Brownian Dynamics with Hydrodynamic Interactions Application to Lipid

Brownian Dynamics with Hydrodynamic Interactions: Application to Lipid Bilayers and Biomembranes Frank L. H. Brown University of California, Santa Barbara

Journal of Chemical Physics, 69, 1352 -1360 (1978). (910 citations)

Limitations of Fully Atomic Molecular Dynamics Simulation A recent “large” membrane simulation (Pitman et. al. , JACS, 127, 4576 (2005)) • 1 rhodopsin, 99 lipids, 24 cholesterols, 7400 waters (43, 222 atoms total) • 5. 5 x 7. 7 x 10. 3 nm periodic box for 118 ns duration Length/time scales relevant to cellular biology • ms, m (and longer) • A 1. 0 x 0. 1 m simulation for 1 ms would be approximately 2 x 109 more expensive than our abilities in 2005 • Moore’s law: this might be possible in 46 yrs.

Outline • • • Elastic membrane model (Energetics) Elastic membrane model (Dynamics) Brownian dynamics of Fourier modes Protein motion on the surface of the red blood cell Fluctuations in intermembrane junctions and active membranes
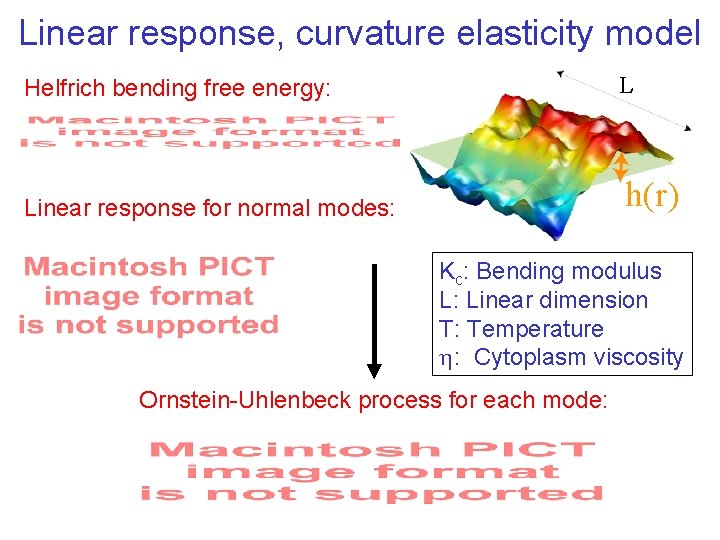
Linear response, curvature elasticity model Helfrich bending free energy: L Linear response for normal modes: h(r) Kc: Bending modulus L: Linear dimension T: Temperature : Cytoplasm viscosity Ornstein-Uhlenbeck process for each mode:

Relaxation frequencies Solve for relaxation of membrane modes coupled to a fluid in the overdamped limit: Non-inertial Navier-Stokes eq: Nonlocal Langevin equation: Oseen tensor (for infinite medium) Bending force R. Granek, J. Phys. II France, 7, 1761 -1788 (1997).

Membrane Dynamics

Harmonic Interactions • Membrane is pinned to the cytoskeleton at discrete points • Add interaction term to Helfrich free energy • When g is large, interaction mimics localized pinning L. Lin and F. Brown, Biophys. J. , 86, 764 (2004).

Pinned Membranes • Can diagonalize the free energy with interactions and find eigenmodes • Eigenmodes are described by Ornstein-Uhlenbeck processes

Extension to non-harmonic systems Helfrich bending free energy + additional interactions: Overdamped dynamics: (or generalized expressions) Solve via Brownian dynamics • Handle bulk of calculation in Fourier Space (FSBD) • Efficient handling of hydrodynamics • Natural way to coarse grain over short length scales L. Lin and F. Brown, Phys. Rev. Lett. , 93, 256001 (2004). L. Lin and F. Brown, Phys. Rev. E, 72, 011910 (2005).

