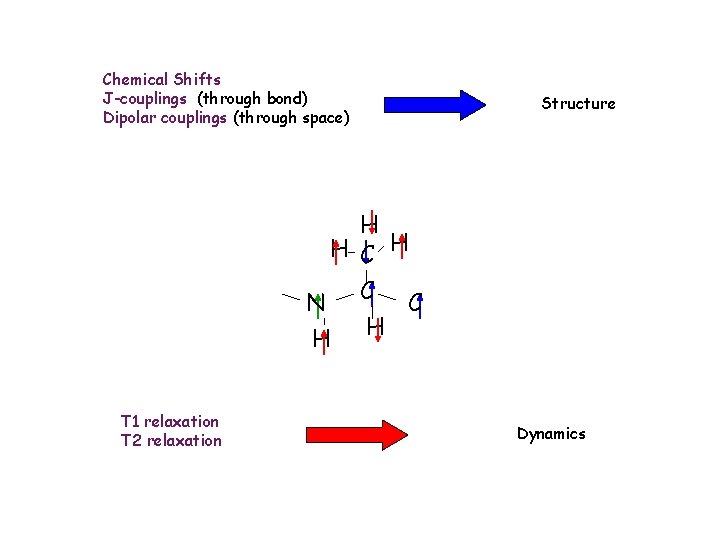
Chemical Shifts J-couplings (through bond) Dipolar couplings (through space) Structure H H C H N C C H H T 1 relaxation T 2 relaxation Dynamics
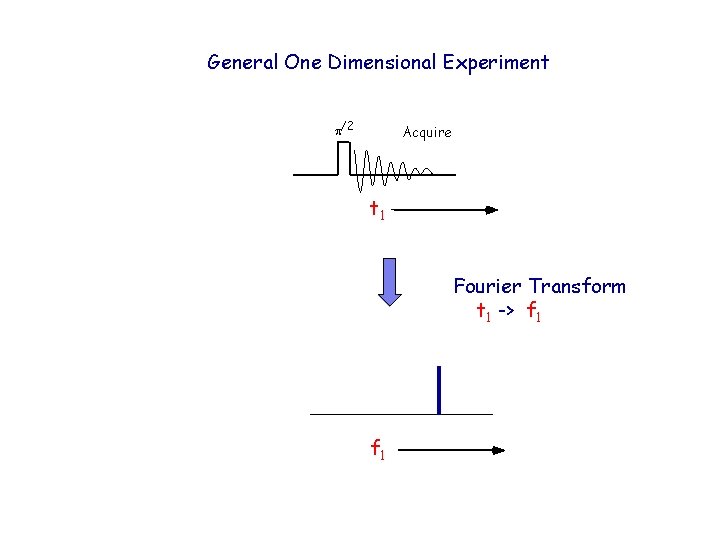
General One Dimensional Experiment p/2 Acquire t 1 Fourier Transform t 1 -> f 1
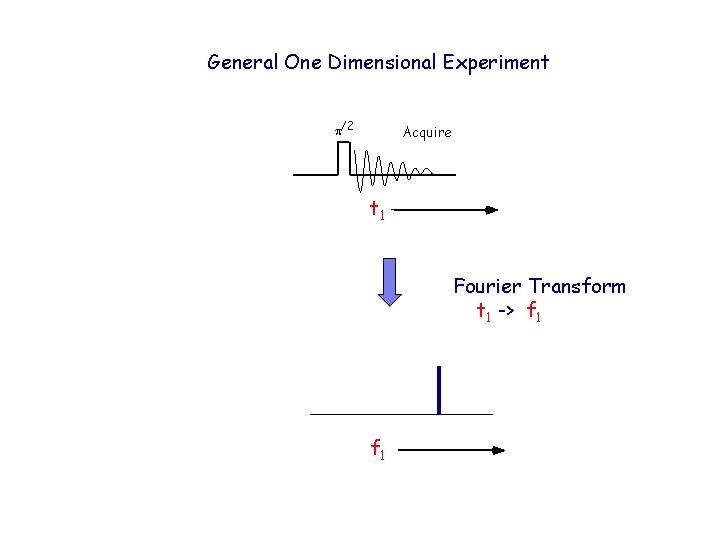
General One Dimensional Experiment p/2 Acquire t 1 Fourier Transform t 1 -> f 1
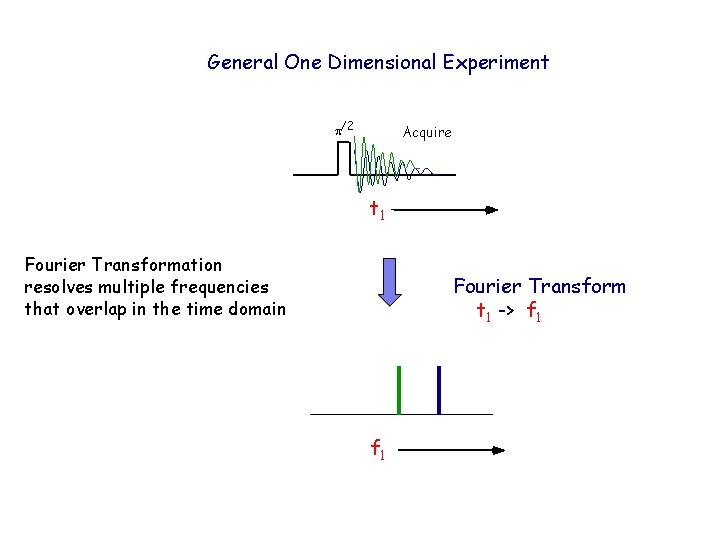
