BRC Science Highlight Genetic architecture of floweringtime variation
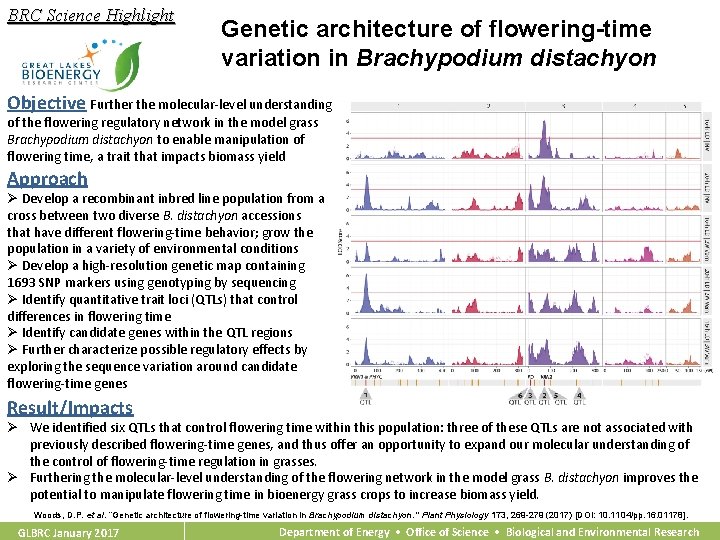
BRC Science Highlight Genetic architecture of flowering-time variation in Brachypodium distachyon Objective Further the molecular-level understanding of the flowering regulatory network in the model grass Brachypodium distachyon to enable manipulation of flowering time, a trait that impacts biomass yield Approach Ø Develop a recombinant inbred line population from a cross between two diverse B. distachyon accessions that have different flowering-time behavior; grow the population in a variety of environmental conditions Ø Develop a high-resolution genetic map containing 1693 SNP markers using genotyping by sequencing Ø Identify quantitative trait loci (QTLs) that control differences in flowering time Ø Identify candidate genes within the QTL regions Ø Further characterize possible regulatory effects by exploring the sequence variation around candidate flowering-time genes Result/Impacts Ø We identified six QTLs that control flowering time within this population: three of these QTLs are not associated with previously described flowering-time genes, and thus offer an opportunity to expand our molecular understanding of the control of flowering-time regulation in grasses. Ø Furthering the molecular-level understanding of the flowering network in the model grass B. distachyon improves the potential to manipulate flowering time in bioenergy grass crops to increase biomass yield. Woods, D. P. et al. “Genetic architecture of flowering-time variation in Brachypodium distachyon. ” Plant Physiology 173, 269 -279 (2017) [DOI: 10. 1104/pp. 16. 01178]. GLBRC January 2017 Department of Energy • Office of Science • Biological and Environmental Research
- Slides: 1