Fourier Space Brownian Dynamics 1. Evaluate F(r) in real space (use h(r) from previous time step). 2. FFT F(r) to obtain Fk. 3. Draw k’s from Gaussian distributions. 4. Compute hk(t+ t) using above e. o. m. . 5. Inverse FFT hk(t+ t) to obtain h(r) for the next iteration.
Protein motion on the surface of red blood cells

S. Liu et al. , J. Cell. Biol. , 104, 527 (1987).
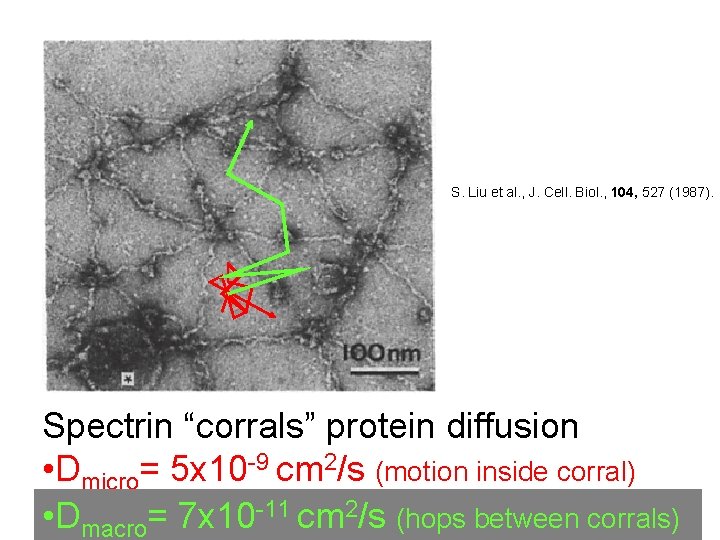
S. Liu et al. , J. Cell. Biol. , 104, 527 (1987). Spectrin “corrals” protein diffusion • Dmicro= 5 x 10 -9 cm 2/s (motion inside corral) • Dmacro= 7 x 10 -11 cm 2/s (hops between corrals)
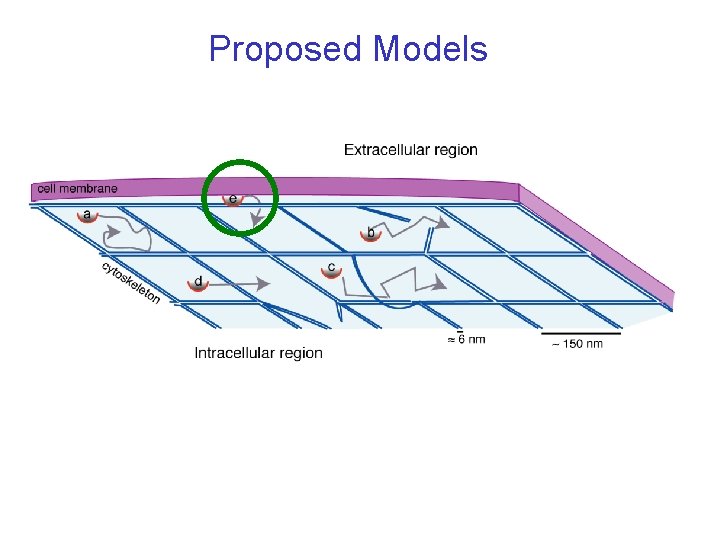
Proposed Models
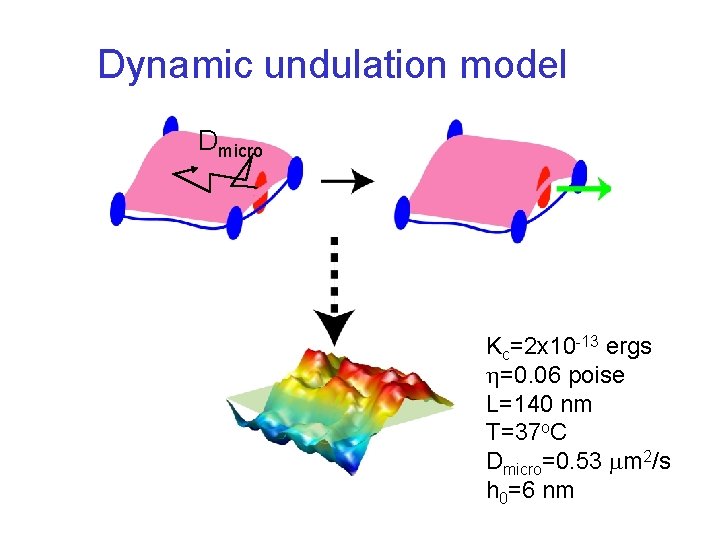
Dynamic undulation model Dmicro Kc=2 x 10 -13 ergs =0. 06 poise L=140 nm T=37 o. C Dmicro=0. 53 m 2/s h 0=6 nm