General One Dimensional Experiment p/2 Acquire t 1 Fourier Transformation resolves multiple frequencies that overlap in the time domain Fourier Transform t 1 -> f 1
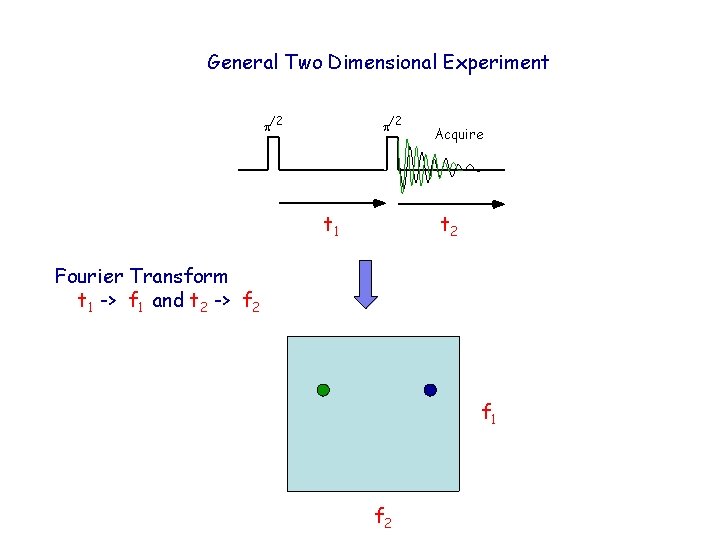
General Two Dimensional Experiment p/2 t 1 Acquire t 2 Fourier Transform t 1 -> f 1 and t 2 -> f 2 f 1 f 2
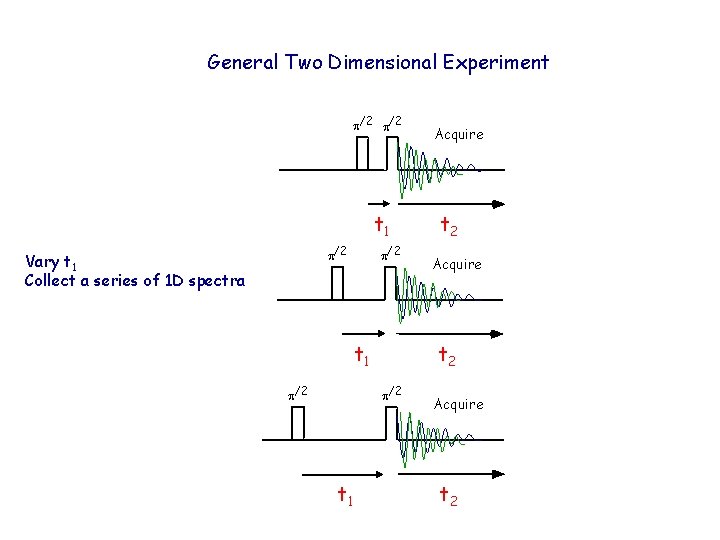
General Two Dimensional Experiment p/2 t 1 p/2 Vary t 1 Collect a series of 1 D spectra p/2 t 1 p/2 t 2 Acquire t 2 p/2 t 1 Acquire t 2
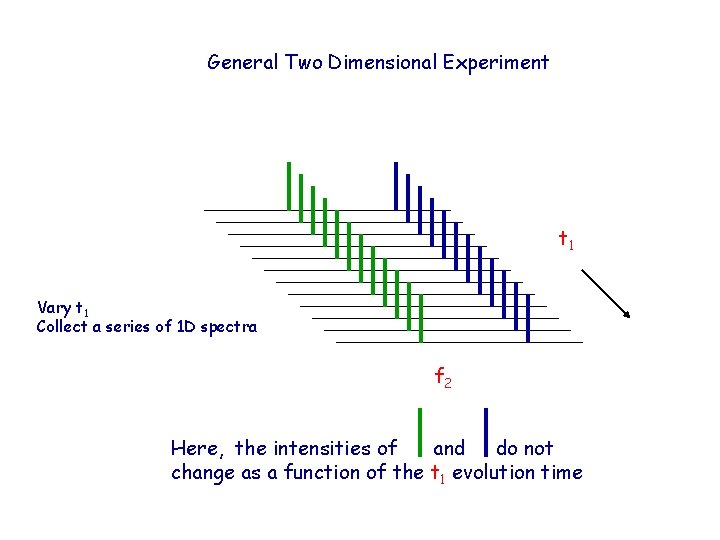
General Two Dimensional Experiment t 1 Vary t 1 Collect a series of 1 D spectra f 2 Here, the intensities of and do not change as a function of the t 1 evolution time
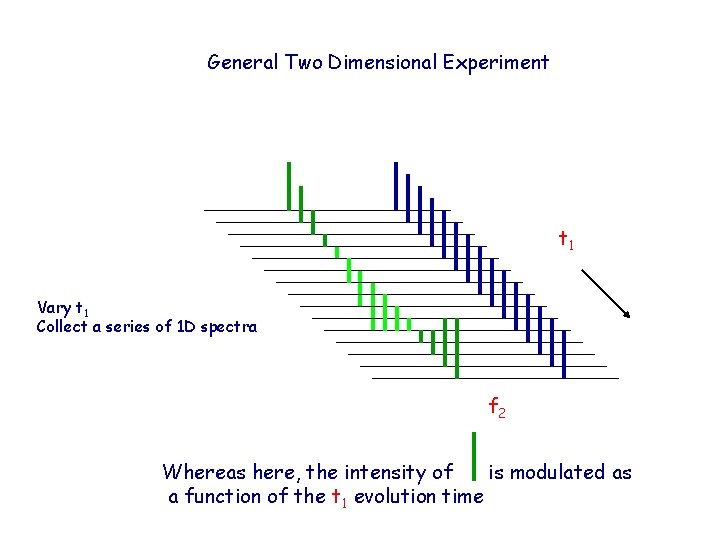
General Two Dimensional Experiment t 1 Vary t 1 Collect a series of 1 D spectra f 2 Whereas here, the intensity of is modulated as a function of the t 1 evolution time
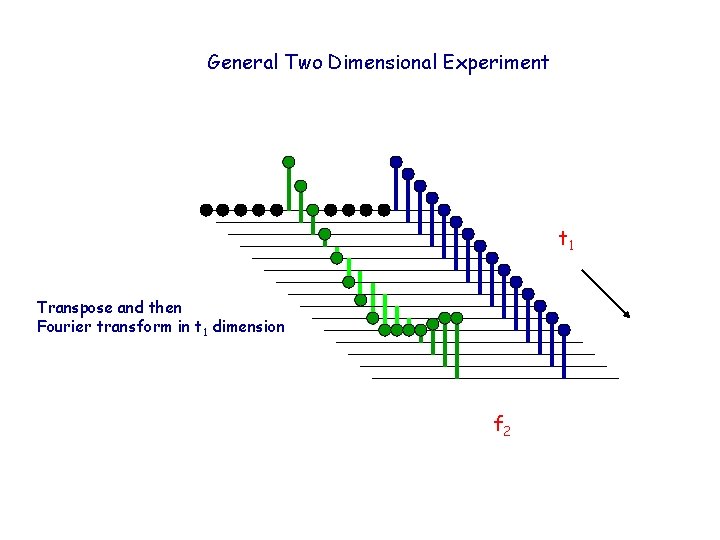
General Two Dimensional Experiment t 1 Transpose and then Fourier transform in t 1 dimension f 2
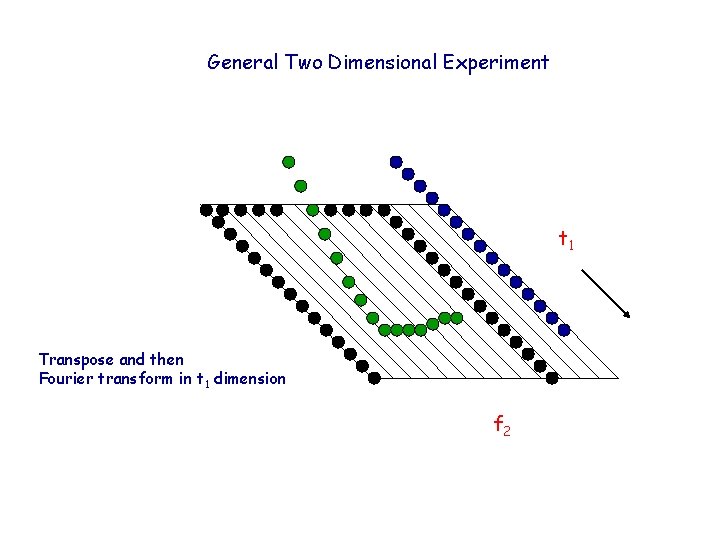