Explicit Cytoskeletal Interactions • Harmonic anchoring of spectrin cytoskeleton to the bilayer • Additional repulsive interaction along the edges of the corral to mimic spectrin L. Lin and F. Brown, Biophys. J. , 86, 764 (2004). L. Lin and F. Brown, Phys. Rev. Lett. , 93, 256001 (2004).

Dynamics with repulsive spectrin

Information extracted from the simulation • Probability that thermal bilayer fluctuation exceeds h 0=6 nm at equilibrium (intracellular domain size) • Probability that such a fluctuation persists longer than t 0=23 s (time to diffuse over spectrin) • Escape rate for protein from a corral • Macroscopic diffusion constant on cell surface (experimentally measured)
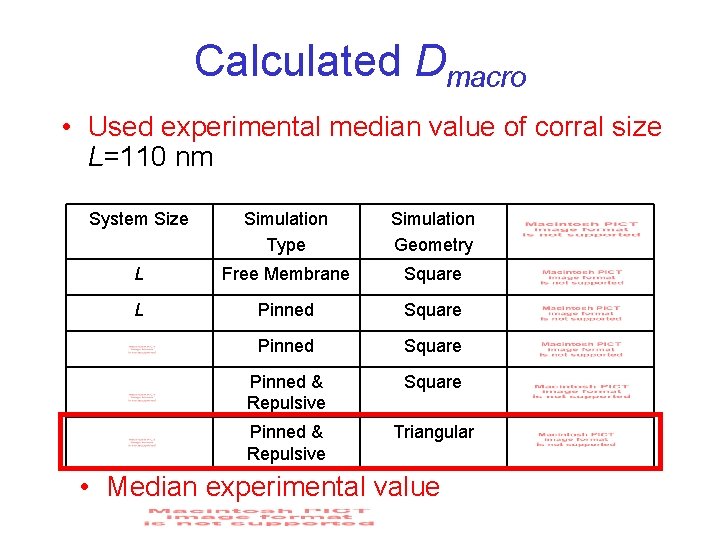
Calculated Dmacro • Used experimental median value of corral size L=110 nm System Size Simulation Type Simulation Geometry L Free Membrane Square L Pinned Square Pinned & Repulsive Triangular • Median experimental value
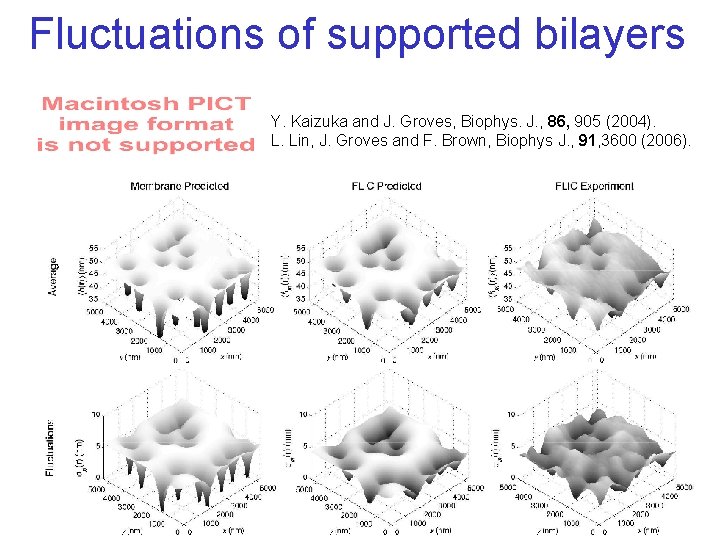
Fluctuations of supported bilayers Y. Kaizuka and J. Groves, Biophys. J. , 86, 905 (2004). L. Lin, J. Groves and F. Brown, Biophys J. , 91, 3600 (2006).