General Two Dimensional Experiment t 1 Transpose and then Fourier transform in t 1 dimension f 2
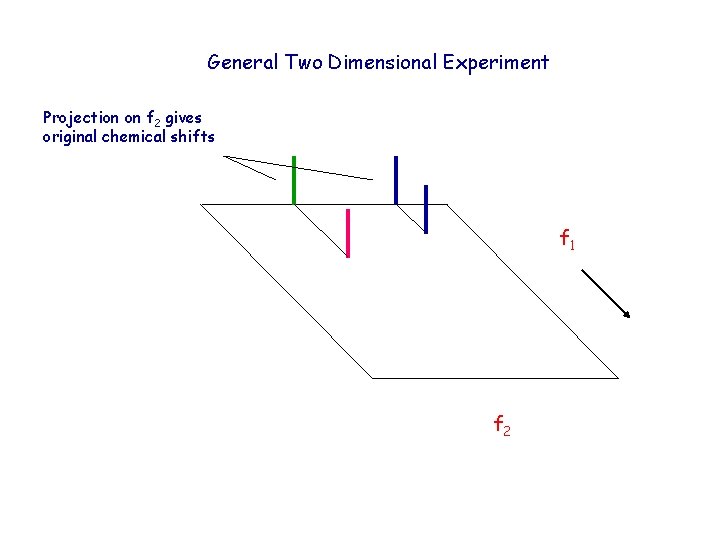
General Two Dimensional Experiment Projection on f 2 gives original chemical shifts f 1 f 2
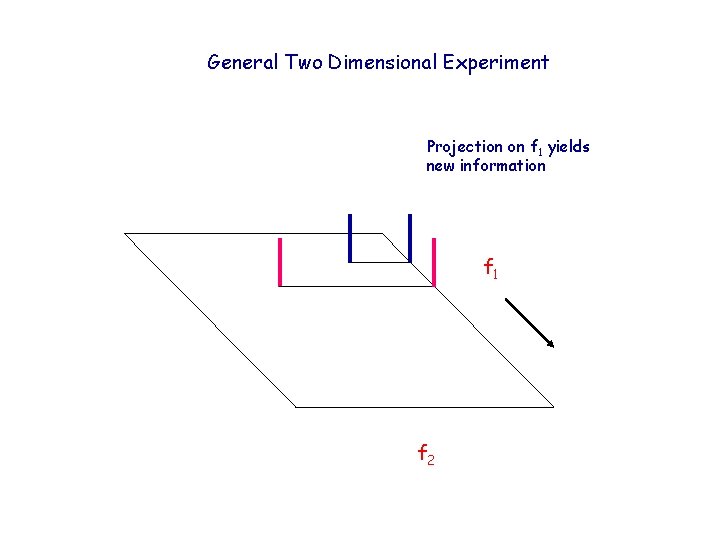
General Two Dimensional Experiment Projection on f 1 yields new information f 1 f 2
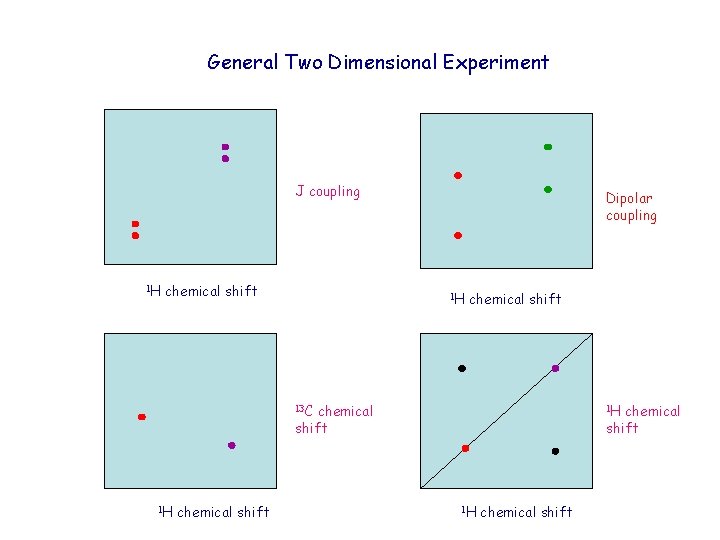
General Two Dimensional Experiment J coupling 1 H chemical shift Dipolar coupling 1 H chemical shift 13 C chemical shift 1 H chemical shift
- Slides: 43