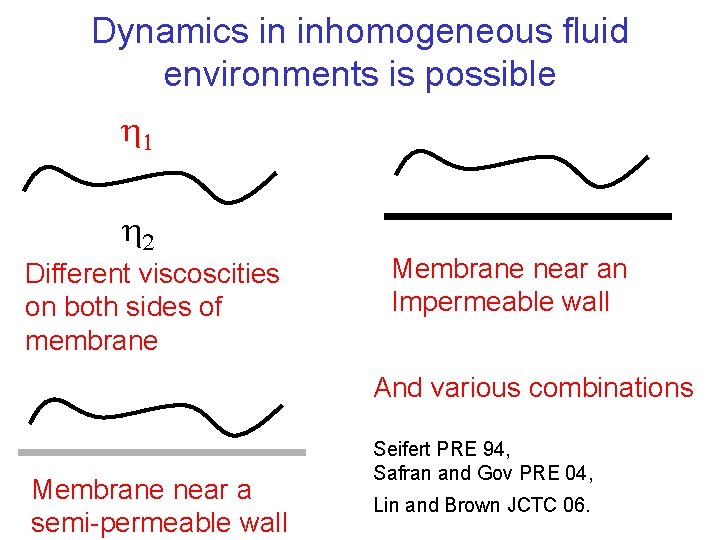
Dynamics in inhomogeneous fluid environments is possible Different viscoscities on both sides of membrane Membrane near an Impermeable wall And various combinations Membrane near a semi-permeable wall Seifert PRE 94, Safran and Gov PRE 04, Lin and Brown JCTC 06.
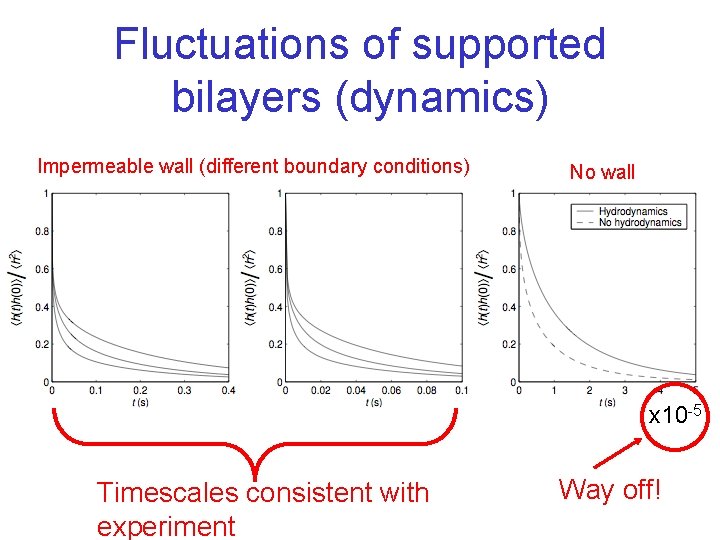
Fluctuations of supported bilayers (dynamics) Impermeable wall (different boundary conditions) No wall x 10 -5 Timescales consistent with experiment Way off!

Fluctuations of “active” bilayers Fluctuations of active membranes (experiments) J. -B. Manneville, P. Bassereau, D. Levy and J. Prost, PRL, 4356, 1999. Off kon koff Push down Push up L. Lin, N. Gov and F. L. H. Brown, JCP, 124, 074903 (2006).

Summary (elastic modeling) • Elastic models for membrane undulations can be extended to complex geometries and potentials via Brownian dynamics simulation. • “Thermal” undulations appear to be able to promote protein mobility on the RBC. • Other biophysical and biochemical systems are well suited to this approach.

Acknowledgements Lawrence Lin Ali Naji NSF, ACS-PRF, Sloan Foundation, UCSB
- Slides: 26